

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.6
(493 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
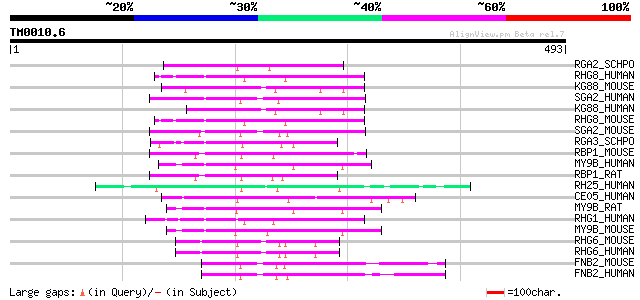
Score E
Sequences producing significant alignments: (bits) Value
RGA2_SCHPO (Q10164) Probable Rho-type GTPase-activating protein 2 78 5e-14
RHG8_HUMAN (Q9NSG0) Rho-GTPase-activating protein 8 (PP610) 70 1e-11
KG88_MOUSE (P59281) Protein KIAA1688 homolog (Fragment) 68 5e-11
SGA2_HUMAN (O43295) SLIT-ROBO Rho GTPase activating protein 2 (s... 66 2e-10
KG88_HUMAN (Q9C0H5) Protein KIAA1688 65 3e-10
RHG8_MOUSE (Q9CXP4) Rho-GTPase-activating protein 8 65 4e-10
SGA2_MOUSE (Q812A2) SLIT-ROBO Rho GTPase activating protein 2 (s... 64 1e-09
RGA3_SCHPO (O14014) Probable Rho-type GTPase-activating protein 3 63 2e-09
RBP1_MOUSE (Q62172) RalA binding protein 1 (RalBP1) (Ral interac... 62 3e-09
MY9B_HUMAN (Q13459) Myosin IXb (Unconventional myosin-9b) 61 6e-09
RBP1_RAT (Q62796) RalA binding protein 1 (RalBP1) (Ral interacti... 61 8e-09
RH25_HUMAN (P42331) Rho-GTPase-activating protein 25 60 1e-08
CE05_HUMAN (Q9NYF5) Protein C5orf5 (GAP-like protein N61) 60 2e-08
MY9B_RAT (Q63358) Myosin IXb (Unconventional myosin-9b) 59 3e-08
RHG1_HUMAN (Q07960) Rho-GTPase-activating protein 1 (GTPase-acti... 59 4e-08
MY9B_MOUSE (Q9QY06) Myosin IXb (Unconventional myosin-9b) 59 4e-08
RHG6_MOUSE (O54834) Rho-GTPase-activating protein 6 (Rho-type GT... 57 8e-08
RHG6_HUMAN (O43182) Rho-GTPase-activating protein 6 (Rho-type GT... 57 1e-07
FNB2_MOUSE (Q91Z67) Formin binding protein 2 (Formin binding pro... 57 1e-07
FNB2_HUMAN (O75044) Formin binding protein 2 (srGAP2) 57 1e-07
>RGA2_SCHPO (Q10164) Probable Rho-type GTPase-activating protein 2
Length = 1275
Score = 78.2 bits (191), Expect = 5e-14
Identities = 51/174 (29%), Positives = 86/174 (49%), Gaps = 14/174 (8%)
Query: 137 TSVFGVS-TESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVRE 195
+ +FG+ E++ +S + +P ++ +L + + EGI+R++ S + ++E
Sbjct: 1060 SGIFGLPLNEAVNISTQFNDSGLPIVVYRCIEYLESCRAEKEEGIYRLSGSASTIKHLKE 1119
Query: 196 QLNRGV------VPNGVDVHCLAGLIKAWFRELPTGILDP-------LSPEEVMQSQSEE 242
Q N GV DVH +AGL+K + R LPT +LD L P S +
Sbjct: 1120 QFNEGVDYDLLSSDEEFDVHVIAGLLKLYLRNLPTNLLDTSMHKLFELLPNVPNDSAALG 1179
Query: 243 ECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMAD 296
E ++ LPP ALLD ++ + + E NKMN RN+ +VF+P + +D
Sbjct: 1180 ELCDVISKLPPENFALLDSLLHHLRRIIAFEKVNKMNIRNVCIVFSPTLNIPSD 1233
>RHG8_HUMAN (Q9NSG0) Rho-GTPase-activating protein 8 (PP610)
Length = 718
Score = 70.5 bits (171), Expect = 1e-11
Identities = 56/196 (28%), Positives = 95/196 (47%), Gaps = 13/196 (6%)
