

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
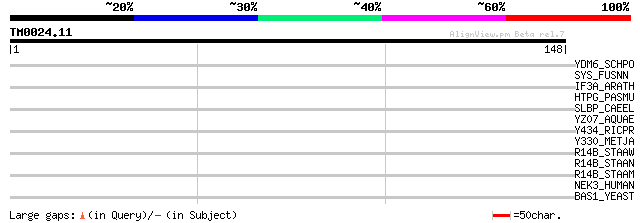
Query= TM0024.11
(148 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YDM6_SCHPO (P87137) Hypothetical protein C57A7.06 in chromosome I 33 0.29
SYS_FUSNN (Q8RH11) Seryl-tRNA synthetase (EC 6.1.1.11) (Serine--... 30 1.5
IF3A_ARATH (Q9LD55) Eukaryotic translation initiation factor 3 s... 30 1.5
HTPG_PASMU (Q9CM20) Chaperone protein htpG (Heat shock protein h... 30 1.5
SLBP_CAEEL (Q09599) Histone RNA hairpin-binding protein (Histone... 30 1.9
YZ07_AQUAE (O66402) Hypothetical protein AA07 30 2.5
Y434_RICPR (O05965) Hypothetical protein RP434 28 9.5
Y330_METJA (Q57776) Hypothetical protein MJ0330 28 9.5
R14B_STAAW (P66416) 30S ribosomal protein S14-2 28 9.5
R14B_STAAN (P66415) 30S ribosomal protein S14-2 28 9.5
R14B_STAAM (P66414) 30S ribosomal protein S14-2 28 9.5
NEK3_HUMAN (P51956) Serine/threonine-protein kinase Nek3 (EC 2.7... 28 9.5
BAS1_YEAST (P22035) MYB-like DNA-binding protein BAS1 28 9.5
>YDM6_SCHPO (P87137) Hypothetical protein C57A7.06 in chromosome I
Length = 929
Score = 32.7 bits (73), Expect = 0.29
Identities = 26/99 (26%), Positives = 49/99 (49%), Gaps = 10/99 (10%)
Query: 34 LREILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEYYRAYSPEFRNETLV 93
+RE++ E + +K+++KI K Y K +K KE +A +P+SE + NE +
Sbjct: 436 MRELMFREERKAKRVAKIKSKTYRKIRK---NRKEKEMALIPKSE------EDLENERIK 486
Query: 94 EGKIKRIFKFSKDSKSHLKRSSKS-EIAYSGHARRKAAN 131
+ + + + ++ K+ + K E A G R+A N
Sbjct: 487 SEEARALERMTQRHKNTSSWTRKMLERASHGEGTREAVN 525
>SYS_FUSNN (Q8RH11) Seryl-tRNA synthetase (EC 6.1.1.11)
(Serine--tRNA ligase) (SerRS)
Length = 421
Score = 30.4 bits (67), Expect = 1.5
Identities = 19/58 (32%), Positives = 32/58 (54%), Gaps = 3/58 (5%)
Query: 65 RSKEISVARLPESEYYRAYSPEFRNETLVEGK-IKRIFKFSKDSKSHLKRSSKSEIAY 121
R + + A LP+ YY AYSP FR E GK +K + + + +K + + + +E +Y
Sbjct: 239 RKEILEQAELPK--YYTAYSPCFRREAGSYGKDVKGLIRLHQFNKVEMVKITDAESSY 294
>IF3A_ARATH (Q9LD55) Eukaryotic translation initiation factor 3
subunit 10 (eIF-3 theta) (Eukaryotic translation
initiation factor 3 large subunit) (eIF3a) (p114)
Length = 987
Score = 30.4 bits (67), Expect = 1.5
Identities = 28/116 (24%), Positives = 53/116 (45%), Gaps = 4/116 (3%)
Query: 31 QGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEYYRAYSPEFRNE 90
Q ILREI E + ++ + + EKR K +K+ E V + +S RA + + +
Sbjct: 618 QRILREIEEKELEEAQALLEETEKRMKKGKKKPLLDGE-KVTK--QSVKERALTEQLKER 674
Query: 91 TLVEGKIKRIFKFSKDSKSHLKRSSKSEIAYSGHARRKAANSCHAYTDSTRESEIN 146
+E K++++ K + D KR + + + + RR + RE E++
Sbjct: 675 QEMEKKLQKLAK-TMDYLERAKREEAAPLIEAAYQRRLVEEREFYEREQQREVELS 729
>HTPG_PASMU (Q9CM20) Chaperone protein htpG (Heat shock protein
htpG) (High temperature protein G)
Length = 631
Score = 30.4 bits (67), Expect = 1.5
Identities = 22/70 (31%), Positives = 37/70 (52%), Gaps = 5/70 (7%)
Query: 24 KFRNHVHQGILREILASEFQN-SKQISKIWEKRYLKSQKRWFRSKEISVARLPESEYYRA 82
K+ +H+ G+ EILA E+ + K+ WEK K+Q W R+K ++ E+Y+
Sbjct: 200 KYSDHI--GLPVEILAKEYDDEGKETGIKWEK-INKAQALWTRAKN-EISEEEYQEFYKH 255
Query: 83 YSPEFRNETL 92
S +F + L
Sbjct: 256 LSHDFTDPLL 265
>SLBP_CAEEL (Q09599) Histone RNA hairpin-binding protein (Histone
stem-loop binding protein)
Length = 367
Score = 30.0 bits (66), Expect = 1.9
Identities = 24/85 (28%), Positives = 40/85 (46%), Gaps = 7/85 (8%)
Query: 65 RSKEISVARLPESEYYRAYSPEFRNETLVEGKIKR----IFKFSKDS-KSHLKRSSKSEI 119
RS+EI A+ E Y+ Y+ E ++G+ R + FS+ S + +K+ +S
