

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.4
(310 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
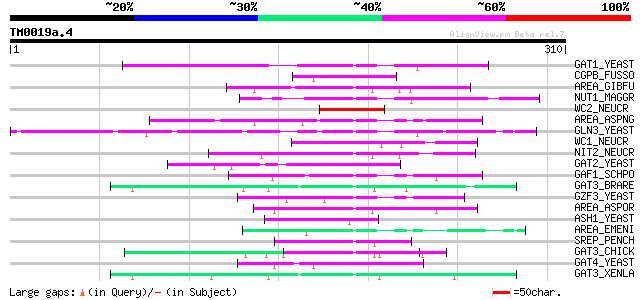
Score E
Sequences producing significant alignments: (bits) Value
GAT1_YEAST (P43574) Transcriptional regulatory protein GAT1 59 2e-08
CGPB_FUSSO (Q00858) Cutinase gene palindrome-binding protein (PBP) 59 2e-08
AREA_GIBFU (P78688) Nitrogen regulatory protein areA 57 6e-08
NUT1_MAGGR (Q01168) Nitrogen regulatory protein NUT1 57 8e-08
WC2_NEUCR (P78714) White collar 2 protein (WC2) 56 1e-07
AREA_ASPNG (O13412) Nitrogen regulatory protein areA 55 2e-07
GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3 54 4e-07
WC1_NEUCR (Q01371) White collar 1 protein (WC1) 54 5e-07
NIT2_NEUCR (P19212) Nitrogen catabolic enzyme regulatory protein 54 5e-07
GAT2_YEAST (P40209) GAT2 protein 54 6e-07
GAF1_SCHPO (Q10280) Transcription factor gaf1 (Gaf-1) 53 1e-06
GAT3_BRARE (Q91428) Transcription factor GATA-3 (GATA binding fa... 52 1e-06
GZF3_YEAST (P42944) GZF3 protein 51 3e-06
AREA_ASPOR (O13415) Nitrogen regulatory protein areA 51 3e-06
ASH1_YEAST (P34233) Daughter cells HO repressor protein ASH1 50 7e-06
AREA_EMENI (P17429) Nitrogen regulatory protein areA 50 9e-06
SREP_PENCH (Q92259) GATA factor SREP 49 1e-05
GAT3_CHICK (P23825) GATA binding factor-3 (Transcription factor ... 49 2e-05
GAT4_YEAST (P40569) GAT4 protein 49 2e-05
GAT3_XENLA (P23773) Transcription factor xGATA-3 (GATA binding f... 49 2e-05
>GAT1_YEAST (P43574) Transcriptional regulatory protein GAT1
Length = 510
Score = 58.9 bits (141), Expect = 2e-08
Identities = 49/213 (23%), Positives = 93/213 (43%), Gaps = 34/213 (15%)
Query: 64 SSSSSSSNHLYNSTFISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRL 123
+++S ++NH +NS+ + P + N +++ + N S S + + S + R
Sbjct: 213 NNNSINNNHSHNSSHNNNSPSIANNTNANTNTNTSASTNTNSPLLRRNPSPSIVKPGSRR 272
Query: 124 MKKMMST-PATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSS 182
+ PA K +ST++ S + +N SSN CS+C TS+
Sbjct: 273 NSSVRKKKPALKKIKSSTSVQSS---------------ATPPSNTSSNPDIKCSNCTTST 317
Query: 183 TPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTAS--------TKTRVHHN 234
TPLWR P G LCNACG+ + +G+ P++ + + T++++N
Sbjct: 318 TPLWRKDPKG-LPLCNACGLFLK---------LHGVTRPLSLKTDIIKKRQRSSTKINNN 367
Query: 235 KEKKPRANHFAQFKNKSKSTSSNAGSSQETKKL 267
P ++ K K+ +++ +S+ L
Sbjct: 368 ITPPPSSSLNPGAAGKKKNYTASVAASKRKNSL 400
Score = 30.0 bits (66), Expect = 7.6
Identities = 27/96 (28%), Positives = 43/96 (44%), Gaps = 16/96 (16%)
