

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.23
(1569 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
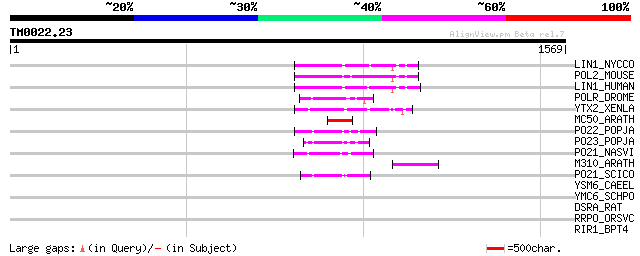
Score E
Sequences producing significant alignments: (bits) Value
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 100 3e-20
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 96 1e-18
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 91 2e-17
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 79 1e-13
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 73 5e-12
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 64 3e-09
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 64 4e-09
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 62 9e-09
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 61 2e-08
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 53 8e-06
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 46 7e-04
YSM6_CAEEL (Q10126) Hypothetical protein F52C9.6 in chromosome III 33 6.2
YMC6_SCHPO (P05511) Hypothetical 91 kDa protein in cob intron 33 6.2
DSRA_RAT (P55266) Double-stranded RNA-specific adenosine deamina... 33 6.2
RRPO_ORSVC (P89659) RNA-directed RNA polymerase (EC 2.7.7.48) (1... 33 8.0
RIR1_BPT4 (P32282) Ribonucleoside-diphosphate reductase alpha ch... 33 8.0
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 100 bits (249), Expect = 3e-20
Identities = 91/364 (25%), Positives = 166/364 (45%), Gaps = 33/364 (9%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEK 863
K++ NR++Q ++++I Q FIPG N ++ H+N + K +++ +D EK
Sbjct: 544 KILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHINKLKNK-DHMILSIDAEK 602
Query: 864 ACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPL 923
A ++ F+ TL G T + LI S + +++ NGVKL G R G PL
Sbjct: 603 AFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFPLRSGTRQGCPL 662
Query: 924 SPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKV 983
SP LF + +E L I+I++ + + I I FADD+++ + K+
Sbjct: 663 SPLLFNIVMEVLAIAIRE-----EKAIKGIHIGSEEIKLSLFADDMIVYLENTRDSTTKL 717
Query: 984 QNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTA--NLERYLGFPIFQ 1041
++ ++ SG +N KS F + +A+K V SIPFT +YLG + +
Sbjct: 718 LEVIKEYSNVSGYKINTHKSVAFIYTNNNQAEKT---VKDSIPFTVVPKKMKYLGVYLTK 774
Query: 1042 GRQT--QEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVL--------NAIPV-YPMQ 1090
+ +E+++ + + WK + G+ + K+ + NAIP+ P+
Sbjct: 775 DVKDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVKMSILPKAIYNFNAIPIKAPLS 834
Query: 1091 LFWFH*AVCDHLDSLAKNFIWSRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLG 1150
F L+ + +FIW++ + + +A ++ GG+ + RL S++
Sbjct: 835 YF-------KDLEKIILHFIWNQKKPQ----IAKTLLSNKNKAGGITLPDLRLYYKSIVI 883
Query: 1151 KLVW 1154
K W
Sbjct: 884 KTAW 887
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 95.5 bits (236), Expect = 1e-18
Identities = 88/362 (24%), Positives = 162/362 (44%), Gaps = 29/362 (8%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEK 863
K++ NR+++ ++ +I P Q FIPG N ++H++N + K +++ LD EK
Sbjct: 571 KILANRIQEHIKAIIHPDQVGFIPGMQGWFNIRKSINVIHYINKLKDK-NHMIISLDAEK 629
Query: 864 ACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPL 923
