

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0016.15
(1469 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
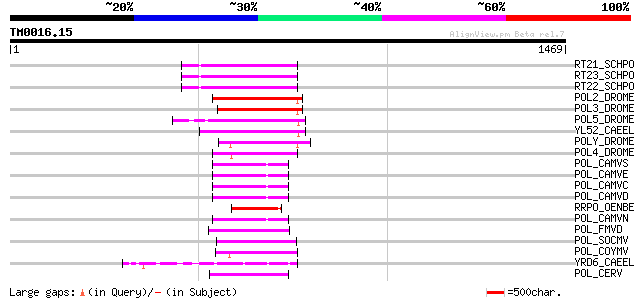
Score E
Sequences producing significant alignments: (bits) Value
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 197 2e-49
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 194 1e-48
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 194 1e-48
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 191 2e-47
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 185 7e-46
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 169 7e-41
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 163 4e-39
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 159 5e-38
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 148 1e-34
POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic pro... 146 4e-34
POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic pro... 146 5e-34
POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic pro... 146 5e-34
POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic pro... 144 2e-33
RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse... 142 9e-33
POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic pro... 140 2e-32
POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic prot... 134 2e-30
POL_SOCMV (P15629) Enzymatic polyprotein [Contains: Aspartic pro... 132 7e-30
POL_COYMV (P19199) Putative polyprotein [Contains: Coat protein;... 132 9e-30
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 125 7e-28
POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic prot... 117 2e-25
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 197 bits (501), Expect = 2e-49
Identities = 113/309 (36%), Positives = 176/309 (56%), Gaps = 5/309 (1%)
Query: 455 KRVVFPDSVLSEYLFANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVVKEFTDVFPG- 513
K+ P ++ L+ N+I +S + + +S K PE+ I KEF D+
Sbjct: 332 KKFSHPAAISFTTLYDNNIEISSSKHTLSQMNKVSNIVK-EPELPDI--YKEFKDITAET 388
Query: 514 DVPGLP-PVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPW 572
+ LP P++ +EF +++ + I Y + P ++ + ++ L G IR S +
Sbjct: 389 NTEKLPKPIKGLEFEVELTQENYRLPIRNYPLPPGKMQAMNDEINQGLKSGIIRESKAIN 448
Query: 573 GAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYH 632
PV+ V KK+G R+ VDY+ LNK N YPLP I+ L+ +++G+ IF+K+DLKS YH
Sbjct: 449 ACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLPLIEQLLAKIQGSTIFTKLDLKSAYH 508
Query: 633 QIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDI 692
IRV+ D K AFR G +EYLVMP+G++ APA F ++N + VV ++DDI
Sbjct: 509 LIRVRKGDEHKLAFRCPRGVFEYLVMPYGISTAPAHFQYFINTILGEAKESHVVCYMDDI 568
Query: 693 LIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVE 752
LI+SK+ EH H++ VLQ L+ L N +KCEF +VKF+G+ IS++G ++
Sbjct: 569 LIHSKSESEHVKHVKDVLQKLKNANLIINQAKCEFHQSQVKFIGYHISEKGFTPCQENID 628
Query: 753 TVLSWERPK 761
VL W++PK
Sbjct: 629 KVLQWKQPK 637
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 194 bits (494), Expect = 1e-48
