

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
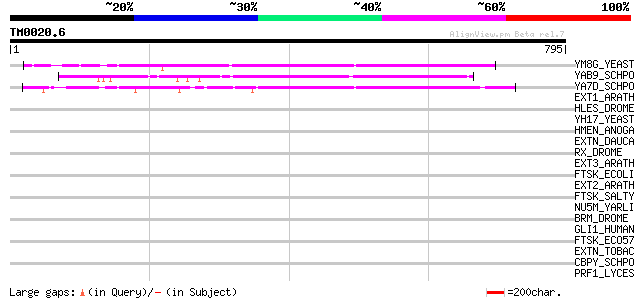
Query= TM0020.6
(795 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YM8G_YEAST (Q03516) Hypothetical 107.7 kDa protein in TSP3-IPP2 ... 178 4e-44
YAB9_SCHPO (Q09809) Hypothetical protein C2G11.09 in chromosome I 173 2e-42
YA7D_SCHPO (Q09766) Hypothetical protein C24H6.13 in chromosome I 160 9e-39
EXT1_ARATH (Q38913) Extensin 1 precursor (AtExt1) (AtExt4) 42 0.008
HLES_DROME (Q02308) Hairless protein 41 0.014
YH17_YEAST (P38898) Hypothetical 17.1 kDa protein in PUR5 3'region 40 0.024
HMEN_ANOGA (O02491) Segmentation polarity homeobox protein engra... 40 0.024
EXTN_DAUCA (P06599) Extensin precursor 40 0.024
RX_DROME (Q9W2Q1) Retinal homeobox protein Rx (DRx1) (DRx) 40 0.032
EXT3_ARATH (Q9FS16) Extensin 3 precursor (AtExt3) (AtExt5) 40 0.032
FTSK_ECOLI (P46889) DNA translocase ftsK 39 0.041
EXT2_ARATH (Q9M1G9) Extensin 2 precursor (AtExt2) (Cell wall hyd... 39 0.041
FTSK_SALTY (Q8ZQD5) DNA translocase ftsK 39 0.054
NU5M_YARLI (Q9B6D3) NADH-ubiquinone oxidoreductase chain 5 (EC 1... 37 0.16
BRM_DROME (P25439) Homeotic gene regulator (Brahma protein) 37 0.16
GLI1_HUMAN (P08151) Zinc finger protein GLI1 (Glioma-associated ... 37 0.20
FTSK_ECO57 (Q8X5H9) DNA translocase ftsK 37 0.20
EXTN_TOBAC (P13983) Extensin precursor (Cell wall hydroxyproline... 37 0.27
CBPY_SCHPO (O13849) Carboxypeptidase Y precursor (EC 3.4.16.5) (... 37 0.27
PRF1_LYCES (Q00451) 36.4 kDa proline-rich protein 36 0.35
>YM8G_YEAST (Q03516) Hypothetical 107.7 kDa protein in TSP3-IPP2
intergenic region
Length = 953
Score = 178 bits (452), Expect = 4e-44
Identities = 168/706 (23%), Positives = 314/706 (43%), Gaps = 62/706 (8%)
Query: 21 FLLAFALLRIQPINDRVYFPKWYIN--GERTSPRSTGNFVGKFVNLNFKTYLTFLNWMPQ 78
F++AF +LRI+ R+Y PK N E P V + W+
Sbjct: 37 FVIAFLILRIKL--KRIYEPKSSFNLINEEKKPEPLPQGVWQ--------------WLKP 80
Query: 79 ALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALL-ILIPVNVSSGALFFLKRDL 137
L+ S+N +I AGLD FLR Y + I+ V++ + IL+ +N S+G
Sbjct: 81 LLKKSDNFVIQQAGLDGYFFLR-YLFIIAIYCAVSMSYIFPILLSINASNGNH------- 132
Query: 138 VVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQR--R 195
S +++L+ NV + R+F H+ ++F +++Y+E M+ LAS R +
Sbjct: 133 -ESGLNQLAYQNVKHRG-RYFAHVFCGWIFFWGFLYIIYRELYFYTSMKQAVLASPRYAK 190
Query: 196 RVEQFTVVVRSVP--HISGHSVSD---------------SVESFFQT--NHPNHYIGHQA 236
++ TV+ ++VP ++S S S+E+ + N G +
Sbjct: 191 KLSSRTVLFQTVPKQYLSEEEFSKLFDGVKRVWIARGSGSIEAMVKARDNMAIQLEGAET 250
Query: 237 VYNANKFAKLVRRRDRLQNWLDYYQLKFQRHPDK-RP--TINTGVLGLWGRKVDAIEYYS 293
Y K +++ ++ L + PDK RP IN +G+KVD I Y
Sbjct: 251 KY-LKAALKKIKKLNKKSPQLSVSDNIAEYVPDKKRPHHKINKVAKFFFGKKVDTISYIK 309
Query: 294 HTIKELD-KVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAP 352
+ +L+ KV L+ P F+ F S++ A V AQ P +
Sbjct: 310 EELPKLNQKVKALQEDHENSSP---FNSVFVEFESQYQAQVAAQITTYHAPLFMTPVYIG 366
Query: 353 -EPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLR 411
EP D+ W NL + + R++ ++ ALV + P+AFV ++N+ L +L+
Sbjct: 367 IEPSDVVWFNLRMFWWERLGREVSAVSAIVALVILWAFPVAFVGMISNITSLTNEVKWLK 426
Query: 412 PVIEL-KFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFM 470
+ +L K + L P +AL + + LP + M+ +G +K +E T Y+ F
Sbjct: 427 FIYKLPKQLLGLLTSLAPTVALAVLMSFLPKFIRGMAITQGAPSKQNVEYFTQQAYFAFQ 486
Query: 471 LVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEI 530
++ VFL + ++ A + + + PT+ + ++P + FFM+Y+++ G + +G +
Sbjct: 487 VIQVFLVTTLSSAATSTVTEIVKE-PTKAMDLLASNLPKASNFFMSYVILQGLSISSGAL 545
Query: 531 LRLKPLVIYHLKNMFIVKTERDR-GKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILL 589
L++ PL+++++ F+ T R + + S+ + P ++ Y++++P++L
Sbjct: 546 LQIVPLILFYVLGAFLDGTVRKKWNRFCGLSSMQWGTAFPVYTNLAVITFSYSIISPLIL 605
Query: 590 PFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKA 649
F V F L Y+ Y + + VY + ++ ++P + I + I Q+ +LGL + K
Sbjct: 606 LFAAVAFFLLYIAYLYNLTYVYQESPDARGIYYPRALFQTIVGIYIGQICLLGLFAVGKG 665
Query: 650 AKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKT 695
L I +T H + F+ + P++ D + T
Sbjct: 666 WGPIVLQVIGICVTVLIHLHLSAAFDHLSKVIPVDTMKPLDGVSDT 711
>YAB9_SCHPO (Q09809) Hypothetical protein C2G11.09 in chromosome I
Length = 796
Score = 173 bits (438), Expect = 2e-42
Identities = 152/675 (22%), Positives = 300/675 (43%), Gaps = 96/675 (14%)
Query: 71 TFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNV----- 125
++ W+ + + ++ + N AGLD VFL + +G+K +++ +LI++PVN
Sbjct: 70 SYYKWLMDLVNIPDDVVQNCAGLDGYVFLLFFKMGIKFLSFASLLGVLIIMPVNKHFRGD 129
Query: 126 ----------SSGALFF----LKRDLVVSDI---------------------DKLSISNV 150
+ FF +K+ +V S I + I +
Sbjct: 130 AFGNITLSMPAKSEYFFSSPLVKKSIVQSPIIANGSELNVGVLGPSLFNPIGNLSDIPGL 189
Query: 151 PSKSSRF-FVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPH 209
P F ++++ Y ++++ ++L+ +IA +R +LA Q R ++ TV + +P+
Sbjct: 190 PQPGDGFLYLYVLFTYFISIFLLYVLFSSTKSIADIRQSYLARQNRLTDR-TVFISGLPN 248
