

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0007.1
(254 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
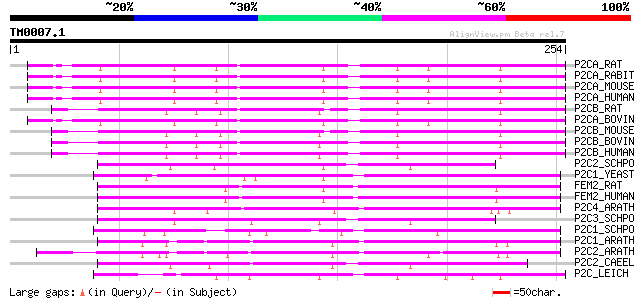
P2CA_RAT (P20650) Protein phosphatase 2C alpha isoform (EC 3.1.3... 124 2e-28
P2CA_RABIT (P35814) Protein phosphatase 2C alpha isoform (EC 3.1... 124 2e-28
P2CA_MOUSE (P49443) Protein phosphatase 2C alpha isoform (EC 3.1... 124 2e-28
P2CA_HUMAN (P35813) Protein phosphatase 2C alpha isoform (EC 3.1... 124 2e-28
P2CB_RAT (P35815) Protein phosphatase 2C beta isoform (EC 3.1.3.... 124 2e-28
P2CA_BOVIN (O62829) Protein phosphatase 2C alpha isoform (EC 3.1... 124 2e-28
P2CB_MOUSE (P36993) Protein phosphatase 2C beta isoform (EC 3.1.... 124 3e-28
P2CB_BOVIN (O62830) Protein phosphatase 2C beta isoform (EC 3.1.... 124 3e-28
P2CB_HUMAN (O75688) Protein phosphatase 2C beta isoform (EC 3.1.... 123 5e-28
P2C2_SCHPO (Q09172) Protein phosphatase 2C homolog 2 (EC 3.1.3.1... 117 2e-26
P2C1_YEAST (P35182) Protein phosphatase 2C homolog 1 (EC 3.1.3.1... 117 3e-26
FEM2_RAT (Q9WVR7) Ca(2+)/calmodulin-dependent protein kinase pho... 115 8e-26
FEM2_HUMAN (P49593) Ca(2+)/calmodulin-dependent protein kinase p... 115 1e-25
P2C4_ARATH (P49598) Protein phosphatase 2C (EC 3.1.3.16) (PP2C) 111 1e-24
P2C3_SCHPO (Q09173) Protein phosphatase 2C homolog 3 (EC 3.1.3.1... 110 4e-24
P2C1_SCHPO (P40371) Protein phosphatase 2C homolog 1 (EC 3.1.3.1... 109 6e-24
P2C1_ARATH (P49597) Protein phosphatase 2C ABI1 (EC 3.1.3.16) (P... 109 7e-24
P2C2_ARATH (O04719) Protein phosphatase 2C ABI2 (EC 3.1.3.16) (P... 108 1e-23
P2C2_CAEEL (P49596) Probable protein phosphatase 2C T23F11.1 (EC... 105 1e-22
P2C_LEICH (P36982) Protein phosphatase 2C (EC 3.1.3.16) (PP2C) 100 6e-21
>P2CA_RAT (P20650) Protein phosphatase 2C alpha isoform (EC
3.1.3.16) (PP2C-alpha) (IA) (Protein phosphatase 1A)
Length = 382
Score = 124 bits (312), Expect = 2e-28
Identities = 93/279 (33%), Positives = 141/279 (50%), Gaps = 44/279 (15%)
Query: 9 HGCYLIQGMMAHGMEDYVFAKHKKLNGYELGL-----YAIFDGHAGRDVAKYLQSHLFEN 63
+G +QG MED H + G GL +A++DGHAG VAKY HL ++
Sbjct: 24 YGLSSMQGWRVE-MED----AHTAVIGLPSGLETWSFFAVYDGHAGSQVAKYCCEHLLDH 78
Query: 64 ILNEPDFWKNP----VHAVKKACKATDEEILENI--------ADSRGGSTAVAAILINGV 111
I N DF + V VK + EI E++ R GSTAV +LI+
Sbjct: 79 ITNNQDFKGSAGAPSVENVKNGIRTGFLEIDEHMRVMSEKKHGADRSGSTAVG-VLISPQ 137
Query: 112 KLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILA 169
N GDSR + C+N T+DH+P E++ +++ GG V+ + RV+G LA
Sbjct: 138 HTYFINCGDSRGLLCRNRKVHFFTQDHKPSNPLEKERIQNAGGSVM-----IQRVNGSLA 192
Query: 170 MTRAFGD---------GNLKDHITAEPDVM-IRKIDEDTEFIILASDGLWKVMANQEACD 219
++RA GD G + ++ EP+V I + +ED +FIILA DG+W VM N+E CD
Sbjct: 193 VSRALGDFDYKCVHGKGPTEQLVSPEPEVHDIERSEEDDQFIILACDGIWDVMGNEELCD 252
Query: 220 CIKD----IEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
++ +D + ++V +GS D++S I++ F
Sbjct: 253 FVRSRLEVTDDLEKVCNEVVDTCLYKGSRDNMSVILICF 291
>P2CA_RABIT (P35814) Protein phosphatase 2C alpha isoform (EC
3.1.3.16) (PP2C-alpha) (IA) (Protein phosphatase 1A)