Identities = 112/309 (36%), Positives = 176/309 (56%), Gaps = 5/309 (1%)
Query: 455 KRVVFPDSVLSEYLFANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVVKEFTDVFPG- 513
K+ P ++ L+ N+I +S + + +S K PE+ I KEF D+
Sbjct: 332 KKFSHPAAISFTTLYDNNIEISSSKHTLSQMNKVSNIVK-EPELPDI--YKEFKDITAET 388
Query: 514 DVPGLP-PVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPW 572
+ LP P++ +EF +++ + I Y + P ++ + ++ L G IR S +
Sbjct: 389 NTEKLPKPIKGLEFEVELTQENYRLPIRNYPLPPGKMQAMNDEINQGLKSGIIRESKAIN 448
Query: 573 GAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYH 632
PV+ V KK+G R+ VDY+ LNK N YPLP I+ L+ +++G+ IF+K+DLKS YH
Sbjct: 449 ACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLPLIEQLLAKIQGSTIFTKLDLKSAYH 508
Query: 633 QIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDI 692
IRV+ D K AFR G +EYLVMP+G++ APA F ++N + VV ++D+I
Sbjct: 509 LIRVRKGDEHKLAFRCPRGVFEYLVMPYGISIAPAHFQYFINTILGEVKESHVVCYMDNI 568
Query: 693 LIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVE 752
LI+SK+ EH H++ VLQ L+ L N +KCEF +VKF+G+ IS++G ++
Sbjct: 569 LIHSKSESEHVKHVKDVLQKLKNANLIINQAKCEFHQSQVKFIGYHISEKGFTPCQENID 628
Query: 753 TVLSWERPK 761
VL W++PK
Sbjct: 629 KVLQWKQPK 637
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 194 bits (494), Expect = 1e-48
Identities = 112/309 (36%), Positives = 176/309 (56%), Gaps = 5/309 (1%)
Query: 455 KRVVFPDSVLSEYLFANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVVKEFTDVFPG- 513
K+ P ++ L+ N+I +S + + +S K PE+ I KEF D+
Sbjct: 332 KKFSHPAAISFTTLYDNNIEISSSKHTLSQMNKVSNIVK-EPELPDI--YKEFKDITAET 388
Query: 514 DVPGLP-PVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPW 572
+ LP P++ +EF +++ + I Y + P ++ + ++ L G IR S +
Sbjct: 389 NTEKLPKPIKGLEFEVELTQENYRLPIRNYPLPPGKMQAMNDEINQGLKSGIIRESKAIN 448
Query: 573 GAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYH 632
PV+ V KK+G R+ VDY+ LNK N YPLP I+ L+ +++G+ IF+K+DLKS YH
Sbjct: 449 ACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLPLIEQLLAKIQGSTIFTKLDLKSAYH 508
Query: 633 QIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDI 692
IRV+ D K AFR G +EYLVMP+G++ APA F ++N + VV ++D+I
Sbjct: 509 LIRVRKGDEHKLAFRCPRGVFEYLVMPYGISIAPAHFQYFINTILGEVKESHVVCYMDNI 568
Query: 693 LIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVE 752
LI+SK+ EH H++ VLQ L+ L N +KCEF +VKF+G+ IS++G ++
Sbjct: 569 LIHSKSESEHVKHVKDVLQKLKNANLIINQAKCEFHQSQVKFIGYHISEKGFTPCQENID 628
Query: 753 TVLSWERPK 761
VL W++PK
Sbjct: 629 KVLQWKQPK 637
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 191 bits (484), Expect = 2e-47
Identities = 95/250 (38%), Positives = 153/250 (61%), Gaps = 11/250 (4%)
Query: 536 PISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKD-----GKSRLCV 590
PI Y +A +E+++Q++++L++G IR S SP+ +P +V KK K R+ +
Sbjct: 206 PIYSKQYPLAQTHEIEVENQVQEMLNQGLIRESNSPYNSPTWVVPKKPDASGANKYRVVI 265
Query: 591 DYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRY 650
DYR+LN++T+ +RYP+P +D+++ +L F+ IDL G+HQI + E I KTAF T+
Sbjct: 266 DYRKLNEITIPDRYPIPNMDEILGKLGKCQYFTTIDLAKGFHQIEMDEESISKTAFSTKS 325