Query: 210 ISGHSVSDSVESFFQTNHPNHYIGHQAVYN-------ANKFAKLVRRRDRL--------- 253
+++++++F N +K +K V++ ++
Sbjct: 249 EL--CSTENLKAYFDKLDVGSIDSLSICRNYSYMDILLSKKSKYVKKLEKYWSIYLSNCK 306
Query: 254 ---------QNWLDYYQLKFQRHPDK-----------RPTINTGVLGLWGRKVDAIEYYS 293
N+L + + + P++ P I T G++G+K+DAI++YS
Sbjct: 307 KLGISTLPPSNYLSPNRAELESTPEQLLEVPWQHHQCHPLIKTHFFGIFGQKIDAIDFYS 366
Query: 294 HTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWL-TDWAP 352
+ ++ + +E R P AF++F S A + AQT + L + AP
Sbjct: 367 AKLYKISQ--QIENARSFDYPTT--GQAFITFESMATAQIVAQTHIDSKSLMGLHIELAP 422
Query: 353 EPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRP 412
DI W N I + I+L F ++ + +P+ + NL+ + R+ P L
Sbjct: 423 AANDIQWHNTYIGRWHKFFQGWFITLVTFMIILLWTVPVGAIAVFINLDTIRRLWPELGR 482
Query: 413 VIE-LKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFML 471
+IE L F+ S L+ FLP L +F+ I P + +S ++G +++ E K Y ++
Sbjct: 483 MIEDLPFLNSLLRTFLPTLVYSLFISISPFLFRWLSSMQGLSSRAEEEIYAVGKNYAYLF 542
Query: 472 VNVFLGSIVTGTA--FEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGE 529
VN FL ++ G+ +E L + T + +P +A FF+ I++ G +
Sbjct: 543 VNFFLVYVIAGSTSIWE-----LAKDTTSFAHFLANRLPHQAQFFIDLIVLQGIGMFPLK 597
Query: 530 ILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILL 589
+++L L Y ++ F+ + + K P S +P L+ + Y++++P++L
Sbjct: 598 LIQLGKLSSYFVRRSFVPYSIASK-KFETPDSFSVGIFLPQPMFIMLICLCYSIISPLIL 656
Query: 590 PFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKA 649
F L++F + +LVY++++I S W + R+I +I QL M+GL+S +KA
Sbjct: 657 VFGLIYFIIGFLVYKYELIYQMEHPQHSTGELWSTIFLRMIFGCVIMQLTMMGLMSLRKA 716
Query: 650 AKSTPLLAILPILTF 664
+ + I P+L F
Sbjct: 717 YWLSTV--IFPLLCF 729
>YA7D_SCHPO (Q09766) Hypothetical protein C24H6.13 in chromosome I
Length = 871
Score = 160 bits (406), Expect = 9e-39
Identities = 175/748 (23%), Positives = 323/748 (42%), Gaps = 86/748 (11%)
Query: 19 FAFLLAFA--LLRIQPINDRVYFPKWYING----ERTSPRSTGNFVGKFVNLNFKTYLTF 72
FA AF L ++P VY P+ I+ E+ P + F G F +
Sbjct: 20 FAIFCAFIGLFLCLRPREKHVYQPRCIIDTQPKEEKPEPSPSSPF-GLFAYV-------- 70
Query: 73 LNWMPQALRMSENEIINHAGLDSAVFLR-IYTLGLKIFVPVTVVALLILIPVNVSSGALF 131
++ SE +I +AG+D F+R ++T G + + +V IL+PVN ++G
Sbjct: 71 -------VKRSETYLIQYAGVDGYFFIRYLFTFGA-LCILGCLVLFPILLPVNATNG--- 119
Query: 132 FLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKE------------- 178
+ D LS SNV + + RF+ H+ L ++F + F++Y+E
Sbjct: 120 -----VGEKGFDILSFSNVKNHN-RFYAHVFLSWLFFGFTIFIIYRELRYYVIFRHAMQS 173
Query: 179 ---YDNIAMMRLHFLASQRRRVEQFTVVVRSV-PHISGHSVSDSVESFFQTNHPNHYIGH 234
Y+N+ L V + + P+ S + ++ + +G+
Sbjct: 174 SGLYNNLPSSSTMLLTELPNSVLNDEETLHELFPNASEFTCVRDLKKLEKKVKKRSDLGN 233
Query: 235 QAVYNAN--------KFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKV 286
+ N K KLV++ L + LDY + KRPT L G+KV
Sbjct: 234 KYESTLNSLINKSVKKHNKLVKKHKPLPSTLDY-----TAYVKKRPTHRLKFL--IGKKV 286
Query: 287 DAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQT-QQSKNPTL 345
D I+Y TI ELD+V+ + Q +++ K + V F+ F S+ Q SK
Sbjct: 287 DTIDYCRDTIAELDEVVD-KLQTSLEERKKVGSV-FIRFRSQTDLQTAYQAFLYSKKFRK 344
Query: 346 W-----LTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANL 400
+ L APE DI W NL + + +K + + + ++ F+ P+A V ++N+
Sbjct: 345 YRFGRALVGIAPE--DIVWSNLDLSMYTRRGKKTISNTILTLMIIFWAFPVAVVGCISNV 402
Query: 401 EGLERVAPFLRPVIELK-FIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLE 459
L FL+ + + + + G LP +AL I + ++P + + K G + +E
Sbjct: 403 NYLIEKVHFLKFIDHMPPKLLGIITGILPSVALSILMSLVPPFIKFLGKFGGALTVQEIE 462
Query: 460 RKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIM 519
YY F +V VFL + +T A + + + P + ++P + F+++Y +
Sbjct: 463 NYCQNWYYAFQVVQVFLVTTMTSAATSAVVQVIKE-PASSMTLLASNLPKASNFYISYFL 521
Query: 520 VDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETI-PSLQLYFLLG 578
+ G + G +L++ L++ + T R + + S T+ P L +
Sbjct: 522 LQGLSIPGGALLQIVTLLLSKVLGRIFDNTPRKKWNRWNQLSAPSWGTVYPVYSLLVTIM 581
Query: 579 IVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQL 638
I Y+++ PI++ F V F L Y Y + +I V ++ +P ++ L ++++
Sbjct: 582 ICYSIIAPIIIGFAAVAFVLIYFAYSYNLIYVLGHNADAKGRNYPRALFQVFVGLYLAEV 641
Query: 639 LMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTE- 697
++GL K +T L A+ T A H Y +++F PL +A+ +E +E
Sbjct: 642 CLIGLFVLAKNWGATVLEAVFLGFTVACHLYFKYKF------LPLMDAVPISAIESVSER 695
Query: 698 PDLNIKAYLADAYLHPIFRSF-EVEEEL 724
P++ L + + + R++ E+ E+L
Sbjct: 696 PEIKYPMDLGTSEMKNVGRAYPEILEKL 723
>EXT1_ARATH (Q38913) Extensin 1 precursor (AtExt1) (AtExt4)
Length = 373
Score = 41.6 bits (96), Expect = 0.008
Identities = 25/76 (32%), Positives = 31/76 (39%), Gaps = 13/76 (17%)
Query: 733 PETQVASPTLSEPSSPSPLHDH-----VHQPSPPQHVNQ---YSHYPT-----SPPGYYY 779
P + P + S P P+H H P PP H + H P SPP Y
Sbjct: 260 PPVHYSPPPVVYHSPPPPVHYSPPPVVYHSPPPPVHYSPPPVVYHSPPPPVHYSPPPVVY 319
Query: 780 HPPPPPHYAYQYQDEP 795
H PPPP Y+Y+ P
Sbjct: 320 HSPPPPKKHYEYKSPP 335
Score = 37.4 bits (85), Expect = 0.16
