

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0009.7
(1310 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
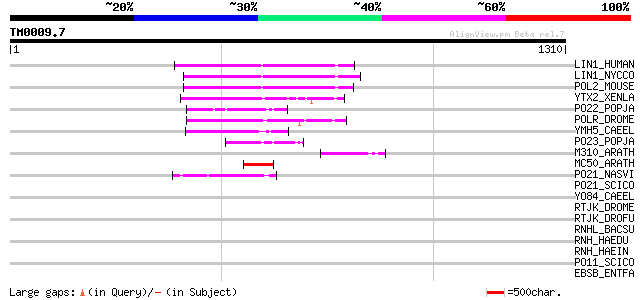
Score E
Sequences producing significant alignments: (bits) Value
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 134 1e-30
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 134 2e-30
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 126 3e-28
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 121 1e-26
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 86 5e-16
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 83 6e-15
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 74 2e-12
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 67 2e-10
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 66 5e-10
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 65 9e-10
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 64 2e-09
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 44 0.002
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 42 0.008
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 42 0.011
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 41 0.019
RNHL_BACSU (P54162) 14.7 kDa ribonuclease H-like protein 41 0.019
RNH_HAEDU (Q7VM15) Ribonuclease HI (EC 3.1.26.4) (RNase HI) 40 0.032
RNH_HAEIN (P43807) Ribonuclease HI (EC 3.1.26.4) (RNase HI) 40 0.042
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 40 0.042
EBSB_ENTFA (P36921) Cell wall enzyme ebsB 39 0.093
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 134 bits (338), Expect = 1e-30
Identities = 108/434 (24%), Positives = 201/434 (45%), Gaps = 18/434 (4%)
Query: 389 PQGLTALI*NSFRNSGAWFEW-MFEDFFSSGRISRGMNAAFITLIPKK-ENASSLNHYRP 446
P+G TA ++ F +F+ G + A I LIPK + + ++RP
Sbjct: 472 PEGFTAEFYQRYKEELVPFLLKLFQSIEKEGILPNSFYEASIILIPKPGRDTTKKENFRP 531
Query: 447 ISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLANEVADFLSK-KSE 505
ISL+ K++ K+LA ++Q+ + KL+ +Q F Q I + + +++ K
Sbjct: 532 ISLMNIDAKILNKILANQIQQHIKKLIHHDQVGFIPAMQGWFNIRKSINIIQHINRTKDT 591
Query: 506 GGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFS 565
+++ +D KA+D++ F++ L+++ ++ +++ + I++NG +
Sbjct: 592 NHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANIILNGQKLEAPP 651
Query: 566 IEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFC 625
++ G RQG PLSPLL N+ L L+ + Q E+ G+++ + + LFADD +++
Sbjct: 652 LKTGTRQGCPLSPLLPNIVLEVLARAIRQ---EKEIKGIQLGKE-EVKLSLFADDMIVYL 707
Query: 626 ENNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISY 685
EN + + L ++ F SG K+N QKS+ ++ +++ + I Y
Sbjct: 708 ENPIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTNNRQTESQIMSELPFTIASKRIKY 767
Query: 686 LGAPLGGNPRRIVFWN--PMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCS--IPLFLMV 741
LG L + + + N P+L ++K T + + S GR+ I K + I F +
Sbjct: 768 LGIQLTRDVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKMAILPKVIYRFNAI 827
Query: 742 VFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLL 801
K P E+EK FIW Q++ I + KAGG+ L + + +
