

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.3.5
(80 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
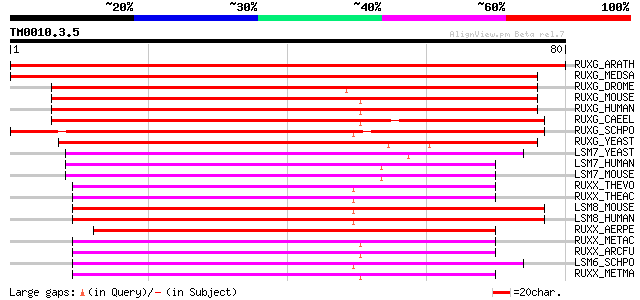
Score E
Sequences producing significant alignments: (bits) Value
RUXG_ARATH (O82221) Probable small nuclear ribonucleoprotein G (... 145 2e-35
RUXG_MEDSA (P24715) Probable small nuclear ribonucleoprotein G (... 138 2e-33
RUXG_DROME (Q9VXE0) Probable small nuclear ribonucleoprotein G (... 92 3e-19
RUXG_MOUSE (P62309) Small nuclear ribonucleoprotein G (snRNP-G) ... 87 6e-18
RUXG_HUMAN (P62308) Small nuclear ribonucleoprotein G (snRNP-G) ... 87 6e-18
RUXG_CAEEL (Q9N4G9) Probable small nuclear ribonucleoprotein G (... 81 4e-16
RUXG_SCHPO (O74966) Probable small nuclear ribonucleoprotein G (... 76 1e-14
RUXG_YEAST (P40204) Small nuclear ribonucleoprotein G (snRNP-G) ... 75 3e-14
LSM7_YEAST (P53905) U6 snRNA-associated Sm-like protein LSm3 60 1e-09
LSM7_HUMAN (Q9UK45) U6 snRNA-associated Sm-like protein LSm7 52 3e-07
LSM7_MOUSE (Q9CQQ8) U6 snRNA-associated Sm-like protein LSm7 51 5e-07
RUXX_THEVO (Q97BU5) Putative snRNP Sm-like protein 48 4e-06
RUXX_THEAC (P57670) Putative snRNP Sm-like protein 48 4e-06
LSM8_MOUSE (Q6ZWM4) U6 snRNA-associated Sm-like protein LSm8 48 5e-06
LSM8_HUMAN (O95777) U6 snRNA-associated Sm-like protein LSm8 48 5e-06
RUXX_AERPE (Q9YEQ5) Putative snRNP Sm-like protein 44 7e-05
RUXX_METAC (Q8TL47) Putative snRNP Sm-like protein 42 3e-04
RUXX_ARCFU (O29386) Putative snRNP Sm-like protein 42 3e-04
LSM6_SCHPO (Q9UUI1) U6 snRNA-associated Sm-like protein LSm6 42 3e-04
RUXX_METMA (Q8PZZ9) Putative snRNP Sm-like protein 42 4e-04
>RUXG_ARATH (O82221) Probable small nuclear ribonucleoprotein G
(snRNP-G) (Sm protein G) (Sm-G) (SmG)
Length = 80
Score = 145 bits (366), Expect = 2e-35
Identities = 70/80 (87%), Positives = 76/80 (94%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANRM+ GTLRG+DQFMN+VVDNTVEVNGN+K DIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRMVTGTLRGFDQFMNLVVDNTVEVNGNDKTDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTRTS 80
VIRGNS+VTVEALEPV R+S
Sbjct: 61 VIRGNSIVTVEALEPVGRSS 80
>RUXG_MEDSA (P24715) Probable small nuclear ribonucleoprotein G
(snRNP-G) (Sm protein G) (Sm-G) (SmG)
Length = 81
Score = 138 bits (348), Expect = 2e-33
Identities = 69/76 (90%), Positives = 71/76 (92%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MS SGQPP LKKYMDK LQI L ANRMIVGTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 1 MSTSGQPPALKKYMDKQLQINLKANRMIVGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 60
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNSVVTVEALEPV
Sbjct: 61 VIRGNSVVTVEALEPV 76
>RUXG_DROME (Q9VXE0) Probable small nuclear ribonucleoprotein G