Identities = 20/61 (32%), Positives = 26/61 (41%), Gaps = 12/61 (19%)
Query: 733 PETQVASPTLSEPSSPSPLHDH-----VHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P + P + S P P+H H P PP H + PP Y+ PPPP HY
Sbjct: 244 PPVHYSPPPVVYHSPPPPVHYSPPPVVYHSPPPPVHYSP-------PPVVYHSPPPPVHY 296
Query: 788 A 788
+
Sbjct: 297 S 297
Score = 37.0 bits (84), Expect = 0.20
Identities = 19/46 (41%), Positives = 20/46 (43%), Gaps = 9/46 (19%)
Query: 746 SSPSPLHDH----VHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
S P P+H H P PP H HY Y Y PPPPHY
Sbjct: 333 SPPPPVHYSPPTVYHSPPPPVH-----HYSPPHQPYLYKSPPPPHY 373
Score = 35.8 bits (81), Expect = 0.46
Identities = 25/70 (35%), Positives = 29/70 (40%), Gaps = 14/70 (20%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSHYPT----------SPPGYYY 779
P + SP S P P+ + P SPP V YS P SPP Y
Sbjct: 196 PPVKYYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPPVYKSPPPPVHYSPPPVVY 255
Query: 780 HPPPPP-HYA 788
H PPPP HY+
Sbjct: 256 HSPPPPVHYS 265
Score = 34.7 bits (78), Expect = 1.0
Identities = 19/64 (29%), Positives = 28/64 (43%), Gaps = 3/64 (4%)
Query: 733 PETQVASPTLSEPSSPSPL-HDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQY 791
P + P + S P P H P PP H + + Y + PP +++ PPH Y Y
Sbjct: 308 PPVHYSPPPVVYHSPPPPKKHYEYKSPPPPVHYSPPTVYHSPPPPVHHY--SPPHQPYLY 365
Query: 792 QDEP 795
+ P
Sbjct: 366 KSPP 369
Score = 33.1 bits (74), Expect = 3.0
Identities = 23/63 (36%), Positives = 28/63 (43%), Gaps = 7/63 (11%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSH---YPTSPPGYYYHPPPP-P 785
P + SP S P P+ + P SPP V YS Y + PP Y PPPP
Sbjct: 44 PPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPPVYKSPPPPVYKSPPPPVK 103
Query: 786 HYA 788
HY+
Sbjct: 104 HYS 106
Score = 31.6 bits (70), Expect = 8.6
Identities = 21/58 (36%), Positives = 24/58 (41%), Gaps = 8/58 (13%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P + SP S P P+ + P SPP V YS PP Y PPPP Y
Sbjct: 116 PPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKHYS-----PPPVYKSPPPPVKY 168
>HLES_DROME (Q02308) Hairless protein
Length = 1077
Score = 40.8 bits (94), Expect = 0.014
Identities = 21/50 (42%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Query: 736 QVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPP 785
+ ASP+ S S PSP D P +H+ Q H S P +YY PPPP
Sbjct: 809 EFASPSASSSSCPSP-GDRSASPPERRHMQQQPHLQRSSPLHYYMYPPPP 857
>YH17_YEAST (P38898) Hypothetical 17.1 kDa protein in PUR5 3'region
Length = 153
Score = 40.0 bits (92), Expect = 0.024
Identities = 28/88 (31%), Positives = 33/88 (36%), Gaps = 37/88 (42%)
Query: 733 PETQVASPTLSEPSSPSPL-HDHVHQPSP-------PQH------------VNQYSHYPT 772
P +PT + P +P+P H H H P P P H +N SHYPT
Sbjct: 26 PHPHPHTPTHTHPHTPTPTPHPHPHTPHPHTTPTPTPHHTHTPHTTLSNLSLNLPSHYPT 85
Query: 773 SP-----------------PGYYYHPPP 783
SP YYYHPPP
Sbjct: 86 SPLVTLPHSTIPLPTTIHLSTYYYHPPP 113
>HMEN_ANOGA (O02491) Segmentation polarity homeobox protein
engrailed
Length = 596
Score = 40.0 bits (92), Expect = 0.024
Identities = 23/63 (36%), Positives = 32/63 (50%), Gaps = 7/63 (11%)
Query: 730 DKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPT-SPPGY-----YYHPPP 783
++Q + SP + EP SP P+H V P PQHV +H+P P Y +Y+ P
Sbjct: 145 EQQQQPHPQSPAIREPISPGPIHPAVLLPY-PQHVLHPAHHPALLHPAYHTGLHHYYQPS 203
Query: 784 PPH 786
P H
Sbjct: 204 PSH 206
>EXTN_DAUCA (P06599) Extensin precursor
Length = 306
Score = 40.0 bits (92), Expect = 0.024
Identities = 25/73 (34%), Positives = 31/73 (42%), Gaps = 16/73 (21%)
Query: 739 SPTLSEPSSPSPLHDHV-----HQPSPPQHVNQYSHYP----TSPPGYYY-------HPP 782
SP E S P P H H P PP V +Y P + PP Y++ H P
Sbjct: 45 SPPPPEHSPPPPYHYESPPPPKHSPPPPTPVYKYKSPPPPMHSPPPPYHFESPPPPKHSP 104
Query: 783 PPPHYAYQYQDEP 795
PPP Y+Y+ P
Sbjct: 105 PPPTPVYKYKSPP 117
Score = 39.3 bits (90), Expect = 0.041
Identities = 22/64 (34%), Positives = 30/64 (46%), Gaps = 15/64 (23%)
Query: 747 SPSPLHDHVHQ-PSPPQHVNQYS------HYPTSPPGYYYHPPPPP--------HYAYQY 791
SP+P+H + ++ P PP V +Y H P Y Y PPPP HY Y+Y
Sbjct: 122 SPAPVHHYKYKSPPPPTPVYKYKSPPPPKHSPAPEHHYKYKSPPPPKHFPAPEHHYKYKY 181
Query: 792 QDEP 795
+ P
Sbjct: 182 KSPP 185
Score = 39.3 bits (90), Expect = 0.041
Identities = 23/66 (34%), Positives = 30/66 (44%), Gaps = 10/66 (15%)
Query: 740 PTLSEPSSPSPLHDH------VHQPSPPQHVNQYSHYPT---SPPGYYYHPPPPP-HYAY 789
P P P+P++ + +H P PP V +Y P SPP Y PPPP HY+Y
Sbjct: 238 PPEHSPPPPTPVYKYKSPPPPMHSPPPPTPVYKYKSPPPPMHSPPPPVYSPPPPKHHYSY 297
Query: 790 QYQDEP 795
P
Sbjct: 298 TSPPPP 303
Score = 37.4 bits (85), Expect = 0.16
Identities = 20/58 (34%), Positives = 26/58 (44%), Gaps = 9/58 (15%)
Query: 747 SPSPLHDHV--------HQPSPPQHVN-QYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP+P H + H P+P H +Y P P Y Y PPPP Y+Y+ P
Sbjct: 152 SPAPEHHYKYKSPPPPKHFPAPEHHYKYKYKSPPPPTPVYKYKSPPPPTPVYKYKSPP 209
Score = 37.0 bits (84), Expect = 0.20
Identities = 20/67 (29%), Positives = 29/67 (42%), Gaps = 11/67 (16%)
