

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0006.8
(1142 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
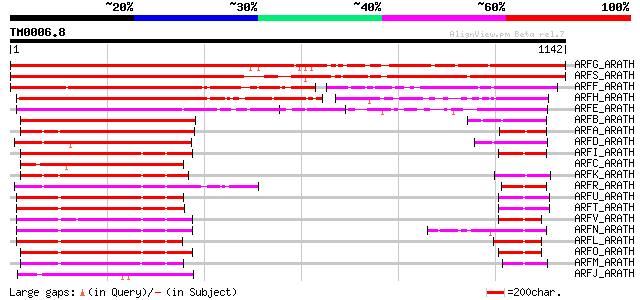
Score E
Sequences producing significant alignments: (bits) Value
ARFG_ARATH (P93022) Auxin response factor 7 (Non-phototropic hyp... 1354 0.0
ARFS_ARATH (Q8RYC8) Auxin response factor 19 (Auxin-responsive p... 1352 0.0
ARFF_ARATH (Q9ZTX8) Auxin response factor 6 630 e-180
ARFH_ARATH (Q9FGV1) Auxin response factor 8 561 e-159
ARFE_ARATH (P93024) Auxin response factor 5 (Transcription facto... 509 e-143
ARFB_ARATH (Q94JM3) Auxin response factor 2 (ARF1-binding protei... 387 e-107
ARFA_ARATH (Q8L7G0) Auxin response factor 1 380 e-104
ARFD_ARATH (Q9ZTX9) Auxin response factor 4 353 2e-96
ARFI_ARATH (Q9XED8) Auxin response factor 9 341 8e-93
ARFC_ARATH (O23661) Auxin response factor 3 (ETTIN protein) 337 1e-91
ARFK_ARATH (Q9ZPY6) Auxin response factor 11 333 1e-90
ARFR_ARATH (Q9C5W9) Auxin response factor 18 332 4e-90
ARFU_ARATH (Q9C8N9) Putative auxin response factor 21 298 6e-80
ARFT_ARATH (Q9C7I9) Putative auxin response factor 20 295 5e-79
ARFV_ARATH (Q9C8N7) Putative auxin response factor 22 294 8e-79
ARFN_ARATH (Q9LQE8) Putative auxin response factor 14 294 8e-79
ARFL_ARATH (Q9XID4) Putative auxin response factor 12 291 5e-78
ARFO_ARATH (Q9LQE3) Putative auxin response factor 15 288 6e-77
ARFM_ARATH (Q9FX25) Putative auxin response factor 13 276 2e-73
ARFJ_ARATH (Q9SKN5) Auxin response factor 10 272 4e-72
>ARFG_ARATH (P93022) Auxin response factor 7 (Non-phototropic
hypocotyl 4) (BIPOSTO protein) (Auxin-responsive protein
IAA21/IAA23/IAA25)
Length = 1164
Score = 1354 bits (3505), Expect = 0.0
Identities = 770/1214 (63%), Positives = 869/1214 (71%), Gaps = 122/1214 (10%)
Query: 1 MKAPS-NCYLPNSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQK 59
MKAPS N PN EGER+ INSELWHACAGPL+SLPP GSLVVYFPQGHSEQVAASMQK
Sbjct: 1 MKAPSSNGVSPNPVEGERRNINSELWHACAGPLISLPPAGSLVVYFPQGHSEQVAASMQK 60
Query: 60 ETDFIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQN 119
+TDFIPSYPNLPSKLICMLHNV L+ADPETDEVYAQMTLQPVNKYD+DA+LASDMGLK N
Sbjct: 61 QTDFIPSYPNLPSKLICMLHNVTLNADPETDEVYAQMTLQPVNKYDRDALLASDMGLKLN 120
Query: 120 RQPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFR 179
RQP EFFCKTLTASDTSTHGGFSVPRRAAEKIFP LD+SMQPP QELVAKD+HD TW+FR
Sbjct: 121 RQPNEFFCKTLTASDTSTHGGFSVPRRAAEKIFPALDFSMQPPCQELVAKDIHDNTWTFR 180
Query: 180 HIYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVI 239
HIYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRD K QLLLGIRRANRQQPALSSSVI
Sbjct: 181 HIYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDGKAQLLLGIRRANRQQPALSSSVI 240
Query: 240 SSDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMM 299