A + F+ L G +++I S ++ NG KL I G R G PL
Sbjct: 630 AFDKIQHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGTRQGCPL 689
Query: 924 SPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKV 983
SPYLF + +E L +I+ Q + + ++I ++ IS L ADD+++ + R++
Sbjct: 690 SPYLFNIVLEVLARAIR---QQKEIKGIQIGKEEVKISLL--ADDMIVYISDPKNSTREL 744
Query: 984 QNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQGR 1043
N++ F G +N KS F +A+K +I T N +YLG + +
Sbjct: 745 LNLINSFGEVVGYKINSNKSMAFLYTKNKQAEK-EIRETTPFSIVTNNIKYLGVTLTKEV 803
Query: 1044 QT--QEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVL--------NAIPV-YPMQLF 1092
+ ++F + + WK + G+ + K+ + NAIP+ P Q F
Sbjct: 804 KDLYDKNFKSLKKEIKEDLRRWKDLPCSWIGRINIVKMAILPKAIYRFNAIPIKIPTQFF 863
Query: 1093 WFH*AVCDHLDSLAKNFIWSRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKL 1152
+ L+ F+W+ + R +A + + +GG+ + +L +++ K
Sbjct: 864 -------NELEGAICKFVWNNKKPR----IAKSLLKDKRTSGGITMPDLKLYYRAIVIKT 912
Query: 1153 VW 1154
W
Sbjct: 913 AW 914
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 90.9 bits (224), Expect = 2e-17
Identities = 83/371 (22%), Positives = 162/371 (43%), Gaps = 33/371 (8%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEK 863
K++ N+++Q +++LI Q FIP N I+ H+N R +++ +D EK
Sbjct: 544 KILANQIQQHIKKLIHHDQVGFIPAMQGWFNIRKSINIIQHIN-RTKDTNHMIISIDAEK 602
Query: 864 ACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPL 923
A + F+ L G T + +I + +++ NG KL G R G PL
Sbjct: 603 AFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANIILNGQKLEAPPLKTGTRQGCPL 662
Query: 924 SPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKV 983
SP L + +E L +I+ + E+ I + FADD+++ + + + +
Sbjct: 663 SPLLPNIVLEVLARAIRQ-----EKEIKGIQLGKEEVKLSLFADDMIVYLENPIVSAQNL 717
Query: 984 QNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLER--YLGFPIFQ 1041
++++F SG +N+ KS+ F + ++ + +++ +PFT +R YLG + +
Sbjct: 718 LKLISNFSKVSGYKINVQKSQAFLYTN---NRQTESQIMSELPFTIASKRIKYLGIQLTR 774
Query: 1042 GRQT--QEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVL--------NAIPV-YPMQ 1090
+ +E++ +L+ + WK + G+ + K+ + NAIP+ PM
Sbjct: 775 DVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKMAILPKVIYRFNAIPIKLPMT 834
Query: 1091 LFWFH*AVCDHLDSLAKNFIWSRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLG 1150
F L+ FIW++ +A T++ GG+ + +L + +
Sbjct: 835 FF-------TELEKTTLKFIWNQKRAH----IAKSTLSQKNKAGGITLPDFKLYYKATVT 883
Query: 1151 KLVWDLVNSHD 1161
K W + D
Sbjct: 884 KTAWYWYQNRD 894
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 79.0 bits (193), Expect = 1e-13
Identities = 63/230 (27%), Positives = 99/230 (42%), Gaps = 31/230 (13%)
Query: 820 PLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEKACGSVSWAFLRDTLCF 879
P Q F+P G DNA I+ ++ H + C + LD+ KA S+S A + DTL
Sbjct: 444 PRQRGFLPTDGCADNATIVDLVLRHSHKHFRSCY--IANLDVSKAFDSLSHASIYDTLRA 501
Query: 880 FGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISI 939
+G P V + SL +G P+RG++ GDPLSP LF L ++RL ++
Sbjct: 502 YGAPKGFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTL 561
Query: 940 QDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVN 999
+ G + + FADD++L A+ ++ ++ + + DF GL +N