Query: 740 PTLSEPSSPSPLHDHV------HQPSPPQHVNQYSHYPTSPPGYYYHPPPP-----PHYA 788
P P P+P++ + H P+P H S P +P Y PPPP P +
Sbjct: 99 PPKHSPPPPTPVYKYKSPPPPKHSPAPVHHYKYKSPPPPTPVYKYKSPPPPKHSPAPEHH 158
Query: 789 YQYQDEP 795
Y+Y+ P
Sbjct: 159 YKYKSPP 165
Score = 37.0 bits (84), Expect = 0.20
Identities = 19/57 (33%), Positives = 28/57 (48%), Gaps = 9/57 (15%)
Query: 748 PSPLHDHVHQ---PSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA------YQYQDEP 795
P+P H + ++ P PP V +Y P P Y Y PPPP ++ Y+Y+ P
Sbjct: 171 PAPEHHYKYKYKSPPPPTPVYKYKSPPPPTPVYKYKSPPPPKHSPAPVHHYKYKSPP 227
Score = 37.0 bits (84), Expect = 0.20
Identities = 25/73 (34%), Positives = 32/73 (43%), Gaps = 8/73 (10%)
Query: 724 LVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYS-HYPTSPPGYYYHPP 782
+V V ++ ET A T S P P H P PP+H HY + PP + PP
Sbjct: 19 VVLVSLNLASET-TAKYTYSSPPPPE------HSPPPPEHSPPPPYHYESPPPPKHSPPP 71
Query: 783 PPPHYAYQYQDEP 795
P P Y Y+ P
Sbjct: 72 PTPVYKYKSPPPP 84
Score = 36.6 bits (83), Expect = 0.27
Identities = 21/76 (27%), Positives = 32/76 (41%), Gaps = 12/76 (15%)
Query: 731 KQPETQVASPTLSEPSSPSPLHDHVHQPSPPQH----VNQYSHYPTSPPGYYYHPPPPPH 786
K P P P+P++ + P PP+H V+ Y + PP Y PPPP
Sbjct: 182 KSPPPPTPVYKYKSPPPPTPVYKY-KSPPPPKHSPAPVHHYKYKSPPPPTPVYKSPPPPE 240
Query: 787 YA-------YQYQDEP 795
++ Y+Y+ P
Sbjct: 241 HSPPPPTPVYKYKSPP 256
Score = 35.8 bits (81), Expect = 0.46
Identities = 16/56 (28%), Positives = 24/56 (42%)
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P P P+P++ + P P H+ + PP + PPP P Y Y+ P
Sbjct: 64 PPKHSPPPPTPVYKYKSPPPPMHSPPPPYHFESPPPPKHSPPPPTPVYKYKSPPPP 119
Score = 34.3 bits (77), Expect = 1.3
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 6/50 (12%)
Query: 747 SPSPLHDHVHQ-PSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP+P+H + ++ P PP V Y + PP + PPP P Y Y+ P
Sbjct: 214 SPAPVHHYKYKSPPPPTPV-----YKSPPPPEHSPPPPTPVYKYKSPPPP 258
Score = 33.5 bits (75), Expect = 2.3
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 2/51 (3%)
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYY-HPPPPPHYAY 789
P + P P+P++ + P PP H Y PP ++Y + PPP + Y
Sbjct: 257 PPMHSPPPPTPVYKY-KSPPPPMHSPPPPVYSPPPPKHHYSYTSPPPPHHY 306
>RX_DROME (Q9W2Q1) Retinal homeobox protein Rx (DRx1) (DRx)
Length = 873
Score = 39.7 bits (91), Expect = 0.032
Identities = 20/54 (37%), Positives = 23/54 (42%), Gaps = 5/54 (9%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPH 786
P +A L+ S + H H H PP HV H PP PPPPPH
Sbjct: 644 PPLSLAPGNLTMSSLAAMGHHHAHNGPPPPHVGHGGHGQPQPP-----PPPPPH 692
>EXT3_ARATH (Q9FS16) Extensin 3 precursor (AtExt3) (AtExt5)
Length = 427
Score = 39.7 bits (91), Expect = 0.032
Identities = 25/83 (30%), Positives = 34/83 (40%), Gaps = 6/83 (7%)
Query: 713 PIFRSFEVEEELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPT 772
P++ S ++ E + P + P + SP P H SPP V YS
Sbjct: 48 PVYHSPPPPKKHYEYKSPPPPVKHYSPPPVYH--SPPPPKKHYVYKSPPPPVKHYS---- 101
Query: 773 SPPGYYYHPPPPPHYAYQYQDEP 795
PP Y+ PPP HY Y+ P
Sbjct: 102 PPPVYHSPPPPKKHYVYKSPPPP 124
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 108 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 152
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 304 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 348
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 276 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 320
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 136 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 180
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 164 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 208
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 248 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 292
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 332 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 376
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 192 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 236
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 4/49 (8%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS PP Y+ PPP HY Y+ P
Sbjct: 220 SPPPPKKHYVYKSPPPPVKHYS----PPPVYHSPPPPKKHYVYKSPPPP 264
Score = 38.1 bits (87), Expect = 0.092
Identities = 21/55 (38%), Positives = 26/55 (47%), Gaps = 2/55 (3%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P + SP S P P +V++ PP V+ YS P P Y PPPP HY
Sbjct: 375 PPVKHYSPPPVYHSPPPPKEKYVYKSPPPPPVHHYS--PPHHPYLYKSPPPPYHY 427
Score = 37.7 bits (86), Expect = 0.12
Identities = 20/49 (40%), Positives = 21/49 (42%), Gaps = 6/49 (12%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SP P H SPP V YS P YH PPPP Y Y+ P
Sbjct: 360 SPPPPKKHYVYKSPPPPVKHYSPPPV------YHSPPPPKEKYVYKSPP 402
Score = 33.5 bits (75), Expect = 2.3
Identities = 19/51 (37%), Positives = 24/51 (46%), Gaps = 12/51 (23%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPP--------PPPHYAY 789