Query: 129 PRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENS 188
P RPP T FGVS + L +G +P +L +L +G L+ EG+FR +A
Sbjct: 468 PPRPP-LPTQQFGVSLQ--YLKDKNQGELIPPVLRFTVTYLREKG-LRTEGLFRRSASVQ 523
Query: 189 QEELVREQLNRGVVPNGVD---VHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEE-- 243
++ N+G N D +H A ++K + RELP +L + E+++ E
Sbjct: 524 TVREIQRLYNQGKPVNFDDYGDIHIPAVILKTFLRELPQPLLTFQAYEQILGITCVESSL 583
Query: 244 ----CDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLT 299
C Q++R LP +L + + + V++ FNKMN+ N+A VF N+ + ++
Sbjct: 584 RVTGCRQILRSLPEHNYVVLRYLMGFLHAVSRESIFNKMNSSNLACVFGLNLIWPSQGVS 643
Query: 300 ALMYAVQVMSFLKTLV 315
+L V + F + L+
Sbjct: 644 SLSALVPLNMFTELLI 659
>KG88_MOUSE (P59281) Protein KIAA1688 homolog (Fragment)
Length = 1097
Score = 68.2 bits (165), Expect = 5e-11
Identities = 55/192 (28%), Positives = 87/192 (44%), Gaps = 16/192 (8%)
Query: 136 STSVFGVSTESMQLSFDAR--GNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELV 193
S S+FG + + + R +P + + + A G Q EGIFR+ + + +
Sbjct: 898 SPSMFGSALQEVMSMQKERYPDRQLPWVQTRLSEEVLALNGDQTEGIFRVPGDIDEVNAL 957
Query: 194 REQLNRGVVPNGV-DVHCLAGLIKAWFRELPTGILDPLSPEE-----VMQSQSEEECDQL 247
+ Q+++ VP G+ D H A L+K W+REL +PL P E + +S E +
Sbjct: 958 KLQVDQWKVPTGLEDPHVPASLLKLWYRELE----EPLIPHEFYEQCIAHYESPEAAVAV 1013
Query: 248 VRLLPPTEAALLDWAINLMADVAQMEH--FNKMNARNIAMVFAPNMTHMA--DPLTALMY 303
V LP +L + I + Q + KM+ N+AMV APN DP
Sbjct: 1014 VHALPRINRMVLCYLIRFLQVFVQPANVAITKMDVSNLAMVMAPNCLRCQSDDPRVIFEN 1073
Query: 304 AVQVMSFLKTLV 315
+ MSFL+ L+
Sbjct: 1074 TRKEMSFLRVLI 1085
>SGA2_HUMAN (O43295) SLIT-ROBO Rho GTPase activating protein 2
(srGAP2) (srGAP3) (WAVE-associated Rac GTPase activating
protein) (WRP) (Mental disorder-associated GAP)
(Rho-GTPase-activating protein 14)
Length = 1099
Score = 65.9 bits (159), Expect = 2e-10
Identities = 47/208 (22%), Positives = 93/208 (44%), Gaps = 21/208 (10%)
Query: 125 EPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRIN 184
+P+ RRP S + SM+ G ++P ++ R++ G LQ +GIFR+
Sbjct: 486 KPQKMRRPRPLSVYSHKLFNGSMEAFIKDSGQAIPLVVESCIRYINLYG-LQQQGIFRVP 544
Query: 185 AENSQEELVREQLNRGVVP-----NGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQ-- 237
+ ++ RG P N D++ +AG++K +FR G+ +PL P+E Q
Sbjct: 545 GSQVEVNDIKNSFERGEDPLVDDQNERDINSVAGVLKLYFR----GLENPLFPKERFQDL 600
Query: 238 ---------SQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFA 288
++ + Q++ LP ++ + + ++Q N M+ N+A+ F
Sbjct: 601 ISTIKLENPAERVHQIQQILVTLPRVVIVVMRYLFAFLNHLSQYSDENMMDPYNLAICFG 660
Query: 289 PNMTHMADPLTALMYAVQVMSFLKTLVV 316
P + H+ D + + +KT+++
Sbjct: 661 PTLMHIPDGQDPVSCQAHINEVIKTIII 688