Length = 382
Score = 124 bits (312), Expect = 2e-28
Identities = 93/279 (33%), Positives = 141/279 (50%), Gaps = 44/279 (15%)
Query: 9 HGCYLIQGMMAHGMEDYVFAKHKKLNGYELGL-----YAIFDGHAGRDVAKYLQSHLFEN 63
+G +QG MED H + G GL +A++DGHAG VAKY HL ++
Sbjct: 24 YGLSSMQGWRVE-MED----AHTAVIGLPSGLETWSFFAVYDGHAGSQVAKYCCEHLLDH 78
Query: 64 ILNEPDFWKNP----VHAVKKACKATDEEILENI--------ADSRGGSTAVAAILINGV 111
I N DF + V VK + EI E++ R GSTAV +LI+
Sbjct: 79 ITNNQDFKGSAGAPSVENVKNGIRTGFLEIDEHMRVMSEKKHGADRSGSTAVG-VLISPQ 137
Query: 112 KLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILA 169
N GDSR + C+N T+DH+P E++ +++ GG V+ + RV+G LA
Sbjct: 138 HTYFINCGDSRGLLCRNRKVHFFTQDHKPSNPLEKERIQNAGGSVM-----IQRVNGSLA 192
Query: 170 MTRAFGD---------GNLKDHITAEPDVM-IRKIDEDTEFIILASDGLWKVMANQEACD 219
++RA GD G + ++ EP+V I + +ED +FIILA DG+W VM N+E CD
Sbjct: 193 VSRALGDFDYKCVHGKGPTEQLVSPEPEVHDIERSEEDDQFIILACDGIWDVMGNEELCD 252
Query: 220 CIKD----IEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
++ +D + ++V +GS D++S I++ F
Sbjct: 253 FVRSRLEVTDDLEKVCNEVVDTCLYKGSRDNMSVILICF 291
>P2CA_MOUSE (P49443) Protein phosphatase 2C alpha isoform (EC
3.1.3.16) (PP2C-alpha) (IA) (Protein phosphatase 1A)
Length = 382
Score = 124 bits (312), Expect = 2e-28
Identities = 93/279 (33%), Positives = 141/279 (50%), Gaps = 44/279 (15%)
Query: 9 HGCYLIQGMMAHGMEDYVFAKHKKLNGYELGL-----YAIFDGHAGRDVAKYLQSHLFEN 63
+G +QG MED H + G GL +A++DGHAG VAKY HL ++
Sbjct: 24 YGLSSMQGWRVE-MED----AHTAVIGLPSGLETWSFFAVYDGHAGSQVAKYCCEHLLDH 78
Query: 64 ILNEPDFWKNP----VHAVKKACKATDEEILENI--------ADSRGGSTAVAAILINGV 111
I N DF + V VK + EI E++ R GSTAV +LI+
Sbjct: 79 ITNNQDFRGSAGAPSVENVKNGIRTGFLEIDEHMRVMSEKKHGADRSGSTAVG-VLISPQ 137
Query: 112 KLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILA 169
N GDSR + C+N T+DH+P E++ +++ GG V+ + RV+G LA
Sbjct: 138 HTYFINCGDSRGLLCRNRKVHFFTQDHKPSNPLEKERIQNAGGSVM-----IQRVNGSLA 192
Query: 170 MTRAFGD---------GNLKDHITAEPDVM-IRKIDEDTEFIILASDGLWKVMANQEACD 219
++RA GD G + ++ EP+V I + +ED +FIILA DG+W VM N+E CD
Sbjct: 193 VSRALGDFDYKCVHGKGPTEQLVSPEPEVHDIERSEEDDQFIILACDGIWDVMGNEELCD 252
Query: 220 CIKD----IEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
++ +D + ++V +GS D++S I++ F
Sbjct: 253 FVRSRLEVTDDLEKVCNEVVDTCLYKGSRDNMSVILICF 291
>P2CA_HUMAN (P35813) Protein phosphatase 2C alpha isoform (EC
3.1.3.16) (PP2C-alpha) (IA) (Protein phosphatase 1A)
Length = 382
Score = 124 bits (312), Expect = 2e-28
Identities = 93/279 (33%), Positives = 141/279 (50%), Gaps = 44/279 (15%)
Query: 9 HGCYLIQGMMAHGMEDYVFAKHKKLNGYELGL-----YAIFDGHAGRDVAKYLQSHLFEN 63
+G +QG MED H + G GL +A++DGHAG VAKY HL ++
Sbjct: 24 YGLSSMQGWRVE-MED----AHTAVIGLPSGLESWSFFAVYDGHAGSQVAKYCCEHLLDH 78
Query: 64 ILNEPDFWKNP----VHAVKKACKATDEEILENI--------ADSRGGSTAVAAILINGV 111
I N DF + V VK + EI E++ R GSTAV +LI+
Sbjct: 79 ITNNQDFKGSAGAPSVENVKNGIRTGFLEIDEHMRVMSEKKHGADRSGSTAVG-VLISPQ 137
Query: 112 KLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILA 169
N GDSR + C+N T+DH+P E++ +++ GG V+ + RV+G LA