SP P++ H P PP+ Y P PP ++Y PP PPP Y Y
Sbjct: 381 SPPPVY---HSPPPPKEKYVYKS-PPPPPVHHYSPPHHPYLYKSPPPPYHY 427
Score = 33.1 bits (74), Expect = 3.0
Identities = 23/76 (30%), Positives = 32/76 (41%), Gaps = 13/76 (17%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQ--PSPPQHVNQYSHYPTSPP---GYYYHPPP---- 783
P + SP S P P +V++ P P +H + Y + PP Y Y PP
Sbjct: 347 PPVKHYSPPPVYHSPPPPKKHYVYKSPPPPVKHYSPPPVYHSPPPPKEKYVYKSPPPPPV 406
Query: 784 ----PPHYAYQYQDEP 795
PPH+ Y Y+ P
Sbjct: 407 HHYSPPHHPYLYKSPP 422
Score = 33.1 bits (74), Expect = 3.0
Identities = 17/41 (41%), Positives = 21/41 (50%), Gaps = 5/41 (12%)
Query: 759 SPPQHVNQYS----HYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SPP V Y+ HY + PP Y+ PPP HY Y+ P
Sbjct: 29 SPPPPVKHYTPPVKHY-SPPPVYHSPPPPKKHYEYKSPPPP 68
>FTSK_ECOLI (P46889) DNA translocase ftsK
Length = 1329
Score = 39.3 bits (90), Expect = 0.041
Identities = 27/96 (28%), Positives = 38/96 (39%), Gaps = 6/96 (6%)
Query: 702 IKAYLADAYLHPIFRSFEVEEELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPP 761
+KA L D P+F + E + + + P+ Q P +P P P + QP P
Sbjct: 719 MKALLDDGPHEPLFTP--IVEPVQQPQQPVAPQQQYQQP--QQPVPPQPQYQQPQQPVAP 774
Query: 762 QHVNQYSHYPTSPPGYYYHPPPP--PHYAYQYQDEP 795
Q Q P +P Y P P P YQ +P
Sbjct: 775 QPQYQQPQQPVAPQQQYQQPQQPVAPQQQYQQPQQP 810
>EXT2_ARATH (Q9M1G9) Extensin 2 precursor (AtExt2) (Cell wall
hydroxyproline-rich glycoprotein 1) (HRGP1)
Length = 743
Score = 39.3 bits (90), Expect = 0.041
Identities = 16/42 (38%), Positives = 23/42 (54%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SP P + + P P + HY + PP Y Y+ PPPP+Y+
Sbjct: 545 SPPPPYVYSSPPPPYYSPSPKVHYKSPPPPYVYNSPPPPYYS 586
Score = 38.9 bits (89), Expect = 0.054
Identities = 20/58 (34%), Positives = 29/58 (49%), Gaps = 5/58 (8%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSH--YPTSPPGYYYHPPPPPHYA 788
P T SP + S P P +V+ PP + + Y + PP Y Y+ PPPP+Y+
Sbjct: 82 PPTYSPSPKVDYKSPPPP---YVYSSPPPPYYSPSPKVDYKSPPPPYVYNSPPPPYYS 136
Score = 38.5 bits (88), Expect = 0.070
Identities = 16/42 (38%), Positives = 22/42 (52%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SP P + + P P + HY + PP Y Y PPPP+Y+
Sbjct: 520 SPPPPYVYSSPPPPYYSPSPKVHYKSPPPPYVYSSPPPPYYS 561
Score = 38.5 bits (88), Expect = 0.070
Identities = 19/56 (33%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P T +P + E SP P + + P P + Y + PP Y Y PPPP+Y+
Sbjct: 57 PPTYTPAPEV-EYKSPPPPYVYSSPPPPTYSPSPKVDYKSPPPPYVYSSPPPPYYS 111
Score = 38.5 bits (88), Expect = 0.070
Identities = 20/58 (34%), Positives = 29/58 (49%), Gaps = 5/58 (8%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYS--HYPTSPPGYYYHPPPPPHYA 788
P T SP + S P P +V+ PP + + +Y + PP Y Y PPPP+Y+
Sbjct: 407 PPTYSPSPKVYYKSPPPP---YVYSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYS 461
Score = 37.7 bits (86), Expect = 0.12
Identities = 16/45 (35%), Positives = 23/45 (50%)
Query: 744 EPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
E SP P + + P P + +Y + PP Y Y PPPP+Y+
Sbjct: 392 EYKSPPPPYVYSSPPPPTYSPSPKVYYKSPPPPYVYSSPPPPYYS 436
Score = 37.0 bits (84), Expect = 0.20
Identities = 20/58 (34%), Positives = 27/58 (46%), Gaps = 5/58 (8%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYS--HYPTSPPGYYYHPPPPPHYA 788
P T SP + S P P +V+ PP + + Y + PP Y Y PPPP Y+
Sbjct: 357 PPTYSPSPKVDYKSPPPP---YVYSSPPPPYYSPSPKVEYKSPPPPYVYSSPPPPTYS 411
Score = 36.2 bits (82), Expect = 0.35
Identities = 18/52 (34%), Positives = 27/52 (51%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYS--HYPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + +Y + PP Y Y PPPP+Y+
Sbjct: 563 SPKVHYKSPPPP---YVYNSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYS 611
Score = 36.2 bits (82), Expect = 0.35
Identities = 15/42 (35%), Positives = 22/42 (51%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SP P + + P P + +Y + PP Y Y PPPP+Y+
Sbjct: 495 SPPPPYVYSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYS 536
Score = 36.2 bits (82), Expect = 0.35
Identities = 15/42 (35%), Positives = 22/42 (51%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SP P + + P P + +Y + PP Y Y PPPP+Y+
Sbjct: 470 SPPPPYVYSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYS 511
Score = 36.2 bits (82), Expect = 0.35
Identities = 15/42 (35%), Positives = 22/42 (51%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SP P + + P P + +Y + PP Y Y PPPP+Y+
Sbjct: 445 SPPPPYVYSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYS 486
Score = 36.2 bits (82), Expect = 0.35
Identities = 15/42 (35%), Positives = 22/42 (51%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SP P + + P P + +Y + PP Y Y PPPP+Y+
Sbjct: 595 SPPPPYVYSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYS 636
Score = 35.8 bits (81), Expect = 0.46
Identities = 16/45 (35%), Positives = 22/45 (48%)
Query: 744 EPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
