

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.14
(597 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
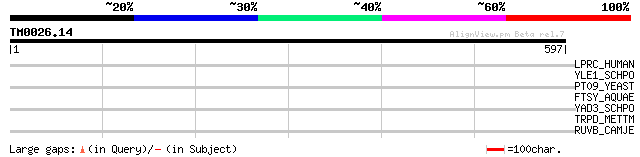
Score E
Sequences producing significant alignments: (bits) Value
LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP13... 44 0.001
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 41 0.010
PT09_YEAST (P32522) PET309 protein, mitochondrial precursor 34 1.2
FTSY_AQUAE (O67066) Cell division protein ftsY homolog 34 1.2
YAD3_SCHPO (Q09829) Hypothetical protein C4G8.03c in chromosome I 33 2.1
TRPD_METTM (P26925) Anthranilate phosphoribosyltransferase (EC 2... 32 3.6
RUVB_CAMJE (Q9PMT7) Holliday junction DNA helicase ruvB 31 8.1
>LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP130)
(Leucine-rich PPR-motif containing protein)
Length = 1273
Score = 44.3 bits (103), Expect = 0.001
Identities = 34/138 (24%), Positives = 60/138 (42%), Gaps = 4/138 (2%)
Query: 369 DLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYF 428
D+ +N ++ Y + Y + KM+E Q N VTY L+ + + G + EG
Sbjct: 40 DVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIASYCNVGDI-EGASKI 98
Query: 429 DKLLKNRSIQVKEDHYACLVDLCGRAGRLKEA---FYIIEGLGVKLSLSVWGPLLAGCNV 485
+K + + V E ++ LV RAG ++ A ++ G++ + LL
Sbjct: 99 LGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAGIEPGPDTYLALLNAYAE 158
Query: 486 HGNADIGKLVAKKILKVE 503
G+ D K +K+ K E
Sbjct: 159 KGDIDHVKQTLEKVEKFE 176
Score = 36.2 bits (82), Expect = 0.25
Identities = 30/145 (20%), Positives = 68/145 (46%), Gaps = 10/145 (6%)
Query: 43 MKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTT--MIDGYIKCGAIKEARKLFDQP 100
+ R + + +L ++G TD +K D + Q + + +++ G KEA+K+ + P
Sbjct: 734 LPRIHDVLCKLVEKGETDLIQKAMDFVSQEQGEMVMLYDLFFAFLQTGNYKEAKKIIETP 793
Query: 101 DAKKSVVTWTTMVNGYVKLNQIEEAESL------FYEMPERNELSWNIMIGGYGQNGQIE 154
+ + V NQ+E E L +E +R+++ +N ++ Y NG +
Sbjct: 794 GIRARSARLQWFCDRCVANNQVETLEKLVELTQKLFEC-DRDQMYYN-LLKLYKINGDWQ 851
Query: 155 KALDLFRRMPKRNLVSWNIIIKALA 179
+A ++ ++ + N++ ++ LA
Sbjct: 852 RADAVWNKIQEENVIPREKTLRLLA 876
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 40.8 bits (94), Expect = 0.010
Identities = 47/247 (19%), Positives = 102/247 (41%), Gaps = 18/247 (7%)
Query: 203 NVVSWNAMITGYAQNRRLDEALELFERMPERDM----ASWNAMLTGFFQNGELNRAEKLF 258
+V +NA+++ + RR E +LF+ M E + ++ ++ + G+ + AEKLF
Sbjct: 925 SVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEKLF 984
Query: 259 AELP-----QKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACS 313
AE+ Q V + +M+ Q + E + ++P++ T+ ++ A
Sbjct: 985 AEMENQPNYQPRVAPYNTMIQFEVQTMFNREKALFYYNRLCATDIEPSSHTYKLLMDAYG 1044
Query: 314 GLASLTEG--QQIHQLISKTGFQENTRVVSALINMYSK-CGELHIARKIFDDGLLR---- 366