Query: 146 PRIQNQGHENRYSQRSPRNNNNSS--NTTRVCSDCNTS-----STPLWRSGPN----GPK 194
P + N N +S S NNN+ S N T ++ NTS ++PL R P+ P
Sbjct: 210 PFLNNNSINNNHSHNSSHNNNSPSIANNTNANTNTNTSASTNTNSPLLRRNPSPSIVKPG 269
Query: 195 SLCNACGIRQRKAR----RAMAEAANGLATPINTAS 226
S N+ +R++K ++ + P NT+S
Sbjct: 270 SRRNS-SVRKKKPALKKIKSSTSVQSSATPPSNTSS 304
>CGPB_FUSSO (Q00858) Cutinase gene palindrome-binding protein (PBP)
Length = 457
Score = 58.5 bits (140), Expect = 2e-08
Identities = 25/62 (40%), Positives = 36/62 (57%), Gaps = 4/62 (6%)
Query: 159 QRSPRNNNNS----SNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEA 214
+R PR + ++ VC+DC T +P WR GP+GPK+LCNACG+R K + +
Sbjct: 382 ERDPRTGDKKKKLKTSEEYVCTDCGTLDSPEWRKGPSGPKTLCNACGLRWAKKEKKRNSS 441
Query: 215 AN 216
N
Sbjct: 442 VN 443
>AREA_GIBFU (P78688) Nitrogen regulatory protein areA
Length = 971
Score = 57.0 bits (136), Expect = 6e-08
Identities = 45/157 (28%), Positives = 67/157 (42%), Gaps = 24/157 (15%)
Query: 122 RLMKKMMSTPATDK-------ANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRV 174
+L + M S+PA D ++ + + P SP + QG ++ N N N
Sbjct: 636 QLAQSMQSSPAGDGNGTMSGFSSVAPSRPSSPPMSKQGSTTNL--QAAAGNGNDGNAPTT 693
Query: 175 CSDCNTSSTPLWRSGPNGPKSLCNACG--------IRQRKARRAMAEAAN---GLATPI- 222
C++C T +TPLWR P G + LCNACG +R + + + N G P+
Sbjct: 694 CTNCFTQTTPLWRRNPEG-QPLCNACGLFLKLHGVVRPLSLKTDVIKKRNRGSGTNVPVG 752
Query: 223 --NTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSN 257
+T S KT N K + N +K SSN
Sbjct: 753 GSSTRSKKTASTLNSRKNSTLSMSTATANSTKPNSSN 789
>NUT1_MAGGR (Q01168) Nitrogen regulatory protein NUT1
Length = 956
Score = 56.6 bits (135), Expect = 8e-08
Identities = 48/170 (28%), Positives = 74/170 (43%), Gaps = 35/170 (20%)
Query: 129 STPATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRS 188
S+P N STT +Q QG NN + C++C T +TPLWR
Sbjct: 634 SSPPPGSKNGSTT-----NLQQQG------------NNQGGDAPTTCTNCATQTTPLWRR 676
Query: 189 GPNGPKSLCNACGIRQRKARRAMAEAANGLATPIN--TASTKTRVHHNKEKKPRANHFAQ 246
P G + LCNACG+ + +G+ P++ T K R + P A ++
Sbjct: 677 NPEG-QPLCNACGLFLK---------LHGVVRPLSLKTDVIKKRNRGSGSNVPGATSGSR 726
Query: 247 FKNKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFPRDEVAEAA 296
K + ST+ + ++++ L AIS ++T QV P +A AA
Sbjct: 727 SKKGATSTAVSGTNTRKNSSL------AISRTASTTNVQVAPTPAIAPAA 770
>WC2_NEUCR (P78714) White collar 2 protein (WC2)
Length = 530
Score = 56.2 bits (134), Expect = 1e-07
Identities = 21/36 (58%), Positives = 27/36 (74%)
Query: 174 VCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARR 209
VC+DC T +P WR GP+GPK+LCNACG+R K +
Sbjct: 467 VCTDCGTLDSPEWRKGPSGPKTLCNACGLRWAKKEK 502
