

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.17
(341 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
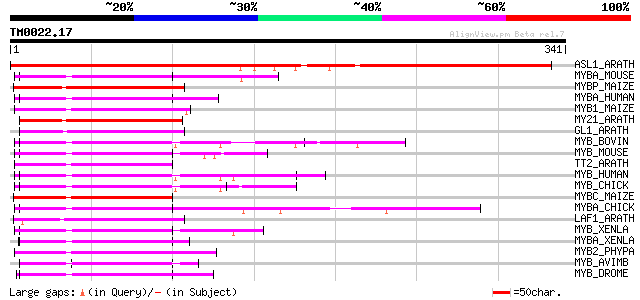
Score E
Sequences producing significant alignments: (bits) Value
ASL1_ARATH (O80931) ASYMMETRIC LEAVES1 protein (Myb-related prot... 400 e-111
MYBA_MOUSE (P51960) Myb-related protein A (A-Myb) 91 4e-18
MYBP_MAIZE (P27898) Myb-related protein P 91 5e-18
MYBA_HUMAN (P10243) Myb-related protein A (A-Myb) 91 5e-18
MYB1_MAIZE (P20024) Myb-related protein Zm1 89 1e-17
MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related prot... 89 1e-17
GL1_ARATH (P27900) Trichome differentiation protein GL1 (GLABROU... 88 3e-17
MYB_BOVIN (P46200) Myb proto-oncogene protein (C-myb) 87 8e-17
MYB_MOUSE (P06876) Myb proto-oncogene protein (C-myb) 86 2e-16
TT2_ARATH (Q9FJA2) TRANSPARENT TESTA 2 protein (Myb-related prot... 85 2e-16
MYB_HUMAN (P10242) Myb proto-oncogene protein (C-myb) 85 2e-16
MYB_CHICK (P01103) Myb proto-oncogene protein (C-myb) 85 2e-16
MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein 85 2e-16
MYBA_CHICK (P52550) Myb-related protein A (A-Myb) 84 5e-16
LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-re... 84 5e-16
MYB_XENLA (Q08759) Myb protein 84 7e-16
MYBA_XENLA (Q05935) Myb-related protein A (A-Myb) (XAMYB) (MYB-r... 83 1e-15
MYB2_PHYPA (P80073) Myb-related protein Pp2 83 1e-15
MYB_AVIMB (P01104) Transforming protein Myb 82 1e-15
MYB_DROME (P04197) Myb protein 82 3e-15
>ASL1_ARATH (O80931) ASYMMETRIC LEAVES1 protein (Myb-related protein
91) (AtMYB91) (PHANTASTICA protein)
Length = 367
Score = 400 bits (1028), Expect = e-111
Identities = 220/364 (60%), Positives = 264/364 (72%), Gaps = 37/364 (10%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRWS EEDALL AYV+Q+GPREW+LVS+RMN PLNRD KSCLERWKNYLKPGIKKG
Sbjct: 1 MKERQRWSGEEDALLRAYVRQFGPREWHLVSERMNKPLNRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQ +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQRE+ E N V P
Sbjct: 61 SLTEEEQRLVIRLQEKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREEKESNKRVEP 120
Query: 121 ISDTKYEHMLEGFAEKLVKE--HTLPSFAMA-----ASSNEAFLHTNS-----SAMLPSW 168
I ++KY+ +LE FAEKLVKE + +P+ A A A+SN FLH+ + ++P W
Sbjct: 121 IDESKYDRILESFAEKLVKERSNVVPAAAAAATVVMANSNGGFLHSEQQVQPPNPVIPPW 180
Query: 169 LSNYDS----TSTPPSSISVTLSLSPSTVAT--------------PRGLENN-APFVLRN 209
L+ ++ + PP SVTL+LSPSTVA P EN VL +
Sbjct: 181 LATSNNGNNVVARPP---SVTLTLSPSTVAAAAPQPPIPWLQQQQPERAENGPGGLVLGS 237
Query: 210 VTAHNGSVPSFSDHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRR 269
+ S S+ + +SELV +ELEEGHRA A HKKEA WRLRRLELQLESEK CR+
Sbjct: 238 MMP---SCSGSSESVFLSELVECCRELEEGHRAWADHKKEAAWRLRRLELQLESEKTCRQ 294
Query: 270 RETVEEFEANIKALQEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTR 329
RE +EE EA +KAL+EEQ A+ +IE REQL GLRRDAE+K+QKLA++WTS+H+RLT+
Sbjct: 295 REKMEEIEAKMKALREEQKNAMEKIEGEYREQLVGLRRDAEAKDQKLADQWTSRHIRLTK 354
Query: 330 LLEQ 333
LEQ
Sbjct: 355 FLEQ 358
>MYBA_MOUSE (P51960) Myb-related protein A (A-Myb)
Length = 751
Score = 90.9 bits (224), Expect = 4e-18
Identities = 50/166 (30%), Positives = 82/166 (49%), Gaps = 10/166 (6%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + VQ+YGP+ W+L+++ + R K C ERW N+L P +KK S T+EE
Sbjct: 90 WTKEEDQRVIELVQKYGPKRWSLIAKHLK---GRIGKQCRERWHNHLNPEVKKSSWTEEE 146
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW-EVYKEKQQREKIEINGIVSPISDTK 125
R++ GN+W +IA +PGRT + W + K ++E +GI S S +K
Sbjct: 147 DRIIYEAHKRLGNRWAEIAKLLPGRTDNSIKNHWNSTMRRKVEQEGYLQDGIKSERSSSK 206
Query: 126 YEHMLEGFAEKLVKEH------TLPSFAMAASSNEAFLHTNSSAML 165
+H + L ++ +P + + H +SA +
Sbjct: 207 LQHKPCATMDHLQTQNQFYIPVQIPGYQYVSPDGNCVEHVQTSAFI 252
Score = 67.8 bits (164), Expect = 4e-11
Identities = 36/98 (36%), Positives = 51/98 (51%), Gaps = 4/98 (4%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R +W+ +ED L V+Q+G +W L++ + NR C RW+ L P + KG T
Sbjct: 35 RVKWTRDEDDKLKKLVEQHGTDDWTLIASHLQ---NRSDFQCQHRWQKVLNPELIKGPWT 91
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW 100
KEE + VI L YG K W IA + GR K+ + W
Sbjct: 92 KEEDQRVIELVQKYGPKRWSLIAKHLKGRIGKQCRERW 129
>MYBP_MAIZE (P27898) Myb-related protein P
Length = 399
Score = 90.5 bits (223), Expect = 5e-18
Identities = 41/105 (39%), Positives = 65/105 (61%), Gaps = 2/105 (1%)
Query: 3 ERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSL 62
+R RW++EED LL Y+ ++G W + + N L R KSC RW NYL+ +K+G++
Sbjct: 13 KRGRWTAEEDQLLANYIAEHGEGSWRSLPK--NAGLLRCGKSCRLRWINYLRADVKRGNI 70
Query: 63 TKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
+KEE+ ++I L A GN+W IA+ +PGRT + +W + +Q
Sbjct: 71 SKEEEDIIIKLHATLGNRWSLIASHLPGRTDNEIKNYWNSHLSRQ 115
>MYBA_HUMAN (P10243) Myb-related protein A (A-Myb)
Length = 752
Score = 90.5 bits (223), Expect = 5e-18
Identities = 45/123 (36%), Positives = 69/123 (55%), Gaps = 4/123 (3%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + VQ+YGP+ W+L+++ + R K C ERW N+L P +KK S T+EE
Sbjct: 90 WTKEEDQRVIELVQKYGPKRWSLIAKHLK---GRIGKQCRERWHNHLNPEVKKSSWTEEE 146
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW-EVYKEKQQREKIEINGIVSPISDTK 125
R++ GN+W +IA +PGRT + W + K ++E +GI S S +K
Sbjct: 147 DRIIYEAHKRLGNRWAEIAKLLPGRTDNSIKNHWNSTMRRKVEQEGYLQDGIKSERSSSK 206
Query: 126 YEH 128
+H
Sbjct: 207 LQH 209
Score = 67.8 bits (164), Expect = 4e-11
Identities = 36/98 (36%), Positives = 51/98 (51%), Gaps = 4/98 (4%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R +W+ +ED L V+Q+G +W L++ + NR C RW+ L P + KG T