Sbjct: 828 PIKLPMTFFTELEKTTLKFIWNQKRAH-----IAKSTLSQKNKAGGITLPDFKLYYKATV 882
Query: 802 LK--WAWRFGREIN 813
K W W R+I+
Sbjct: 883 TKTAWYWYQNRDID 896
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 134 bits (337), Expect = 2e-30
Identities = 112/427 (26%), Positives = 200/427 (46%), Gaps = 20/427 (4%)
Query: 410 MFEDFFSSGRISRGMNAAFITLIPKK-ENASSLNHYRPISLIRSSYKLIAKVLATRLQKV 468
+F++ G + A ITLIPK ++ + +YRPISL+ K++ K+L R+Q+
Sbjct: 494 LFQNIEKEGILPNTFYEANITLIPKPGKDPTRKENYRPISLMNIDAKILNKILTNRIQQH 553
Query: 469 MPKLLSQNQFAFTKERQIADCILLANEVADFLSK-KSEGGLMLKVDLAKAYDQVDWEFLI 527
+ K++ +Q F Q I + V ++K K++ ++L +D KA+D + F+I
Sbjct: 554 IKKIIHHDQVGFIPGSQGWFNIRKSINVIQHINKLKNKDHMILSIDAEKAFDNIQHPFMI 613
Query: 528 NMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNG 587
L ++ ++ +++ S + I++NG F + G RQG PLSPLLFN+ +
Sbjct: 614 RTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFPLRSGTRQGCPLSPLLFNIVMEV 673
Query: 588 LSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASG 647
L+ + + E+ G+ I I LFADD +++ EN L V+ + SG
Sbjct: 674 LAIAIRE---EKAIKGIHIGSE-EIKLSLFADDMIVYLENTRDSTTKLLEVIKEYSNVSG 729
Query: 648 LKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWN--PMLE 705
K+N KS + + S ++V + YLG L + + + N + +
Sbjct: 730 YKINTHKSVAFIYTNNNQAEKTVKDSIPFTVVPKKMKYLGVYLTKDVKDLYKENYETLRK 789
Query: 706 KLKCKTRSYNSKFMSLSGRLVITKSVLC--SIPLFLMVVFKAPKGVILEVEKLFHAFIWG 763
++ + + S GR+ I K + +I F + KAP ++EK+ FIW
Sbjct: 790 EIAEDVNKWKNIPCSWLGRINIVKMSILPKAIYNFNAIPIKAPLSYFKDLEKIILHFIWN 849
Query: 764 QEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLK--WAWRFGREINSLWRFFVI 821
Q++ + I + KAGG+ L + + +++K W W RE++ +W I
Sbjct: 850 QKKPQ-----IAKTLLSNKNKAGGITLPDLRLYYKSIVIKTAWYWHKNREVD-VWN--RI 901
Query: 822 ERYNLDP 828
E +DP
Sbjct: 902 ENQEMDP 908
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 126 bits (317), Expect = 3e-28
Identities = 101/408 (24%), Positives = 192/408 (46%), Gaps = 15/408 (3%)
Query: 410 MFEDFFSSGRISRGMNAAFITLIPK-KENASSLNHYRPISLIRSSYKLIAKVLATRLQKV 468
+F G + A ITLIPK +++ + + ++RPISL+ K++ K+LA R+Q+
Sbjct: 521 LFHKIEVEGTLPNSFYEATITLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRIQEH 580
Query: 469 MPKLLSQNQFAFTKERQIADCILLANEVADFLSK-KSEGGLMLKVDLAKAYDQVDWEFLI 527
+ ++ +Q F Q I + V +++K K + +++ +D KA+D++ F+I
Sbjct: 581 IKAIIHPDQVGFIPGMQGWFNIRKSINVIHYINKLKDKNHMIISLDAEKAFDKIQHPFMI 640
Query: 528 NMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNG 587
+L +++ +K+ S I VNG + ++ G RQG PLSP LFN+ L
Sbjct: 641 KVLERSGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLFNIVLEV 700
Query: 588 LSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASG 647
L+ + Q ++ G++I + + L ADD +++ + + L N++ +F G
Sbjct: 701 LARAIRQ---QKEIKGIQIGKE-EVKISLLADDMIVYISDPKNSTRELLNLINSFGEVVG 756
Query: 648 LKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWN--PMLE 705
K+N+ KS + + + +S+ T I YLG L + + N + +
Sbjct: 757 YKINSNKSMAFLYTKNKQAEKEIRETTPFSIVTNNIKYLGVTLTKEVKDLYDKNFKSLKK 816
Query: 706 KLKCKTRSYNSKFMSLSGRLVITKSVLC--SIPLFLMVVFKAPKGVILEVEKLFHAFIWG 763
++K R + S GR+ I K + +I F + K P E+E F+W