L + G + + +L+ +T + +A I++ ++ A + + L +
Sbjct: 1045 TLKPVNVGSVKAVLELMERTDVPILSMHYAAYIHILGNVVSDVQAATSCYMNALAKHDAG 1104
Query: 367 --QRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEG 424
Q D + I + + E I + + M+ N L+ + G++ +
Sbjct: 1105 EIQLDANLFQSQIESLIANDRIVEGIQIVSDMKRYNVSLNAYIVNALIKGFTKVGMISKA 1164
Query: 425 IQYFDKL 431
YFD L
Sbjct: 1165 RYYFDLL 1171
Score = 33.9 bits (76), Expect = 1.2
Identities = 39/205 (19%), Positives = 89/205 (43%), Gaps = 11/205 (5%)
Query: 256 KLFAELPQKDVITWTSMMTGYAQHGLSEEA---LKMFTKMQANGGLKPNNGTFVTVLGAC 312
K+ +P+ T+ ++ + G +++A L +F + + + +KP+ + VL
Sbjct: 880 KMLGYIPRAS--TFAHLINNSTRRGDTDDATTALNIFEETKRHN-VKPSVFLYNAVLSKL 936
Query: 313 SGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIF---DDGLLRQRD 369
TE ++ Q + ++G + +IN + G+ +A K+F ++ Q
Sbjct: 937 GRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEMENQPNYQPR 996
Query: 370 LISWNGMIAAYAHHGYGNE-AINLFNKMQELGFQANDVTYVELLTACSHAGLVDEG-IQY 427
+ +N MI + E A+ +N++ + + TY L+ A V+ G ++
Sbjct: 997 VAPYNTMIQFEVQTMFNREKALFYYNRLCATDIEPSSHTYKLLMDAYGTLKPVNVGSVKA 1056
Query: 428 FDKLLKNRSIQVKEDHYACLVDLCG 452
+L++ + + HYA + + G
Sbjct: 1057 VLELMERTDVPILSMHYAAYIHILG 1081
Score = 31.2 bits (69), Expect = 8.1
Identities = 26/106 (24%), Positives = 49/106 (45%), Gaps = 6/106 (5%)
Query: 318 LTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFD----DGLLRQRDLISW 373
+ EG QI + + N +V+ALI ++K G + AR FD +G + ++ ++
Sbjct: 1126 IVEGIQIVSDMKRYNVSLNAYIVNALIKGFTKVGMISKARYYFDLLECEG-MSGKEPSTY 1184
Query: 374 NGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAG 419
M+ AY G +A+ + +++ + V + L SH G
Sbjct: 1185 ENMVRAYLSVNDGRKAMEIVEQLKRKRYPLPVVNRISSLVN-SHMG 1229
>PT09_YEAST (P32522) PET309 protein, mitochondrial precursor
Length = 965
Score = 33.9 bits (76), Expect = 1.2
Identities = 24/113 (21%), Positives = 53/113 (46%), Gaps = 6/113 (5%)
Query: 343 LINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAA--YAHHGYGNEA--INLFNKMQE 398
L+ ++ GEL K++ L +R +I ++ + YAH+ G+ A + F ++
Sbjct: 352 LMYTMARIGELDSVNKLYTQ--LLRRGMIPTYAVLQSLLYAHYKVGDFAACFSHFELFKK 409
Query: 399 LGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLC 451
+ T+ +L +D + +L ++ S+++ E H+A L+ +C
Sbjct: 410 YDITPSTATHTIMLKVYRGLNDLDGAFRILKRLSEDPSVEITEGHFALLIQMC 462
>FTSY_AQUAE (O67066) Cell division protein ftsY homolog
Length = 461
Score = 33.9 bits (76), Expect = 1.2
Identities = 60/289 (20%), Positives = 119/289 (40%), Gaps = 31/289 (10%)
Query: 50 ISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTW 109
I L K+G+ + A K+ + + + L + + Y++ G + +A L + K
Sbjct: 22 ILELIKQGKKEKAEKILKKYASQREDLAELLFELYVQEGKLVQAYPLLKKYGDKIGKAKE 81
Query: 110 TTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLV 169