Sbjct: 35 RVKWTRDEDDKLKKLVEQHGTDDWTLIASHLQ---NRSDFQCQHRWQKVLNPELIKGPWT 91
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW 100
KEE + VI L YG K W IA + GR K+ + W
Sbjct: 92 KEEDQRVIELVQKYGPKRWSLIAKHLKGRIGKQCRERW 129
>MYB1_MAIZE (P20024) Myb-related protein Zm1
Length = 340
Score = 89.4 bits (220), Expect = 1e-17
Identities = 46/110 (41%), Positives = 65/110 (58%), Gaps = 4/110 (3%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R W+ +ED L AY+Q++G W + ++ L R KSC RW NYL+P +K+G+ T
Sbjct: 16 RGSWTPQEDMRLIAYIQKHGHTNWRALPKQAG--LLRCGKSCRLRWINYLRPDLKRGNFT 73
Query: 64 KEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ--QREK 111
EE+ +I L GNKW KIAA +PGRT + W + +K+ QREK
Sbjct: 74 DEEEEAIIRLHGLLGNKWSKIAACLPGRTDNEIKNVWNTHLKKKVAQREK 123
>MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related protein
21) (AtMYB21) (Myb homolog 3) (AtMyb3)
Length = 226
Score = 89.4 bits (220), Expect = 1e-17
Identities = 39/100 (39%), Positives = 62/100 (62%), Gaps = 2/100 (2%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L Y+ +G WN +++ + L R KSC RW NYL+P +++G++T EE
Sbjct: 25 WTMEEDLILINYIANHGDGVWNSLAK--SAGLKRTGKSCRLRWLNYLRPDVRRGNITPEE 82
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEK 106
Q +++ L A +GN+W KIA +PGRT + +W +K
Sbjct: 83 QLIIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQK 122
>GL1_ARATH (P27900) Trichome differentiation protein GL1 (GLABROUS1
protein)
Length = 228
Score = 88.2 bits (217), Expect = 3e-17
Identities = 41/101 (40%), Positives = 60/101 (58%), Gaps = 2/101 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L YV +G +WN + ++ T L R KSC RW NYL P + KG+ T++E
Sbjct: 19 WTVEEDNILMDYVLNHGTGQWNRIVRK--TGLKRCGKSCRLRWMNYLSPNVNKGNFTEQE 76
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
+ L+I L GN+W IA VPGRT ++ +W + K+
Sbjct: 77 EDLIIRLHKLLGNRWSLIAKRVPGRTDNQVKNYWNTHLSKK 117
>MYB_BOVIN (P46200) Myb proto-oncogene protein (C-myb)
Length = 640
Score = 86.7 bits (213), Expect = 8e-17
Identities = 71/243 (29%), Positives = 103/243 (42%), Gaps = 28/243 (11%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + VQ+YGP+ W+++++ + R K C ERW N+L P +KK S T+EE
Sbjct: 95 WTKEEDQRVIELVQKYGPKRWSVIAKHLK---GRIGKQCRERWHNHLNPEVKKTSWTEEE 151
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
R++ GN+W +IA +PGRT + W R K+E G + S
Sbjct: 152 DRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHW----NSTMRRKVEQEGYLQESSKASQ 207
Query: 127 EHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPSSISVTL 186
+ F + S F H SA LP ++ P IS
Sbjct: 208 PAVTTSFQKN--------------SHLMGFTHAPPSAQLPPAGQPSVNSDYPYYHISEAQ 253
Query: 187 SLSPSTVATPRGLENNAPFVLRNVTA-----HNGSVPSFSDHILMSELVGFSKELE-EGH 240