>AREA_ASPNG (O13412) Nitrogen regulatory protein areA
Length = 882
Score = 55.5 bits (132), Expect = 2e-07
Identities = 54/196 (27%), Positives = 81/196 (40%), Gaps = 27/196 (13%)
Query: 79 ISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDK--- 135
I P +MD PS +L + + V + + + + + STP T +
Sbjct: 573 IGPTDMMDTPSEWAQSGSLGRTHGSAASVSEVRNREQ--DPRRQKIARTTSTPNTAQLLR 630
Query: 136 ----ANNSTTIPISPRIQNQGHENRYSQRSP---RNNNNSSNTTRVCSDCNTSSTPLWRS 188
AN S T P +P SP +N SS T C++C T +TPLWR
Sbjct: 631 QSMNANTSHTSPNTPPESGLSSAVPSRPASPGGSKNGEQSSGPT-TCTNCFTQTTPLWRR 689
Query: 189 GPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFK 248
P G + LCNACG+ + +G+ P+ S KT V K + AN A
Sbjct: 690 NPEG-QPLCNACGLFLK---------LHGVVRPL---SLKTDV-IKKRNRSSANSLAVGT 735
Query: 249 NKSKSTSSNAGSSQET 264
+++ S+ S Q+T
Sbjct: 736 SRASKKSARKNSVQQT 751
>GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3
Length = 730
Score = 54.3 bits (129), Expect = 4e-07
Identities = 71/303 (23%), Positives = 124/303 (40%), Gaps = 43/303 (14%)
Query: 1 MTPVSLNPPGPSIQGQNHLFNSLNNQDYHASLFNILDRRQGIGIGELRENDHQDDKLVVW 60
MTP N G + + F S Q H SL N + + N + + +
Sbjct: 144 MTPS--NSSGSATISAPNSFTSDIPQYNHGSLGNSVSKSSLFPYNSSTSNSNINQPSI-- 199
Query: 61 HDGSSSSSSSNHLYN-------STFISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHGS 113
++ S++++ S+H +N ++ S + +N +S+ + N+ +++ D + S
Sbjct: 200 NNNSNTNAQSHHSFNIYKLQNNNSSSSAMNITNNNNSN--NSNIQHPFLKKSDSIGLSSS 257
Query: 114 AKWMSSKMRLMKKMMSTPATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTR 173
S + + K MS+ T AN +S I N P N
Sbjct: 258 NTTNSVRKNSLIKPMSS--TSLANFKRAASVSSSISNM---------EPSGQNKKPLIQ- 305
Query: 174 VCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTAS--TKTRV 231
C +C T TPLWR P G +LCNACG+ Q+ +G P++ S K R+
Sbjct: 306 -CFNCKTFKTPLWRRSPEG-NTLCNACGLFQK---------LHGTMRPLSLKSDVIKKRI 354
Query: 232 HHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFPRDE 291
+ K+ N AQ + +T+S + ++ K + K SL+ NS +V P +
Sbjct: 355 SKKRAKQTDPN-IAQNTPSAPATASTSVTTTNAKPIRSRKK---SLQQNS-LSRVIPEEI 409
Query: 292 VAE 294
+ +
Sbjct: 410 IRD 412
>WC1_NEUCR (Q01371) White collar 1 protein (WC1)
Length = 1167
Score = 53.9 bits (128), Expect = 5e-07
Identities = 31/111 (27%), Positives = 54/111 (47%), Gaps = 12/111 (10%)
Query: 158 SQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRK-----ARRAMA 212
+++ + N R C++C+T +TP WR GP+G + LCN+CG+R K + R +
Sbjct: 917 NKKKRKRRKGGGNMVRDCANCHTRNTPEWRRGPSGNRDLCNSCGLRWAKQTGRVSPRTSS 976
Query: 213 EAANG--LATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSS 261
NG ++ N+ S + +H + N +K++ S GSS