Query: 651 GHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVL 710
GHYEYL MPFG+ NAPA F MN L++ +V++DDI+I+S ++ EH ++ V
Sbjct: 326 GHYEYLRMPFGLRNAPATFQRCMNNILRPLLNKHCLVYLDDIIIFSTSLTEHLNSIQLVF 385
Query: 711 QVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP------KTVL 764
L + L KCEF +E FLGH+++ +GI +P KV+ ++S+ P + L
Sbjct: 386 TKLADANLKLQLDKCEFLKKEANFLGHIVTPDGIKPNPIKVKAIVSYPIPTKDKEIRAFL 445
Query: 765 GVSLVWRDII 774
G++ +R I
Sbjct: 446 GLTGYYRKFI 455
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 185 bits (470), Expect = 7e-46
Identities = 93/235 (39%), Positives = 147/235 (61%), Gaps = 11/235 (4%)
Query: 551 ELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDGKS-----RLCVDYRQLNKVTVKNRYP 605
E++SQ++D+L++G IR S SP+ +P+ +V KK S R+ +DYR+LN++TV +R+P
Sbjct: 222 EVESQIQDMLNQGIIRTSNSPYNSPIWVVPKKQDASGKQKFRIVIDYRKLNEITVGDRHP 281
Query: 606 LPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNA 665
+P +D+++ +L F+ IDL G+HQI + E + KTAF T++GHYEYL MPFG+ NA
Sbjct: 282 IPNMDEILGKLGRCNYFTTIDLAKGFHQIEMDPESVSKTAFSTKHGHYEYLRMPFGLKNA 341
Query: 666 PAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKC 725
PA F MN L++ +V++DDI+++S ++ EH L V + L + L KC
Sbjct: 342 PATFQRCMNDILRPLLNKHCLVYLDDIIVFSTSLDEHLQSLGLVFEKLAKANLKLQLDKC 401
Query: 726 EFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP------KTVLGVSLVWRDII 774
EF +E FLGHV++ +GI +P K+E + + P K LG++ +R I
Sbjct: 402 EFLKQETTFLGHVLTPDGIKPNPEKIEAIQKYPIPTKPKEIKAFLGLTGYYRKFI 456
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 169 bits (427), Expect = 7e-41
Identities = 112/365 (30%), Positives = 185/365 (50%), Gaps = 27/365 (7%)
Query: 431 LKNLDVILGMDWLSHYHCLLDCN*KRVVFPDSVLSEYLFANHISVSLNGGSQEYSVLLSL 490
L + D I+G D L ++D ++ + L I+V+ LL+
Sbjct: 28 LHSFDGIIGDDTLKDLKAIVDRKNNCLIITPGIKIPLLARASINVN---------PLLAA 78
Query: 491 ESKGNPEVDSIPVVKEFTDVFPGDVPGLPPVRDIEFAIDIVPGTG---PISIAPYRMAPA 547
E + ++ EF +F + G+ +E A+ T PI Y
Sbjct: 79 EHPDGTQEILNSLLGEFPRIFEPPLSGM----SVETAVKAEIRTNTQDPIYAKSYPYPVN 134
Query: 548 ELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKK-----DGKSRLCVDYRQLNKVTVKN 602
E++ Q+++LL G IRPS SP+ +P+ +V KK + + R+ VD+++LN VT+ +
Sbjct: 135 MRGEVERQIDELLQDGIIRPSNSPYNSPIWIVPKKPKPNGEKQYRMVVDFKRLNTVTIPD 194
Query: 603 RYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGV 662
YP+P I+ + L A F+ +DL SG+HQI +K DI KTAF T G YE+L +PFG+
Sbjct: 195 TYPIPDINATLASLGNAKYFTTLDLTSGFHQIHMKESDIPKTAFSTLNGKYEFLRLPFGL 254
Query: 663 TNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANP 722
NAPAIF ++ + + V+IDDI+++S++ H +LR VL L + L N
Sbjct: 255 KNAPAIFQRMIDDILREHIGKVCYVYIDDIIVFSEDYDTHWKNLRLVLASLSKANLQVNL 314
Query: 723 SKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERPKTV------LGVSLVWRDIIVA 776
K F +V+FLG++++ +GI DP KV + P +V LG++ +R I
Sbjct: 315 EKSHFLDTQVEFLGYIVTADGIKADPKKVRAISEMPPPTSVKELKRFLGMTSYYRKFIQD 374
Query: 777 LSRIS 781
++++
Sbjct: 375 YAKVA 379
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 163 bits (412), Expect = 4e-39