Sbjct: 138 HTYFINCGDSRGLLCRNRKVHFFTQDHKPSNPLEKERIQNAGGSVM-----IQRVNGSLA 192
Query: 170 MTRAFGD---------GNLKDHITAEPDVM-IRKIDEDTEFIILASDGLWKVMANQEACD 219
++RA GD G + ++ EP+V I + +ED +FIILA DG+W VM N+E CD
Sbjct: 193 VSRALGDFDYKCVHGKGPTEQLVSPEPEVHDIERSEEDDQFIILACDGIWDVMGNEELCD 252
Query: 220 CIKD----IEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
++ +D + ++V +GS D++S I++ F
Sbjct: 253 FVRSRLEVTDDLEKVCNEVVDTCLYKGSRDNMSVILICF 291
>P2CB_RAT (P35815) Protein phosphatase 2C beta isoform (EC 3.1.3.16)
(PP2C-beta) (IA) (Protein phosphatase 1B)
Length = 390
Score = 124 bits (311), Expect = 2e-28
Identities = 88/269 (32%), Positives = 136/269 (49%), Gaps = 55/269 (20%)
Query: 20 HGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDF--------- 70
HG+ED+ F +A++DGHAG VA Y +HL E+I DF
Sbjct: 48 HGLEDWSF-------------FAVYDGHAGSRVANYCSTHLLEHITTNEDFRAADKSGFA 94
Query: 71 WKNPVHAVKKACKA---TDEEILENIAD-----SRGGSTAVAAILINGVKLLVFNVGDSR 122
+ V VK + +E + N +D R GSTAV ++I+ + N GDSR
Sbjct: 95 LEPSVENVKTGIRTGFLKIDEYMRNFSDLRNGMDRSGSTAVG-VMISPTHIYFINCGDSR 153
Query: 123 AISCKNGVAKQLTEDHEP----EKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGD-- 176
A+ C+NG T+DH+P EKER +++ GG V+ + RV+G LA++RA GD
Sbjct: 154 AVLCRNGQVCFSTQDHKPCNPMEKER--IQNAGGSVM-----IQRVNGSLAVSRALGDYD 206
Query: 177 -------GNLKDHITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKD----IE 225
G + ++ EP+V E+ EF++LA DG+W VM+N+E C+ + +
Sbjct: 207 YKCVDGKGPTEQLVSPEPEVYEILRAEEDEFVVLACDGIWDVMSNEELCEFVNSRLEVSD 266
Query: 226 DAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
D +N +V +GS D++S ++V F
Sbjct: 267 DLENVCNWVVDTCLHKGSRDNMSIVLVCF 295
>P2CA_BOVIN (O62829) Protein phosphatase 2C alpha isoform (EC
3.1.3.16) (PP2C-alpha)
Length = 382
Score = 124 bits (311), Expect = 2e-28
Identities = 93/279 (33%), Positives = 141/279 (50%), Gaps = 44/279 (15%)
Query: 9 HGCYLIQGMMAHGMEDYVFAKHKKLNGYELGL-----YAIFDGHAGRDVAKYLQSHLFEN 63
+G +QG MED H + G GL +A++DGHAG VAKY HL ++
Sbjct: 24 YGLSSMQGWRVE-MED----AHTAVIGLPSGLETWSFFAVYDGHAGSQVAKYCCEHLLDH 78
Query: 64 ILNEPDFWKNP----VHAVKKACKATDEEILENI--------ADSRGGSTAVAAILINGV 111
I N DF + V VK + EI E++ R GSTAV +LI+
Sbjct: 79 ITNNQDFKGSAGAPSVENVKNGIRTGFLEIDEHMRVMSEKKHGADRSGSTAVG-VLISPQ 137
Query: 112 KLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILA 169
N GDSR + C+N T+DH+P E++ +++ GG V+ + RV+G LA
Sbjct: 138 HTYFINCGDSRGLLCRNRKVYFFTQDHKPSNPLEKERIQNAGGSVM-----IQRVNGSLA 192
Query: 170 MTRAFGD---------GNLKDHITAEPDVM-IRKIDEDTEFIILASDGLWKVMANQEACD 219
++RA GD G + ++ EP+V I + +ED +FIILA DG+W VM N+E CD
Sbjct: 193 VSRALGDFDYKCVHGKGPTEQLVSPEPEVHDIERSEEDDQFIILACDGIWDVMGNEELCD 252
Query: 220 CIKD----IEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
++ +D + ++V +GS D++S I++ F
Sbjct: 253 FVRSRLEVTDDLEKVCNEVVDTCLYKGSRDNMSVILICF 291
>P2CB_MOUSE (P36993) Protein phosphatase 2C beta isoform (EC
3.1.3.16) (PP2C-beta) (IA) (Protein phosphatase 1B)
Length = 390
Score = 124 bits (310), Expect = 3e-28
Identities = 86/269 (31%), Positives = 138/269 (50%), Gaps = 55/269 (20%)