++S S V P L N V + A +N P I EL+ S E E +G
Sbjct: 254 NVS-SHVPYPVALHVNIVNVPQPAAAAIQRHYNDEDPEKEKRIKELELLLMSTENELKGQ 312
Query: 241 RAL 243
+AL
Sbjct: 313 QAL 315
Score = 67.8 bits (164), Expect = 4e-11
Identities = 55/195 (28%), Positives = 82/195 (41%), Gaps = 21/195 (10%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
+ RW+ EED L V+Q G +W +++ + NR C RW+ L P + KG T
Sbjct: 40 KTRWTREEDEKLKKLVEQNGTDDWKVIANYLP---NRTDVQCQHRWQKVLNPELIKGPWT 96
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW------EVYKEKQQREKIEING 116
KEE + VI L YG K W IA + GR K+ + W EV K E+ I
Sbjct: 97 KEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEEEDRIIY 156
Query: 117 IVSPISDTKYEH---MLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYD 173
++ +L G + +K H S E +L +S A P+ +++
Sbjct: 157 QAHKRLGNRWAEIAKLLPGRTDNAIKNH-WNSTMRRKVEQEGYLQESSKASQPAVTTSFQ 215
Query: 174 S-------TSTPPSS 181
T PPS+
Sbjct: 216 KNSHLMGFTHAPPSA 230
>MYB_MOUSE (P06876) Myb proto-oncogene protein (C-myb)
Length = 636
Score = 85.5 bits (210), Expect = 2e-16
Identities = 49/162 (30%), Positives = 79/162 (48%), Gaps = 18/162 (11%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + VQ+YGP+ W+++++ + R K C ERW N+L P +KK S T+EE
Sbjct: 95 WTKEEDQRVIELVQKYGPKRWSVIAKHLK---GRIGKQCRERWHNHLNPEVKKTSWTEEE 151
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIV-------- 118
R++ GN+W +IA +PGRT + W R K+E G +
Sbjct: 152 DRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHW----NSTMRRKVEQEGYLQEPSKASQ 207
Query: 119 SPISDT--KYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLH 158
+P++ + K H++ GF PS + +S + H
Sbjct: 208 TPVATSFQKNNHLM-GFGHASPPSQLSPSGQSSVNSEYPYYH 248
Score = 65.5 bits (158), Expect = 2e-10
Identities = 36/98 (36%), Positives = 50/98 (50%), Gaps = 4/98 (4%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
+ RW+ EED L V+Q G +W +++ + NR C RW+ L P + KG T
Sbjct: 40 KTRWTREEDEKLKKLVEQNGTDDWKVIANYLP---NRTDVQCQHRWQKVLNPELIKGPWT 96
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW 100
KEE + VI L YG K W IA + GR K+ + W
Sbjct: 97 KEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERW 134
>TT2_ARATH (Q9FJA2) TRANSPARENT TESTA 2 protein (Myb-related protein
123) (AtMYB123) (Myb-related transcription factor
LBM2-like)
Length = 258
Score = 85.1 bits (209), Expect = 2e-16
Identities = 38/97 (39%), Positives = 57/97 (58%), Gaps = 2/97 (2%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R W+ ED +L Y+ +G +W+ + + L R KSC RWKNYL+PGIK+G+++
Sbjct: 16 RGAWTDHEDKILRDYITTHGEGKWSTLPNQAG--LKRCGKSCRLRWKNYLRPGIKRGNIS 73
Query: 64 KEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW 100
+E+ L+I L GN+W IA +PGRT + W
Sbjct: 74 SDEEELIIRLHNLLGNRWSLIAGRLPGRTDNEIKNHW 110
