

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.11
(334 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
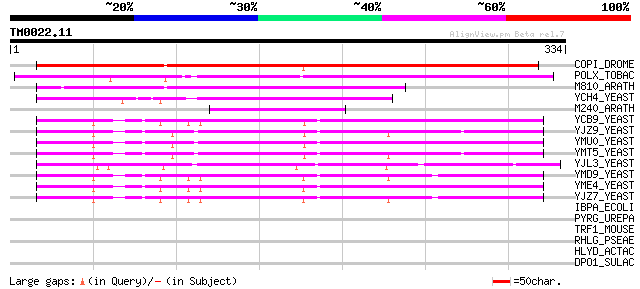
Score E
Sequences producing significant alignments: (bits) Value
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 220 4e-57
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 187 4e-47
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 158 2e-38
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 91 4e-18
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 64 5e-10
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 57 8e-08
YJZ9_YEAST (P47100) Transposon Ty1 protein B 56 1e-07
YMU0_YEAST (Q04670) Transposon Ty1 protein B 55 2e-07
YMT5_YEAST (Q04214) Transposon Ty1 protein B 55 2e-07
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 55 2e-07
YMD9_YEAST (Q03434) Transposon Ty1 protein B 52 2e-06
YME4_YEAST (Q04711) Transposon Ty1 protein B 51 4e-06
YJZ7_YEAST (P47098) Transposon Ty1 protein B 51 4e-06
IBPA_ECOLI (P29209) 16 kDa heat shock protein A 32 1.7
PYRG_UREPA (Q9PQK7) CTP synthase (EC 6.3.4.2) (UTP--ammonia liga... 32 2.2
TRF1_MOUSE (P70371) Telomeric repeat binding factor 1 (TTAGGG re... 31 5.0
RHLG_PSEAE (Q9RPT1) Rhamnolipids biosynthesis 3-oxoacyl-[acyl-ca... 30 8.5
HLYD_ACTAC (P18790) Leukotoxin secretion protein D 30 8.5
DPO1_SULAC (P95690) DNA polymerase I (EC 2.7.7.7) 30 8.5
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 220 bits (560), Expect = 4e-57
Identities = 116/307 (37%), Positives = 187/307 (60%), Gaps = 6/307 (1%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTKE 76
+YVDD++ + + F + + +F M+ + E+K+F+GI+++ Q + Y+ QS Y K+
Sbjct: 1086 LYVDDVVIATGDMTRMNNFKRYLMEKFRMTDLNEIKHFIGIRIEMQEDKIYLSQSAYVKK 1145
Query: 77 FLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLT-ASRPDILFSVH 135
L KFNM C TP+ P+ I + S + C R +IG L+Y+ +RPD+ +V+
Sbjct: 1146 ILSKFNMENCNAVSTPL-PSKINYELLNSDEDCNTPCRSLIGCLMYIMLCTRPDLTTAVN 1204
Query: 136 LCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*Y--KLSGYCDADYAGDRTERK 193
+ +++ S +KR+L+YLKGT ++ L++KK+ + K+ GY D+D+AG +RK
Sbjct: 1205 ILSRYSSKNNSELWQNLKRVLRYLKGTIDMKLIFKKNLAFENKIIGYVDSDWAGSEIDRK 1264
Query: 194 STYGNC-QFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LES 251
ST G + NL+ W +KRQ+++A S+ EAEY++ + LW+K L I LE+
Sbjct: 1265 STTGYLFKMFDFNLICWNTKRQNSVAASSTEAEYMALFEAVREALWLKFLLTSINIKLEN 1324
Query: 252 NISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSK 311
I IY DN IS++ N H RAKHI++KYHF R+ VQ V+ L+++ T++Q ADIF+K
Sbjct: 1325 PIKIYEDNQGCISIANNPSCHKRAKHIDIKYHFAREQVQNNVICLEYIPTENQLADIFTK 1384
Query: 312 PLAEDRF 318
PL RF
Sbjct: 1385 PLPAARF 1391
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 187 bits (474), Expect = 4e-47
Identities = 114/337 (33%), Positives = 181/337 (52%), Gaps = 18/337 (5%)
Query: 4 FYHFSYLQLVSEQIYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQV--DQ 61
F FS + +YVDD++ + L + + F+M +G + LG+++ ++
Sbjct: 994 FKRFSENNFIILLLYVDDMLIVGKDKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRER 1053
Query: 62 QPEATYIHQSKYTKEFLKKFNMTECTPAKTP----------MHPTCILEKEDVSAKVCQK 111
++ Q KY + L++FNM P TP M PT + EK ++ AKV
Sbjct: 1054 TSRKLWLSQEKYIERVLERFNMKNAKPVSTPLAGHLKLSKKMCPTTVEEKGNM-AKVP-- 1110
Query: 112 LYRGMIGSLLY-LTASRPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYK 170
Y +GSL+Y + +RPDI +V + ++F +P + H VK IL+YL+GTT L +