Query: 20 HGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDF--------- 70
HG++++ F +A++DGHAG VA Y +HL E+I DF
Sbjct: 48 HGLDNWSF-------------FAVYDGHAGSRVANYCSTHLLEHITTNEDFRAADKSGSA 94
Query: 71 WKNPVHAVKKACKA---TDEEILENIAD-----SRGGSTAVAAILINGVKLLVFNVGDSR 122
+ V +VK + +E + N +D R GSTAV ++++ + N GDSR
Sbjct: 95 LEPSVESVKTGIRTGFLKIDEYMRNFSDLRNGMDRSGSTAVG-VMVSPTHMYFINCGDSR 153
Query: 123 AISCKNGVAKQLTEDHEP----EKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGD-- 176
A+ C+NG T+DH+P EKER +++ GG V+ + RV+G LA++RA GD
Sbjct: 154 AVLCRNGQVCFSTQDHKPCNPVEKER--IQNAGGSVM-----IQRVNGSLAVSRALGDYD 206
Query: 177 -------GNLKDHITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKD----IE 225
G + ++ EP+V E+ EF++LA DG+W VM+N+E C+ +K +
Sbjct: 207 YKCVDGKGPTEQLVSPEPEVYEIVRAEEDEFVVLACDGIWDVMSNEELCEFVKSRLEVSD 266
Query: 226 DAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
D +N +V +GS D++S ++V F
Sbjct: 267 DLENVCNWVVDTCLHKGSRDNMSVVLVCF 295
>P2CB_BOVIN (O62830) Protein phosphatase 2C beta isoform (EC
3.1.3.16) (PP2C-beta)
Length = 387
Score = 124 bits (310), Expect = 3e-28
Identities = 87/267 (32%), Positives = 137/267 (50%), Gaps = 51/267 (19%)
Query: 20 HGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDF--------- 70
HG+ED+ F +A++DGHAG VA Y +HL E+I N DF
Sbjct: 48 HGLEDWSF-------------FAVYDGHAGSRVANYCSTHLLEHITNNEDFRAAGKSGSA 94
Query: 71 WKNPVHAVKKACKA---TDEEILENIAD-----SRGGSTAVAAILINGVKLLVFNVGDSR 122
+ V VK + +E + N +D R GSTAV ++I+ + N GDSR
Sbjct: 95 LEPSVENVKNGIRTGFLKIDEYMRNFSDLRNGMDRSGSTAVG-VMISPKHIYFINCGDSR 153
Query: 123 AISCKNGVAKQLTEDHEP--EKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGD---- 176
A+ ++G T+DH+P +E++ +++ GG V+ + RV+G LA++RA GD
Sbjct: 154 AVLYRSGQVCFSTQDHKPCNPREKERIQNAGGSVM-----IQRVNGSLAVSRALGDYDYK 208
Query: 177 -----GNLKDHITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKD----IEDA 227
G + ++ EP+V E+ EFIILA DG+W VM+N+E C+ +K +D
Sbjct: 209 CVDGKGPTEQLVSPEPEVYEILRAEEDEFIILACDGIWDVMSNEELCEFVKSRLEVSDDL 268
Query: 228 KNAAKKLVKEAKSQGSYDDISCIVVKF 254
+N +V +GS D++S ++V F
Sbjct: 269 ENVCNWVVDTCLHKGSRDNMSIVLVCF 295
>P2CB_HUMAN (O75688) Protein phosphatase 2C beta isoform (EC
3.1.3.16) (PP2C-beta)
Length = 479
Score = 123 bits (308), Expect = 5e-28
Identities = 87/267 (32%), Positives = 136/267 (50%), Gaps = 51/267 (19%)
Query: 20 HGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDF--------- 70
HG+ED+ F +A++DGHAG VA Y +HL E+I DF
Sbjct: 48 HGLEDWSF-------------FAVYDGHAGSRVANYCSTHLLEHITTNEDFRAAGKSGSA 94
Query: 71 WKNPVHAVKKACKA---TDEEILENIAD-----SRGGSTAVAAILINGVKLLVFNVGDSR 122
+ V VK + +E + N +D R GSTAV ++I+ + N GDSR
Sbjct: 95 LELSVENVKNGIRTGFLKIDEYMRNFSDLRNGMDRSGSTAVG-VMISPKHIYFINCGDSR 153
Query: 123 AISCKNGVAKQLTEDHEP--EKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGD---- 176
A+ +NG T+DH+P +E++ +++ GG V+ + RV+G LA++RA GD
Sbjct: 154 AVLYRNGQVCFSTQDHKPCNPREKERIQNAGGSVM-----IQRVNGSLAVSRALGDYDYK 208
Query: 177 -----GNLKDHITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKD----IEDA 227