>MYB_HUMAN (P10242) Myb proto-oncogene protein (C-myb)
Length = 640
Score = 85.1 bits (209), Expect = 2e-16
Identities = 53/190 (27%), Positives = 86/190 (44%), Gaps = 9/190 (4%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + VQ+YGP+ W+++++ + R K C ERW N+L P +KK S T+EE
Sbjct: 95 WTKEEDQRVIELVQKYGPKRWSVIAKHLK---GRIGKQCRERWHNHLNPEVKKTSWTEEE 151
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
R++ GN+W +IA +PGRT + W R K+E G + S
Sbjct: 152 DRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHW----NSTMRRKVEQEGYLQESSKASQ 207
Query: 127 EHMLEGFAEK--LVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPSSISV 184
+ F + L+ P A ++ + ++ + S S N S P ++ V
Sbjct: 208 PAVATSFQKNSHLMGFAQAPPTAQLPATGQPTVNNDYSYYHISEAQNVSSHVPYPVALHV 267
Query: 185 TLSLSPSTVA 194
+ P A
Sbjct: 268 NIVNVPQPAA 277
Score = 67.4 bits (163), Expect = 5e-11
Identities = 52/183 (28%), Positives = 78/183 (42%), Gaps = 14/183 (7%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
+ RW+ EED L V+Q G +W +++ + NR C RW+ L P + KG T
Sbjct: 40 KTRWTREEDEKLKKLVEQNGTDDWKVIANYLP---NRTDVQCQHRWQKVLNPELIKGPWT 96
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW------EVYKEKQQREKIEING 116
KEE + VI L YG K W IA + GR K+ + W EV K E+ I
Sbjct: 97 KEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEEEDRIIY 156
Query: 117 IVSPISDTKYEH---MLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYD 173
++ +L G + +K H S E +L +S A P+ +++
Sbjct: 157 QAHKRLGNRWAEIAKLLPGRTDNAIKNH-WNSTMRRKVEQEGYLQESSKASQPAVATSFQ 215
Query: 174 STS 176
S
Sbjct: 216 KNS 218
>MYB_CHICK (P01103) Myb proto-oncogene protein (C-myb)
Length = 641
Score = 85.1 bits (209), Expect = 2e-16
Identities = 43/127 (33%), Positives = 64/127 (49%), Gaps = 7/127 (5%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + VQ+YGP+ W+++++ + R K C ERW N+L P +KK S T+EE
Sbjct: 95 WTKEEDQRVIELVQKYGPKRWSVIAKHLK---GRIGKQCRERWHNHLNPEVKKTSWTEEE 151
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
R++ GN+W +IA +PGRT + W R K+E G + S
Sbjct: 152 DRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHW----NSTMRRKVEQEGYLQESSKAGL 207
Query: 127 EHMLEGF 133
GF
Sbjct: 208 PSATTGF 214
Score = 70.5 bits (171), Expect = 6e-12
Identities = 54/183 (29%), Positives = 79/183 (42%), Gaps = 14/183 (7%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
+ RW+ EED L V+Q G +W +++ + NR C RW+ L P + KG T
Sbjct: 40 KTRWTREEDEKLKKLVEQNGTEDWKVIASFLP---NRTDVQCQHRWQKVLNPELIKGPWT 96
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW------EVYKEKQQREKIEING 116
KEE + VI L YG K W IA + GR K+ + W EV K E+ I
Sbjct: 97 KEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEEEDRIIY 156
Query: 117 IVSPISDTKYEH---MLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYD 173