>KG88_HUMAN (Q9C0H5) Protein KIAA1688
Length = 1083
Score = 65.5 bits (158), Expect = 3e-10
Identities = 50/168 (29%), Positives = 77/168 (45%), Gaps = 14/168 (8%)
Query: 158 VPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGV-DVHCLAGLIK 216
+P + + + A G Q EGIFR+ + + ++ Q+++ VP G+ D H A L+K
Sbjct: 908 LPWVQTRLSEEVLALNGDQTEGIFRVPGDIDEVNALKLQVDQWKVPTGLEDPHVPASLLK 967
Query: 217 AWFRELPTGILDPLSPEE-----VMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQ 271
W+REL +PL P E + S E +V LP +L + I + Q
Sbjct: 968 LWYRELE----EPLIPHEFYEQCIAHYDSPEAAVAVVHALPRINRMVLCYLIRFLQVFVQ 1023
Query: 272 MEH--FNKMNARNIAMVFAPNMTHMA--DPLTALMYAVQVMSFLKTLV 315
+ KM+ N+AMV APN DP + MSFL+ L+
Sbjct: 1024 PANVAVTKMDVSNLAMVMAPNCLRCQSDDPRVIFENTRKEMSFLRVLI 1071
>RHG8_MOUSE (Q9CXP4) Rho-GTPase-activating protein 8
Length = 425
Score = 65.1 bits (157), Expect = 4e-10
Identities = 56/196 (28%), Positives = 94/196 (47%), Gaps = 13/196 (6%)
Query: 129 PRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENS 188
P RPP T FGVS + L +G +P +L +L +G L+ EG+FR +A
Sbjct: 183 PPRPP-LPTQQFGVSLQ--YLRDKNQGELIPPVLRWTVTYLREKG-LRTEGLFRRSASAQ 238
Query: 189 QEELVREQLNRGVVPNGVD---VHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEE-- 243
V+ ++G N D +H A ++K + RELP +L + E+++ S E
Sbjct: 239 TVRQVQRLYDQGKPVNFDDYGDMHLPAVILKTFLRELPQPLLTFQAYEQILGITSVESSL 298
Query: 244 ----CDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLT 299
C ++R LP A+L + + + +V+ NKMN+ N+A VF N+ + +
Sbjct: 299 RVTHCRLILRSLPEHNYAVLRYLMGFLHEVSLESISNKMNSSNLACVFGLNLIWPSQGVA 358
Query: 300 ALMYAVQVMSFLKTLV 315
+L V + F + L+
Sbjct: 359 SLSALVPLNLFTELLI 374
>SGA2_MOUSE (Q812A2) SLIT-ROBO Rho GTPase activating protein 2
(srGAP2) (WAVE-associated Rac GTPase activating protein)
(WRP) (Rho-GTPase-activating protein 14)
Length = 1099
Score = 63.5 bits (153), Expect = 1e-09
Identities = 52/210 (24%), Positives = 93/210 (43%), Gaps = 25/210 (11%)
Query: 125 EPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQR--HLYARGGLQAEGIFR 182
+P+ RRP S + SM+ G ++P + R +LY GLQ +GIFR
Sbjct: 486 KPQKMRRPRPLSVYSHKLFNGSMEAFIKDSGQAIPLVAESCIRFINLY---GLQQQGIFR 542
Query: 183 INAENSQEELVREQLNRGVVP-----NGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQ 237
+ + ++ RG P N D++ +AG++K +FR G+ +PL P+E Q
Sbjct: 543 VPGSQVEVNDIKNSFERGEDPLVDDQNERDINSVAGVLKLYFR----GLENPLFPKERFQ 598
Query: 238 S-----QSEEECD------QLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMV 286
+ E D Q++ LP ++ + + ++Q N M+ N+A+
Sbjct: 599 DLISTIKLENPADRVHPIQQILITLPRVVIVVMRYLFAFLNHLSQYSDENMMDPYNLAIC 658
Query: 287 FAPNMTHMADPLTALMYAVQVMSFLKTLVV 316
F P + H+ D + V +KT+++