G + ++ EP+V E+ EFIILA DG+W VM+N+E C+ +K +D
Sbjct: 209 CVDGKGPTEQLVSPEPEVYEILRAEEDEFIILACDGIWDVMSNEELCEYVKSRLEVSDDL 268
Query: 228 KNAAKKLVKEAKSQGSYDDISCIVVKF 254
+N +V +GS D++S ++V F
Sbjct: 269 ENVCNWVVDTCLHKGSRDNMSIVLVCF 295
>P2C2_SCHPO (Q09172) Protein phosphatase 2C homolog 2 (EC 3.1.3.16)
(PP2C-2)
Length = 370
Score = 117 bits (294), Expect = 2e-26
Identities = 68/196 (34%), Positives = 108/196 (54%), Gaps = 19/196 (9%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWK-NPVHAVKKACKATDEEILEN--IADSR 97
+ +FDGH G VAKY + HL + I ++P FWK N A+K A D ++++ + +
Sbjct: 59 FGVFDGHGGDRVAKYCRQHLPDIIKSQPSFWKGNYDEALKSGFLAADNALMQDRDMQEDP 118
Query: 98 GGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVL 155
G TA A++++ + N GDSR + + G A+ L+ DH+P E+ + + GGF+
Sbjct: 119 SGCTATTALIVDHQVIYCANAGDSRTVLGRKGTAEPLSFDHKPNNDVEKARITAAGGFI- 177
Query: 156 NRPGSVPRVDGILAMTRAFGDGNLKDH---------ITAEPDVMIRKIDEDTEFIILASD 206
RV+G LA++RA GD K +TA PDV+I ID D EF+ILA D
Sbjct: 178 ----DFGRVNGSLALSRAIGDFEYKKDSSLPPEKQIVTAFPDVVIHNIDPDDEFLILACD 233
Query: 207 GLWKVMANQEACDCIK 222
G+W ++Q+ + ++
Sbjct: 234 GIWDCKSSQQVVEFVR 249
>P2C1_YEAST (P35182) Protein phosphatase 2C homolog 1 (EC 3.1.3.16)
(PP2C-1)
Length = 281
Score = 117 bits (292), Expect = 3e-26
Identities = 81/235 (34%), Positives = 122/235 (51%), Gaps = 28/235 (11%)
Query: 39 GLYAIFDGHAGRDVAKYLQSHLF----ENILNEPDFWKNPVHAVKKACKATDEEILENIA 94
G +A+FDGHAG +K+ HL +NIL D ++ + + A DEEI +
Sbjct: 52 GYFAVFDGHAGIQASKWCGKHLHTIIEQNIL--ADETRDVRDVLNDSFLAIDEEINTKLV 109
Query: 95 DSRGGSTAVAAI---LINGV------------KLLVFNVGDSRAISCKNGVAKQLTEDHE 139
+ G + AV + L + V KL NVGDSR + +NG + +LT DH+
Sbjct: 110 GNSGCTAAVCVLRWELPDSVSDDSMDLAQHQRKLYTANVGDSRIVLFRNGNSIRLTYDHK 169
Query: 140 PEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDED 197
E VE GG ++ RV+G+LA+TR+ GD + P +I +
Sbjct: 170 ASDTLEMQRVEQAGGLIMKS-----RVNGMLAVTRSLGDKFFDSLVVGSPFTTSVEITSE 224
Query: 198 TEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGSYDDISCIVV 252
+F+ILA DGLW V+ +Q+AC+ IKDI + AAK LV+ A G+ D+++ +VV
Sbjct: 225 DKFLILACDGLWDVIDDQDACELIKDITEPNEAAKVLVRYALENGTTDNVTVMVV 279
>FEM2_RAT (Q9WVR7) Ca(2+)/calmodulin-dependent protein kinase
phosphatase (EC 3.1.3.16) (CaM-kinase phosphatase)
(CaMKPase) (Partner of PIX 2) (Protein phosphatase 1F)
Length = 450
Score = 115 bits (289), Expect = 8e-26
Identities = 74/221 (33%), Positives = 123/221 (55%), Gaps = 12/221 (5%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNPVHAVKKACKATDEEILENIADSR--G 98
+A+FDGH G D A+Y H+ N ++P+ +P A+K+A + TD+ L+ R
Sbjct: 190 FAVFDGHGGVDAARYASVHVHTNASHQPELLTDPAAALKEAFRHTDQMFLQKAKRERLQS 249
Query: 99 GSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLN 156
G+T V A LI G L V +GDS+ I + G +L E H+PE+ E+ +E+ GGFV
Sbjct: 250 GTTGVCA-LITGAALHVAWLGDSQVILVQQGQVVKLMEPHKPERQDEKSRIEALGGFVSL 308
Query: 157 RPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDEDTEFIILASDGLWKVMANQE 216
RV+G LA++RA GD K +++ E D R++ ++++LA DG + V+ + E
Sbjct: 309 M--DCWRVNGTLAVSRAIGDVFQKPYVSGEADAASRELTGLEDYLLLACDGFFDVVPHHE 366