++ +L G + +K H S E +L +S A LPS + +
Sbjct: 157 QAHKRLGNRWAEIAKLLPGRTDNAIKNH-WNSTMRRKVEQEGYLQESSKAGLPSATTGFQ 215
Query: 174 STS 176
+S
Sbjct: 216 KSS 218
>MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein
Length = 273
Score = 85.1 bits (209), Expect = 2e-16
Identities = 40/98 (40%), Positives = 60/98 (60%), Gaps = 2/98 (2%)
Query: 3 ERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSL 62
+R W+S+ED L AYV+ +G +W V Q+ L R KSC RW NYL+P I++G++
Sbjct: 13 KRGAWTSKEDDALAAYVKAHGEGKWREVPQKAG--LRRCGKSCRLRWLNYLRPNIRRGNI 70
Query: 63 TKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW 100
+ +E+ L+I L GN+W IA +PGRT + +W
Sbjct: 71 SYDEEDLIIRLHRLLGNRWSLIAGRLPGRTDNEIKNYW 108
>MYBA_CHICK (P52550) Myb-related protein A (A-Myb)
Length = 757
Score = 84.0 bits (206), Expect = 5e-16
Identities = 65/302 (21%), Positives = 124/302 (40%), Gaps = 34/302 (11%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + VQ+YGP+ W+L+++ + R K C ERW N+L P +KK S T+ E
Sbjct: 90 WTKEEDQRVIELVQKYGPKRWSLIAKHLK---GRIGKQCRERWHNHLNPEVKKSSWTEAE 146
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW-EVYKEKQQREKIEINGIVSPISDTK 125
R++ GN+W +IA +PGRT + W + K ++E +G S T
Sbjct: 147 DRVIYEAHKRLGNRWAEIAKLLPGRTDNSIKNHWNSTMRRKVEQEGYLQDGTKSSSERTG 206
Query: 126 YEHMLEGFAEKLVKEHT-----------LPSFAMAASSNEAFLHTNSSAML--PSWLSNY 172
+ + + HT +P + A+ + H ++SA L S++ +
Sbjct: 207 SSTLAQKPCVTMEHLHTQNQFYIPVQTHIPVYQYASPEDSCIEHASASANLVQQSFIDDD 266
Query: 173 DSTSTPPSSISVTLSLSPSTVATPRGLENNAPFVLRNVTAHNGSVPSFSDHILMSELV-- 230
+ + L + + + R +++ GS+P +S +M + V
Sbjct: 267 PDKEKKIKELELLLMSTENEIRRKR------------LSSQAGSLPGWSGSFVMEDCVPN 314
Query: 231 ---GFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQ 287
++ E + + L +E+ +T+ EF ++ ++ +
Sbjct: 315 TLNSLGEQTSEFYSMDETQGTSVQQNSPTKYLAVEANAVLSSLQTIPEFAETLELIESDP 374
Query: 288 TA 289
A
Sbjct: 375 LA 376
Score = 64.3 bits (155), Expect = 4e-10
Identities = 35/98 (35%), Positives = 49/98 (49%), Gaps = 4/98 (4%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R +W+ +ED L V+Q G +W ++ + NR C RW+ L P + KG T
Sbjct: 35 RVKWTRDEDEKLKKLVEQNGTDDWAFIASHLQ---NRSDFQCQHRWQKVLNPELIKGPWT 91
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW 100
KEE + VI L YG K W IA + GR K+ + W
Sbjct: 92 KEEDQRVIELVQKYGPKRWSLIAKHLKGRIGKQCRERW 129
>LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-red
light protein 1) (Myb-related protein 18) (AtMYB18)
Length = 283
Score = 84.0 bits (206), Expect = 5e-16
Identities = 40/108 (37%), Positives = 62/108 (57%), Gaps = 5/108 (4%)
Query: 3 ERQR---WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKK 59
ER R WS EED L +++ YG W V + L R+ KSC RW NYL+PG+K+
Sbjct: 8 ERHRKGLWSPEEDEKLRSFILSYGHSCWTTVP--IKAGLQRNGKSCRLRWINYLRPGLKR 65