E SP P + + P P + Y + PP Y Y PPPP+Y+
Sbjct: 167 EYKSPPPPYVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYS 211
Score = 35.8 bits (81), Expect = 0.46
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYS--HYPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + Y + PP Y Y PPPP+Y+
Sbjct: 138 SPKVDYKSPPPP---YVYSSPPPPYYSPSPKVEYKSPPPPYVYSSPPPPYYS 186
Score = 35.8 bits (81), Expect = 0.46
Identities = 15/42 (35%), Positives = 21/42 (49%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SP P + + P P + Y + PP Y Y PPPP+Y+
Sbjct: 345 SPPPPYVYSSPPPPTYSPSPKVDYKSPPPPYVYSSPPPPYYS 386
Score = 35.8 bits (81), Expect = 0.46
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYS--HYPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + Y + PP Y Y PPPP+Y+
Sbjct: 188 SPKVDYKSPPPP---YVYSSPPPPYYSPSPKVEYKSPPPPYVYSSPPPPYYS 236
Score = 35.8 bits (81), Expect = 0.46
Identities = 16/45 (35%), Positives = 22/45 (48%)
Query: 744 EPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
E SP P + + P P + Y + PP Y Y PPPP+Y+
Sbjct: 217 EYKSPPPPYVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYS 261
Score = 35.4 bits (80), Expect = 0.60
Identities = 18/43 (41%), Positives = 22/43 (50%), Gaps = 3/43 (6%)
Query: 749 SPLHDHVHQPSPPQHVNQYSH---YPTSPPGYYYHPPPPPHYA 788
SP H HV PP S Y + PP Y Y+ PPPP+Y+
Sbjct: 661 SPPHPHVCVCPPPPPCYSPSPKVVYKSPPPPYVYNSPPPPYYS 703
Score = 35.4 bits (80), Expect = 0.60
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYSH--YPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + Y + PP Y Y PPPP+Y+
Sbjct: 288 SPKVDYKSPPPP---YVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYS 336
Score = 35.4 bits (80), Expect = 0.60
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYSH--YPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + Y + PP Y Y PPPP+Y+
Sbjct: 113 SPKVDYKSPPPP---YVYNSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYS 161
Score = 35.4 bits (80), Expect = 0.60
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYSH--YPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + Y + PP Y Y PPPP+Y+
Sbjct: 238 SPKVDYKSPPPP---YVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYS 286
Score = 35.4 bits (80), Expect = 0.60
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYSH--YPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + Y + PP Y Y PPPP+Y+
Sbjct: 263 SPKVDYKSPPPP---YVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPYYS 311
Score = 34.3 bits (77), Expect = 1.3
Identities = 20/63 (31%), Positives = 27/63 (42%), Gaps = 6/63 (9%)
Query: 732 QPETQVASPTLSEPSSP-----SPLHDHVHQPSPPQHVNQYS-HYPTSPPGYYYHPPPPP 785
+PET + P L P +P +V PP + Y + PP Y Y PPPP
Sbjct: 24 EPETYASPPPLYSSPLPEVEYKTPPLPYVDSSPPPTYTPAPEVEYKSPPPPYVYSSPPPP 83
Query: 786 HYA 788
Y+
Sbjct: 84 TYS 86
Score = 33.9 bits (76), Expect = 1.7
Identities = 18/52 (34%), Positives = 25/52 (47%), Gaps = 5/52 (9%)
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYSH--YPTSPPGYYYHPPPPPHYA 788
SP + S P P +V+ PP + + Y + PP Y Y PPPP Y+
Sbjct: 313 SPKVDYKSPPPP---YVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPPTYS 361
Score = 32.7 bits (73), Expect = 3.9
Identities = 19/56 (33%), Positives = 25/56 (43%), Gaps = 9/56 (16%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P SP + S P P +V+ PP + Y SP YY PPPP +Y+
Sbjct: 674 PPCYSPSPKVVYKSPPPP---YVYNSPPPPY------YSPSPKVYYKSPPPPSYYS 720
Score = 32.0 bits (71), Expect = 6.6
Identities = 15/42 (35%), Positives = 21/42 (49%)
Query: 747 SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
SPSP + P P + + Y + P YY PPPP+Y+
Sbjct: 611 SPSPKVYYKSPPPPYVYSSPPPPYYSPSPKVYYKSPPPPYYS 652
Score = 32.0 bits (71), Expect = 6.6
Identities = 26/77 (33%), Positives = 30/77 (38%), Gaps = 17/77 (22%)
Query: 726 EVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYY------- 778
EV P V S SPSP D+ + PP +V Y + PP YY
Sbjct: 65 EVEYKSPPPPYVYSSPPPPTYSPSPKVDY--KSPPPPYV-----YSSPPPPYYSPSPKVD 117
Query: 779 YHPPPPPHYAYQYQDEP 795
Y PPPP Y Y P
Sbjct: 118 YKSPPPP---YVYNSPP 131
>FTSK_SALTY (Q8ZQD5) DNA translocase ftsK
Length = 1351
Score = 38.9 bits (89), Expect = 0.054
Identities = 24/77 (31%), Positives = 31/77 (40%), Gaps = 4/77 (5%)
Query: 721 EEELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYH 780
E V+ V QP+ Q P +P +P P + QP PQ Q P +P Y
Sbjct: 721 ESTPVQQPVAPQPQPQYQQP--QQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQ 778
Query: 781 PPPP--PHYAYQYQDEP 795
P P P YQ +P
Sbjct: 779 PQQPVAPQPQYQQPQQP 795
Score = 35.4 bits (80), Expect = 0.60
Identities = 20/65 (30%), Positives = 26/65 (39%), Gaps = 4/65 (6%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP--PHYAYQ 790
P+ Q P +P +P P + QP PQ Q P +P Y P P P YQ
Sbjct: 772 PQPQYQQP--QQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQ 829
Query: 791 YQDEP 795
+P
Sbjct: 830 QPQQP 834
Score = 35.4 bits (80), Expect = 0.60
