

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.15
(313 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
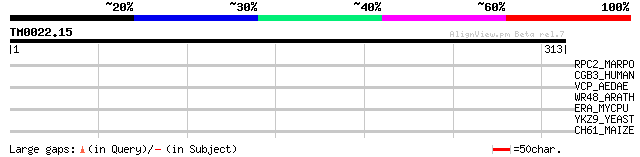
Score E
Sequences producing significant alignments: (bits) Value
RPC2_MARPO (P06274) DNA-directed RNA polymerase beta'' chain (EC... 35 0.18
CGB3_HUMAN (Q8WWL7) G2/mitotic-specific cyclin B3 33 1.2
VCP_AEDAE (P42660) Vitellogenic carboxypeptidase precursor (EC 3... 32 1.6
WR48_ARATH (Q9FGZ4) Probable WRKY transcription factor 48 (WRKY ... 32 2.0
ERA_MYCPU (Q98QI1) GTP-binding protein era homolog 31 4.5
YKZ9_YEAST (P36115) Hypothetical 68.9 kDa protein in YPT52-DBP7 ... 30 5.9
CH61_MAIZE (P29185) Chaperonin CPN60-1, mitochondrial precursor ... 30 7.8
>RPC2_MARPO (P06274) DNA-directed RNA polymerase beta'' chain (EC
2.7.7.6) (PEP) (Plastid-encoded RNA polymerase beta''
subunit) (RNA polymerase beta'' subunit)
Length = 1386
Score = 35.4 bits (80), Expect = 0.18
Identities = 25/76 (32%), Positives = 39/76 (50%), Gaps = 6/76 (7%)
Query: 14 QSETQHSILTKVTRSSIYSEQLSEQRSSKKRR------KSSSPLLLLKLTGDGKMRENEN 67
+S+ ++IL K+ S Y+ L E KK+ K+ + L LK+ +G ++ NE
Sbjct: 535 KSQLTNNILNKINNSKNYNFILQEYNIKKKKNFYFLKNKNLTCPLFLKIKKNGVLKNNEI 594
Query: 68 FAILIFPAPSSVNSKI 83
FAIL P+ NS I
Sbjct: 595 FAILDDPSYKVKNSGI 610
>CGB3_HUMAN (Q8WWL7) G2/mitotic-specific cyclin B3
Length = 1395
Score = 32.7 bits (73), Expect = 1.2
Identities = 29/116 (25%), Positives = 55/116 (47%), Gaps = 13/116 (11%)
Query: 17 TQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFPAP 76
T+ ++LTK T S+ +++++ + S ++ PL+L K+T E E+F + P
Sbjct: 576 TEETVLTK-TSLSLQEKKITQGKMSHLKK----PLVLQKITS-----EEESFYKKLLPFK 625
Query: 77 SSVNSKIGFGDRIP---KLKRGFLELSGCRKRKLPSEERLETSREMGVRVWNLHLK 129
++ F + P K K L+ K L +E+ T E +++LH+K
Sbjct: 626 MKSTTEEKFLSQEPSALKEKHTTLQEVSLSKESLAIQEKATTEEEFSQELFSLHVK 681
>VCP_AEDAE (P42660) Vitellogenic carboxypeptidase precursor (EC
3.4.16.-)
Length = 471
Score = 32.3 bits (72), Expect = 1.6
Identities = 20/55 (36%), Positives = 26/55 (46%)
Query: 31 YSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFPAPSSVNSKIGF 85
Y + + S + +S PL L L DGK+ E N A + P SSV S GF
Sbjct: 26 YKKLMRGSASPPRPGESGEPLFLTPLLQDGKIEEARNKARVNHPMLSSVESYSGF 80
>WR48_ARATH (Q9FGZ4) Probable WRKY transcription factor 48 (WRKY
DNA-binding protein 48)
Length = 399
Score = 32.0 bits (71), Expect = 2.0
Identities = 27/97 (27%), Positives = 36/97 (36%)
Query: 3 LRAKTTEFHHNQSETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKM 62
+ K E HH+Q + Q K T + I EQ EQ+ + SSS + L + D
Sbjct: 1 MEKKKEEDHHHQQQQQQQKEIKNTETKIEQEQEQEQKQEISQASSSSNMANLVTSSDHHP 60
Query: 63 RENENFAILIFPAPSSVNSKIGFGDRIPKLKRGFLEL 99
E IF S F D FL+L
Sbjct: 61 LELAGNLSSIFDTSSLPFPYSYFEDHSSNNPNSFLDL 97
>ERA_MYCPU (Q98QI1) GTP-binding protein era homolog
Length = 293
Score = 30.8 bits (68), Expect = 4.5
Identities = 15/41 (36%), Positives = 27/41 (65%), Gaps = 1/41 (2%)
Query: 101 GCRKRKLPSEERLETSREMGVRVWNLHLKLRIEKVFINQEK 141
G + +K+ R + R++GV V L LK++++K ++NQEK
Sbjct: 247 GSKIKKIGISARKKIQRKLGVNV-KLFLKIKVKKNWVNQEK 286
>YKZ9_YEAST (P36115) Hypothetical 68.9 kDa protein in YPT52-DBP7
intergenic region
Length = 615
Score = 30.4 bits (67), Expect = 5.9
Identities = 19/71 (26%), Positives = 33/71 (45%), Gaps = 5/71 (7%)
Query: 15 SETQHSILTKVTRSSIYSEQLSEQRSSKKRRKS-----SSPLLLLKLTGDGKMRENENFA 69
+ +Q ++LT + S E ++K K+ SP L L G G++ ENEN +
Sbjct: 178 NRSQENLLTSFSSGRRLSSSSMEPATNKDSNKALPKRRPSPPLQSSLVGSGQLHENENLS 237
Query: 70 ILIFPAPSSVN 80
+ + S+N
Sbjct: 238 SISIDSRHSLN 248
>CH61_MAIZE (P29185) Chaperonin CPN60-1, mitochondrial precursor
(HSP60-1)
Length = 577
Score = 30.0 bits (66), Expect = 7.8
Identities = 20/71 (28%), Positives = 40/71 (56%), Gaps = 8/71 (11%)
Query: 91 KLKRGFLE---LSGCRKRKLPSEERLETSREMGVRVWNLHLKLRIEKVFINQEKPLTGKL 147
KL RG++ ++ + +K E+ L + +V N+H +++ ++ + ++KPL L
Sbjct: 227 KLDRGYISPYFITNSKTQKCELEDPLILIHDK--KVTNMHAVVKVLEMALKKQKPL---L 281
Query: 148 VRAEDVQDEPL 158
+ AEDV+ E L
Sbjct: 282 IVAEDVESEAL 292
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,686,995
Number of Sequences: 164201
Number of extensions: 1483958
Number of successful extensions: 3329
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 3326
Number of HSP's gapped (non-prelim): 8
length of query: 313
length of database: 59,974,054
effective HSP length: 110
effective length of query: 203
effective length of database: 41,911,944
effective search space: 8508124632
effective search space used: 8508124632
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0022.15