Query: 60 GSLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
++ EE+ ++ ++ GNKW +IA +PGRT + +W + +K+
Sbjct: 66 DMISAEEEETILTFHSSLGNKWSQIAKFLPGRTDNEIKNYWHSHLKKK 113
>MYB_XENLA (Q08759) Myb protein
Length = 624
Score = 83.6 bits (205), Expect = 7e-16
Identities = 47/153 (30%), Positives = 73/153 (46%), Gaps = 10/153 (6%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + V +YGP+ W+++++ + R K C ERW N+L P +KK S T+EE
Sbjct: 92 WTKEEDQRVIELVHKYGPKRWSVIAKHLK---GRIGKQCRERWHNHLNPEVKKSSWTEEE 148
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
R + GN+W +IA +PGRT + W R K E G + S T
Sbjct: 149 DRTIYEAHKRLGNRWAEIAKLLPGRTDNAIKNHW----NSTMRRKEEQEGYLQNSSKTNQ 204
Query: 127 EHMLEGFAEK---LVKEHTLPSFAMAASSNEAF 156
++ F + + HT S + +S +F
Sbjct: 205 HTIVTNFPKSNHLMTFTHTRASAEHSQASTSSF 237
Score = 66.6 bits (161), Expect = 8e-11
Identities = 37/98 (37%), Positives = 50/98 (50%), Gaps = 4/98 (4%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
+ RW+ EED L V+Q G EW +++ + NR C RW+ L P + KG T
Sbjct: 37 KTRWTREEDEKLKKLVEQNGTEEWKVIASFLP---NRTDVQCQHRWQKVLNPELIKGPWT 93
Query: 64 KEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLGKWW 100
KEE + VI L YG K W IA + GR K+ + W
Sbjct: 94 KEEDQRVIELVHKYGPKRWSVIAKHLKGRIGKQCRERW 131
>MYBA_XENLA (Q05935) Myb-related protein A (A-Myb) (XAMYB)
(MYB-related protein 2) (XMYB2)
Length = 728
Score = 82.8 bits (203), Expect = 1e-15
Identities = 38/105 (36%), Positives = 60/105 (56%), Gaps = 4/105 (3%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + V +YGP++W+++++ + R K C ERW N+L P +KK S T+EE
Sbjct: 89 WTKEEDQRVIELVHKYGPKKWSIIAKHLK---GRIGKQCRERWHNHLNPDVKKSSWTEEE 145
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW-EVYKEKQQRE 110
R++ GN+W +IA +PGRT + W K K ++E
Sbjct: 146 DRIIYSAHKRMGNRWAEIAKLLPGRTDNSIKNHWNSTMKRKVEQE 190
Score = 61.6 bits (148), Expect = 3e-09
Identities = 34/96 (35%), Positives = 50/96 (51%), Gaps = 5/96 (5%)
Query: 6 RWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKE 65
RW+ +ED + V+++G +W +V++ +NR C RW L P + KG TKE
Sbjct: 37 RWTKDEDDKVKKLVEKHG-EDWGVVARHF---INRSEVQCQHRWHKVLSPELVKGPWTKE 92
Query: 66 EQRLVILLQANYG-NKWKKIAAEVPGRTAKRLGKWW 100
E + VI L YG KW IA + GR K+ + W
Sbjct: 93 EDQRVIELVHKYGPKKWSIIAKHLKGRIGKQCRERW 128
>MYB2_PHYPA (P80073) Myb-related protein Pp2
Length = 421
Score = 82.8 bits (203), Expect = 1e-15
Identities = 42/124 (33%), Positives = 68/124 (53%), Gaps = 2/124 (1%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R W+SEED L +++ G W + + L R KSC RW NYL+P +K+G +
Sbjct: 14 RGPWTSEEDQKLVSHITNNGLSCWRAIPKLAG--LLRCGKSCRLRWTNYLRPDLKRGIFS 71
Query: 64 KEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISD 123
+ E+ L++ L A GN+W +IAA++PGRT + +W +K+ R + P+ D