Identities = 90/285 (31%), Positives = 155/285 (53%), Gaps = 6/285 (2%)
Query: 503 VVKEFTDVFPGDVPGLPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSK 562
V+++F DVF L E I++ G PI P + A E++ ++ +L++
Sbjct: 909 VIEQFQDVFAISDDELGRNSGTECVIELKEGAEPIRQKPRPIPLALKPEIRKMIQKMLNQ 968
Query: 563 GFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIF 622
IR S SPW +PV+LVKKKDG R+C+DYR++NKV N +PLP I+ + L G ++
Sbjct: 969 KVIRESKSPWSSPVVLVKKKDGSIRMCIDYRKVNKVVKNNAHPLPNIEATLQSLAGKKLY 1028
Query: 623 SKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLD 682
+ D+ +G+ QI + + + TAF +E+ V+PFG+ +PA+F M L
Sbjct: 1029 TVFDMIAGFWQIPLDEKSKEITAFAIGSELFEWNVLPFGLVISPALFQGTMEEIIGDLLG 1088
Query: 683 RFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKE 742
V++DD+LI SK++ +H +++ L +R+ + SKC +EV++LGH ++ +
Sbjct: 1089 VCAFVYVDDLLIASKDMEQHLQDVKEALTRIRKSGMKLRASKCHIAKKEVEYLGHKVTLD 1148
Query: 743 GIAVDPAKVETVLSWERPKTV------LGVSLVWRDIIVALSRIS 781
G+ K + + + RP V LG+ +R I+ ++I+
Sbjct: 1149 GVETQEVKTDKMKQFSRPTNVKELQSFLGLVGYYRKFILNFAQIA 1193
Score = 33.5 bits (75), Expect = 4.4
Identities = 26/98 (26%), Positives = 43/98 (43%), Gaps = 14/98 (14%)
Query: 281 CFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYTM 340
CF+C + GH A +NC K ++ P A+ V T G R +
Sbjct: 591 CFRCNEMGHIA----------WNCPKKNENTSEKEAPVAK--VETIEGVRMKDCLLMVKS 638
Query: 341 DGDDTEGVDGLIRGDCEIDGNLLSVLFDSGATHSFISR 378
+ ++E L +G +I + +L DSGA+ S +S+
Sbjct: 639 EKSESEVTRSLEKG--QIGKANVEILLDSGASISLMSK 674
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 159 bits (402), Expect = 5e-38
Identities = 83/257 (32%), Positives = 145/257 (56%), Gaps = 12/257 (4%)
Query: 552 LKSQLEDLLSKGFIRPSVSPWGAPVLLVKKK------DGKSRLCVDYRQLNKVTVKNRYP 605
+ ++++ LL G IRPS SP+ +P +V KK + RL +D+R+LN+ T+ +RYP
Sbjct: 197 VNNEVKQLLKDGIIRPSRSPYNSPTWVVDKKGTDAFGNPNKRLVIDFRKLNEKTIPDRYP 256
Query: 606 LPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNA 665
+P I ++ L A F+ +DLKSGYHQI + D +KT+F G YE+ +PFG+ NA
Sbjct: 257 MPSIPMILANLGKAKFFTTLDLKSGYHQIYLAEHDREKTSFSVNGGKYEFCRLPFGLRNA 316
Query: 666 PAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKC 725
+IF ++ + + V++DD++I+S+N +H H+ VL+ L + + + K
Sbjct: 317 SSIFQRALDDVLREQIGKICYVYVDDVIIFSENESDHVRHIDTVLKCLIDANMRVSQEKT 376
Query: 726 EFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP------KTVLGVSLVWRDIIVALSR 779
F+ E V++LG ++SK+G DP KV+ + + P ++ LG++ +R I +
Sbjct: 377 RFFKESVEYLGFIVSKDGTKSDPEKVKAIQEYPEPDCVYKVRSFLGLASYYRVFIKDFAA 436
Query: 780 ISPRSLDL*HNSRGRIN 796
I+ D+ G ++
Sbjct: 437 IARPITDILKGENGSVS 453
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 148 bits (373), Expect = 1e-34
Identities = 80/231 (34%), Positives = 127/231 (54%), Gaps = 6/231 (2%)
Query: 536 PISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDG------KSRLC 589
P+ YR +++ E+++Q++ L+ + PSVS + +P+LLV KK K RL
Sbjct: 314 PVYTKNYRSPHSQVEEIQAQVQKLIKDKIVEPSVSQYNSPLLLVPKKSSPNSDKKKWRLV 373
Query: 590 VDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTR 649