V Y + + ++A F ++ + L +N+ Y Q + +KAL+ +R K LV
Sbjct: 82 RAKV--YQAVGEYQKAIEEFLKVGDFESL-YNV-AKIYHQLEKPDKALEYAQRAEK--LV 135
Query: 170 SW-------NIIIKALARCGRIEDAQWHLI-RCRKGMMPLRNVVSWNAMITGYAQNRRLD 221
+ N I + G IE+ + ++ + R+G+ + V + + G R++D
Sbjct: 136 PYEKKEELENFITQLKKELGLIEEKKESILDKLRRGLQKTKEAVEFGVLFRG----RKVD 191
Query: 222 EALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGL 281
E E FE + E + + + T + EKL E +K++ + + L
Sbjct: 192 E--EFFEELEEMLVKADVGVKTA------VELTEKLRKEAIRKNIKEGEKI-----KELL 238
Query: 282 SEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISK 330
+E ++ Q + G + +G + T G+ HQL K
Sbjct: 239 KKELKELLKNCQGELKIPEKVGAVLLFVGVNGSGKTTTIGKLAHQLKQK 287
>YAD3_SCHPO (Q09829) Hypothetical protein C4G8.03c in chromosome I
Length = 780
Score = 33.1 bits (74), Expect = 2.1
Identities = 35/141 (24%), Positives = 60/141 (41%), Gaps = 16/141 (11%)
Query: 7 PPLSFI------LMHAPKLNTHPTYILRTRLASTFSPNPNSEMKRCNSDISRLCKEGRTD 60
PPL + L+H L+ HP + ST S N +++ + SD S L + R
Sbjct: 232 PPLPVLKNSERNLLHRQLLSNHPFFSQNNVSLSTNSKNYSTDFTKIQSDSSLL--QNRQQ 289
Query: 61 HARKLFDQMPQRDKHLWTTMIDGYIKCGAIK--EARKLFDQPDAKKSVVTWTTMVNGYVK 118
+ R DQ+ HL + I + +++ E+RKL + D + +N +
Sbjct: 290 NHRIETDQLSHFPDHLDPSRIPSPYQPSSLQPLESRKLHSKVDVH------SKKLNALSQ 343
Query: 119 LNQIEEAESLFYEMPERNELS 139
LN I +E++ + LS
Sbjct: 344 LNPILRSENVLQNDNHHSSLS 364
>TRPD_METTM (P26925) Anthranilate phosphoribosyltransferase (EC
2.4.2.18)
Length = 350
Score = 32.3 bits (72), Expect = 3.6
Identities = 29/97 (29%), Positives = 42/97 (42%), Gaps = 10/97 (10%)
Query: 402 QANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAG-RLKE- 459
+ DV LLTA + G + I F + ++ R+++VK +VD CG G R K
Sbjct: 32 EMGDVQIAALLTALAMKGETVDEITGFARAMRERAVKVKIPESMRVVDACGTGGDRFKSY 91
Query: 460 -----AFYIIEGLGVKLSLSVWGPLLAGCNVHGNADI 491
A I GVK++ + C G ADI
Sbjct: 92 NVSTAAAIIAAAAGVKVAKHGNRAVTGSC---GGADI 125
>RUVB_CAMJE (Q9PMT7) Holliday junction DNA helicase ruvB
Length = 335
Score = 31.2 bits (69), Expect = 8.1
Identities = 16/28 (57%), Positives = 20/28 (71%), Gaps = 1/28 (3%)
Query: 387 NEAINLFNKMQELGFQANDVTYVELLTA 414
NEA+N + ELGF A D+ Y+ELLTA
Sbjct: 245 NEALNSLG-VNELGFDAMDLRYLELLTA 271
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,766,306
Number of Sequences: 164201
Number of extensions: 2979858
Number of successful extensions: 6682
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 6665
Number of HSP's gapped (non-prelim): 20
length of query: 597
length of database: 59,974,054
effective HSP length: 116
effective length of query: 481
effective length of database: 40,926,738
effective search space: 19685760978
effective search space used: 19685760978
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0026.14