Sbjct: 817 EIKEDLRRWKDLPCSWIGRINIVKMAILPKAIYRFNAIPIKIPTQFFNELEGAICKFVWN 876
Query: 764 QEQGRKKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGRE 811
++ R I + R +GG+ + ++ R +++K AW + R+
Sbjct: 877 NKKPR-----IAKSLLKDKRTSGGITMPDLKLYYRAIVIKTAWYWYRD 919
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 121 bits (304), Expect = 1e-26
Identities = 96/397 (24%), Positives = 189/397 (47%), Gaps = 29/397 (7%)
Query: 404 GAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLAT 463
G F + + F G + A ++L+PKK + + ++RP+SL+ + YK++AK ++
Sbjct: 485 GPDFHRVLTEAFKKGELPLSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYKIVAKAISL 544
Query: 464 RLQKVMPKLLSQNQFAFTKERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDW 523
RL+ V+ +++ +Q R I D + L ++ F + L +D KA+D+VD
Sbjct: 545 RLKSVLAEVIHPDQSYTVPGRTIFDNVFLIRDLLHFARRTGLSLAFLSLDQEKAFDRVDH 604
Query: 524 EFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNL 583
++LI L +FG +++ ++K+ +SA + +N S T + +G+RQG PLS L++L
Sbjct: 605 QYLIGTLQAYSFGPQFVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSL 664
Query: 584 CLNGLSCLLNQFLGEENYCGVRISR-NLTINHLLFADDTLLFCENNELQLETLCNVLITF 642
+ CLL + L G+ + ++ + +ADD +L + + + LE +
Sbjct: 665 AIEPFLCLLRKRL-----TGLVLKEPDMRVVLSAYADDVILVAQ-DLVDLERAQECQEVY 718
Query: 643 LFASGLKVNNQKSELIGCNVQTVEVERLAHSF---GWSVGTLPISYLGAPLGGNPRRIVF 699
AS ++N KS G +++V+ L +F W + I YLG L +
Sbjct: 719 AAASSARINWSKSS--GLLEGSLKVDFLPPAFRDISWE--SKIIKYLGVYLSAEEYPVS- 773
Query: 700 WNPMLEKLKC------KTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEV 753
+E +C K + + +K +S+ GR ++ ++ S + ++ + I ++
Sbjct: 774 -QNFIELEECVLTRLGKWKGF-AKVLSMRGRALVINQLVASQIWYRLICLSPTQEFIAKI 831
Query: 754 EKLFHAFIWGQEQGRKKINWIGWDQICLNRKAGGLGL 790
++ F+W + +W+ L K GG G+
Sbjct: 832 QRRLLDFLWIGK------HWVSAGVSSLPLKEGGQGV 862
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 86.3 bits (212), Expect = 5e-16
Identities = 66/239 (27%), Positives = 116/239 (47%), Gaps = 9/239 (3%)
Query: 418 GRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQ 477
G + A TLIPK + + +++RPI++ + +L+ ++LA RL+ + +Q
Sbjct: 50 GHVPTPWTAMRTTLIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVELHPAQKG 109
Query: 478 FAFTKERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGD 537
+A + + + +LL ++ ++ ++ +D+ KA+D V + L + +
Sbjct: 110 YARI-DGTLVNSLLLDTYISSRREQRKTYNVV-SLDVRKAFDTVSHSSICRALQRLGIDE 167
Query: 538 KWISWMKSCISSASLAILVN-GSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFL 596
+++ +S ++ I V GS T I +G++QGDPLSP LFN L+ L C L
Sbjct: 168 GTSNYITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDELLCSLQSTP 227
Query: 597 GEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKS 655
G G I L FADD LL E+N++ L T + F G+ +N +KS
Sbjct: 228 GIGGTIGEE-----KIPVLAFADD-LLLLEDNDVLLPTTLATVANFFRLRGMSLNAKKS 280
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 82.8 bits (203), Expect = 6e-15
Identities = 91/390 (23%), Positives = 159/390 (40%), Gaps = 31/390 (7%)
Query: 418 GRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQ 477
G + + A IPK A +RPIS+ + + +LATRL + Q