Sbjct: 659 FGPTLMHIPDGQDPVSCQAHVNEVIKTIII 688
>RGA3_SCHPO (O14014) Probable Rho-type GTPase-activating protein 3
Length = 969
Score = 63.2 bits (152), Expect = 2e-09
Identities = 47/178 (26%), Positives = 92/178 (51%), Gaps = 16/178 (8%)
Query: 126 PEVPRR--PPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRI 183
PE+P R PP S ++FG S E+ QL + G+ +P ++ + + A G L+ EGI+RI
Sbjct: 762 PEMPTRMPPPGPSPTMFGRSLEN-QLKIE--GSVLPQVIAMCVSCVDAHG-LEVEGIYRI 817
Query: 184 NAENSQEELVREQLNRGVVPNG---VDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQS 240
+ SQ ++ ++ G + D+ ++K + LP ++ EE+++++
Sbjct: 818 SGSASQVRVLVDEFENGSIRMEHLTSDLFACTSVLKTYLHRLPEPVIPGTQYEELLEAEK 877
Query: 241 ----EEECDQLVRL---LPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNM 291
EE+ +++V + L P ++ + I + V + N MN++N++ VFAP +
Sbjct: 878 IEKEEEKIERVVEVMKTLHPAHLSVFRFLIAHLGRVCKHAEKNLMNSKNVSTVFAPTL 935
>RBP1_MOUSE (Q62172) RalA binding protein 1 (RalBP1) (Ral
interacting protein 1) (Dinitrophenyl S-glutathione
ATPase) (DNP-SG ATPase)
Length = 647
Score = 62.0 bits (149), Expect = 3e-09
Identities = 54/207 (26%), Positives = 98/207 (47%), Gaps = 21/207 (10%)
Query: 125 EPEVPRRPPSASTSVFGVS-TESMQLSFDARGNSVPTILL----LMQRHLYARGGLQAEG 179
EPEVP+ + +FGV ++++ + G +P + M++H G++ EG
Sbjct: 174 EPEVPQMDAPSVKPIFGVPLVDAVERTMMYDGVRLPAVFRECVDYMEKH-----GMKCEG 228
Query: 180 IFRINAENSQEELVREQLNRGVVPN--GVDVHCLAGLIKAWFRELPTGILDP-LSPE--- 233
++R++ S+ + ++ +R PN + + +A L+K + R+LP +L L P
Sbjct: 229 VYRVSGIKSKVDELKAAYDREESPNLEEYEPNTVASLLKQYLRDLPENLLTKELMPRFEE 288
Query: 234 ---EVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPN 290
+ + + +E +L+R LP LL W I + V E KMN +NI++V +P
Sbjct: 289 ACGKTTEMEKVQEFQRLLRELPECNHLLLSWLIVHLDHVIAKELETKMNIQNISIVLSPT 348
Query: 291 MTHMADPLTALMYAVQVMSFLKTLVVK 317
+ L L VQ T+V+K
Sbjct: 349 VQISNRVLYVLFTHVQ--ELFGTVVLK 373
>MY9B_HUMAN (Q13459) Myosin IXb (Unconventional myosin-9b)
Length = 2158
Score = 61.2 bits (147), Expect = 6e-09
Identities = 49/202 (24%), Positives = 98/202 (48%), Gaps = 18/202 (8%)
Query: 133 PSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEEL 192
P A FGV +S+ + SVP +L + H+ G L EG++R + ++
Sbjct: 1691 PGAEPGHFGVCVDSLT----SDKASVPIVLEKLLEHVEMHG-LYTEGLYRKSGAANRTRE 1745
Query: 193 VREQLNR---GVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQS-QSEEECDQLV 248
+R+ L V +H + G++K W RELP ++ + +++ + E+ +QL
Sbjct: 1746 LRQALQTDPAAVKLENFPIHAITGVLKQWLRELPEPLMTFAQYGDFLRAVELPEKQEQLA 1805
Query: 249 RL------LPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHM---ADPLT 299
+ LP L+ I + VA +E N+M+ +A++FAP + +DPLT
Sbjct: 1806 AIYAVLEHLPEANHNSLERLIFHLVKVALLEDVNRMSPGALAIIFAPCLLRCPDNSDPLT 1865