(snRNP-G) (Sm protein G) (Sm-G) (SmG)
Length = 76
Score = 91.7 bits (226), Expect = 3e-19
Identities = 44/71 (61%), Positives = 56/71 (77%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGN 65
PP++KKYMDK + +KLN R + G LRG+D FMN+V+D+TV E N KN+IGMVVIRGN
Sbjct: 6 PPEVKKYMDKRMMLKLNGGRAVTGILRGFDPFMNVVLDDTVEECKDNTKNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S+V VEAL+ V
Sbjct: 66 SIVMVEALDRV 76
>RUXG_MOUSE (P62309) Small nuclear ribonucleoprotein G (snRNP-G)
(Sm protein G) (Sm-G) (SmG)
Length = 76
Score = 87.4 bits (215), Expect = 6e-18
Identities = 41/71 (57%), Positives = 55/71 (76%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+V+D VE+ ++N+IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVIDECVEMATSGQQNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>RUXG_HUMAN (P62308) Small nuclear ribonucleoprotein G (snRNP-G)
(Sm protein G) (Sm-G) (SmG)
Length = 76
Score = 87.4 bits (215), Expect = 6e-18
Identities = 41/71 (57%), Positives = 55/71 (76%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+V+D VE+ ++N+IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVIDECVEMATSGQQNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>RUXG_CAEEL (Q9N4G9) Probable small nuclear ribonucleoprotein G
(snRNP-G) (Sm protein G) (Sm-G) (SmG)
Length = 77
Score = 81.3 bits (199), Expect = 4e-16
Identities = 41/73 (56%), Positives = 52/73 (71%), Gaps = 3/73 (4%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV--NGNEKNDIGMVVIRG 64
PP+LKKYMDK + +KLN NR + G LRG+D FMNMV+D VE +G N +GM VIRG
Sbjct: 6 PPELKKYMDKEMDLKLNGNRRVSGILRGFDPFMNMVIDEAVEYQKDGGSVN-LGMTVIRG 64
Query: 65 NSVVTVEALEPVT 77
NSVV +E E ++
Sbjct: 65 NSVVIMEPKERIS 77
>RUXG_SCHPO (O74966) Probable small nuclear ribonucleoprotein G
(snRNP-G) (Sm protein G) (Sm-G) (SmG)
Length = 77
Score = 76.3 bits (186), Expect = 1e-14
Identities = 39/79 (49%), Positives = 60/79 (75%), Gaps = 4/79 (5%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE--VNGNEKNDIG 58
MS++G P DLKKY+D+ + ++LN +R + G LRGYD F+N+V+++++E V+G EK IG
Sbjct: 1 MSKAGAP-DLKKYLDRQVFVQLNGSRKVYGVLRGYDIFLNIVLEDSIEEKVDG-EKVKIG 58
Query: 59 MVVIRGNSVVTVEALEPVT 77
V IRGNSV+ +E L+ +T
Sbjct: 59 SVAIRGNSVIMIETLDKMT 77
>RUXG_YEAST (P40204) Small nuclear ribonucleoprotein G (snRNP-G)
(Sm protein G) (Sm-G) (SmG)
Length = 77
Score = 75.1 bits (183), Expect = 3e-14
Identities = 33/73 (45%), Positives = 55/73 (75%), Gaps = 4/73 (5%)
Query: 8 PDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNE---KNDIGM-VVIR 63
P+LKKYMDK + + +N +R + G LRGYD F+N+V+D+ +E+NG + + +G+ VIR
Sbjct: 5 PELKKYMDKKILLNINGSRKVAGILRGYDIFLNVVLDDAMEINGEDPANNHQLGLQTVIR 64
Query: 64 GNSVVTVEALEPV 76
GNS++++EAL+ +
Sbjct: 65 GNSIISLEALDAI 77
>LSM7_YEAST (P53905) U6 snRNA-associated Sm-like protein LSm3