SSDSMHIG+LAAAAHA ANNSPFTIFYNPRA+P+EFV+PLAK+ KAMY QVSLGMRFRM+
Sbjct: 241 SSDSMHIGVLAAAAHANANNSPFTIFYNPRAAPAEFVVPLAKYTKAMYAQVSLGMRFRMI 300
Query: 300 FETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVT 359
FETEE GVRR+MGT+TGISDLD RWK+SQWRNLQ+GWDES AG+RPSRVS+W+IEPV+T
Sbjct: 301 FETEECGVRRYMGTVTGISDLDPVRWKNSQWRNLQIGWDESAAGDRPSRVSVWDIEPVLT 360
Query: 360 PFYICPPPFFRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFGMKDASGSIFPGLSLVQW 419
PFYICPPPFFRP+F QPGMPDDE+D+E+A KRAMPWL + MKD S +IFPGLSLVQW
Sbjct: 361 PFYICPPPFFRPRFSGQPGMPDDETDMESALKRAMPWLDNSLEMKDPSSTIFPGLSLVQW 420
Query: 420 MSMQQNN---QFSAAQSGCFPPMLSPN-TLHSNL-GTDDPSKLLSFQAP--ALSAPSLQF 472
M+MQQ N +AAQ G FP MLSP LH+NL GTDDPSKLLSFQ P +S+ +LQF
Sbjct: 421 MNMQQQNGQLPSAAAQPGFFPSMLSPTAALHNNLGGTDDPSKLLSFQTPHGGISSSNLQF 480
Query: 473 NKPNLPNQINQLHQSPVSWPQ--------------QQQQQQQQQQQQQQQL--------- 509
NK N ++QL Q P + Q QQQQ Q QQQQQQQQL
Sbjct: 481 NKQNQQAPMSQLPQPPTTLSQQQQLQQLLHSSLNHQQQQSQSQQQQQQQQLLQQQQQLQS 540
Query: 510 -------QLQHQQQQQQLQQQQQQQQQQQLQQQQQQQLQQQQSQQQQLQSLLQIPMNQLQ 562
Q Q QQQQQ LQQQQQQQ QQQ QQ QQQ QQQQ + Q LQS QLQ
Sbjct: 541 QQHSNNNQSQSQQQQQLLQQQQQQQLQQQHQQPLQQQTQQQQLRTQPLQSHSHPQPQQLQ 600
Query: 563 QPRQQQLPEPKNLPLPQQQLPQQLGQQSQKAI-------MSNGVVASNQIT------NQL 609
Q + QQL P+N QQ QQ QSQ+A + +G +AS+ IT NQ
Sbjct: 601 QHKLQQLQVPQNQLYNGQQAAQQ--HQSQQASTHHLQPQLVSGSMASSVITPPSSSLNQS 658
Query: 610 VLQQQQLLS---------GGISPQQSIHSASKTTFPLTSL-PQESQFLQQIDQQASLLQR 659
QQQQ G + Q S+ SK++ L S PQE+QF +Q++QQ
Sbjct: 659 FQQQQQQSKQLQQAHHHLGASTSQSSVIETSKSSSNLMSAPPQETQFSRQVEQQ------ 712
Query: 660 QQLPQQTQLQQSSLHLLQQNQPQRAPQQPPVATMSQQTPSEQ-QLHLQLLQKLQQHQQHQ 718
Q Q QQ+ LLQQ Q QQ + QQ+ EQ + QLLQ+LQQ QQ Q
Sbjct: 713 QPPGLNGQNQQT---LLQQKAHQAQAQQ-----IFQQSLLEQPHIQFQLLQRLQQ-QQQQ 763
Query: 719 QLLSTSSPLLQPQLLQQQSTPQQNQQLTQLP--ASQHQ-TQQLGNNAFSTEKVLNSNSFS 775
Q LS S L QL Q+QQL QLP + HQ NN ST
Sbjct: 764 QFLSPQSQLPHHQL--------QSQQLQQLPTLSQGHQFPSSCTNNGLST---------- 805
Query: 776 SSLMQPHQLPVNHPQNTQKSL--AITRAPSTLTD-GDAPSCSTSPSTNNCQISPSNLLKR 832
+QP Q+ V+ PQ Q +A S +TD GDAPS STSPSTNNCQIS S L R
Sbjct: 806 ---LQPPQMLVSRPQEKQNPPVGGGVKAYSGITDGGDAPSSSTSPSTNNCQISSSGFLNR 862
Query: 833 NQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTITD-QMEASS 891
+Q PA L ++ + NL+Q+L SKSDM++K E Q K K ++TD Q+EAS+
Sbjct: 863 SQSGPAILIPDAAIDMSGNLVQDLYSKSDMRLKQEL-----VGQQKSKASLTDHQLEASA 917
Query: 892 SGTSYCLDPG--NVQQSLPLSNFCMEGDVQSHSRSNLPFDSNLD-GLTPDPVLMRGYDSQ 948
SGTSY LD G N QQ+ F ++GD SR++L +N+D G PD +L RGYDSQ
Sbjct: 918 SGTSYGLDGGENNRQQNFLAPTFGLDGD----SRNSLLGGANVDNGFVPDTLLSRGYDSQ 973
Query: 949 KDLQNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGVLNNGLRA 1008
KDLQNMLSNYGG DI TE+ST+ + +QSFGVPN+ P S D+ +ND GVL GL
Sbjct: 974 KDLQNMLSNYGGVTNDIGTEMSTSAVRTQSFGVPNV---PAISNDLAVNDAGVLGGGLWP 1030