+DYRQ+NK + +++PLPRIDD++DQL A FS +DL SG+HQI + T+F T
Sbjct: 374 IDYRQINKKLLADKFPLPRIDDILDQLGRAKYFSCLDLMSGFHQIELDEGSRDITSFSTS 433
Query: 650 YGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQV 709
G Y + +PFG+ AP F M F +++DD+++ + +L +V
Sbjct: 434 NGSYRFTRLPFGLKIAPNSFQRMMTIAFSGIEPSQAFLYMDDLIVIGCSEKHMLKNLTEV 493
Query: 710 LQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP 760
RE L +P KC F++ EV FLGH + +GI D K + + ++ P
Sbjct: 494 FGKCREYNLKLHPEKCSFFMHEVTFLGHKCTDKGILPDDKKYDVIQNYPVP 544
>POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 146 bits (369), Expect = 4e-34
Identities = 76/204 (37%), Positives = 119/204 (58%), Gaps = 5/204 (2%)
Query: 537 ISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLV----KKKDGKSRLCVDY 592
I + P + +P + E Q+++LL I+PS SP AP LV +K+ GK R+ V+Y
Sbjct: 247 IKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNY 306
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
+ +NK TV + Y LP D+L+ +RG IFS D KSG+ Q+ + E TAF GH
Sbjct: 307 KAMNKATVGDAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGH 366
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
YE+ V+PFG+ AP+IF +M+ F F +F V++DDIL++S N +H H+ +LQ
Sbjct: 367 YEWNVVPFGLKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQK 425
Query: 713 LREKELYANPSKCEFWLEEVKFLG 736
+ + + K + + +++ FLG
Sbjct: 426 CNQHGIILSKKKAQLFKKKINFLG 449
>POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 146 bits (368), Expect = 5e-34
Identities = 75/204 (36%), Positives = 119/204 (57%), Gaps = 5/204 (2%)
Query: 537 ISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLV----KKKDGKSRLCVDY 592
I + P + +P + E Q+++LL I+PS SP AP LV +K+ GK R+ V+Y
Sbjct: 247 IKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNY 306
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
+ +NK T+ + Y LP D+L+ +RG IFS D KSG+ Q+ + E TAF GH
Sbjct: 307 KAMNKATIGDAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGH 366
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
YE+ V+PFG+ AP+IF +M+ F F +F V++DDIL++S N +H H+ +LQ
Sbjct: 367 YEWNVVPFGLKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQK 425
Query: 713 LREKELYANPSKCEFWLEEVKFLG 736
+ + + K + + +++ FLG
Sbjct: 426 CNQHGIILSKKKAQLFKKKINFLG 449
>POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 146 bits (368), Expect = 5e-34
Identities = 75/204 (36%), Positives = 119/204 (57%), Gaps = 5/204 (2%)
Query: 537 ISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLV----KKKDGKSRLCVDY 592
I + P + +P + E Q+++LL I+PS SP AP LV +K+ GK R+ V+Y
Sbjct: 247 IKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNY 306
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
+ +NK T+ + Y LP D+L+ +RG IFS D KSG+ Q+ + E TAF GH
Sbjct: 307 KAMNKATIGDAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGH 366
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
YE+ V+PFG+ AP+IF +M+ F F +F V++DDIL++S N +H H+ +LQ
Sbjct: 367 YEWNVVPFGLKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQK 425
Query: 713 LREKELYANPSKCEFWLEEVKFLG 736