Sbjct: 1111 -YSSAVGSLMYAMVCTRPDIAHAVGVVSRFLENPGKEHWEAVKWILRYLRGTTGDCLCFG 1169
Query: 171 KSS*YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAA 230
S L GY DAD AGD RKS+ G +SW SK Q +ALST EAEYI+A
Sbjct: 1170 GSDPI-LKGYTDADMAGDIDNRKSSTGYLFTFSGGAISWQSKLQKCVALSTTEAEYIAAT 1228
Query: 231 ICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQ 290
+M+W+K L++ + + +YCD+ +AI LSKN + H+R KHI+V+YH+IR+ V
Sbjct: 1229 ETGKEMIWLKRFLQELGLHQKEYVVYCDSQSAIDLSKNSMYHARTKHIDVRYHWIREMVD 1288
Query: 291 KGVLLLKFVDTDHQWADIFSKPLAEDRFKFILKNLNM 327
L + + T+ AD+ +K + ++F+ + + M
Sbjct: 1289 DESLKVLKISTNENPADMLTKVVPRNKFELCKELVGM 1325
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 158 bits (400), Expect = 2e-38
Identities = 80/223 (35%), Positives = 130/223 (57%), Gaps = 3/223 (1%)
Query: 17 IYVDDIIF-GSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTK 75
+YVDDI+ GS+NT L + + F M +G + YFLGIQ+ P ++ Q+KY +
Sbjct: 5 LYVDDILLTGSSNTLL-NMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTKYAE 63
Query: 76 EFLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASRPDILFSVH 135
+ L M +C P TP+ P + + +R ++G+L YLT +RPDI ++V+
Sbjct: 64 QILNNAGMLDCKPMSTPL-PLKLNSSVSTAKYPDPSDFRSIVGALQYLTLTRPDISYAVN 122
Query: 136 LCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTERKST 195
+ Q +P +KR+L+Y+KGT GL K+S + +CD+D+AG + R+ST
Sbjct: 123 IVCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNSKLNVQAFCDSDWAGCTSTRRST 182
Query: 196 YGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLW 238
G C FLG N++SW++KRQ T++ S+ E EY + A+ + ++ W
Sbjct: 183 TGFCTFLGCNIISWSAKRQPTVSRSSTETEYRALALTAAELTW 225
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 90.9 bits (224), Expect = 4e-18
Identities = 63/225 (28%), Positives = 111/225 (49%), Gaps = 19/225 (8%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEAT-------YIH 69
+YVDD++ + + + + + + M +G++ FLG+ + Q YI
Sbjct: 86 VYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKVDKFLGLNIHQSTNGDITLSLQDYIA 145
Query: 70 QSKYTKEFLKKFNMTECTPA--KTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLT-AS 126
++ E + F +T+ TP P+ T +D++ Y+ ++G LL+
Sbjct: 146 KAASESE-INTFKLTQ-TPLCNSKPLFETTSPHLKDITP------YQSIVGQLLFCANTG 197
Query: 127 RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYA 186
RPDI + V L ++F +PR HL + +R+L+YL T ++ L Y+ S L+ YCDA +
Sbjct: 198 RPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTTRSMCLKYRSGSQVALTVYCDASHG 257
Query: 187 GDRTERKSTYGNCQFLGSNLVSWASKR-QSTIALSTAEAEYISAA 230
ST G L V+W+SK+ + I + + EAEYI+A+
Sbjct: 258 AIHDLPHSTGGYVTLLAGAPVTWSSKKLKGVIPVPSTEAEYITAS 302
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 63.9 bits (154), Expect = 5e-10
Identities = 30/82 (36%), Positives = 47/82 (56%)
Query: 121 LYLTASRPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGY 180
+YLT +RPD+ F+V+ +QF S R + V ++L Y+KGT GL Y +S +L +
Sbjct: 1 MYLTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDLQLKAF 60
Query: 181 CDADYAGDRTERKSTYGNCQFL 202
D+D+A R+S G C +
Sbjct: 61 ADSDWASCPDTRRSVTGFCSLV 82
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 56.6 bits (135), Expect = 8e-08
Identities = 72/335 (21%), Positives = 147/335 (43%), Gaps = 39/335 (11%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMG------ELKY-FLGIQVDQQPEATYIH 69
++VDD+I S + K+ ++ +++ ++ E++Y LG+++ Q
Sbjct: 1439 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQ------- 1491