Sbjct: 215 RSREIDRAK--EKAVYQRYTSEVPLRDRIKGQHPRTPNKLINFSRRSWDTQIKKWKRSLY 272
Query: 120 AYSGHARRKAANSCHAYTDSTRESE 144
Y G + N+ Y+DS SE
Sbjct: 273 EYCGEEPSDSVNTSFCYSDSDALSE 297
>YZ07_AQUAE (O66402) Hypothetical protein AA07
Length = 317
Score = 29.6 bits (65), Expect = 2.5
Identities = 18/58 (31%), Positives = 31/58 (53%), Gaps = 4/58 (6%)
Query: 13 GTKSDYDFSTIKFRNHVHQGILREILASEFQ--NSKQISKIWEKRY--LKSQKRWFRS 66
GT++ YDF F V + LRE+L E + +++ +K +E++ +K WF S
Sbjct: 200 GTQTPYDFLLSSFLEEVLEETLRELLEKEIEKIEAERRAKEYEEKMEKIKEIVEWFES 257
>Y434_RICPR (O05965) Hypothetical protein RP434
Length = 212
Score = 27.7 bits (60), Expect = 9.5
Identities = 13/41 (31%), Positives = 24/41 (57%)
Query: 18 YDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLK 58
Y F +KF N+ I A+ ++ K++S++ +KRYL+
Sbjct: 97 YSFDPVKFINNAKTAFQMIIEAAYKKDVKELSELIDKRYLE 137
>Y330_METJA (Q57776) Hypothetical protein MJ0330
Length = 549
Score = 27.7 bits (60), Expect = 9.5
Identities = 24/76 (31%), Positives = 30/76 (38%), Gaps = 4/76 (5%)
Query: 7 VTTIPKGTKSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRS 66
V T G K DY S KF NH+ + FQ+ I K +R K K W
Sbjct: 387 VLTPTLGHKIDY-LSLEKFYNHLQNHFSVKFTFDRFQSEYFIQKFKGERLSKHVKLWTTF 445
Query: 67 KEISVARLPESEYYRA 82
+E+ EYY A
Sbjct: 446 QELVEG---TKEYYDA 458
>R14B_STAAW (P66416) 30S ribosomal protein S14-2
Length = 89
Score = 27.7 bits (60), Expect = 9.5
Identities = 11/26 (42%), Positives = 15/26 (57%)
Query: 7 VTTIPKGTKSDYDFSTIKFRNHVHQG 32
VT P+G ++ S I FR H H+G
Sbjct: 54 VTGRPRGVLRKFEMSRIAFREHAHKG 79
>R14B_STAAN (P66415) 30S ribosomal protein S14-2
Length = 89
Score = 27.7 bits (60), Expect = 9.5
Identities = 11/26 (42%), Positives = 15/26 (57%)
Query: 7 VTTIPKGTKSDYDFSTIKFRNHVHQG 32
VT P+G ++ S I FR H H+G
Sbjct: 54 VTGRPRGVLRKFEMSRIAFREHAHKG 79
>R14B_STAAM (P66414) 30S ribosomal protein S14-2
Length = 89
Score = 27.7 bits (60), Expect = 9.5
Identities = 11/26 (42%), Positives = 15/26 (57%)
Query: 7 VTTIPKGTKSDYDFSTIKFRNHVHQG 32
VT P+G ++ S I FR H H+G
Sbjct: 54 VTGRPRGVLRKFEMSRIAFREHAHKG 79
>NEK3_HUMAN (P51956) Serine/threonine-protein kinase Nek3 (EC
2.7.1.37) (NimA-related protein kinase 3) (HSPK 36)
Length = 506
Score = 27.7 bits (60), Expect = 9.5
Identities = 24/86 (27%), Positives = 43/86 (49%), Gaps = 4/86 (4%)
Query: 36 EILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEYYRAYSP-EFRNETLVE 94
E + E +NSK + K+ S+ R E S + E + +++ E NE LVE
Sbjct: 275 EEVLEEIKNSKHNTP--RKKTNPSRIRIALGNEASTVQEEEQDRKGSHTDLESINENLVE 332
Query: 95 GKIKRIFKFSKDSKS-HLKRSSKSEI 119
++R+ + K +KS HL+++S +
Sbjct: 333 SALRRVNREEKGNKSVHLRKASSPNL 358
>BAS1_YEAST (P22035) MYB-like DNA-binding protein BAS1
Length = 811
Score = 27.7 bits (60), Expect = 9.5
Identities = 14/41 (34%), Positives = 25/41 (60%)
Query: 19 DFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKS 59
D TI+ NH+ + I ++LA+ F+++ + SK KR+ S
Sbjct: 69 DVKTIQQSNHLSKCIAWDVLATRFKHTVRTSKDVRKRWTGS 109
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,428,645
Number of Sequences: 164201
Number of extensions: 555013
Number of successful extensions: 1892
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1887
Number of HSP's gapped (non-prelim): 13
length of query: 148
length of database: 59,974,054
effective HSP length: 100
effective length of query: 48
effective length of database: 43,553,954
effective search space: 2090589792
effective search space used: 2090589792
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0024.11