+ + + K + + +++ FLG
Sbjct: 426 CNQHGIILSKKKAQLFKKKINFLG 449
>POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 674
Score = 144 bits (362), Expect = 2e-33
Identities = 75/204 (36%), Positives = 118/204 (57%), Gaps = 5/204 (2%)
Query: 537 ISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLV----KKKDGKSRLCVDY 592
I + P + +P + E Q+++LL I+PS SP AP LV +K+ GK R+ V+Y
Sbjct: 242 IKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNY 301
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
+ +NK TV + Y P D+L+ +RG IFS D KSG+ Q+ + E TAF GH
Sbjct: 302 KAMNKATVGDAYNPPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGH 361
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
YE+ V+PFG+ AP+IF +M+ F F +F V++DDIL++S N +H H+ +LQ
Sbjct: 362 YEWNVVPFGLKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQK 420
Query: 713 LREKELYANPSKCEFWLEEVKFLG 736
+ + + K + + +++ FLG
Sbjct: 421 CNQHGIILSKKKAQLFKKKINFLG 444
>RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse
transcriptase homolog)
Length = 142
Score = 142 bits (357), Expect = 9e-33
Identities = 69/136 (50%), Positives = 96/136 (69%), Gaps = 6/136 (4%)
Query: 587 RLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAF 646
R+C+DYR L KVT+KN+YP+PR+DDL D+L A F+K+DL+SGY Q+R+ D KT
Sbjct: 7 RMCIDYRALTKVTIKNKYPIPRVDDLFDRLAQATWFTKLDLRSGYWQVRIAKGDEPKTTC 66
Query: 647 RTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDIL---IYSKNVVEHE 703
TRYG +E+ VMPFG+TNA A F + MN + +LD FVVV++DD++ IYS ++ EH
Sbjct: 67 VTRYGSFEFRVMPFGLTNALATFCNLMNNVLYEYLDHFVVVYLDDLVVYTIYSNSLHEHI 126
Query: 704 GHLRQVLQVLREKELY 719
HLR V + KE++
Sbjct: 127 KHLRVVRE---SKEIF 139
>POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 680
Score = 140 bits (354), Expect = 2e-32
Identities = 73/204 (35%), Positives = 116/204 (56%), Gaps = 5/204 (2%)
Query: 537 ISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKD----GKSRLCVDY 592
I + P + +P + E Q+++LL I+PS SP AP LV + G R+ V+Y
Sbjct: 248 IKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAENGRGNKRMVVNY 307
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
+ +NK TV + Y LP D+L+ +RG IFS D KSG+ Q+ + E TAF GH
Sbjct: 308 KAMNKATVGDAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGH 367
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
YE+ V+PFG+ AP+IF +M+ F F +F V++DDI+++S N +H H+ +LQ
Sbjct: 368 YEWNVVPFGLKQAPSIFQRHMDEAFRVF-RKFCCVYVDDIVVFSNNEEDHLLHVAMILQK 426
Query: 713 LREKELYANPSKCEFWLEEVKFLG 736
+ + + K + + +++ FLG
Sbjct: 427 CNQHGIILSKKKAQLFKKKINFLG 450
>POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 666
Score = 134 bits (337), Expect = 2e-30
Identities = 70/219 (31%), Positives = 127/219 (57%), Gaps = 5/219 (2%)
Query: 527 AIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVK----KK 582
+I ++ I + P +P + Q+++LL G I PS S +P LV+ ++
Sbjct: 230 SIKLIDPLKVIRVKPMSYSPQDREGFAKQIKELLDLGLIIPSKSQHMSPAFLVENEAERR 289
Query: 583 DGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQ 642
GK R+ V+Y+ +N+ T+ + + LP + +L+ LRG +IFS D KSG+ Q+ + E +