Sbjct: 977 RGGNGDSMSKKSNSPSHSSPLH-----REVGNDSPSTTTATKNSPSLRGSS 1022
>NIT2_NEUCR (P19212) Nitrogen catabolic enzyme regulatory protein
Length = 1036
Score = 53.9 bits (128), Expect = 5e-07
Identities = 45/163 (27%), Positives = 74/163 (44%), Gaps = 22/163 (13%)
Query: 112 GSAKWMSSKMRLMKKMMSTPATDKANNS--TTIPIS-PRIQNQGHENRYSQRSPRNNNNS 168
G + M+ RL + +P D +S +++P S P G + + NS
Sbjct: 677 GLSARMNPFERLAQSASHSPPADVGRSSGLSSVPASRPSSPPPGAKQGSTTNLQGAAGNS 736
Query: 169 SNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACG--------IRQRKARRAMAEAAN---G 217
++T C++C T +TPLWR P+G + LCNACG +R + + + N G
Sbjct: 737 TDTPTTCTNCFTQTTPLWRRNPDG-QPLCNACGLFLKLHGVVRPLSLKTDVIKKRNRGSG 795
Query: 218 LATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGS 260
+ P+ ST++ KK + A KN + S +SNA +
Sbjct: 796 ASLPVGGTSTRS-------KKNASMSAAARKNSTLSITSNANN 831
>GAT2_YEAST (P40209) GAT2 protein
Length = 560
Score = 53.5 bits (127), Expect = 6e-07
Identities = 37/136 (27%), Positives = 61/136 (44%), Gaps = 12/136 (8%)
Query: 89 SSSTCDPNLSFSKMEEEDIKNVHGS---AKWMSSKMR---LMKKMMSTPATDKANNSTTI 142
+S T PN + S M + +KN+ S K +++ R + ++ T ++ NNS
Sbjct: 386 NSITYIPNTTLSPMVQTQLKNLTTSNLNTKKKNNRGRPRAIQRQPTLTTSSHFINNSN-- 443
Query: 143 PISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGI 202
P + + + S N N+ C C + TP WR GP G ++LCNACG+
Sbjct: 444 PGAAAVST----TTPAANSDEKNPNAKKIIEFCFHCGETETPEWRKGPYGTRTLCNACGL 499
Query: 203 RQRKARRAMAEAANGL 218
RK + ++ L
Sbjct: 500 FYRKVTKKFGSKSSNL 515
>GAF1_SCHPO (Q10280) Transcription factor gaf1 (Gaf-1)
Length = 855
Score = 52.8 bits (125), Expect = 1e-06
Identities = 42/161 (26%), Positives = 69/161 (42%), Gaps = 34/161 (21%)
Query: 123 LMKKMMSTPATDKANNSTTIPIS---------------PRIQNQGHENRYSQRSPRNNNN 167
L K+ +T AT +A TT+ P + N + + + R N
Sbjct: 570 LKKRTTNTAATPQAALPTTLDTKKDRSVSFNINKNAEKPTVSNAAEDKKGDANTRRAN-- 627
Query: 168 SSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTAST 227
++N T C++C T +TPLWR P+G + LCNACG+ + NG+ P+ S
Sbjct: 628 ATNPTPTCTNCQTRTTPLWRRSPDG-QPLCNACGLFMK---------INGVVRPL---SL 674
Query: 228 KTRVHHNKEK----KPRANHFAQFKNKSKSTSSNAGSSQET 264
KT V + + K ++ +SS + S++ T
Sbjct: 675 KTDVIKKRNRGVGTSATPKQSGGRKGSTRKSSSKSSSAKST 715
>GAT3_BRARE (Q91428) Transcription factor GATA-3 (GATA binding
factor-3)
Length = 438
Score = 52.4 bits (124), Expect = 1e-06
Identities = 71/273 (26%), Positives = 102/273 (37%), Gaps = 52/273 (19%)
Query: 57 LVVWHDGSSSS----SSSNHLYNSTFISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHG 112
L V+ SSSS SS HL+ P+ V +P+ ST S + ++E IK