Query: 70 QSKYTKEFLKKFNMTECTPA------------KTPMHPTCILEKEDVSA---KVCQKLY- 113
+SKY K ++K ++TE P + P P ++++++ + +K++
Sbjct: 1492 RSKYMKLGMEK-SLTEKLPKLNVPLNPKGKKLRAPGQPGHYIDQDELEIDEDEYKEKVHE 1550
Query: 114 -RGMIGSLLYLTAS-RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKK 171
+ +IG Y+ R D+L+ ++ AQ P L +++++ T + L++ K
Sbjct: 1551 MQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHK 1610
Query: 172 SS*YK----LSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYI 227
+ K L DA Y G++ KS GN L ++ S + S ST EAE
Sbjct: 1611 NKPTKPDNKLVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIH 1669
Query: 228 SAAICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHS-RAKHIEVKYHFIR 286
+ + + + H +++ + D+ + IS+ K+ R + K +R
Sbjct: 1670 AVSEAIPLLNNLSHLVQELNKKPIIKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRLR 1729
Query: 287 DYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFI 321
D V L + +++T AD+ +KPL FK +
Sbjct: 1730 DEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLL 1764
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 56.2 bits (134), Expect = 1e-07
Identities = 79/338 (23%), Positives = 144/338 (42%), Gaps = 45/338 (13%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMG------ELKY-FLGIQVDQQPEATYIH 69
++VDD++ S N K + ++ +++ ++ E++Y LG+++ Q
Sbjct: 1424 LFVDDMVLFSKNLNSNKRIIEKLKMQYDTKIINLGESDEEIQYDILGLEIKYQ------- 1476
Query: 70 QSKYTKEFLKKFNMTECTPA-KTPMHPT------------------CILEKEDVSAKVCQ 110
+ KY K ++ ++TE P P++P LE++D KV +
Sbjct: 1477 RGKYMKLGMEN-SLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHE 1535
Query: 111 KLYRGMIGSLLYLTAS-RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMY 169
+ +IG Y+ R D+L+ ++ AQ P + L +++++ T + L++
Sbjct: 1536 M--QKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIW 1593
Query: 170 KKSS*YK----LSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAE 225
KS K L DA Y G++ KS GN L ++ S + S ST EAE
Sbjct: 1594 HKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1652
Query: 226 Y--ISAAICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHSRAKHIEVKYH 283
IS ++ L Q D + + + +T +I +S N R + K
Sbjct: 1653 IHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNEEKF-RNRFFGTKAM 1711
Query: 284 FIRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFI 321
+RD V L + +++T AD+ +KPL FK +
Sbjct: 1712 RLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1749
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 55.5 bits (132), Expect = 2e-07
Identities = 75/337 (22%), Positives = 143/337 (42%), Gaps = 43/337 (12%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMG------ELKY-FLGIQVDQQPEATYIH 69
++VDD+I S + K+ ++ +++ ++ E++Y LG+++ Q
Sbjct: 997 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQ------- 1049
Query: 70 QSKYTKEFLKKFNMTECTPA-KTPMHPT------------------CILEKEDVSAKVCQ 110
+ KY K ++ ++TE P P++P LE++D KV +
Sbjct: 1050 RGKYMKLGMEN-SLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHE 1108
Query: 111 KLYRGMIGSLLYLTAS-RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMY 169
+ +IG Y+ R D+L+ ++ AQ P + L +++++ T + L++
Sbjct: 1109 M--QKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIW 1166
Query: 170 KKSS*YK----LSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAE 225
KS K L DA Y G++ KS GN L ++ S + S ST EAE
Sbjct: 1167 HKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1225