Sbjct: 290 RGKKRMVVNYKAINQATIGDSHNLPNMQELLTLLRGKSIFSSFDCKSGFWQVVLDEESQK 349
Query: 643 KTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEH 702
TAF GH+++ V+PFG+ AP+IF +M +T D+F +V++DDI+++S + ++H
Sbjct: 350 LTAFTCPQGHFQWKVVPFGLKQAPSIFQRHM-QTALNGADKFCMVYVDDIIVFSNSELDH 408
Query: 703 EGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISK 741
H+ VL+++ + + + K + E++ FLG I K
Sbjct: 409 YNHVYAVLKIVEKYGIILSKKKANLFKEKINFLGLEIDK 447
>POL_SOCMV (P15629) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 692
Score = 132 bits (332), Expect = 7e-30
Identities = 77/217 (35%), Positives = 128/217 (58%), Gaps = 8/217 (3%)
Query: 548 ELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKK----KDGKSRLCVDYRQLNKVTVKNR 603
++ E K + EDLL KG IR S SP AP V+ K GK R+ ++Y+++N+ T+ +
Sbjct: 215 DVQEFKEECEDLLKKGLIRESQSPHSAPAFYVENHNEIKRGKRRMVINYKKMNEATIGDS 274
Query: 604 YPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFR-TRYGHYEYLVMPFGV 662
Y LPR D ++++++G+ FS +D KSGY+Q+R+ TAF HYE+ V+ FG+
Sbjct: 275 YKLPRKDFILEKIKGSLWFSSLDAKSGYYQLRLHENTKPLTAFSCPPQKHYEWNVLSFGL 334
Query: 663 TNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVE-HEGHLRQVLQVLREKELYAN 721
AP+I+ +M+++ L+ + +IDDILI++K E H +R VLQ ++EK + +
Sbjct: 335 KQAPSIYQRFMDQSLKG-LEHICLAYIDDILIFTKGSKEQHVNDVRIVLQRIKEKGIIIS 393
Query: 722 PSKCEFWLEEVKFLGHVISKEG-IAVDPAKVETVLSW 757
K + +E+++LG I G I + P E +L +
Sbjct: 394 KKKSKLIQQEIEYLGLKIQGNGEIDLSPHTQEKILQF 430
>POL_COYMV (P19199) Putative polyprotein [Contains: Coat protein;
Protease (EC 3.4.23.-); Reverse transcriptase (EC
2.7.7.49); Ribonuclease H (EC 3.1.26.4)]
Length = 1886
Score = 132 bits (331), Expect = 9e-30
Identities = 76/229 (33%), Positives = 124/229 (53%), Gaps = 12/229 (5%)
Query: 544 MAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLV-----------KKKDGKSRLCVDY 592
+ P + + Q+ LL IRPS S + +V K+K GK R+ +Y
Sbjct: 1410 VTPGDEEAMTRQINLLLQMKVIRPSESKHRSTAFIVRSGTEIDPITGKEKKGKERMVFNY 1469
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
+ LN+ T ++Y LP I+ ++ ++ + I+SK DLKSG+ Q+ ++ E + TAF
Sbjct: 1470 KLLNENTESDQYSLPGINTIISKVGRSKIYSKFDLKSGFWQVAMEEESVPWTAFLAGNKL 1529
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
YE+LVMPFG+ NAPAIF M+ F ++F+ V+IDDIL++S+ +H HL +LQ+
Sbjct: 1530 YEWLVMPFGLKNAPAIFQRKMDNVFKG-TEKFIAVYIDDILVFSETAEQHSQHLYTMLQL 1588
Query: 713 LREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERPK 761
+E L +P+K + E+ FLG + I + P + + + K
Sbjct: 1589 CKENGLILSPTKMKIGTPEIDFLGASLGCTKIKLQPHIISKICDFSDEK 1637
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 125 bits (315), Expect = 7e-28
Identities = 129/481 (26%), Positives = 201/481 (40%), Gaps = 74/481 (15%)
Query: 299 PQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYTMDGDDTEGVDGLI------ 352
P C+ CNK GH A CR + P G + +K G D+ VDGL
Sbjct: 239 PTCFYCNKKGHYATNCR---SNPKTGNQGGNKGKSK-------GCDSVHVDGLDVKTEHQ 288
Query: 353 ---RGDCEIDGNLLSVLFDSGATHSFISRECAIRLRLPITALSFDLIVTTPAKTLLANTA 409
R E+ G ++ D+G+ + IS +C +L P + P + AN