Sbjct: 123 LSVYPPASSSSLSAGHSSPHLFTFPPTPPKDVSPDPAISTSGSGSSVRQEDKECIKYQVS 182
Query: 113 SAKWMSSKMRLMKKMMS--TPATDKANNSTTIP----------ISPRIQNQGHENRYSQR 160
A+ M + M S A+ + T P P G + Y +
Sbjct: 183 LAESMKLDSAHSRSMASIGAGASSAHHPIATYPSYVPDYGPGLFPPSSLIGGSSSSYGSK 242
Query: 161 SPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGI----------------RQ 204
+ R SS+ R C +C +STPLWR G LCNACG+ R
Sbjct: 243 T-RPKTRSSSEGRECVNCGATSTPLWRRDGTG-HYLCNACGLYHKMNGQNRPLIKPKRRL 300
Query: 205 RKARRAMAEAANGLAT------------PI-NTASTKTRVHHNKEKKPRANHFAQFKNKS 251
ARRA AN T P+ N ++H+ Q +N+
Sbjct: 301 SAARRAGTSCANCQTTTTTLWRRNANGDPVCNACGLYYKLHNINRPLTMKKEGIQTRNRK 360
Query: 252 KSTSSNAGSSQETKKLECFKDFAISL-RNNSTF 283
S+ S + K + +DF+ SL NS+F
Sbjct: 361 MSSK----SKKSKKSHDSMEDFSKSLMEKNSSF 389
>GZF3_YEAST (P42944) GZF3 protein
Length = 551
Score = 51.2 bits (121), Expect = 3e-06
Identities = 44/137 (32%), Positives = 59/137 (42%), Gaps = 23/137 (16%)
Query: 128 MSTPATDKA-NNSTTIPISPRIQNQGH-------ENRYSQRSPRNNNNSSNTTRV--CSD 177
M+ A DK+ + ST +SP + H +N NNN SN V C +
Sbjct: 74 MNFKANDKSFSTSTAGRMSPDTNSLHHILPKNQVKNNGQTMDANCNNNVSNDANVPVCKN 133
Query: 178 CNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEK 237
C TS+TPLWR +G LCNACG+ + +G PI S KT V ++ +
Sbjct: 134 CLTSTTPLWRRDEHG-AMLCNACGLFLK---------LHGKPRPI---SLKTDVIKSRNR 180
Query: 238 KPRANHFAQFKNKSKST 254
K NH N T
Sbjct: 181 KSNTNHAHNLDNFRNQT 197
>AREA_ASPOR (O13415) Nitrogen regulatory protein areA
Length = 866
Score = 51.2 bits (121), Expect = 3e-06
Identities = 40/148 (27%), Positives = 63/148 (42%), Gaps = 24/148 (16%)
Query: 137 NNSTTIPISP---RIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGP 193
N S T P +P + + S +N + SN C++C T +TPLWR P G
Sbjct: 623 NTSHTSPNTPPESALSSAVPSRPASPGGSKNGDQGSNGPTTCTNCFTQTTPLWRRNPEG- 681
Query: 194 KSLCNACG-------------IRQRKARRAMAEAANGLATPINTASTKTRVHHNKEK--- 237
+ LCNACG ++ ++ +AN LA + AS KT ++ ++
Sbjct: 682 QPLCNACGLFLKLHGVVRPLSLKTDVIKKRNRSSANSLAVGTSRASKKTARKNSVQQASV 741
Query: 238 ----KPRANHFAQFKNKSKSTSSNAGSS 261
RA + F++ S+ AG S
Sbjct: 742 TTPTSSRAQNGTSFESPPAGFSAAAGRS 769
>ASH1_YEAST (P34233) Daughter cells HO repressor protein ASH1
Length = 588
Score = 50.1 bits (118), Expect = 7e-06
Identities = 23/66 (34%), Positives = 35/66 (52%), Gaps = 2/66 (3%)
Query: 143 PISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRS--GPNGPKSLCNAC 200
P SPR + + R ++ + +TTRVC C++S +P WR P LCN+C
Sbjct: 467 PQSPRRSSNSSITKKGSRRSSGSSPTRHTTRVCVSCHSSDSPCWRPSWSPRKQDQLCNSC 526
Query: 201 GIRQRK 206
G+R +K
Sbjct: 527 GLRYKK 532
>AREA_EMENI (P17429) Nitrogen regulatory protein areA