Sbjct: 72 EAEENLILDLHATLGNRWSRIAAQLPGRTDNEIKNYWNTRLKKRLRSQGLDPNTHLPLED 131
Query: 124 TKYE 127
+K +
Sbjct: 132 SKLD 135
>MYB_AVIMB (P01104) Transforming protein Myb
Length = 382
Score = 82.4 bits (202), Expect = 1e-15
Identities = 40/110 (36%), Positives = 60/110 (54%), Gaps = 7/110 (6%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED + +VQ+YGP+ W+ +++ + R K C ERW N+L P +KK S T+EE
Sbjct: 24 WTKEEDQRVIEHVQKYGPKRWSDIAKHLK---GRIGKQCRERWHNHLNPEVKKTSWTEEE 80
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEING 116
R++ GN+W +IA +PGRT + W R K+E G
Sbjct: 81 DRIIYQAHKRLGNRWAEIAKLLPGRTDNAVKNHW----NSTMRRKVEQEG 126
Score = 47.0 bits (110), Expect = 7e-05
Identities = 25/63 (39%), Positives = 31/63 (48%), Gaps = 1/63 (1%)
Query: 39 NRDTKSCLERWKNYLKPGIKKGSLTKEEQRLVILLQANYGNK-WKKIAAEVPGRTAKRLG 97
NR C RW+ L P + KG TKEE + VI YG K W IA + GR K+
Sbjct: 1 NRTDVQCQHRWQKVLNPELNKGPWTKEEDQRVIEHVQKYGPKRWSDIAKHLKGRIGKQCR 60
Query: 98 KWW 100
+ W
Sbjct: 61 ERW 63
>MYB_DROME (P04197) Myb protein
Length = 657
Score = 81.6 bits (200), Expect = 3e-15
Identities = 38/120 (31%), Positives = 65/120 (53%), Gaps = 4/120 (3%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ +ED ++ V+ +GP++W L+++ +N R K C ERW N+L P IKK + T++E
Sbjct: 139 WTRDEDDMVIKLVRNFGPKKWTLIARYLN---GRIGKQCRERWHNHLNPNIKKTAWTEKE 195
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW-EVYKEKQQREKIEINGIVSPISDTK 125
++ GN+W KIA +PGRT + W + K E+ +N S + ++
Sbjct: 196 DEIIYQAHLELGNQWAKIAKRLPGRTDNAIKNHWNSTMRRKYDVERRSVNASGSDLKSSR 255
Score = 57.4 bits (137), Expect = 5e-08
Identities = 32/97 (32%), Positives = 48/97 (48%), Gaps = 5/97 (5%)
Query: 5 QRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTK 64
+RWS ED LL V+ +G W ++ L + + +RW L P + KG T+
Sbjct: 86 KRWSKSEDVLLKQLVETHG-ENWEIIGPHFKDRLEQQVQ---QRWAKVLNPELIKGPWTR 141
Query: 65 EEQRLVILLQANYG-NKWKKIAAEVPGRTAKRLGKWW 100
+E +VI L N+G KW IA + GR K+ + W
Sbjct: 142 DEDDMVIKLVRNFGPKKWTLIARYLNGRIGKQCRERW 178
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.127 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,680,834
Number of Sequences: 164201
Number of extensions: 1612124
Number of successful extensions: 5729
Number of sequences better than 10.0: 228
Number of HSP's better than 10.0 without gapping: 57
Number of HSP's successfully gapped in prelim test: 171
Number of HSP's that attempted gapping in prelim test: 5310
Number of HSP's gapped (non-prelim): 519
length of query: 341
length of database: 59,974,054
effective HSP length: 111
effective length of query: 230
effective length of database: 41,747,743
effective search space: 9601980890
effective search space used: 9601980890
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0022.17