Query: 217 ACDCI-----KDIEDAKNAAKKLVKEAKSQGSYDDISCIVV 252
+ + + A++LV A+ +GS+D+I+ +VV
Sbjct: 367 IPGLVHGHLLRQKGSGMHVAEELVAVARDRGSHDNITVMVV 407
>FEM2_HUMAN (P49593) Ca(2+)/calmodulin-dependent protein kinase
phosphatase (EC 3.1.3.16) (CaM-kinase phosphatase)
(CaMKPase) (Partner of PIX 2) (hFEM-2) (Protein
phosphatase 1F)
Length = 454
Score = 115 bits (288), Expect = 1e-25
Identities = 74/221 (33%), Positives = 120/221 (53%), Gaps = 12/221 (5%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNPVHAVKKACKATDEEILENIADSR--G 98
+A+FDGH G D A+Y H+ N +P+ +P A+++A + TD+ L R
Sbjct: 194 FAVFDGHGGVDAARYAAVHVHTNAARQPELPTDPEGALREAFRRTDQMFLRKAKRERLQS 253
Query: 99 GSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLN 156
G+T V A LI G L V +GDS+ I + G +L E H PE+ E+ +E+ GGFV +
Sbjct: 254 GTTGVCA-LIAGATLHVAWLGDSQVILVQQGQVVKLMEPHRPERQDEKARIEALGGFVSH 312
Query: 157 RPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDEDTEFIILASDGLWKVMANQE 216
RV+G LA++RA GD K +++ E D R + ++++LA DG + V+ +QE
Sbjct: 313 M--DCWRVNGTLAVSRAIGDVFQKPYVSGEADAASRALTGSEDYLLLACDGFFDVVPHQE 370
Query: 217 ACDCI-----KDIEDAKNAAKKLVKEAKSQGSYDDISCIVV 252
+ + A++LV A+ +GS+D+I+ +VV
Sbjct: 371 VVGLVQSHLTRQQGSGLRVAEELVAAARERGSHDNITVMVV 411
>P2C4_ARATH (P49598) Protein phosphatase 2C (EC 3.1.3.16) (PP2C)
Length = 399
Score = 111 bits (278), Expect = 1e-24
Identities = 81/253 (32%), Positives = 131/253 (51%), Gaps = 44/253 (17%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNP--VHAVKKACKATDEEI--------- 89
Y +FDGH VA+ + L + + E + + + K+ + D+E+
Sbjct: 138 YGVFDGHGCSHVAEKCRERLHDIVKKEVEVMASDEWTETMVKSFQKMDKEVSQRECNLVV 197
Query: 90 --------------LENIADSRGGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLT 135
L++ GSTAV ++ + K++V N GDSRA+ C+NGVA L+
Sbjct: 198 NGATRSMKNSCRCELQSPQCDAVGSTAVVSV-VTPEKIIVSNCGDSRAVLCRNGVAIPLS 256
Query: 136 EDHEPEKERDLV--ESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRK 193
DH+P++ +L+ + GG V+ G+ RV G+LAM+RA GD LK ++ +P+V +
Sbjct: 257 VDHKPDRPDELIRIQQAGGRVIYWDGA--RVLGVLAMSRAIGDNYLKPYVIPDPEVTVTD 314
Query: 194 IDEDTEFIILASDGLWKVMANQEACD----CIK-----DIEDA-----KNAAKKLVKEAK 239
++ E +ILASDGLW V+ N+ AC C++ D DA +AA L K A
Sbjct: 315 RTDEDECLILASDGLWDVVPNETACGVARMCLRGAGAGDDSDAAHNACSDAALLLTKLAL 374
Query: 240 SQGSYDDISCIVV 252
++ S D++S +VV
Sbjct: 375 ARQSSDNVSVVVV 387
>P2C3_SCHPO (Q09173) Protein phosphatase 2C homolog 3 (EC 3.1.3.16)
(PP2C-3)
Length = 414
Score = 110 bits (274), Expect = 4e-24
Identities = 68/194 (35%), Positives = 108/194 (55%), Gaps = 17/194 (8%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNP-VHAVKKACKATDEEILENIADSRGG 99
+A++DGH G VAK+ S+L + + PDF K V+A+K + D+ IL++
Sbjct: 58 FAVYDGHGGDKVAKWCGSNLPQILEKNPDFQKGDFVNALKSSFLNADKAILDDDQFHTDP 117
Query: 100 STAVAAILIN-GVKLLVFNVGDSRAISCKNGVAKQLTEDHEP--EKERDLVESRGGFVLN 156
S A +++ G KL N GDSR + G+AK L+ DH+P E E+ + + GGFV
Sbjct: 118 SGCTATVVLRVGNKLYCANAGDSRTVLGSKGIAKPLSADHKPSNEAEKARICAAGGFV-- 175
Query: 157 RPGSVPRVDGILAMTRAFGDGNLKDH--------ITAEPDVMIRKIDEDTEFIILASDGL 208