Length = 876
Score = 49.7 bits (117), Expect = 9e-06
Identities = 47/178 (26%), Positives = 71/178 (39%), Gaps = 56/178 (31%)
Query: 131 PATDKANNSTTIPISPRIQNQGHENRYSQRSPRN--------------------NNNSSN 170
P K +++ P + ++ Q +N+ S SP N N
Sbjct: 609 PRRQKIARTSSTPNTAQLLRQSMQNQSSHTSPNTPPESGLNSAAPSRPASPGGTKNGEQN 668
Query: 171 TTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTR 230
C++C T +TPLWR P G + LCNACG+ + +G+ P+ S KT
Sbjct: 669 GPTTCTNCFTQTTPLWRRNPEG-QPLCNACGLFLK---------LHGVVRPL---SLKTD 715
Query: 231 VHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFP 288
V + +N++ + S GSS+ +KK S R NS QQV P
Sbjct: 716 V-------------IKKRNRNSANSLAVGSSRVSKK---------SARKNSV-QQVTP 750
>SREP_PENCH (Q92259) GATA factor SREP
Length = 532
Score = 49.3 bits (116), Expect = 1e-05
Identities = 29/85 (34%), Positives = 41/85 (48%), Gaps = 10/85 (11%)
Query: 149 QNQGHENRYSQR-SPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGI----- 202
Q GH S++ SP + +++ CS+C T STPLWR P G +CNACG+
Sbjct: 67 QWNGHNETPSEKTSPSSQKDTTFLGHSCSNCGTKSTPLWRRSPTG-AMICNACGLYLKAR 125
Query: 203 ---RQRKARRAMAEAANGLATPINT 224
R K R +E A P ++
Sbjct: 126 NVARPTKRNRMQSEGAEKPPPPASS 150
Score = 41.2 bits (95), Expect = 0.003
Identities = 17/36 (47%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Query: 167 NSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGI 202
++SNT C +C T+ TPLWR G +CNACG+
Sbjct: 230 DASNTLVACQNCGTTVTPLWRRDEQG-HPICNACGL 264
>GAT3_CHICK (P23825) GATA binding factor-3 (Transcription factor
NF-E1C) (GATA-3)
Length = 444
Score = 48.9 bits (115), Expect = 2e-05
Identities = 52/187 (27%), Positives = 76/187 (39%), Gaps = 24/187 (12%)
Query: 65 SSSSSSNHLYNSTFISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRLM 124
S+ SS HL+ P+ V +PS ST S + E+E IK A M +
Sbjct: 142 SAGHSSPHLFTFPPTPPKDVSPDPSISTPGSTGSTRQDEKECIKYQVSLADTMKLESSHS 201
Query: 125 KKMMST--PATDKANNSTTI--PISPRIQNQ---------GHENRYSQRSPRNNNNSSNT 171
+ M++ AT A++ T P P + G + +S R SS
Sbjct: 202 RSSMASLGGATSSAHHPITTYPPYVPEYSSGLFPPSSLLGGSPTGFGCKS-RPKARSSTE 260
Query: 172 TRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQR---------KARRAMAEAANGLATPI 222
R C +C +STPLWR G LCNACG+ + K +R ++ A +
Sbjct: 261 GRECVNCGATSTPLWRRDGTG-HYLCNACGLYHKMNGQNRPLIKPKRRLSAARRAGTSCA 319
Query: 223 NTASTKT 229
N +T T
Sbjct: 320 NCQTTTT 326
Score = 46.2 bits (108), Expect = 1e-04
Identities = 28/97 (28%), Positives = 47/97 (47%), Gaps = 7/97 (7%)
Query: 154 ENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGI--RQRKARRAM 211
+NR + R + + C++C T++T LWR NG +CNACG+ + R +