Query: 300 ALMYAVQVMSFLKTLVVKTLRE 321
++ +++ + ++ L+ + +R+
Sbjct: 1866 SMKDVLKITTCVEMLIKEQMRK 1887
>RBP1_RAT (Q62796) RalA binding protein 1 (RalBP1) (Ral interacting
protein 1) (Cytocentrin) (Dinitrophenyl S-glutathione
ATPase) (DNP-SG ATPase)
Length = 646
Score = 60.8 bits (146), Expect = 8e-09
Identities = 51/181 (28%), Positives = 89/181 (48%), Gaps = 19/181 (10%)
Query: 125 EPEVPRRPPSASTSVFGVS-TESMQLSFDARGNSVPTILL----LMQRHLYARGGLQAEG 179
EPEVP+ + +FG ++++ + G +P + M++H G++ EG
Sbjct: 174 EPEVPQTDAPSLRPIFGAPFADAVERTMMYDGIRLPAVFRECVDYMEKH-----GMKCEG 228
Query: 180 IFRINAENSQEELVREQLNRGVVPN--GVDVHCLAGLIKAWFRELPTGILDP-LSP--EE 234
I+R++ S+ + ++ +R PN + + +A L+K + R+LP +L L P EE
Sbjct: 229 IYRVSGIKSKVDELKAAYDREESPNLEEYEPNTVASLLKQYLRDLPENLLTKELMPRFEE 288
Query: 235 VMQSQSE----EECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPN 290
+E +E +L+R LP LL W I M V E KMN +NI++V +P
Sbjct: 289 ACGRTTEVEKVQEFQRLLRELPEYNHLLLSWLIVHMDHVIAKELETKMNIQNISIVLSPT 348
Query: 291 M 291
+
Sbjct: 349 V 349
>RH25_HUMAN (P42331) Rho-GTPase-activating protein 25
Length = 638
Score = 60.1 bits (144), Expect = 1e-08
Identities = 73/350 (20%), Positives = 140/350 (39%), Gaps = 39/350 (11%)
Query: 77 IGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVP--RRPPS 134
I +T+P ++G + W N + D + E E V RR
Sbjct: 90 IKEIATNPEEAGKFVFEIIPASWDQN-----RMGQDSYVLMASSQAEMEEWVKFLRRVAG 144
Query: 135 ASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVR 194
VFG + G + IL+ G EGIFR+ +++ + +R
Sbjct: 145 TPCGVFGQRLDETVAYEQKFGPHLVPILVEKCAEFILEHGRNEEGIFRLPGQDNLVKQLR 204
Query: 195 EQLNRGVVPN---GVDVHCLAGLIKAWFRELPTGILDPLSPEEVM----------QSQSE 241
+ + G P+ DVH +A L+K + R+LP ++ P S E +++++
Sbjct: 205 DAFDAGERPSFDRDTDVHTVASLLKLYLRDLPEPVV-PWSQYEGFLLCGQLTNADEAKAQ 263
Query: 242 EECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNM--THMADPLT 299
+E + + +LP +LL + + ++ NKM+ N+A V N+ + + DP
Sbjct: 264 QELMKQLSILPRDNYSLLSYICRFLHEIQLNCAVNKMSVDNLATVIGVNLIRSKVEDPAV 323
Query: 300 ALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGN 359
+ Q+ + ++ R+ E KS +P ++D + P S G
Sbjct: 324 IMRGTPQIQRVMTMMI----RDHEVLFPKSKDIP----LSPPAQKNDPKKAPVARSSVGW 375
Query: 360 DCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRL 409
D +++ + +S S S T ++++ S T P++ F S++
Sbjct: 376 DATED--LRISRTDSFSSMT------SDSDTTSPTGQQPSDAFPEDSSKV 417
>CE05_HUMAN (Q9NYF5) Protein C5orf5 (GAP-like protein N61)
Length = 915
Score = 59.7 bits (143), Expect = 2e-08
Identities = 56/246 (22%), Positives = 110/246 (43%), Gaps = 23/246 (9%)
Query: 136 STSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVRE 195