Length = 107
Score = 60.1 bits (144), Expect = 1e-09
Identities = 28/78 (35%), Positives = 47/78 (59%), Gaps = 12/78 (15%)
Query: 9 DLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKND------------ 56
DL KY D +++KL ++++G L+GYDQ MN+V+D+TVE N ++
Sbjct: 21 DLAKYKDSKIRVKLMGGKLVIGVLKGYDQLMNLVLDDTVEYMSNPDDENNTELISKNARK 80
Query: 57 IGMVVIRGNSVVTVEALE 74
+G+ VIRG +V++ + E
Sbjct: 81 LGLTVIRGTILVSLSSAE 98
>LSM7_HUMAN (Q9UK45) U6 snRNA-associated Sm-like protein LSm7
Length = 103
Score = 52.0 bits (123), Expect = 3e-07
Identities = 25/71 (35%), Positives = 40/71 (56%), Gaps = 9/71 (12%)
Query: 9 DLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGN---------EKNDIGM 59
DL KY+DK +++K R G L+G+D +N+V+D T+E + + +G+
Sbjct: 14 DLSKYIDKTIRVKFQGGREASGILKGFDPLLNLVLDGTIEYMRDPDDQYKLTEDTRQLGL 73
Query: 60 VVIRGNSVVTV 70
VV RG SVV +
Sbjct: 74 VVCRGTSVVLI 84
>LSM7_MOUSE (Q9CQQ8) U6 snRNA-associated Sm-like protein LSm7
Length = 103
Score = 51.2 bits (121), Expect = 5e-07
Identities = 25/71 (35%), Positives = 40/71 (56%), Gaps = 9/71 (12%)
Query: 9 DLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGN---------EKNDIGM 59
DL KY+DK +++K R G L+G+D +N+V+D T+E + + +G+
Sbjct: 14 DLSKYIDKTIRVKFQGGREASGILKGFDPLLNLVLDGTMEYMRDPDDQYKLTEDTRQLGL 73
Query: 60 VVIRGNSVVTV 70
VV RG SVV +
Sbjct: 74 VVCRGTSVVLI 84
>RUXX_THEVO (Q97BU5) Putative snRNP Sm-like protein
Length = 83
Score = 48.1 bits (113), Expect = 4e-06
Identities = 23/62 (37%), Positives = 37/62 (59%), Gaps = 1/62 (1%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-VNGNEKNDIGMVVIRGNSVV 68
LK + +N+ I + NR G L GYD +MN+V+ N E +NG K +++RG++V+
Sbjct: 14 LKNALSRNVLIDVKGNREYSGILEGYDVYMNVVLQNASEIINGENKGVFDRILVRGDNVI 73
Query: 69 TV 70
V
Sbjct: 74 FV 75
>RUXX_THEAC (P57670) Putative snRNP Sm-like protein
Length = 83
Score = 48.1 bits (113), Expect = 4e-06
Identities = 24/62 (38%), Positives = 37/62 (58%), Gaps = 1/62 (1%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-VNGNEKNDIGMVVIRGNSVV 68
LK + +N+ I + NR G L GYD +MN+V+ N E +NG K V++RG++V+
Sbjct: 14 LKSALSRNVLIDVKGNREYSGILEGYDVYMNIVLQNASEIINGENKGVYDRVLVRGDNVI 73
Query: 69 TV 70
V
Sbjct: 74 FV 75
>LSM8_MOUSE (Q6ZWM4) U6 snRNA-associated Sm-like protein LSm8
Length = 95
Score = 47.8 bits (112), Expect = 5e-06
Identities = 22/73 (30%), Positives = 46/73 (62%), Gaps = 5/73 (6%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-----VNGNEKNDIGMVVIRG 64
L+ Y+++ + + + RMIVGTL+G+DQ +N+++D + E G E+ +G+ ++RG
Sbjct: 4 LENYINRTVAVITSDGRMIVGTLKGFDQTINLILDESHERVFSSSQGVEQVVLGLYIVRG 63
Query: 65 NSVVTVEALEPVT 77
++V + ++ T
Sbjct: 64 DNVAVIGEIDEET 76
>LSM8_HUMAN (O95777) U6 snRNA-associated Sm-like protein LSm8
Length = 95
Score = 47.8 bits (112), Expect = 5e-06