Sbjct: 389 GNLPHSIRLARTVFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSIN--WDPRQ 446
Query: 478 FAFTKERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGD 537
F AD + + V K + +D++KA+D + + + L
Sbjct: 447 RGFLPTDGCADNATIVDLVLRHSHKHFRSCYIANLDVSKAFDSLSHASIYDTLRAYGAPK 506
Query: 538 KWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLG 597
++ ++++ ++ +G ++ F +G++QGDPLSP+LFNL ++ L L +G
Sbjct: 507 GFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTLPSEIG 566
Query: 598 EENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSEL 657
+ N N FADD +LF E + L+ L + + FL GLK+N K
Sbjct: 567 AK-------VGNAITNAAAFADDLVLFAE-TRMGLQVLLDKTLDFLSIVGLKLNADKCFT 618
Query: 658 IGCNVQTVEVERLAHSFGWSVGTLPI---------SYLGAPLGGNPRRIVFWNPMLEKLK 708
+G Q + + + + VG+ I YLG R V NP E +
Sbjct: 619 VGIKGQPKQKCTVLEAQSFYVGSSEIPSLKRTDEWKYLGINFTATGR--VRCNP-AEDIG 675
Query: 709 CKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGR 768
K + + RL ++VL + + GV+ + +KL ++ R
Sbjct: 676 PKLQRLTKAPLKPQQRLFALRTVLIPQLYHKLALGSVAIGVLRKTDKLIRYYV------R 729
Query: 769 KKINW---IGWDQICLNRKAGGLGLGFIEW 795
+ +N + + K+GGLG+ + W
Sbjct: 730 RWLNLPLDVPIAFVHAPPKSGGLGIPSLRW 759
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 74.3 bits (181), Expect = 2e-12
Identities = 61/245 (24%), Positives = 109/245 (43%), Gaps = 18/245 (7%)
Query: 415 FSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPKLLS 474
FS I A I IPKK N SS ++YRPISL +++ +++ +R++ LLS
Sbjct: 656 FSDSAIPNRWKHAVIIPIPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLS 715
Query: 475 QNQFAFTKERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMN 534
+Q F R ++ + + + K + +L D AKA+D+V L+ L
Sbjct: 716 PHQHGFLNFRSCPSSLVRSISLYHSILKNEKSLDILFFDFAKAFDKVSHPILLKKLALFG 775
Query: 535 FGDKWISWMKSCISSASLAILVNG-SPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLN 593
SW K + + ++ +N ++ + I G+ QG PLLF L +N L
Sbjct: 776 LDKLTCSWFKEFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFILFINDLL---- 831
Query: 594 QFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQ 653
+ + N+ ++ FADD +F +N L+ + ++ + + L +
Sbjct: 832 ----------IDLEPNIHVS--CFADDIKIF-HHNPSTLQNSIDTIVKWSKKNKLPLAPA 878
Query: 654 KSELI 658
KS ++
Sbjct: 879 KSSVL 883
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 67.4 bits (163), Expect = 2e-10
Identities = 53/185 (28%), Positives = 86/185 (45%), Gaps = 13/185 (7%)
Query: 509 MLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEK 568
++ +D+ KA+D V ++ + D ++ S I+ A I+V G T+ I
Sbjct: 25 VVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTNIVVGGRTTNKIYIRN 84
Query: 569 GLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENN 628
G++QGDPLSP+LFN+ L+ L LN + G ++ I L FADD LL E+
Sbjct: 85 GVKQGDPLSPVLFNIVLDELVTRLN-----DEQPGASMTPACKIASLAFADD-LLLLEDR 138
Query: 629 ELQLETLCNVLITFLFASGLKVNNQK-SELIGCNVQTVEVERLAHSFGWSVGTLPISYLG 687
++ + + G+ +N +K + + V V R SF T+ Y+
Sbjct: 139 DIDVPNSLATTCAYFRTRGMTLNPEKCASISAATVSGRSVPRSKPSF-----TIDGRYI- 192
Query: 688 APLGG 692
PLGG
Sbjct: 193 KPLGG 197