Identities = 20/65 (30%), Positives = 26/65 (39%), Gaps = 4/65 (6%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP--PHYAYQ 790
P+ Q P +P +P P + QP PQ Q P +P Y P P P YQ
Sbjct: 759 PQPQYQQP--QQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQ 816
Query: 791 YQDEP 795
+P
Sbjct: 817 QPQQP 821
Score = 35.4 bits (80), Expect = 0.60
Identities = 20/65 (30%), Positives = 26/65 (39%), Gaps = 4/65 (6%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP--PHYAYQ 790
P+ Q P +P +P P + QP PQ Q P +P Y P P P YQ
Sbjct: 746 PQPQYQQP--QQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQ 803
Query: 791 YQDEP 795
+P
Sbjct: 804 QPQQP 808
Score = 32.0 bits (71), Expect = 6.6
Identities = 16/49 (32%), Positives = 21/49 (42%), Gaps = 2/49 (4%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHP 781
P+ Q P +P +P P + QP PQ Q PT+P HP
Sbjct: 798 PQPQYQQP--QQPVAPQPQYQQPQQPVAPQPQYQQPQQPTAPQDSLIHP 844
>NU5M_YARLI (Q9B6D3) NADH-ubiquinone oxidoreductase chain 5 (EC
1.6.5.3)
Length = 655
Score = 37.4 bits (85), Expect = 0.16
Identities = 54/265 (20%), Positives = 111/265 (41%), Gaps = 37/265 (13%)
Query: 369 LSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLP 428
L + + I LS + L F+++ AF ++L + + L +++ L +LP
Sbjct: 313 LGMMTIAIGLSAYNLALFHLLGHAFFKALLFMSAGSIIHSILNESQDIRTYGGLLS-YLP 371
Query: 429 GLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYY---YFMLVNVFLGSIVTGT-A 484
+ I + L LM M + G+ K + T Y Y + +L +++T +
Sbjct: 372 YTYICITIASLS--LMAMPGLTGYYTKDIIIESTYGSYSISNYVVYWIAYLSAVLTCVYS 429
Query: 485 FEQLHSFLHQSPTQIPRT--------IGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPL 536
+ L+ + +P T I +++PM + M GW
Sbjct: 430 MKILYLTFYSNPNNNTITYYNAHESNIYITLPM--FILAIFAMFAGWI------------ 475
Query: 537 VIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSL-QLYFLLGIVYAVVTPILLPFILVF 595
LK++++ G + P + + +T S+ Q Y LL ++ A++ IL+ + F
Sbjct: 476 ----LKDIYLGVGTDFVGTHILPNNFSYFDTEFSITQFYKLLPLISAILVSILIVVLNEF 531
Query: 596 FALAYLV---YRHQIINVYNQQYES 617
FA+ + + Y + + +++NQ+ S
Sbjct: 532 FAIVFNLNNKYINTVYSIFNQKLVS 556
>BRM_DROME (P25439) Homeotic gene regulator (Brahma protein)
Length = 1638
Score = 37.4 bits (85), Expect = 0.16
Identities = 21/61 (34%), Positives = 27/61 (43%), Gaps = 7/61 (11%)
Query: 738 ASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPH---YAYQYQDE 794
A+ + P +PSP+ P+P H S YP PG PPPP H Y + Q
Sbjct: 7 ANSPMPPPQAPSPMAPPSQSPAPSPH----SPYPHQQPGPLQGPPPPGHPGAYGHPMQHG 62
Query: 795 P 795
P
Sbjct: 63 P 63
>GLI1_HUMAN (P08151) Zinc finger protein GLI1 (Glioma-associated
oncogene) (Oncogene GLI)
Length = 1106
Score = 37.0 bits (84), Expect = 0.20
Identities = 18/40 (45%), Positives = 20/40 (50%), Gaps = 1/40 (2%)
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP 784
P P PL H QPSPPQ++ Q Y PP Y P P
Sbjct: 841 PPHPQPLFSHYPQPSPPQYL-QSGPYTQPPPDYLPSEPRP 879
>FTSK_ECO57 (Q8X5H9) DNA translocase ftsK
Length = 1342
Score = 37.0 bits (84), Expect = 0.20
Identities = 28/105 (26%), Positives = 42/105 (39%), Gaps = 11/105 (10%)
Query: 702 IKAYLADAYLHPIFRSF-----EVEEELVEVRVDKQPETQVA-SPTLSEPS---SPSPLH 752
+KA L D P+F + ++ + + +QP+ VA P +P +P P +
Sbjct: 719 MKALLDDGPHEPLFTPIVEPVQQPQQPVAPQQQYQQPQQPVAPQPQYQQPQQQVAPQPQY 778
Query: 753 DHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP--PHYAYQYQDEP 795
QP PQ Q P +P Y P P P YQ +P
Sbjct: 779 QQPQQPVAPQQQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQPQQP 823
>EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein)
Length = 620
Score = 36.6 bits (83), Expect = 0.27
Identities = 28/77 (36%), Positives = 30/77 (38%), Gaps = 18/77 (23%)
Query: 733 PETQVASPTLSEPSS-----------PSPLHDHV---HQPSPPQHVNQYSHYPTSPPGYY 778
P T V PT S PS P H H HQPSP +H+ PP Y
Sbjct: 208 PPTHV-QPTPSPPSRGHQPQPPTHRHAPPTHRHAPPTHQPSPLRHLPPSPRRQPQPPTY- 265
Query: 779 YHPPPPPHYAYQYQDEP 795
PPPP YA Q P
Sbjct: 266 --SPPPPAYAQSPQPSP 280
Score = 36.2 bits (82), Expect = 0.35
Identities = 20/56 (35%), Positives = 22/56 (38%), Gaps = 2/56 (3%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P T P L SP P P PP + Y PP Y PPPPP Y+
Sbjct: 420 PPTYAQPPPLPPTYSPPP--PAYSPPPPPTYSPPPPTYSPPPPAYAQPPPPPPTYS 473
Score = 36.2 bits (82), Expect = 0.35
Identities = 21/53 (39%), Positives = 26/53 (48%), Gaps = 7/53 (13%)
Query: 739 SPTLSEPS---SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
+PT S P SP P + P PP ++ S SPP Y PPPPP Y+
Sbjct: 314 TPTFSPPPPAYSPPP----TYSPPPPTYLPLPSSPIYSPPPPVYSPPPPPSYS 362
Score = 35.8 bits (81), Expect = 0.46
Identities = 24/76 (31%), Positives = 31/76 (40%), Gaps = 22/76 (28%)
Query: 733 PETQVASPTLSEPS--SPSPLHDHVHQPS--------------------PPQHVNQYSHY 770
P + V SP+ PS +P P H H PS PP+ + SH
Sbjct: 108 PPSPVISPSHPPPSYGAPPPSHGPGHLPSHGQRPPSPSHGHAPPSGGHTPPRGQHPPSHR 167
Query: 771 PTSPPGYYYHPPPPPH 786
SPP + HPPPP +
Sbjct: 168 RPSPPSRHGHPPPPTY 183
Score = 35.0 bits (79), Expect = 0.78