RV+G LA++RA GD K+ +TA PDV++ +I +D EF++LA DG+
Sbjct: 176 ---DFGRVNGNLALSRAIGDFEFKNSNLEPEKQIVTALPDVVVHEITDDDEFVVLACDGI 232
Query: 209 WKVMANQEACDCIK 222
W +Q+ + ++
Sbjct: 233 WDCKTSQQVIEFVR 246
>P2C1_SCHPO (P40371) Protein phosphatase 2C homolog 1 (EC 3.1.3.16)
(PP2C-1)
Length = 347
Score = 109 bits (273), Expect = 6e-24
Identities = 82/235 (34%), Positives = 123/235 (51%), Gaps = 37/235 (15%)
Query: 39 GLYAIFDGHAGRDVAKYLQSHL----FENILNEPD------FWKNPVHAVKKACKATDEE 88
G A++DGHAG + Y Q +L E + NEPD + V K KAT +
Sbjct: 103 GFVAVYDGHAGIQASDYCQKNLHKVLLEKVRNEPDRLVTDLMDETFVEVNSKIAKATHND 162
Query: 89 ILENIADSRGGSTAVAAILI---NGVKLLVF--NVGDSRAISCKNGVAKQLTEDHEPEKE 143
I G TA A N + +++ N GD+R + C++G A +L+ DH K
Sbjct: 163 IC--------GCTAAVAFFRYEKNRTRRVLYTANAGDARIVLCRDGKAIRLSYDH---KG 211
Query: 144 RDLVESR-----GGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKI-DED 197
D ESR GG ++ R++G+LA+TRA GD LK+ ++A P +I +
Sbjct: 212 SDANESRRVTQLGGLMVQN-----RINGVLAVTRALGDTYLKELVSAHPFTTETRIWNGH 266
Query: 198 TEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGSYDDISCIVV 252
EF I+A DGLW V+++QEA D +++ + AA +LV+ A + S D+I+CIVV
Sbjct: 267 DEFFIIACDGLWDVVSDQEAVDFVRNFVSPREAAVRLVEFALKRLSTDNITCIVV 321
>P2C1_ARATH (P49597) Protein phosphatase 2C ABI1 (EC 3.1.3.16)
(PP2C) (Abscisic acid-insensitive 1)
Length = 434
Score = 109 bits (272), Expect = 7e-24
Identities = 85/255 (33%), Positives = 128/255 (49%), Gaps = 50/255 (19%)
Query: 41 YAIFDGHAGRDVAKYLQSH----LFENILNEPDF----------WKNPVHAVKKACKATD 86
+ ++DGH G VA Y + L E I E WK A+ + D
Sbjct: 173 FGVYDGHGGSQVANYCRERMHLALAEEIAKEKPMLCDGDTWLEKWKK---ALFNSFLRVD 229
Query: 87 EEILENIADSRGGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEPEKERDL 146
EI E++A GST+V A++ + V N GDSRA+ C+ A L+ DH+P++E +
Sbjct: 230 SEI-ESVAPETVGSTSVVAVVFPS-HIFVANCGDSRAVLCRGKTALPLSVDHKPDREDEA 287
Query: 147 --VESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDEDTEFIILA 204
+E+ GG V+ G+ RV G+LAM+R+ GD LK I +P+V K ++ + +ILA
Sbjct: 288 ARIEAAGGKVIQWNGA--RVFGVLAMSRSIGDRYLKPSIIPDPEVTAVKRVKEDDCLILA 345
Query: 205 SDGLWKVMANQEACDCI-------------------------KDIED--AKNAAKKLVKE 237
SDG+W VM ++EAC+ K+ +D A +AA+ L K
Sbjct: 346 SDGVWDVMTDEEACEMARKRILLWHKKNAVAGDASLLADERRKEGKDPAAMSAAEYLSKL 405
Query: 238 AKSQGSYDDISCIVV 252
A +GS D+IS +VV
Sbjct: 406 AIQRGSKDNISVVVV 420
>P2C2_ARATH (O04719) Protein phosphatase 2C ABI2 (EC 3.1.3.16)
(PP2C) (Abscisic acid-insensitive 2)
Length = 423
Score = 108 bits (270), Expect = 1e-23
Identities = 91/285 (31%), Positives = 142/285 (48%), Gaps = 63/285 (22%)
Query: 13 LIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSH----LFENILNE- 67
L+ G + +G ++ A + ++DGH G VA Y + L E I+ E
Sbjct: 143 LLDGRVTNGFNPHLSAH----------FFGVYDGHGGSQVANYCRERMHLALTEEIVKEK 192
Query: 68 PDF---------WKNPVHAVKKACKATDEEILENIADSRG--GSTAVAAILINGVKLLVF 116
P+F WK A+ + D EI E +A + GST+V A++ + V
Sbjct: 193 PEFCDGDTWQEKWKK---ALFNSFMRVDSEI-ETVAHAPETVGSTSVVAVVFP-THIFVA 247
Query: 117 NVGDSRAISCKNGVAKQLTEDHEPEKERDL--VESRGGFVLNRPGSVPRVDGILAMTRAF 174