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 66.2 bits (160), Expect = 5e-10
Identities = 42/154 (27%), Positives = 76/154 (49%), Gaps = 12/154 (7%)
Query: 734 SIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICLNRK-AGGLGLGF 792
++P++ M F+ K + ++ F W + ++KI+W+ W ++C +++ GGLG
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 793 IEWRNRGLLLKWAWRFGREINSLWRFFVIERYNLDPDSLIWHMASTTPAMGSRAIQDIWR 852
+ W N+ LL K ++R + ++L + RY S++ T P+ R+I
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRY-FPHSSMMECSVGTRPSYAWRSI----- 115
Query: 853 VLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWV 886
++ LS G LL +IGDG K W D W+
Sbjct: 116 -IHGRELLSRG----LLRTIGDGIHTKVWLDRWI 144
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 65.5 bits (158), Expect = 9e-10
Identities = 35/71 (49%), Positives = 44/71 (61%), Gaps = 1/71 (1%)
Query: 552 LAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNL- 610
L ++NG+P + +GLRQGDPLSP LF LC LS L + + G+R+S N
Sbjct: 10 LLFIINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSP 69
Query: 611 TINHLLFADDT 621
INHLLFADDT
Sbjct: 70 RINHLLFADDT 80
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 64.3 bits (155), Expect = 2e-09
Identities = 61/249 (24%), Positives = 107/249 (42%), Gaps = 20/249 (8%)
Query: 385 TAIKPQGLTALI*NSFRNSGAWFEWMFEDFFSSGRISRGMNAAFITLIPKKENASSLNHY 444
TA P G+T NS + +F G+ R + LIPK+ +
Sbjct: 333 TAAGPDGMTTTAWNSIDEC---IKSLFNMIMYHGQCPRRYLDSRTVLIPKEPGTMDPACF 389
Query: 445 RPISLIRSSYKLIAKVLATRLQKVMPKLLSQNQFAFTKERQIADCILLANEVADFLSKKS 504
RP+S+ + + ++LA R+ + LL Q AF +A+ L + + K
Sbjct: 390 RPLSIASVALRHFHRILANRIGE--HGLLDTRQRAFIVADGVAENTSLLSAMIKEARMKI 447
Query: 505 EGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFF 564
+G + +D+ KA+D V+ +++ L + +++ ++ + V + +
Sbjct: 448 KGLYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEVVKTKGRWI 507
Query: 565 SIEKGLRQGDPLSPLLFNLCLNGLSCLLNQ----FLGEENYCGVRISRNLTINHLLFADD 620
+G+RQGDPLSPLLFN ++ + L + +G E I L+FADD
Sbjct: 508 RPARGVRQGDPLSPLLFNCVMDAVLRRLPENTGFLMGAEK-----------IGALVFADD 556
Query: 621 TLLFCENNE 629
+L E E
Sbjct: 557 LVLLAETRE 565
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 44.3 bits (103), Expect = 0.002
Identities = 57/252 (22%), Positives = 108/252 (42%), Gaps = 17/252 (6%)
Query: 410 MFEDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVM 469
++ F GR+ + + PK E +RP+S+ + K+LA R V
Sbjct: 196 LYNIFVFYGRVPSPIKGSRTVFTPKIEGGPDPGVFRPLSICSVILREFNKILARRF--VS 253
Query: 470 PKLLSQNQFAFTK-ERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLIN 528
+ Q A+ + + +L +A+ + E + + +DL KA++ V LI+
Sbjct: 254 CYTYDERQTAYLPIDGVCINVSMLTAIIAEAKRLRKELHIAI-LDLVKAFNSVYHSALID 312
Query: 529 MLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCL-NG 587
+ E + ++ ++ + G + SI G+ QGDPLS LF L
Sbjct: 313 AITEAGCPPGVVDYIADMYNNVITEMQFEGK-CELASILAGVYQGDPLSGPLFTLAYEKA 371
Query: 588 LSCLLNQFLGEENYCGVRISRNLTINHLLFADDTLLFCENNELQL-ETLCNVLITFLFAS 646
L L N+ G + VR++ + + L T++ ++N + ETL +
Sbjct: 372 LRALNNE--GRFDIADVRVNASAYSDDGLLLAMTVIGLQHNLDKFGETLAKI-------- 421
Query: 647 GLKVNNQKSELI 658