Identities = 22/73 (30%), Positives = 46/73 (62%), Gaps = 5/73 (6%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-----VNGNEKNDIGMVVIRG 64
L+ Y+++ + + + RMIVGTL+G+DQ +N+++D + E G E+ +G+ ++RG
Sbjct: 4 LENYINRTVAVITSDGRMIVGTLKGFDQTINLILDESHERVFSSSQGVEQVVLGLYIVRG 63
Query: 65 NSVVTVEALEPVT 77
++V + ++ T
Sbjct: 64 DNVAVIGEIDEET 76
>RUXX_AERPE (Q9YEQ5) Putative snRNP Sm-like protein
Length = 77
Score = 43.9 bits (102), Expect = 7e-05
Identities = 18/58 (31%), Positives = 35/58 (60%)
Query: 13 YMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMVVIRGNSVVTV 70
Y+D + +KL + I G L+ YDQ +N+++ + E+ +G+ ++RG+SVV +
Sbjct: 16 YVDTPVLVKLKSGLRIKGVLKTYDQHLNIILGDAEEIGETSIRRLGLTLVRGDSVVVI 73
>RUXX_METAC (Q8TL47) Putative snRNP Sm-like protein
Length = 72
Score = 42.0 bits (97), Expect = 3e-04
Identities = 24/62 (38%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGNSVV 68
L +D + ++L R G L+GYD MN+V+DN E+ +G + VVIRG++VV
Sbjct: 9 LNNALDTPVIVRLKGAREFRGELKGYDIHMNLVLDNAEELRDGEVVSKFSSVVIRGDNVV 68
Query: 69 TV 70
V
Sbjct: 69 YV 70
>RUXX_ARCFU (O29386) Putative snRNP Sm-like protein
Length = 77
Score = 42.0 bits (97), Expect = 3e-04
Identities = 24/62 (38%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGNSVV 68
L + + + ++L R GTL GYD MN+V+ + E+ NG +G VVIRG++VV
Sbjct: 9 LNRSLKSPVIVRLKGGREFRGTLDGYDIHMNLVLLDAEEIQNGEVVRKVGSVVIRGDTVV 68
Query: 69 TV 70
V
Sbjct: 69 FV 70
>LSM6_SCHPO (Q9UUI1) U6 snRNA-associated Sm-like protein LSm6
Length = 75
Score = 42.0 bits (97), Expect = 3e-04
Identities = 25/66 (37%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-VNGNEKNDIGMVVIRGNSVV 68
L K + K + I+L++ G L D +MN+ ++ T E VNG + N G IRGN+V+
Sbjct: 9 LNKVIGKKVLIRLSSGVDYKGILSCLDGYMNLALERTEEYVNGKKTNVYGDAFIRGNNVL 68
Query: 69 TVEALE 74
V AL+
Sbjct: 69 YVSALD 74
>RUXX_METMA (Q8PZZ9) Putative snRNP Sm-like protein
Length = 72
Score = 41.6 bits (96), Expect = 4e-04
Identities = 24/62 (38%), Positives = 35/62 (55%), Gaps = 1/62 (1%)
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGNSVV 68
L +D + ++L R G L+GYD MN+V+DN E+ G + VVIRG++VV
Sbjct: 9 LNNALDTPVIVRLKGAREFRGELKGYDIHMNLVLDNAEELREGEVVSKFSSVVIRGDNVV 68
Query: 69 TV 70
V
Sbjct: 69 YV 70
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.133 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,673,803
Number of Sequences: 164201
Number of extensions: 281543
Number of successful extensions: 632
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 552
Number of HSP's gapped (non-prelim): 75
length of query: 80
length of database: 59,974,054
effective HSP length: 56
effective length of query: 24
effective length of database: 50,778,798
effective search space: 1218691152
effective search space used: 1218691152
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0010.3.5


