

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
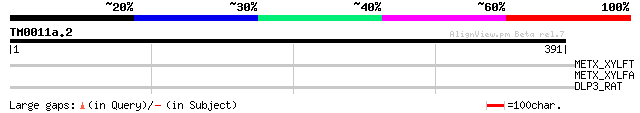
Query= TM0011a.2
(391 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
METX_XYLFT (Q87BG9) Homoserine O-acetyltransferase (EC 2.3.1.31)... 32 3.6
METX_XYLFA (Q9PAN0) Homoserine O-acetyltransferase (EC 2.3.1.31)... 32 3.6
DLP3_RAT (P97838) Disks large-associated protein 3 (DAP-3) (SAP9... 31 4.7
>METX_XYLFT (Q87BG9) Homoserine O-acetyltransferase (EC 2.3.1.31)
(Homoserine O-trans-acetylase) (Homoserine
transacetylase) (HTA)
Length = 370
Score = 31.6 bits (70), Expect = 3.6
Identities = 28/95 (29%), Positives = 43/95 (44%), Gaps = 11/95 (11%)
Query: 8 RSHIERNREEGHVRLWNDYFSTNPVYTEEQFQRRYRMRKHMFLRIVEA-----ISNTDDY 62
RS +E + G VRL +D + P E Q + H F+ + +S + D+
Sbjct: 226 RSALEWDGRFGRVRLDSDQTNDTPFGLEFQIENYLESHAHRFVHTFDPNCYLYLSRSMDW 285
Query: 63 FQMRNDATGRM--GLSPLQKQTAAIRMLAYGSSTD 95
F + A G + GL+ ++ Q R LA GS TD
Sbjct: 286 FDVAEYANGDIIAGLARIRIQ----RALAIGSHTD 316
>METX_XYLFA (Q9PAN0) Homoserine O-acetyltransferase (EC 2.3.1.31)
(Homoserine O-trans-acetylase) (Homoserine
transacetylase) (HTA)
Length = 370
Score = 31.6 bits (70), Expect = 3.6
Identities = 28/95 (29%), Positives = 43/95 (44%), Gaps = 11/95 (11%)
Query: 8 RSHIERNREEGHVRLWNDYFSTNPVYTEEQFQRRYRMRKHMFLRIVEA-----ISNTDDY 62
RS +E + G VRL +D + P E Q + H F+ + +S + D+
Sbjct: 226 RSALEWDGRFGRVRLDSDQTNDTPFGLEFQIENYLESHAHRFVHTFDPNCYLYLSRSMDW 285
Query: 63 FQMRNDATGRM--GLSPLQKQTAAIRMLAYGSSTD 95
F + A G + GL+ ++ Q R LA GS TD
Sbjct: 286 FDVAEYANGDILAGLARIRIQ----RALAIGSHTD 316
>DLP3_RAT (P97838) Disks large-associated protein 3 (DAP-3)
(SAP90/PSD-95-associated protein 3) (SAPAP3)
(PSD-95/SAP90 binding protein 3)
Length = 977
Score = 31.2 bits (69), Expect = 4.7
Identities = 25/84 (29%), Positives = 33/84 (38%), Gaps = 14/84 (16%)
Query: 42 YRMRKHMFLRIVEAISNTDD----YFQMRNDATGRMGLS----------PLQKQTAAIRM 87
+RMR H +LR ++A + DD +GR G S P Q
Sbjct: 494 FRMRSHSYLRAIQAGCSQDDDCLPLLAAPASVSGRPGSSFNFRKAPPPIPPGSQAPPRIS 553
Query: 88 LAYGSSTDSVDEYVCIGESTARKC 111
+ SSTDS E E AR+C
Sbjct: 554 ITAQSSTDSAHESFTAAEGPARRC 577
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,894,265
Number of Sequences: 164201
Number of extensions: 2110917
Number of successful extensions: 4379
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 4378
Number of HSP's gapped (non-prelim): 3
length of query: 391
length of database: 59,974,054
effective HSP length: 112
effective length of query: 279
effective length of database: 41,583,542
effective search space: 11601808218
effective search space used: 11601808218
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0011a.2