GL++N++KS+ +
Sbjct: 422 GLRINSRKSKTV 433
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 42.4 bits (98), Expect = 0.008
Identities = 43/180 (23%), Positives = 79/180 (43%), Gaps = 19/180 (10%)
Query: 512 VDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLR 571
+D +KA+D+V + L++ L + I W+ +++ S + V + ++ G+
Sbjct: 23 LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKVKVGNTLSEPKKTVCGVP 82
Query: 572 QGDPLSPLLFNLCLNGLSCLL------NQFLGEENYCGVRISRNLTINHLLFADDTLLFC 625
QG +SP+LF + +N +S L QF + I+ L D +
Sbjct: 83 QGSVISPVLFGIFVNEISANLPVGVYCKQFADDIKLYAAIPKNQSQISLQLAIDVVTDWA 142
Query: 626 ENNELQLE-------TL--CNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGW 676
NN+L L TL CN T+ F +GL + K E++ ++ + E+L + W
Sbjct: 143 LNNKLLLNPSKTFHLTLGKCNTNYTY-FLNGLPI---KKEVLTRDLGFLVSEKLDFTDHW 198
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 42.0 bits (97), Expect = 0.011
Identities = 39/177 (22%), Positives = 79/177 (44%), Gaps = 4/177 (2%)
Query: 410 MFEDFFSSGRISRGMNAAFITLIPKK-ENASSLNHYRPISLIRSSYKLIAKVLATRLQ-- 466
+F G + +A I +I K + + ++ YRP SL+ S K++ +++ RL
Sbjct: 480 IFNSVLDVGYFPKAWKSASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRLLTC 539
Query: 467 KVMPKLLSQNQFAFTKERQIADCILLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFL 526
K + K + + QF F + + + A + E + +D+ +A+D+V W
Sbjct: 540 KDVTKAIPKFQFGFRLQHGTPEQLHRVVNFALEAMENKEYAVGAFLDIQQAFDRV-WHPG 598
Query: 527 INMLHEMNFGDKWISWMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNL 583
+ + F + +KS + + + V+G + I G+ QG L P L+++
Sbjct: 599 LLYKAKRLFPPQLYLVVKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGPTLYSV 655
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 41.2 bits (95), Expect = 0.019
Identities = 34/161 (21%), Positives = 74/161 (45%), Gaps = 5/161 (3%)
Query: 426 AAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRL--QKVMPKLLSQNQFAFTKE 483
A+ I ++ + ++ YRP SL+ S K++ +++ R+ + + + + + QF F +
Sbjct: 497 ASIIMILKPGKQPLDVDSYRPTSLLPSLGKMLERLILNRILTSEEVTRAIPKFQFGFRLQ 556
Query: 484 RQIADCI-LLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISW 542
+ + + N + L KK G +D+ +A+D+V W + + +
Sbjct: 557 HGTPEQLHRVVNFALEALEKKEYAGSCF-LDIQQAFDRV-WHPGLLYKAKSLLSPQLFQL 614
Query: 543 MKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNL 583
+KS ++ +G + IE G+ QG L P L+++
Sbjct: 615 IKSFWEGRKFSVTADGCRSSVKFIEAGVPQGSVLGPTLYSI 655
>RNHL_BACSU (P54162) 14.7 kDa ribonuclease H-like protein
Length = 132
Score = 41.2 bits (95), Expect = 0.019
Identities = 24/72 (33%), Positives = 38/72 (52%), Gaps = 2/72 (2%)
Query: 1178 KINVDGAAKGSPGRCGIGGILRNNANVILGFFSKSVGEGYAFEAEVLAIHEALLFCVQRQ 1237
+I VDGA+ G+PG GIG +++ FS +G EAE LA+ E + C R
Sbjct: 4 EIYVDGASAGNPGPSGIGIFIKHEGKA--ESFSIPIGVHTNQEAEFLALIEGMKLCATRG 61
Query: 1238 LRNIIIESDSSL 1249
+++ +DS +
Sbjct: 62 YQSVSFRTDSDI 73
>RNH_HAEDU (Q7VM15) Ribonuclease HI (EC 3.1.26.4) (RNase HI)
Length = 153
Score = 40.4 bits (93), Expect = 0.032
Identities = 41/149 (27%), Positives = 67/149 (44%), Gaps = 29/149 (19%)
Query: 1177 IKINVDGAAKGSPGRCGIGGILRNNANVILGFFSKSVGEGYAFEA-----EVLAIHEALL 1231
