

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0023.5
(266 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
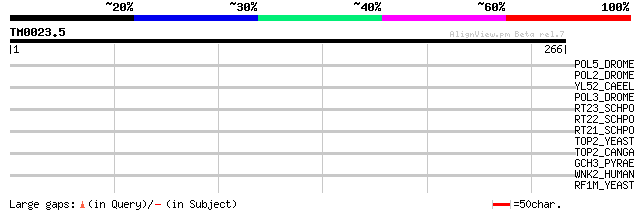
Score E
Sequences producing significant alignments: (bits) Value
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 39 0.010
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 39 0.010
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 35 0.14
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 35 0.25
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 33 0.55
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 33 0.55
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 33 0.55
TOP2_YEAST (P06786) DNA topoisomerase II (EC 5.99.1.3) 33 0.94
TOP2_CANGA (O93794) DNA topoisomerase II (EC 5.99.1.3) 32 1.2
GCH3_PYRAE (Q8ZU20) GTP cyclohydrolase III (EC 3.5.4.29) 31 3.6
WNK2_HUMAN (Q9Y3S1) Serine/threonine-protein kinase WNK2 (EC 2.7... 30 4.7
RF1M_YEAST (P30775) Peptide chain release factor 1, mitochondria... 30 7.9
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 39.3 bits (90), Expect = 0.010
Identities = 25/74 (33%), Positives = 40/74 (53%), Gaps = 1/74 (1%)
Query: 11 PLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLR 70
P L ++ A+ A+L Q++ + + I ++S +L E Y IEK LAI+ + LR
Sbjct: 434 PFHLTTDASNWAIGAVLSQDDQGRDRPIAYISRSLNKTEENYATIEKEMLAIIWSLDNLR 493
Query: 71 PY-FQSFQIKVKTD 83
Y + + IKV TD
Sbjct: 494 AYLYGAGTIKVYTD 507
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 39.3 bits (90), Expect = 0.010
Identities = 26/80 (32%), Positives = 37/80 (45%), Gaps = 4/80 (5%)
Query: 4 QKPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
Q P E +L ++ A+ A+L Q H I F+S TL E+ Y IEK LAI+
Sbjct: 500 QLPDFEKKFVLTTDASNLALGAVLSQNGHP----ISFISRTLNDHELNYSAIEKELLAIV 555
Query: 64 KTARRLRPYFQSFQIKVKTD 83
+ R Y Q + +D
Sbjct: 556 WATKTFRHYLLGRQFLIASD 575
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 35.4 bits (80), Expect = 0.14
Identities = 32/119 (26%), Positives = 46/119 (37%), Gaps = 8/119 (6%)
Query: 11 PLILYLAVTSTAVSAMLLQE-EHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRL 69
P ++Y + + A+L QE + I F S L AE RY + ALA++ RR
Sbjct: 1243 PFMIYTDASRKGIGAVLAQEGPDGQQHPIAFASKALSPAETRYHITDLEALAMMFALRRF 1302
Query: 70 RPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADILGLFSVAKA 128
+ I V TD KP +S L W I D+ ++ KA
Sbjct: 1303 KTIIYGTAITVFTD-------HKPLISLLKGSPLADRLWRWSIEILEFDVKIVYLAGKA 1354
Score = 34.7 bits (78), Expect = 0.25
Identities = 23/74 (31%), Positives = 40/74 (53%), Gaps = 2/74 (2%)
Query: 167 STGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIA-NLIMLPELQREAHLR-N 224
+TGETP L +G D M P+E+ + I + + +L+ L+ + ++ +E +R
Sbjct: 1675 NTGETPMFLMHGRDVMGPLEMSGEDAVGINYADMDEYKHLLTQELLKVQKIAKEHAMREQ 1734
Query: 225 EAGKGRVARKYASK 238
E+ K +KYASK
Sbjct: 1735 ESYKSLFDQKYASK 1748
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 34.7 bits (78), Expect = 0.25
Identities = 19/65 (29%), Positives = 32/65 (49%), Gaps = 4/65 (6%)
Query: 19 TSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRPYFQSFQI 78
+ A+ A+L Q+ H + ++S TL E+ Y IEK LAI+ + R Y
Sbjct: 516 SDVALGAVLSQDGHP----LSYISRTLNEHEINYSTIEKELLAIVWATKTFRHYLLGRHF 571
Query: 79 KVKTD 83
++ +D
Sbjct: 572 EISSD 576
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 33.5 bits (75), Expect = 0.55
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Query: 12 LILYLAVTSTAVSAMLLQE-EHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLR 70
++L + AV A+L Q+ + KY + + S + A++ Y +K LAI+K+ + R
Sbjct: 708 ILLETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWR 767
Query: 71 PYFQSF--QIKVKTD 83
Y +S K+ TD