Query: 226 YISAAICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLS-KNLILHSRAKHIEVKYHF 284
+ + + + H +++ + D+ + IS+ N R + K
Sbjct: 1226 IHAISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMR 1285
Query: 285 IRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFI 321
+RD V L + +++T AD+ +KPL FK +
Sbjct: 1286 LRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 55.5 bits (132), Expect = 2e-07
Identities = 79/338 (23%), Positives = 143/338 (41%), Gaps = 45/338 (13%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMG------ELKY-FLGIQVDQQPEATYIH 69
++VDD++ S N K ++ +++ ++ E++Y LG+++ Q
Sbjct: 997 LFVDDMVLFSKNLNSNKRIIDKLKMQYDTKIINLGESDEEIQYDILGLEIKYQ------- 1049
Query: 70 QSKYTKEFLKKFNMTECTPA-KTPMHPT------------------CILEKEDVSAKVCQ 110
+ KY K ++ ++TE P P++P LE++D KV +
Sbjct: 1050 RGKYMKLGMEN-SLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHE 1108
Query: 111 KLYRGMIGSLLYLTAS-RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMY 169
+ +IG Y+ R D+L+ ++ AQ P + L +++++ T + L++
Sbjct: 1109 M--QKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIW 1166
Query: 170 KKSS*YK----LSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAE 225
KS K L DA Y G++ KS GN L ++ S + S ST EAE
Sbjct: 1167 HKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1225
Query: 226 Y--ISAAICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHSRAKHIEVKYH 283
IS ++ L Q D + + + +T +I +S N R + K
Sbjct: 1226 IHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNEEKF-RNRFFGTKAM 1284
Query: 284 FIRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFI 321
+RD V L + +++T AD+ +KPL FK +
Sbjct: 1285 RLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 55.5 bits (132), Expect = 2e-07
Identities = 76/342 (22%), Positives = 141/342 (41%), Gaps = 34/342 (9%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGEL------KYFLGIQ---------VDQ 61
+YVDD + ++N EF +++ FE+ + G L LG+ +D
Sbjct: 1460 VYVDDCVIAASNEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMDLVYNKRLGTIDL 1519
Query: 62 QPEATYIHQSKYTKEFLKKFNMTECTPAKT----PMHPTCILEKEDVSAKVCQKLYRGMI 117
++ K E LKK + T P + +E+ V + + ++
Sbjct: 1520 TLKSFINRMDKKYNEELKKIRKSSIPHMSTYKIDPKKDVLQMSEEEFRQGVLK--LQQLL 1577
Query: 118 GSLLYLT-ASRPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKK--SS* 174
G L Y+ R DI F+V A+ + P E + +I++YL ++G+ Y + +
Sbjct: 1578 GELNYVRHKCRYDIEFAVKKVARLVNYPHERVFYMIYKIIQYLVRYKDIGIHYDRDCNKD 1637
Query: 175 YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAE----YISAA 230
K+ DA G + +S G + G N+ + S + + +S+ EAE Y A
Sbjct: 1638 KKVIAITDAS-VGSEYDAQSRIGVILWYGMNIFNVYSNKSTNRCVSSTEAELHAIYEGYA 1696
Query: 231 ICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYV- 289
T + +K E ++I + D+ AI + K +K I++ +
Sbjct: 1697 DSETLKVTLKELGEGD---NNDIVMITDSKPAIQGLNRSYQQPKEKFTWIKTEIIKEKIK 1753
Query: 290 QKGVLLLKFVDTDHQWADIFSKPLAEDRFKFILKNLNMDLCS 331
+K + LLK + AD+ +KP++ FK ++ L + S
Sbjct: 1754 EKSIKLLKITGKGN-IADLLTKPVSASDFKRFIQVLKNKITS 1794
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 52.0 bits (123), Expect = 2e-06
Identities = 73/338 (21%), Positives = 149/338 (43%), Gaps = 45/338 (13%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMG------ELKY-FLGIQVDQQPEATYIH 69
++VDD+I S + K+ ++ +++ ++ E++Y LG+++ Q
Sbjct: 997 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQ------- 1049