Sbjct: 562 PSEIGAKVGNAI--------TNAAAFADDLVLFAETRMG-LQVLLDKTLDFLSIVGLKLN 612
Query: 1000 I--------------------AKSRDFSSVSVPRAKKADIFVVTSIPFTA 1029
A+S S +P K+ D + I FTA
Sbjct: 613 ADKCFTVGIKGQPKQKCTVLEAQSFYVGSSEIPSLKRTDEWKYLGINFTA 662
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein (ORF
2)
Length = 1308
Score = 73.2 bits (178), Expect = 5e-12
Identities = 85/346 (24%), Positives = 150/346 (42%), Gaps = 34/346 (9%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEK 863
K + RL+ L E+I P QS +PGR DN +++ ++H +RR LD EK
Sbjct: 540 KAISLRLKSVLAEVIHPDQSYTVPGRTIFDNVFLIRDLLHF--ARRTGLSLAFLSLDQEK 597
Query: 864 ACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPL 923
A V +L TL + F P V + +S+ + N P+A RG+R G PL
Sbjct: 598 AFDRVDHQYLIGTLQAYSFGPQFVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPL 657
Query: 924 SPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKV 983
S L+ L +E ++ + + + + +ADDV+L+A+ V + +
Sbjct: 658 SGQLYSLAIEPFLCLLRKRLTG-----LVLKEPDMRVVLSAYADDVILVAQDLV-DLERA 711
Query: 984 QNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIF--VVTSIPFTANLERYLG-FPIF 1040
Q + AS +N +KS S+ K D I + + + +YLG +
Sbjct: 712 QECQEVYAAASSARINWSKSSGLLEGSL----KVDFLPPAFRDISWESKIIKYLGVYLSA 767
Query: 1041 QGRQTQEHFDFVLDRVNRKFASWKG--RLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AV 1098
+ ++F + + V + WKG ++L+ G+ LV+N + Q+++ +
Sbjct: 768 EEYPVSQNFIELEECVLTRLGKWKGFAKVLSMRGR----ALVINQL--VASQIWYRLICL 821
Query: 1099 CDHLDSLAK------NFIWSRGERRGMNMVAWRTVTLPKCNGGLGI 1138
+ +AK +F+W G + V+ +LP GG G+
Sbjct: 822 SPTQEFIAKIQRRLLDFLWI-----GKHWVSAGVSSLPLKEGGQGV 862
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 63.9 bits (154), Expect = 3e-09
Identities = 33/70 (47%), Positives = 44/70 (62%)
Query: 899 LSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGS 958
L + NG + PSRGLR GDPLSPYLF+LC E L+ + A +Q + +R+S +
Sbjct: 10 LLFIINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSP 69
Query: 959 PISHLFFADD 968
I+HL FADD
Sbjct: 70 RINHLLFADD 79
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 63.5 bits (153), Expect = 4e-09
Identities = 68/241 (28%), Positives = 112/241 (46%), Gaps = 19/241 (7%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEK 863
+++ RL ++ + P Q + GT+ N+++L + +R K NVV LD+ K
Sbjct: 91 RILAKRLEAAVE--LHPAQKGYARIDGTLVNSLLLDTYISSRREQR-KTYNVV-SLDVRK 146
Query: 864 ACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSL-LWNGVKLPPIAPSRGLR*GDP 922
A +VS + + L G + + I +S S+ ++ + G + I RG++ GDP
Sbjct: 147 AFDTVSHSSICRALQRLGIDEGTSNYITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDP 206
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
LSP+LF ++ L S+Q G I + P+ L FADD+LLL V
Sbjct: 207 LSPFLFNAVLDELLCSLQ----STPGIGGTIGEEKIPV--LAFADDLLLLEDNDVLLPTT 260
Query: 983 VQNILADFCMASGLNVNIAKSRDFS-----SVSVPRAKKADIFVVTSIPFTANLE--RYL 1035
+ + A+F G+++N KS S V +PR K +P T ++ RYL
Sbjct: 261 LATV-ANFFRLRGMSLNAKKSVSISVAASGGVCIPRTKPFLRVDNVLLPVTDRMQTFRYL 319
Query: 1036 G 1036
G
Sbjct: 320 G 320
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 62.4 bits (150), Expect = 9e-09
Identities = 60/196 (30%), Positives = 91/196 (45%), Gaps = 24/196 (12%)