+ +FG+ + +Q VP I+ + ++ GGL+ +G+F++N E +R+
Sbjct: 17 ANKIFGIPLDELQQGGHP-DIEVPFIVRHVVDYIEEHGGLEQQGLFQVNGNAETVEWLRQ 75
Query: 196 QLNRGV---VPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQ-SQSEEECDQ----- 246
+ + G + DV L++ + +ELP ++ +MQ SQ D+
Sbjct: 76 RYDSGEEVDLVKEADVPSAISLLRFFLQELPEPVIPGSLHIHLMQLSQDYNNEDEFGRKL 135
Query: 247 --LVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYA 304
L++ LPP +LL + +A+VA H +A ++A VF P++ H+ + +
Sbjct: 136 RFLLQQLPPVNYSLLKFLCRFLANVAS-HHEEIWSANSLAAVFGPDVFHIYTDVEDMKEQ 194
Query: 305 VQVMSFLKTLVVKTLR--EREESIVKSNPVPNL----NSFDDDGHQSDS----QVLPKDG 354
V + L+ E EE SN + ++ N ++ + + + LP++G
Sbjct: 195 EIVSRIMAGLLENYYEFFENEEEDFSSNDLSSITEQVNELSEEEEEDEKLEHIEELPEEG 254
Query: 355 SENGND 360
+E ND
Sbjct: 255 AEKSND 260
>MY9B_RAT (Q63358) Myosin IXb (Unconventional myosin-9b)
Length = 1980
Score = 58.9 bits (141), Expect = 3e-08
Identities = 48/204 (23%), Positives = 100/204 (48%), Gaps = 18/204 (8%)
Query: 140 FGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNR 199
FGV +S+ + SVP +L + H+ G L EG++R + ++ +R+ L
Sbjct: 1658 FGVCVDSLT----SDKASVPIVLEKLLEHVEMHG-LYTEGLYRKSGAANRTRELRQALQT 1712
Query: 200 G---VVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQS-QSEEECDQLVRL----- 250
V +H + G++K W RELP ++ + +++ + E+ +QL +
Sbjct: 1713 DPATVKLEDFPIHAITGVLKQWLRELPEPLMTFAQYGDFLRAVELPEKQEQLAAIYAVLD 1772
Query: 251 -LPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHM---ADPLTALMYAVQ 306
LP L+ I + VA +E N+M+ +A++FAP + +DPLT++ ++
Sbjct: 1773 HLPEANHTSLERLIFHLVKVALLEDVNRMSPGALAIIFAPCLLRCPDNSDPLTSMKDVLK 1832
Query: 307 VMSFLKTLVVKTLREREESIVKSN 330
+ + ++ L+ + +R+ + + + N
Sbjct: 1833 ITTCVEMLIKEQMRKYKVKMEEIN 1856
>RHG1_HUMAN (Q07960) Rho-GTPase-activating protein 1
(GTPase-activating protein rhoOGAP) (Rho-related small
GTPase protein activator) (CDC42 GTPase-activating
protein) (p50-rhoGAP)
Length = 439
Score = 58.5 bits (140), Expect = 4e-08
Identities = 56/205 (27%), Positives = 90/205 (43%), Gaps = 14/205 (6%)
Query: 121 PVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGI 180
P +P RPP + FGVS + +Q + +P +L +L A L EGI
Sbjct: 224 PATAPKPMPPRPPLPNQQ-FGVSLQHLQEK-NPEQEPIPIVLRETVAYLQAHA-LTTEGI 280
Query: 181 FRINAENSQEELVREQLNRGVVPNGVD----VHCLAGLIKAWFRELPTGILD-PLSPE-- 233
FR +A V+++ N G+ P D +H A ++K + RELP +L L P
Sbjct: 281 FRRSANTQVVREVQQKYNMGL-PVDFDQYNELHLPAVILKTFLRELPEPLLTFDLYPHVV 339
Query: 234 ---EVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPN 290
+ +SQ Q+++ LP +L + + ++ NKM N+A+VF PN
Sbjct: 340 GFLNIDESQRVPATLQVLQTLPEENYQVLRFLTAFLVQISAHSDQNKMTNTNLAVVFGPN 399
Query: 291 MTHMADPLTALMYAVQVMSFLKTLV 315