Sbjct: 289 AKHRMSVEVCGKDVAFQLDTGSMITLISVKCWEKLGSPP-------LEKVPHRISCANGT 341
Query: 410 CMY----CSVTYNNRVYHANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVFPDSVLS 465
M C V + + +LG WL+ + P +
Sbjct: 342 PMAVKGRCLVKFKLKGIEYTEYVYVRDRQTNLLGTSWLN-------------LCPQMRSA 388
Query: 466 EYLFANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVVKEFTDVFPGDVPGLPPVRDIE 525
N +S S S+ LE + + +F +VF D GL E
Sbjct: 389 LAQIVNQVSTSETEASR-------LE---------VMLKNDFPEVFK-DGLGLCTKEKAE 431
Query: 526 FAID--IVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRP-SVSPWGAPVLLVKKK 582
F + VP PY L ++++L L G I P + + W AP++++KKK
Sbjct: 432 FRTEENAVPVFKRARPVPY----GSLEAVETELNRLQEMGVIVPITYAKWAAPIVVIKKK 487
Query: 583 D-GKSRLCVDYR--QLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTE 639
GK R+C D++ LN +PLP +D+ +L+G ++S+IDLK Y Q+ + E
Sbjct: 488 GTGKIRVCADFKCSGLNAALKDEFHPLPTSEDIFSRLKGT-VYSQIDLKDAYLQVELDEE 546
Query: 640 DIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNV 699
+ T G ++YL M FG+ APA F M++ V V+ DDI+I + ++
Sbjct: 547 AQKLAVINTHRGIFKYLRMTFGLKPAPASFQKIMDKMVSGLTG--VAVYWDDIIISASSI 604
Query: 700 VEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWER 759
EHE LR++ + +E + KC F ++V FLG V + G D K E + S +
Sbjct: 605 EEHEKILRELFERFKEYGFRVSAEKCAFAQKQVTFLGFV-DEHGRRPDSKKTEAIRSMKA 663
Query: 760 P 760
P
Sbjct: 664 P 664
>POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 659
Score = 117 bits (293), Expect = 2e-25
Identities = 65/214 (30%), Positives = 115/214 (53%), Gaps = 5/214 (2%)
Query: 528 IDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVK----KKD 583
I+++ + + P +P++ E Q+++LL I+PS S +P LV+ ++
Sbjct: 220 IELIDPKTVVKVKPMSYSPSDREEFDRQIKELLELKVIKPSKSTHMSPAFLVENEAERRR 279
Query: 584 GKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQK 643
GK R+ V+Y+ +NK T + + LP D+L+ +RG I+S D KSG Q+ + E
Sbjct: 280 GKKRMVVNYKAMNKATKGDAHNLPNKDELLTLVRGKKIYSSFDCKSGLWQVLLDKESQLL 339
Query: 644 TAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSK-NVVEH 702
TAF GHY++ V+PFG+ AP+IF + ++ V++DDIL++S EH
Sbjct: 340 TAFTCPQGHYQWNVVPFGLKQAPSIFPKTYANSHSNQYSKYCCVYVDDILVFSNTGRKEH 399
Query: 703 EGHLRQVLQVLREKELYANPSKCEFWLEEVKFLG 736
H+ +L+ + + + K + + E++ FLG
Sbjct: 400 YIHVLNILRRCEKLGIILSKKKAQLFKEKINFLG 433
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.345 0.152 0.510
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 154,791,957
Number of Sequences: 164201
Number of extensions: 6220074
Number of successful extensions: 26589
Number of sequences better than 10.0: 240
Number of HSP's better than 10.0 without gapping: 137
Number of HSP's successfully gapped in prelim test: 104
Number of HSP's that attempted gapping in prelim test: 25844
Number of HSP's gapped (non-prelim): 527
length of query: 1469
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1346
effective length of database: 39,777,331
effective search space: 53540287526
effective search space used: 53540287526
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0016.15