Identities = 23/64 (35%), Positives = 29/64 (44%), Gaps = 12/64 (18%)
Query: 732 QPETQVASPTLSEPSSP----SPLHDHVHQPSPPQHVNQ-----YSHYPTSPPGYYYHPP 782
QP T +PT +P SP +P +H P PP + Y P SPP Y P
Sbjct: 553 QPRTP--TPTYGQPPSPPTFSAPPPRQIHSPPPPHRQPRPPTPTYGQ-PPSPPTTYSPPS 609
Query: 783 PPPH 786
PPP+
Sbjct: 610 PPPY 613
Score = 33.9 bits (76), Expect = 1.7
Identities = 22/63 (34%), Positives = 25/63 (38%), Gaps = 11/63 (17%)
Query: 733 PETQVASPTLSEPSS-----PSPLHDHVHQPSPPQHVNQYSHYPTS---PPGYYYHPPPP 784
P+ Q PT S P P P H QP PP P + PP H PPP
Sbjct: 493 PQVQPLPPTFSPPPPRRIHLPPPPH---RQPRPPTPTYGQPPSPPTFSPPPPRQIHSPPP 549
Query: 785 PHY 787
PH+
Sbjct: 550 PHW 552
Score = 33.9 bits (76), Expect = 1.7
Identities = 20/54 (37%), Positives = 23/54 (42%), Gaps = 5/54 (9%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPH 786
P T P P PSP ++ P PPQ V + PP H PPPPH
Sbjct: 469 PPTYSPPPPAYSPPPPSP----IYSPPPPQ-VQPLPPTFSPPPPRRIHLPPPPH 517
Score = 33.5 bits (75), Expect = 2.3
Identities = 21/63 (33%), Positives = 26/63 (40%), Gaps = 16/63 (25%)
Query: 739 SPTLSEPSSPS----PLHDHVHQPSPPQHVNQYSHYPT-----------SPPGYYYHPPP 783
+PT +P SP P +H P PP H + PT +PP H PP
Sbjct: 524 TPTYGQPPSPPTFSPPPPRQIHSP-PPPHWQPRTPTPTYGQPPSPPTFSAPPPRQIHSPP 582
Query: 784 PPH 786
PPH
Sbjct: 583 PPH 585
Score = 32.7 bits (73), Expect = 3.9
Identities = 19/56 (33%), Positives = 22/56 (38%), Gaps = 3/56 (5%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P V SP SP P + P PP + PP Y PPPPP Y+
Sbjct: 348 PPPPVYSPPPPPSYSPPP---PTYLPPPPPSSPPPPSFSPPPPTYEQSPPPPPAYS 400
Score = 32.3 bits (72), Expect = 5.0
Identities = 18/43 (41%), Positives = 18/43 (41%), Gaps = 4/43 (9%)
Query: 748 PSPLHDHVHQPSP----PQHVNQYSHYPTSPPGYYYHPPPPPH 786
PSP H H P P P YS P P Y PPPP H
Sbjct: 169 PSPPSRHGHPPPPTYAQPPPTPIYSPSPQVQPPPTYSPPPPTH 211
Score = 32.0 bits (71), Expect = 6.6
Identities = 20/59 (33%), Positives = 24/59 (39%), Gaps = 11/59 (18%)
Query: 748 PSPLHDHV-----HQPSPPQHVNQY------SHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
PSP H H H P QH + S + PP Y PPP P Y+ Q +P
Sbjct: 142 PSPSHGHAPPSGGHTPPRGQHPPSHRRPSPPSRHGHPPPPTYAQPPPTPIYSPSPQVQP 200
Score = 31.6 bits (70), Expect = 8.6
Identities = 22/54 (40%), Positives = 28/54 (51%), Gaps = 16/54 (29%)
Query: 742 LSEPSSPS---PLHDHVHQP--SPPQHVNQYSHYPTSPPGYYYHPP----PPPH 786
LS PS+P+ P HV P +PP+H YP PP + + PP PPPH
Sbjct: 49 LSPPSAPTTTPPSRGHVPSPRHAPPRHA-----YP--PPSHGHLPPSVGGPPPH 95
>CBPY_SCHPO (O13849) Carboxypeptidase Y precursor (EC 3.4.16.5)
(CPY)
Length = 1002
Score = 36.6 bits (83), Expect = 0.27
Identities = 24/87 (27%), Positives = 33/87 (37%), Gaps = 10/87 (11%)
Query: 719 EVEEELVEVRVDKQPETQVASPTL----SEPSSPSPLHDHV--HQPSPPQHVNQYSHYPT 772
E ++E E +P + P + E P P+H H P PP H H P
Sbjct: 189 EFDDEDREFPAHHEPGEHMPPPPMHHKPGEHMPPPPMHHEPGEHMPPPPMHHEPGEHMPP 248
Query: 773 SP----PGYYYHPPPPPHYAYQYQDEP 795
P PG + PPP H ++ P
Sbjct: 249 PPMHHEPGEHMPPPPMHHEPGEHMPPP 275
Score = 36.2 bits (82), Expect = 0.35
Identities = 19/58 (32%), Positives = 24/58 (40%), Gaps = 6/58 (10%)
Query: 744 EPSSPSPLHDHV--HQPSPPQHVNQYSHYPTSP----PGYYYHPPPPPHYAYQYQDEP 795
E P P+H H P PP H H P P PG + PPP H+ + + P
Sbjct: 296 EHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPPPFKHHELEEHEGP 353
Score = 35.0 bits (79), Expect = 0.78
Identities = 19/58 (32%), Positives = 23/58 (38%), Gaps = 6/58 (10%)
Query: 744 EPSSPSPLHDHV--HQPSPPQHVNQYSHYPTSP----PGYYYHPPPPPHYAYQYQDEP 795
E P P+H H P PP H H P P PG + PPP H ++ P
Sbjct: 270 EHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPP 327
Score = 35.0 bits (79), Expect = 0.78
Identities = 19/58 (32%), Positives = 23/58 (38%), Gaps = 6/58 (10%)
Query: 744 EPSSPSPLHDHV--HQPSPPQHVNQYSHYPTSP----PGYYYHPPPPPHYAYQYQDEP 795
E P P+H H P PP H H P P PG + PPP H ++ P
Sbjct: 244 EHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPPPMHHEPGEHMPPP 301
>PRF1_LYCES (Q00451) 36.4 kDa proline-rich protein
Length = 346
Score = 36.2 bits (82), Expect = 0.35
Identities = 20/59 (33%), Positives = 26/59 (43%), Gaps = 4/59 (6%)
Query: 731 KQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPP---PPH 786
K P T P + P++P HV PS P+H P PP ++ PP PPH
Sbjct: 52 KPPSTPKQPPYVKPPTTPKH-PPHVKPPSTPKHPKHPPQKPCPPPSHHGPKPPIVKPPH 109
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 92,444,375
Number of Sequences: 164201
Number of extensions: 4037901
Number of successful extensions: 17794
Number of sequences better than 10.0: 108
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 81
Number of HSP's that attempted gapping in prelim test: 16649
Number of HSP's gapped (non-prelim): 591
length of query: 795
length of database: 59,974,054
effective HSP length: 118
effective length of query: 677
effective length of database: 40,598,336
effective search space: 27485073472
effective search space used: 27485073472
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0020.6