+ D L + +F K L+
Sbjct: 400 LLWAKDAAITLKAINPINTFTKFLL 424
>MY9B_MOUSE (Q9QY06) Myosin IXb (Unconventional myosin-9b)
Length = 2114
Score = 58.5 bits (140), Expect = 4e-08
Identities = 49/204 (24%), Positives = 98/204 (48%), Gaps = 18/204 (8%)
Query: 140 FGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNR 199
FGV +S+ + SVP +L + H+ G L EG++R + ++ +R+ L
Sbjct: 1656 FGVCVDSLT----SDKASVPIVLEKLLEHVEMHG-LYTEGLYRKSGAANRTRELRQALQT 1710
Query: 200 ---GVVPNGVDVHCLAGLIKAWFRELPTGIL------DPLSPEEVMQSQSE-EECDQLVR 249
V +H + G++K W RELP ++ D L E+ + Q + ++
Sbjct: 1711 DPAAVKLEDFPIHAITGVLKQWLRELPEPLMTFAQYGDFLRAVELPEKQEQLSAIYAVLD 1770
Query: 250 LLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHM---ADPLTALMYAVQ 306
LP L+ I + VA +E N+M+ +A++FAP + +DPLT++ ++
Sbjct: 1771 HLPEANHTSLERLIFHLVKVALLEDVNRMSPGALAIIFAPCLLRCPDNSDPLTSMKDVLK 1830
Query: 307 VMSFLKTLVVKTLREREESIVKSN 330
+ + ++ L+ + +R+ + + + N
Sbjct: 1831 ITTCVEMLIKEQMRKYKMKMEEIN 1854
>RHG6_MOUSE (O54834) Rho-GTPase-activating protein 6 (Rho-type
GTPase-activating protein RhoGAPX-1)
Length = 986
Score = 57.4 bits (137), Expect = 8e-08
Identities = 49/170 (28%), Positives = 79/170 (45%), Gaps = 29/170 (17%)
Query: 148 QLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGV---VPN 204
+LS + VP ++ +HL + GLQ GIFR+ + + +RE+ +RGV +
Sbjct: 401 KLSLNPIYRQVPRLVDSCCQHL-EKHGLQTVGIFRVGSSKKRVRQLREEFDRGVDVCLEE 459
Query: 205 GVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQS-------QSEEE---CDQLVRLLPPT 254
VH +A L+K + R++P DPL E+ + + EE+ L+ LLPP
Sbjct: 460 EHSVHDVAALLKEFLRDMP----DPLLTRELYTAFINTLLLEPEEQLGTLQLLIYLLPPC 515
Query: 255 EAALLDWAINLMADVA-----------QMEHFNKMNARNIAMVFAPNMTH 293
L + ++ VA Q NKM + N+A +F PN+ H
Sbjct: 516 NCDTLHRLLQFLSIVARHADDNVSKDGQEVTGNKMTSLNLATIFGPNLLH 565
>RHG6_HUMAN (O43182) Rho-GTPase-activating protein 6 (Rho-type
GTPase-activating protein RhoGAPX-1)
Length = 974
Score = 57.0 bits (136), Expect = 1e-07
Identities = 48/170 (28%), Positives = 79/170 (46%), Gaps = 29/170 (17%)
Query: 148 QLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGV---VPN 204
+LS + VP ++ +HL + GLQ GIFR+ + + +RE+ +RG+ +
Sbjct: 400 KLSLNPIYRQVPRLVDSCCQHL-EKHGLQTVGIFRVGSSKKRVRQLREEFDRGIDVSLEE 458
Query: 205 GVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQS-------QSEEE---CDQLVRLLPPT 254
VH +A L+K + R++P DPL E+ + + EE+ L+ LLPP
Sbjct: 459 EHSVHDVAALLKEFLRDMP----DPLLTRELYTAFINTLLLEPEEQLGTLQLLIYLLPPC 514
Query: 255 EAALLDWAINLMADVA-----------QMEHFNKMNARNIAMVFAPNMTH 293
L + ++ VA Q NKM + N+A +F PN+ H
Sbjct: 515 NCDTLHRLLQFLSIVARHADDNISKDGQEVTGNKMTSLNLATIFGPNLLH 564