+ I DG+ G+PG GIG +LR N + K V +GY F+ E+ A+ E L
Sbjct: 4 VNIFTDGSCLGNPGPGGIGVVLRYNQH------QKKVSQGY-FQTTNNRMELRAVIEGL- 55
Query: 1232 FCVQRQLRNIIIESDSSL----AVGWVNNRCNRPWKLLN--KLNQIDRWLVETNCLKVNH 1285
+ ++ N+ + SDS W+ WK N + D WL+ +++++
Sbjct: 56 -SMLKEACNVTLYSDSQYMKNGITKWIFKWKKSNWKTANGKAVKNKDLWLLLDEKIQIHY 114
Query: 1286 I-YREANGEA--------DKLAKRGAEYP 1305
I ++ G + D+LAK GA P
Sbjct: 115 IEWKWVKGHSGHYENEICDELAKLGANNP 143
>RNH_HAEIN (P43807) Ribonuclease HI (EC 3.1.26.4) (RNase HI)
Length = 154
Score = 40.0 bits (92), Expect = 0.042
Identities = 44/149 (29%), Positives = 65/149 (43%), Gaps = 29/149 (19%)
Query: 1177 IKINVDGAAKGSPGRCGIGGILRNNANVILGFFSKSVGEGYAFEA-----EVLAIHEALL 1231
I+I DG+ G+PG GIG +LR + K++ +GY F+ E+ A+ EAL
Sbjct: 5 IEIFTDGSCLGNPGAGGIGAVLRYKQH------EKTLSKGY-FQTTNNRMELRAVIEALN 57
Query: 1232 FCVQRQLRNIIIESDSSL----AVGWVNNRCNRPWKLLN--KLNQIDRWLVETNCL---K 1282
+ L I + SDS W+ N WK + + D W+ + K
Sbjct: 58 TLKEPCL--ITLYSDSQYMKNGITKWIFNWKKNNWKASSGKPVKNQDLWIALDESIQRHK 115
Query: 1283 VN------HIYREANGEADKLAKRGAEYP 1305
+N H N D+LAK+GAE P
Sbjct: 116 INWQWVKGHAGHRENEICDELAKKGAENP 144
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 40.0 bits (92), Expect = 0.042
Identities = 37/148 (25%), Positives = 61/148 (41%), Gaps = 10/148 (6%)
Query: 444 YRPISLIRSSYKLIAKVLATRL-QKVMPKLLSQNQFAFTKERQIADCILLANEVADFLSK 502
YRPI L+ + K++ ++ RL QK+M +S QFAFT + D +
Sbjct: 483 YRPICLLDNLGKVLEGIMVKRLDQKLMDVEVSPYQFAFTYGKSTEDAWRCVQRHVECSEM 542
Query: 503 KSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGD--KWISWMKSCISSASLAILVNGSP 560
K G L +D A+D + W ++ L E + W+S+ V +
Sbjct: 543 KYVIG--LNIDFQGAFDNLGWLSMLLKLDEAQSNEFGLWMSYF-----GGRKVYYVGKTG 595
Query: 561 TDFFSIEKGLRQGDPLSPLLFNLCLNGL 588
+ +G QG P ++ L +N L
Sbjct: 596 IVRKDVTRGCPQGSKSGPAMWKLVMNEL 623
>EBSB_ENTFA (P36921) Cell wall enzyme ebsB
Length = 135
Score = 38.9 bits (89), Expect = 0.093
Identities = 38/141 (26%), Positives = 61/141 (42%), Gaps = 18/141 (12%)
Query: 1176 LIKINVDGAAKGSPGRCGIGGIL------RNNANVILGFFSKSVGEGYAFEAEVLAIHEA 1229
+++I VD A KG+PG G GGI+ R +V LG S EAE + EA
Sbjct: 1 MLRIYVDAATKGNPGESG-GGIVYLTDQSRQQLHVPLGIVSN-------HEAEFKVLIEA 52
Query: 1230 LLFCVQRQ--LRNIIIESDSSLAVGWVNNRCNRPWKLLNKLNQIDRWLVETNCLKVNHIY 1287
L + + + +++ SDS + V + + K L + + L + +
Sbjct: 53 LKKAIANEDNQQTVLLHSDSKIVVQTIEKNYAKNEKYQPYLAEYQQLEKNFPLLLIEWLP 112
Query: 1288 REANGEADKLAKRGAE--YPN 1306
N AD LA++ + YPN
Sbjct: 113 ESQNKAADMLARQALQKFYPN 133
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.335 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 148,977,872
Number of Sequences: 164201
Number of extensions: 6096607
Number of successful extensions: 18929
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 18878
Number of HSP's gapped (non-prelim): 50
length of query: 1310
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1188
effective length of database: 39,941,532
effective search space: 47450540016
effective search space used: 47450540016
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0009.7