Sbjct: 297 QNRPLIKPKRRLSAARRAGTSCANCQTTTTTLWRRNANG-DPVCNACGLYYKLHNINRPL 355
Query: 212 AEAANGLATPINTASTKT----RVHHNKEKKPRANHF 244
G+ T S+K+ +VH N E P+++ F
Sbjct: 356 TMKKEGIQTRNRKMSSKSKKCKKVHDNLEDFPKSSSF 392
>GAT4_YEAST (P40569) GAT4 protein
Length = 121
Score = 48.5 bits (114), Expect = 2e-05
Identities = 39/113 (34%), Positives = 48/113 (41%), Gaps = 11/113 (9%)
Query: 128 MSTPATDKANNSTTIPISP--RIQNQGHENRYSQRSPRNNNNS-----SNTTRVCSDCNT 180
MST +N T P + N G E ++ PRN ++ + TR C C
Sbjct: 1 MSTKLPIVISNGTAFKKVPVQLLLNSGSEAQHGL--PRNADSQPARPRTGITRTCGQCGE 58
Query: 181 SSTPL-WRSGPNGPKSLCNACGIRQRK-ARRAMAEAANGLATPINTASTKTRV 231
T L WR GPNG LCNACG+ RK R AA I TK R+
Sbjct: 59 IKTSLQWREGPNGAACLCNACGLFFRKLILRFGRAAAKRYMEQIKGTGTKRRI 111
>GAT3_XENLA (P23773) Transcription factor xGATA-3 (GATA binding
factor-3)
Length = 435
Score = 48.5 bits (114), Expect = 2e-05
Identities = 63/263 (23%), Positives = 93/263 (34%), Gaps = 38/263 (14%)
Query: 57 LVVWHDGSSSS----SSSNHLYNSTFISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVH- 111
L V+ SS+S SS HL+ P+ V +PS ST S E+E + + +
Sbjct: 124 LSVYPPASSASLATGHSSPHLFTFPPTPPKDVSPDPSISTSGSTSSSRHEEKECVVSKYQ 183
Query: 112 ---GSAKWMSSKMRLMKKMMSTPATDKANNSTTIP-------ISPRIQNQGHENRYSQRS 161
G + M + TT P P G +S +
Sbjct: 184 VSLGDTGMKLESLHPRNSMSGIGGVSAHHPITTYPDYYGAGLFQPGSILDGSPTHFSSNA 243
Query: 162 PRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQR---------KARRAMA 212
R SS R C +C +STPLWR G LCNACG+ + K +R ++
Sbjct: 244 -RPKTRSSTEGRECVNCGATSTPLWRRDGTG-HYLCNACGLYHKMNGQNRPLIKPKRRLS 301
Query: 213 EAANGLATPINTASTKTRV-HHNKEKKPRANHFAQF-----------KNKSKSTSSNAGS 260
A + N +T T + N P N + K + N
Sbjct: 302 AARRAGTSCANCQTTTTTLWRRNANGDPVCNACGLYYKLHNINRPLTMKKEGIQTRNRKM 361
Query: 261 SQETKKLECFKDFAISLRNNSTF 283
S ++KK + F D +S+F
Sbjct: 362 SSKSKKSKKFHDSLEDYPKSSSF 384
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.125 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,278,191
Number of Sequences: 164201
Number of extensions: 1504377
Number of successful extensions: 4822
Number of sequences better than 10.0: 172
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 120
Number of HSP's that attempted gapping in prelim test: 4306
Number of HSP's gapped (non-prelim): 343
length of query: 310
length of database: 59,974,054
effective HSP length: 110
effective length of query: 200
effective length of database: 41,911,944
effective search space: 8382388800
effective search space used: 8382388800
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0019a.4