Sbjct: 768 HYLESTIEPFKILTD 782
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 33.5 bits (75), Expect = 0.55
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Query: 12 LILYLAVTSTAVSAMLLQE-EHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLR 70
++L + AV A+L Q+ + KY + + S + A++ Y +K LAI+K+ + R
Sbjct: 708 ILLETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWR 767
Query: 71 PYFQSF--QIKVKTD 83
Y +S K+ TD
Sbjct: 768 HYLESTIEPFKILTD 782
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 33.5 bits (75), Expect = 0.55
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Query: 12 LILYLAVTSTAVSAMLLQE-EHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLR 70
++L + AV A+L Q+ + KY + + S + A++ Y +K LAI+K+ + R
Sbjct: 708 ILLETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWR 767
Query: 71 PYFQSF--QIKVKTD 83
Y +S K+ TD
Sbjct: 768 HYLESTIEPFKILTD 782
>TOP2_YEAST (P06786) DNA topoisomerase II (EC 5.99.1.3)
Length = 1428
Score = 32.7 bits (73), Expect = 0.94
Identities = 26/100 (26%), Positives = 47/100 (47%), Gaps = 5/100 (5%)
Query: 9 EVPLILYLAVTSTAVSAMLLQE-EHKKYKMIYFVSHTLQGAEVRY--QKIEKAALAILKT 65
++P ILY + + A + + ++ + ++ T+ G V Y +I K ILK
Sbjct: 275 DIPTILYERINNRWEVAFAVSDISFQQISFVNSIATTMGGTHVNYITDQIVKKISEILK- 333
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTT 105
++ + +SFQIK I + + + P + EQLTT
Sbjct: 334 -KKKKKSVKSFQIKNNMFIFINCLIENPAFTSQTKEQLTT 372
>TOP2_CANGA (O93794) DNA topoisomerase II (EC 5.99.1.3)
Length = 1406
Score = 32.3 bits (72), Expect = 1.2
Identities = 26/100 (26%), Positives = 44/100 (44%), Gaps = 8/100 (8%)
Query: 10 VPLILYLAVTSTAVSAMLLQE-EHKKYKMIYFVSHTLQGAEVRY---QKIEKAALAILKT 65
+P ILY V+ A + + K+ + ++ T G V Y Q + K + + K
Sbjct: 275 IPTILYERVSDRWEIAFAVSDISFKQVSFVNSIATTTGGTHVNYIADQIVRKVSDILKKK 334
Query: 66 ARRLRPYFQSFQIKVKTDILLRQVWQKPDLSGAPPEQLTT 105
+ ++PY QIK I + + + P + EQLTT
Sbjct: 335 KKNIKPY----QIKNNMFIFINCLIENPAFTSQTKEQLTT 370
>GCH3_PYRAE (Q8ZU20) GTP cyclohydrolase III (EC 3.5.4.29)
Length = 221
Score = 30.8 bits (68), Expect = 3.6
Identities = 17/63 (26%), Positives = 28/63 (43%), Gaps = 5/63 (7%)
Query: 131 KFIVVAVDYFTKWIEA-----EAVETITVAKIRNFLWRRITSTGETPYKLTYGSDAMLPV 185
K ++ + + +W E+ E + AKI + LW+ TS G P+ L Y L
Sbjct: 3 KITLIRLRGYREWTESLGPRREHIIQTVQAKIHSALWKYFTSIGALPHHLRYDFSLALTT 62
Query: 186 EVE 188
+E
Sbjct: 63 NIE 65
>WNK2_HUMAN (Q9Y3S1) Serine/threonine-protein kinase WNK2 (EC
2.7.1.37) (Protein kinase with no lysine 2) (Protein
kinase, lysine-deficient 2) (P/OKcl.13)
Length = 2297
Score = 30.4 bits (67), Expect = 4.7
Identities = 17/39 (43%), Positives = 18/39 (45%)
Query: 93 PDLSGAPPEQLTTMTAPWPFAIWGADILGLFSVAKAQMK 131
P+ S P LT PW A GA L LFS AQ K
Sbjct: 1340 PEASQGPCRGLTLPCLPWRRAACGAVFLSLFSAESAQSK 1378
>RF1M_YEAST (P30775) Peptide chain release factor 1, mitochondrial
precursor (MRF-1)
Length = 413
Score = 29.6 bits (65), Expect = 7.9
Identities = 16/51 (31%), Positives = 27/51 (52%)
Query: 192 WRVITMNEEENNDNLIANLIMLPELQREAHLRNEAGKGRVARKYASKVDSR 242
+R+I+ NE E+ +I ++ + E LR EAG RV R +++ R
Sbjct: 190 YRIISKNENESGSGIIDAILSIEEAGSYDRLRFEAGVHRVQRIPSTETKGR 240
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,024,568
Number of Sequences: 164201
Number of extensions: 1104608
Number of successful extensions: 2614
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 2607
Number of HSP's gapped (non-prelim): 14
length of query: 266
length of database: 59,974,054
effective HSP length: 108
effective length of query: 158
effective length of database: 42,240,346
effective search space: 6673974668
effective search space used: 6673974668
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0023.5