N GDSRA+ C+ L+ DH+P+++ + +E+ GG V+ G+ RV G+LAM+R+
Sbjct: 248 NCGDSRAVLCRGKTPLALSVDHKPDRDDEAARIEAAGGKVIRWNGA--RVFGVLAMSRSI 305
Query: 175 GDGNLKDHITAEPDVM-IRKIDEDTEFIILASDGLWKVMANQEACDCIK----------- 222
GD LK + +P+V +R++ ED + +ILASDGLW VM N+E CD +
Sbjct: 306 GDRYLKPSVIPDPEVTSVRRVKED-DCLILASDGLWDVMTNEEVCDLARKRILLWHKKNA 364
Query: 223 -------------DIED--AKNAAKKLVKEAKSQGSYDDISCIVV 252
+ +D A +AA+ L K A +GS D+IS +VV
Sbjct: 365 MAGEALLPAEKRGEGKDPAAMSAAEYLSKMALQKGSKDNISVVVV 409
>P2C2_CAEEL (P49596) Probable protein phosphatase 2C T23F11.1 (EC
3.1.3.16) (PP2C)
Length = 356
Score = 105 bits (262), Expect = 1e-22
Identities = 70/211 (33%), Positives = 114/211 (53%), Gaps = 20/211 (9%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWK-NPVHAVKKACKATDEEIL--ENIADSR 97
+A++DGH G V++Y +L + ++ + +F + N A++K D+++ E D
Sbjct: 55 FAVYDGHGGSKVSQYSGINLHKKVVAQKEFSEGNMKEAIEKGFLELDQQMRVDEETKDDV 114
Query: 98 GGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEPEKERDL--VESRGGFVL 155
G+TAV ++ G + N GDSRA+S G A+ L+ DH+P E + + + GG+V
Sbjct: 115 SGTTAVVVLIKEG-DVYCGNAGDSRAVSSVVGEARPLSFDHKPSHETEARRIIAAGGWV- 172
Query: 156 NRPGSVPRVDGILAMTRAFGDGNLKDH---------ITAEPDVMIRKIDEDTEFIILASD 206
RV+G LA++RA GD K+ +TA PDV+ K+ D EFI+LA D
Sbjct: 173 ----EFNRVNGNLALSRALGDFAFKNCDTKPAEEQIVTAFPDVITDKLTPDHEFIVLACD 228
Query: 207 GLWKVMANQEACDCIKDIEDAKNAAKKLVKE 237
G+W VM NQE D +++ K + + +E
Sbjct: 229 GIWDVMTNQEVVDFVREKLAEKRDPQSICEE 259
>P2C_LEICH (P36982) Protein phosphatase 2C (EC 3.1.3.16) (PP2C)
Length = 406
Score = 99.8 bits (247), Expect = 6e-21
Identities = 78/244 (31%), Positives = 120/244 (48%), Gaps = 46/244 (18%)
Query: 39 GLYAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNPVHAVKKACKATDEEILENI----- 93
G + +FDGH ++YL+ W++ + K++ TDE + E
Sbjct: 49 GFFGVFDGHVNDQCSQYLERA-----------WRSAIE--KESIPMTDERMKELALRIDQ 95
Query: 94 ----ADSRGGSTAVAAILI---NGVKLLVFNVGDSRAISCKNGVAKQLTEDHEP--EKER 144
+ GGST + + N V L V NVGDSR ++C +GV LTEDH+P E ER
Sbjct: 96 EWMDSGREGGSTGTFFVALKEGNKVHLQVGNVGDSRVVACIDGVCVPLTEDHKPNNEGER 155
Query: 145 DLVESRGGFVLNRPGSVPRVDGILAMTRAFGD--------GNLKDHITAEPDVMIRKIDE 196
+E+ G V N RVDG LA++RAFGD L+ + A DV +
Sbjct: 156 QRIENCAGRVENN-----RVDGSLAVSRAFGDREYKLGSGSQLEQKVIALADVQHKDFTF 210
Query: 197 DT-EFIILASDGLWK-VMANQEACDCIKD----IEDAKNAAKKLVKEAKSQGSYDDISCI 250
D+ +F++L DG+++ N+E +K D A ++ +EA +GS D+ISC+
Sbjct: 211 DSNDFVLLCCDGVFEGNFPNEEVVAYVKQQLETCNDLAEVAGRVCEEAIERGSRDNISCM 270
Query: 251 VVKF 254
+V+F
Sbjct: 271 IVQF 274
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,234,781
Number of Sequences: 164201
Number of extensions: 1232337
Number of successful extensions: 3306
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 3137
Number of HSP's gapped (non-prelim): 86
length of query: 254
length of database: 59,974,054
effective HSP length: 108
effective length of query: 146
effective length of database: 42,240,346
effective search space: 6167090516
effective search space used: 6167090516
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0007.1