>FNB2_MOUSE (Q91Z67) Formin binding protein 2 (Formin binding
protein 27) (FBP 27) (srGAP2) (Fragment)
Length = 405
Score = 57.0 bits (136), Expect = 1e-07
Identities = 55/235 (23%), Positives = 100/235 (42%), Gaps = 39/235 (16%)
Query: 171 ARGGLQAEGIFRINAENSQEELVREQLNRGVVP-----NGVDVHCLAGLIKAWFRELPTG 225
+R GLQ EGIFR++ + ++ RG P N D+ +AG++K +FR G
Sbjct: 85 SRHGLQHEGIFRVSGSQVEVNDIKNAFERGEDPLAGDQNDHDMDSIAGVLKLYFR----G 140
Query: 226 ILDPLSPEEV---------MQSQSEE--ECDQLVRLLPPTEAALLDWAINLMADVAQMEH 274
+ PL P+++ M + E +++ +LP ++ + + ++Q
Sbjct: 141 LEHPLFPKDIFHDLIACVTMDNLQERVVHIRKVLLVLPKPSLIIMRYLFAFLNHLSQFSE 200
Query: 275 FNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPN 334
N M+ N+A+ F P++ + + + V +KT++++ N PN
Sbjct: 201 ENMMDPYNLAICFGPSLMSVPEGHDQVSCQSHVNELIKTIIIQ----------HENIFPN 250
Query: 335 LNSFDDDGHQSDSQVLPKDGSENGNDCSDED-TVFVSAEPSQPSPTHHT-EDGCE 387
+ + + S +G S ED T V+AE HHT +D CE
Sbjct: 251 PRELEGPIYSRGGSMEDYCDSTHGETTSAEDSTQDVTAE-------HHTSDDECE 298
>FNB2_HUMAN (O75044) Formin binding protein 2 (srGAP2)
Length = 1071
Score = 57.0 bits (136), Expect = 1e-07
Identities = 54/235 (22%), Positives = 101/235 (42%), Gaps = 39/235 (16%)
Query: 171 ARGGLQAEGIFRINAENSQEELVREQLNRGVVP-----NGVDVHCLAGLIKAWFRELPTG 225
+R GLQ EGIFR++ + ++ RG P N D+ +AG++K +FR G
Sbjct: 516 SRHGLQHEGIFRVSGSQVEVNDIKNAFERGEDPLAGDQNDHDMDSIAGVLKLYFR----G 571
Query: 226 ILDPLSPEEV---------MQSQSEEECD--QLVRLLPPTEAALLDWAINLMADVAQMEH 274
+ PL P+++ M + E +++ +LP T ++ + + ++Q
Sbjct: 572 LEHPLFPKDIFHDLMACVTMDNLQERALHIRKVLLVLPKTTLIIMRYLFAFLNHLSQFSE 631
Query: 275 FNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPN 334
N M+ N+A+ F P++ + + + V +KT+++ + E+I S
Sbjct: 632 ENMMDPYNLAICFGPSLMSVPEGHDQVSCQAHVNELIKTIII-----QHENIFPS----- 681
Query: 335 LNSFDDDGHQSDSQVLPKDGS-ENGNDCSDEDTVFVSAEPSQPSPTHHT-EDGCE 387
+ + V + GS E+ D +T V + HHT +D CE
Sbjct: 682 -------PRELEGPVYSRGGSMEDYCDSPHGETTPVEDSTQDVTAEHHTSDDECE 729
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.130 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,781,957
Number of Sequences: 164201
Number of extensions: 2441163
Number of successful extensions: 9487
Number of sequences better than 10.0: 111
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 84
Number of HSP's that attempted gapping in prelim test: 9318
Number of HSP's gapped (non-prelim): 158
length of query: 493
length of database: 59,974,054
effective HSP length: 114
effective length of query: 379
effective length of database: 41,255,140
effective search space: 15635698060
effective search space used: 15635698060
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0010.6