Query: 830 GTIDNAIILQKIVHHMNSRR--GKCQNVVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSV 887
GT+ N I+L+ H++ RR GK NVV LD+ KA +VS + + FG
Sbjct: 1 GTLANIIMLE---HYIKLRRLKGKTYNVV-SLDIRKAFDTVSHPAILRAMRAFGIDDGMQ 56
Query: 888 SLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQ 947
IM ++ + +++ G I G++ GDPLSP LF + ++ L + D +Q
Sbjct: 57 DFIMSTITDAYTNIVVGGRTTNKIYIRNGVKQGDPLSPVLFNIVLDELVTRLND--EQPG 114
Query: 948 GELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFC---MASGLNVNIAKSR 1004
+ + I+ L FADD+LLL + V N LA C G+ +N K
Sbjct: 115 ASMTPACK----IASLAFADDLLLLEDRDID----VPNSLATTCAYFRTRGMTLNPEKCA 166
Query: 1005 DFSSV-----SVPRAK 1015
S+ SVPR+K
Sbjct: 167 SISAATVSGRSVPRSK 182
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 61.2 bits (147), Expect = 2e-08
Identities = 65/228 (28%), Positives = 103/228 (44%), Gaps = 17/228 (7%)
Query: 802 FTKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFK-LD 860
F +++ NR+ + L+ Q +FI G +N +L ++ R K + + LD
Sbjct: 402 FHRILANRIGE--HGLLDTRQRAFIVADGVAENTSLLSAMI---KEARMKIKGLYIAILD 456
Query: 861 LEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*G 920
++KA SV + D L P + IM+ +S L K I P+RG+R G
Sbjct: 457 VKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEVVKTKGRWIRPARGVRQG 516
Query: 921 DPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQV 980
DPLSP LF ++ ++ + +N G L+ + G+ L FADD++LLA+ +
Sbjct: 517 DPLSPLLFNCVMD----AVLRRLPENTGFLMGAEKIGA----LVFADDLVLLAETREGLQ 568
Query: 981 RKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFT 1028
+ I A GL + K + VP K+ I V T PFT
Sbjct: 569 ASLSRIEAGL-QEQGLEMMPRKCHTLA--LVPSGKEKKIKVETHKPFT 613
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 52.8 bits (125), Expect = 8e-06
Identities = 38/132 (28%), Positives = 62/132 (46%), Gaps = 3/132 (2%)
Query: 1083 AIPVYPMQLFWFH*AVCDHLDSLAKNFIWSRGE-RRGMNMVAWRTVTLPK-CNGGLGIRK 1140
A+PVY M F +C L S F WS E +R ++ VAW+ + K +GGLG R
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 1141 ARLINTSMLGKLVWDLVNSHDKLWV*VLLDLYCTNSDFLH-APRRRGSCVWNAVMKTRDI 1199
N ++L K + +++ L +L Y +S + + R S W +++ R++
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGREL 121
Query: 1200 IQDGFTWRIGRG 1211
+ G IG G
Sbjct: 122 LSRGLLRTIGDG 133
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 46.2 bits (108), Expect = 7e-04
Identities = 53/198 (26%), Positives = 82/198 (40%), Gaps = 12/198 (6%)
Query: 822 QSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEKACGSVSWAFLRDTLCFFG 881
Q++++P G N +L I+ R + + LDL KA SV + L D + G
Sbjct: 261 QTAYLPIDGVCINVSMLTAIIAEAKRLRKELHIAI--LDLVKAFNSVYHSALIDAITEAG 318
Query: 882 FPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQD 941
PP V I ++ + + G K + G+ GDPLS LF L E+ ++
Sbjct: 319 CPPGVVDYIADMYNNVITEMQFEG-KCELASILAGVYQGDPLSGPLFTLAYEKALRAL-- 375
Query: 942 AVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIA 1001
N+G R ++ ++DD LLLA + + + GL +N
Sbjct: 376 ---NNEG---RFDIADVRVNASAYSDDGLLLAMTVIGLQHNLDK-FGETLAKIGLRINSR 428
Query: 1002 KSRDFSSVSVPRAKKADI 1019
KS+ S V R KK I
Sbjct: 429 KSKTVSLVPSGREKKMKI 446
>YSM6_CAEEL (Q10126) Hypothetical protein F52C9.6 in chromosome III
Length = 279