Query: 70 QSKYTKEFLKKFNMTECTPA------------KTPMHPTCILEKEDVSA---KVCQKLY- 113
+ KY K ++ ++TE P P P ++++++ + +K++
Sbjct: 1050 RGKYMKLGMEN-SLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHE 1108
Query: 114 -RGMIGSLLYLTAS-RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKK 171
+ +IG Y+ R D+L+ ++ AQ P L +++++ T + L++ K
Sbjct: 1109 MQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHK 1168
Query: 172 SS*Y----KLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEY- 226
+ KL DA Y G++ KS GN L ++ S + S ST EAE
Sbjct: 1169 NKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIH 1227
Query: 227 -ISAAI-CSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHS-RAKHIEVKYH 283
IS ++ + ++ +L I++ ++ D+ + IS+ K+ R + K
Sbjct: 1228 AISESVPLLNNLSYLIQELNKKPIIKGLLT---DSRSTISIIKSTNEEKFRNRFFGTKAM 1284
Query: 284 FIRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFI 321
+RD V L + +++T AD+ +KPL FK +
Sbjct: 1285 RLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLL 1322
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 51.2 bits (121), Expect = 4e-06
Identities = 70/335 (20%), Positives = 143/335 (41%), Gaps = 39/335 (11%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMG------ELKY-FLGIQVDQQPEATYIH 69
++VDD+I S + K+ ++ +++ ++ E++Y LG+++ Q
Sbjct: 997 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQ------- 1049
Query: 70 QSKYTKEFLKKFNMTECTPA------------KTPMHPTCILEKEDVSA---KVCQKLY- 113
+ KY K ++ ++TE P P P ++++++ + +K++
Sbjct: 1050 RGKYMKLGMEN-SLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHE 1108
Query: 114 -RGMIGSLLYLTAS-RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKK 171
+ +IG Y+ R D+L+ ++ AQ P L +++++ T + L++ K
Sbjct: 1109 MQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHK 1168
Query: 172 SS*Y----KLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYI 227
+ KL DA Y G++ KS GN L ++ S + S ST EAE
Sbjct: 1169 NKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIH 1227
Query: 228 SAAICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLS-KNLILHSRAKHIEVKYHFIR 286
+ + + + H +++ + D+ + IS+ N R + K +R
Sbjct: 1228 AISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLR 1287
Query: 287 DYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFI 321
D V L + +++T AD+ +KPL FK +
Sbjct: 1288 DEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 51.2 bits (121), Expect = 4e-06
Identities = 73/338 (21%), Positives = 149/338 (43%), Gaps = 45/338 (13%)
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMG------ELKY-FLGIQVDQQPEATYIH 69
++VDD+I S + K+ ++ +++ ++ E++Y LG+++ Q
Sbjct: 1424 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQ------- 1476
Query: 70 QSKYTKEFLKKFNMTECTPA------------KTPMHPTCILEKEDVSA---KVCQKLY- 113
+ KY K ++ ++TE P P P ++++++ + +K++
Sbjct: 1477 RGKYMKLGMEN-SLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHE 1535
Query: 114 -RGMIGSLLYLTAS-RPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKK 171
+ +IG Y+ R D+L+ ++ AQ P L +++++ T + L++ K
Sbjct: 1536 MQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHK 1595
Query: 172 SS*Y----KLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEY- 226
+ KL DA Y G++ KS GN L ++ S + S ST EAE
Sbjct: 1596 NKPTEPDNKLVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIH 1654
Query: 227 -ISAAI-CSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHS-RAKHIEVKYH 283
IS ++ + ++ +L I++ ++ D+ + IS+ K+ R + K