Query: 1009 NQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYV 1068
QTQRMRTYTKVQKRGSVGR IDV RY+GYDELR+DLARMFGIEGQLEDPQ +DWKLVYV
Sbjct: 1031 AQTQRMRTYTKVQKRGSVGRSIDVNRYRGYDELRHDLARMFGIEGQLEDPQTSDWKLVYV 1090
Query: 1069 DHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLGNVPIPNQACSGTDSGNAW 1128
DHENDILLVGDDPWEEFV+CVQSIKILS EVQQMSLDG+ VP+ NQACSG DSGNAW
Sbjct: 1091 DHENDILLVGDDPWEEFVNCVQSIKILSSAEVQQMSLDGNFAGVPVTNQACSGGDSGNAW 1150
Query: 1129 RGPYDDNSAASFNR 1142
RG YDDNSA SFNR
Sbjct: 1151 RGHYDDNSATSFNR 1164
>ARFS_ARATH (Q8RYC8) Auxin response factor 19 (Auxin-responsive
protein IAA22)
Length = 1086
Score = 1352 bits (3500), Expect = 0.0
Identities = 736/1176 (62%), Positives = 851/1176 (71%), Gaps = 124/1176 (10%)
Query: 1 MKAPSNCYLPNSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKE 60
MKAPSN +LP+S EGE+K INS+LWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQK+
Sbjct: 1 MKAPSNGFLPSSNEGEKKPINSQLWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKQ 60
Query: 61 TDFIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNR 120
TDFIP+YPNLPSKLIC+LH+V LHAD ETDEVYAQMTLQPVNKYD++A+LASDMGLK NR
Sbjct: 61 TDFIPNYPNLPSKLICLLHSVTLHADTETDEVYAQMTLQPVNKYDREALLASDMGLKLNR 120
Query: 121 QPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRH 180
QPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLD+SMQPPAQE+VAKDLHDTTW+FRH
Sbjct: 121 QPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDFSMQPPAQEIVAKDLHDTTWTFRH 180
Query: 181 IYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVIS 240
IYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLF+RDEK QL+LGIRRANRQ P LSSSVIS
Sbjct: 181 IYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFVRDEKSQLMLGIRRANRQTPTLSSSVIS 240
Query: 241 SDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMF 300
SDSMHIGILAAAAHA AN+SPFTIF+NPRASPSEFV+PLAK+NKA+Y QVSLGMRFRMMF
Sbjct: 241 SDSMHIGILAAAAHANANSSPFTIFFNPRASPSEFVVPLAKYNKALYAQVSLGMRFRMMF 300
Query: 301 ETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTP 360
ETE+ GVRR+MGT+TGISDLD RWK SQWRNLQVGWDESTAG+RPSRVSIWEIEPV+TP
Sbjct: 301 ETEDCGVRRYMGTVTGISDLDPVRWKGSQWRNLQVGWDESTAGDRPSRVSIWEIEPVITP 360
Query: 361 FYICPPPFFRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFGMKDASGSIFPGLSLVQWM 420
FYICPPPFFRPK+PRQPGMPDDE D+ENAFKRAMPW+G+DFGMKDA S+FPGLSLVQWM
Sbjct: 361 FYICPPPFFRPKYPRQPGMPDDELDMENAFKRAMPWMGEDFGMKDAQSSMFPGLSLVQWM 420
Query: 421 SMQQNNQFSAAQSGCFPPMLSPNTLHSNLGTDDPSKLLSFQAPALSAPSLQFNKPNLPNQ 480
SMQQNN S + + P LS L +N ++DPSKLL+FQ+P LS+ + QFNKPN N
Sbjct: 421 SMQQNNPLSGSATPQLPSALSSFNLPNNFASNDPSKLLNFQSPNLSSANSQFNKPNTVNH 480
Query: 481 INQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQL 540
I+Q Q Q Q + QQQQQQQ
Sbjct: 481 ISQ--------------------------------------QMQAQPAMVKSQQQQQQQQ 502