Score = 33.1 bits (74), Expect = 6.2
Identities = 29/113 (25%), Positives = 55/113 (48%), Gaps = 16/113 (14%)
Query: 956 DGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAK 1015
+G +++L FADD++L+A + + +Q ++ C GL +N K++ V R +
Sbjct: 4 NGRNLTNLRFADDIVLIANHPNTASKMLQELVQK-CSEVGLEINTGKTK------VLRNR 56
Query: 1016 KADIFVV-----TSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASW 1063
AD V +S +++ Y I+ GRQ H + + + R+ A+W
Sbjct: 57 LADPSKVYFGSPSSTTQLDDVDEY----IYLGRQINAHNNLMPEIHRRRRAAW 105
>YMC6_SCHPO (P05511) Hypothetical 91 kDa protein in cob intron
Length = 807
Score = 33.1 bits (74), Expect = 6.2
Identities = 27/123 (21%), Positives = 57/123 (45%), Gaps = 6/123 (4%)
Query: 960 ISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADI 1019
+ ++ +ADD ++ +Q +++ + FC + GL V+ K++ +S + +
Sbjct: 506 LMYVRYADDWIVAVNGSYTQTKEILAKITCFCSSIGLTVSPTKTKITNSYT-----DKIL 560
Query: 1020 FVVTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKL 1079
F+ T+I + N+ F I Q +DR+ +K G +LN G+ ++ L
Sbjct: 561 FLGTNISHSKNVTFSRHFGILQRNSGFILLSAPMDRIAKKLRE-TGLMLNHKGRSVIRWL 619
Query: 1080 VLN 1082
L+
Sbjct: 620 PLD 622
>DSRA_RAT (P55266) Double-stranded RNA-specific adenosine deaminase
(EC 3.5.4.-) (DRADA)
Length = 1175
Score = 33.1 bits (74), Expect = 6.2
Identities = 19/74 (25%), Positives = 37/74 (49%)
Query: 947 QGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDF 1006
+GE S DG P S L ++L +A+ K + N+ + N+ +AK+RD
Sbjct: 221 RGEPDLDSEDGDPASDLEGPSELLDMAEIKEKICDYLFNVSKSSALNLAKNIGLAKARDV 280
Query: 1007 SSVSVPRAKKADIF 1020
++V + ++ D++
Sbjct: 281 NAVLIDLERQGDVY 294
>RRPO_ORSVC (P89659) RNA-directed RNA polymerase (EC 2.7.7.48) (183
kDa protein) [Contains: Methyltransferase/RNA helicase
(MT/HEL) (126 kDa protein)]
Length = 1612
Score = 32.7 bits (73), Expect = 8.0
Identities = 26/70 (37%), Positives = 34/70 (48%), Gaps = 9/70 (12%)
Query: 1459 ADPGRAEVGGVVRDAVGCWLFGYSGFI---GMAEILKAELLAVLF----GLQSCWDRGFR 1511
ADP AE GV G WL G G+AE+ E + VL G C D +R
Sbjct: 728 ADPESAEKSGVYDVVKGKWLLKPKGKCHAWGVAELNNGEKVIVLLEWADGFPICGD--WR 785
Query: 1512 QLVVSSDSMV 1521
++ VSSDS++
Sbjct: 786 RVAVSSDSLI 795
>RIR1_BPT4 (P32282) Ribonucleoside-diphosphate reductase alpha chain
(EC 1.17.4.1) (Ribonucleotide reductase) (B1 protein)
Length = 754
Score = 32.7 bits (73), Expect = 8.0
Identities = 18/44 (40%), Positives = 26/44 (58%), Gaps = 1/44 (2%)
Query: 870 WAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAP 913
W+ LR+ L FG +++S +M C SSS +S NG + PP P
Sbjct: 593 WSALREDLKLFGIRNSTLSALMPCESSSQVSNSTNGYE-PPRGP 635
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.338 0.147 0.507
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 176,691,785
Number of Sequences: 164201
Number of extensions: 7230082
Number of successful extensions: 20520
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 20503
Number of HSP's gapped (non-prelim): 19
length of query: 1569
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1445
effective length of database: 39,613,130
effective search space: 57240972850
effective search space used: 57240972850
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0022.23