Sbjct: 1655 AISESVPLLNNLSYLIQELNKKPIIKGLLT---DSRSTISIIKSTNEEKFRNRFFGTKAM 1711
Query: 284 FIRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFI 321
+RD V L + +++T AD+ +KPL FK +
Sbjct: 1712 RLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLL 1749
>IBPA_ECOLI (P29209) 16 kDa heat shock protein A
Length = 137
Score = 32.3 bits (72), Expect = 1.7
Identities = 14/46 (30%), Positives = 24/46 (51%)
Query: 40 QAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTKEFLKKFNMTE 85
++E E++ L G D+Q E TY++Q + F +KF + E
Sbjct: 58 ESELEITAQDNLLVVKGAHADEQKERTYLYQGIAERNFERKFQLAE 103
>PYRG_UREPA (Q9PQK7) CTP synthase (EC 6.3.4.2) (UTP--ammonia ligase)
(CTP synthetase)
Length = 542
Score = 32.0 bits (71), Expect = 2.2
Identities = 19/77 (24%), Positives = 36/77 (46%), Gaps = 11/77 (14%)
Query: 238 WMKHQLEDYQILESNISIYCDN------TAAISLSKNLILHSRAKHIEVKYHFIRDYVQK 291
++K+++E+Y+I E YC N T+ I L +N + H E H+I +Y
Sbjct: 429 FVKNEIENYRIGE-----YCSNIQANTITSQIYLKQNQLNERHRHHFEFNNHYINNYFLN 483
Query: 292 GVLLLKFVDTDHQWADI 308
+ + D+ + D+
Sbjct: 484 QNWKIGAISVDNNYIDV 500
>TRF1_MOUSE (P70371) Telomeric repeat binding factor 1 (TTAGGG
repeat binding factor 1)
Length = 421
Score = 30.8 bits (68), Expect = 5.0
Identities = 15/40 (37%), Positives = 26/40 (64%), Gaps = 1/40 (2%)
Query: 4 FYHFSYLQLVSE-QIYVDDIIFGSANTFLCKEFSKMMQAE 42
F HFSY ++ + Q YV D++ ++TFL K +K+++ E
Sbjct: 215 FQHFSYSCMMEKIQSYVGDVLSEKSSTFLMKAATKVVENE 254
>RHLG_PSEAE (Q9RPT1) Rhamnolipids biosynthesis
3-oxoacyl-[acyl-carrier-protein] reductase (EC
1.1.1.100) (3-ketoacyl-acyl carrier protein reductase)
Length = 256
Score = 30.0 bits (66), Expect = 8.5
Identities = 16/43 (37%), Positives = 22/43 (50%)
Query: 182 DADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEA 224
DA+ D R S YG+CQ + ++L S A R+ AL A
Sbjct: 42 DAEACADTATRLSAYGDCQAIPADLSSEAGARRLAQALGELSA 84
>HLYD_ACTAC (P18790) Leukotoxin secretion protein D
Length = 477
Score = 30.0 bits (66), Expect = 8.5
Identities = 26/90 (28%), Positives = 42/90 (45%), Gaps = 14/90 (15%)
Query: 253 ISIYCDNTAAISLSKNLILHSRAKHIE------VKYHFIRD--YVQKGVLLLKF------ 298
+SI+ + + S L R+K I+ VK+ F+++ YV+KG LLLK
Sbjct: 74 VSIFSNVEIIATASGKFALSGRSKEIKPIENSLVKHIFVKEGEYVKKGELLLKLTALGAE 133
Query: 299 VDTDHQWADIFSKPLAEDRFKFILKNLNMD 328
DT + L E R+K +L+ + D
Sbjct: 134 ADTLKTKTSLSQAKLEEFRYKSLLEAVEKD 163
>DPO1_SULAC (P95690) DNA polymerase I (EC 2.7.7.7)
Length = 875
Score = 30.0 bits (66), Expect = 8.5
Identities = 23/90 (25%), Positives = 43/90 (47%), Gaps = 6/90 (6%)
Query: 246 YQILESNISIYCDNTAAISLSKNLILHSRAKHIEV----KYHFIRDYVQKGVLLLKFV-- 299
Y +++ + ++ + T + ++N L++ A V +Y Y Q L LK +
Sbjct: 592 YDVVQRAMKVFINATYGVFGAENFPLYAPAVAESVTAIGRYIITTTYKQAEKLNLKVIYG 651
Query: 300 DTDHQWADIFSKPLAEDRFKFILKNLNMDL 329
DTD + +K E+ KF+ +N N+DL
Sbjct: 652 DTDSLFLYNPTKDKLEELIKFVKQNFNLDL 681
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,605,472
Number of Sequences: 164201
Number of extensions: 1433287
Number of successful extensions: 3873
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 3845
Number of HSP's gapped (non-prelim): 20
length of query: 334
length of database: 59,974,054
effective HSP length: 111
effective length of query: 223
effective length of database: 41,747,743
effective search space: 9309746689
effective search space used: 9309746689
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0022.11