Query: 541 QQQQSQQQQLQSLLQIPMNQLQQPRQQQLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGVV 600
QQ Q QQQQLQ QQQL Q Q+ I +NG +
Sbjct: 503 QQHQHQQQQLQQ--------------------------QQQLQMSQQQVQQQGIYNNGTI 536
Query: 601 A-SNQIT----------NQLVLQQQQLL-SGGISPQQSIHS-ASKTTFPLTSLPQESQFL 647
A +NQ++ +Q LQQQ +L +G Q+I+S +K +TS QE QF
Sbjct: 537 AVANQVSCQSPNQPTGFSQSQLQQQSMLPTGAKMTHQNINSMGNKGLSQMTSFAQEMQFQ 596
Query: 648 QQIDQQAS--LLQRQQLPQQTQLQQSSLHLLQQNQPQRAPQQPPVATMSQQTPSEQQLHL 705
QQ++ S LL+ QQ +QSSLH LQQN Q PQQ + S + QQL L
Sbjct: 597 QQLEMHNSSQLLRNQQ-------EQSSLHSLQQNLSQN-PQQLQMQQQSSKPSPSQQLQL 648
Query: 706 QLLQKLQQHQQHQQLLSTSSPLLQPQLLQQQSTPQQNQQLTQLPASQHQTQQL-GNNAFS 764
QLLQKLQQ QQ Q + SS L QPQL Q T Q+ QL QL +SQ+Q GNN+F
Sbjct: 649 QLLQKLQQQQQQQSIPPVSSSL-QPQLSALQQT--QSHQLQQLLSSQNQQPLAHGNNSFP 705
Query: 765 TEKVLNSNSFSSSLMQPHQLPVNHPQNTQKS----LAITRAPSTLTDGDAPSCSTSPSTN 820
+S+ MQP Q+ V+ Q Q S +A R+ S TDG+APSCSTSPS N
Sbjct: 706 ----------ASTFMQPPQIQVSPQQQGQMSNKNLVAAGRSHSGHTDGEAPSCSTSPSAN 755
Query: 821 NC---QISPSNLLKRNQQ---VPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGF 874
N +SP+N L RNQQ + V E SN +QEL +K++ +I N+K
Sbjct: 756 NTGHDNVSPTNFLSRNQQQGQAASVSASDSVFERASNPVQELYTKTESRISQGMMNMKSA 815
Query: 875 -DQLKYKGTITDQMEASSSGTSYCLD---PGNVQQSLPLSNFCMEGDVQSHS-RSNLPFD 929
+ ++K +TDQ++ S++GT+YC D P QQ+ PL +F +GD QSH R+NL F
Sbjct: 816 GEHFRFKSAVTDQIDVSTAGTTYCPDVVGPVQQQQTFPLPSFGFDGDCQSHHPRNNLAFP 875
Query: 930 SNLDGLTPDPVLMRGYDSQKDLQNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPG 989
NL+ +T DP+ SQKD QN++ NYG PRDIETELS+A ISSQSFG+P++ F PG
Sbjct: 876 GNLEAVTSDPLY-----SQKDFQNLVPNYGNTPRDIETELSSAAISSQSFGIPSIPFKPG 930
Query: 990 CSGDVG-INDTGVLNNG-LRANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLAR 1047
CS +VG IND+G++N G L NQTQRMRTYTKVQKRGSVGR IDVTRY GYDELR+DLAR
Sbjct: 931 CSNEVGGINDSGIMNGGGLWPNQTQRMRTYTKVQKRGSVGRSIDVTRYSGYDELRHDLAR 990
Query: 1048 MFGIEGQLEDPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDG 1107
MFGIEGQLEDP +DWKLVY DHENDILLVGDDPWEEFV+CVQ+IKILS EVQQMSLDG
Sbjct: 991 MFGIEGQLEDPLTSDWKLVYTDHENDILLVGDDPWEEFVNCVQNIKILSSVEVQQMSLDG 1050
Query: 1108 DLGNVPIPNQACSGTDSGNAWRGPYDDNS-AASFNR 1142
DL +P NQACS TDSGNAW+ Y+D S AASFNR
Sbjct: 1051 DLAAIPTTNQACSETDSGNAWKVHYEDTSAAASFNR 1086
>ARFF_ARATH (Q9ZTX8) Auxin response factor 6
Length = 933
Score = 630 bits (1625), Expect = e-180
Identities = 356/637 (55%), Positives = 425/637 (65%), Gaps = 45/637 (7%)
Query: 1 MKAPSNCYLPNSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKE 60
M+ S + P EGE++ +NSELWHACAGPLVSLPPVGS VVYFPQGHSEQVAAS KE
Sbjct: 1 MRLSSAGFNPQPHEGEKRVLNSELWHACAGPLVSLPPVGSRVVYFPQGHSEQVAASTNKE 60
Query: 61 TD-FIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYD-KDAILASDMGLKQ 118
D IP+YP+L +LIC LHNV +HAD ETDEVYAQMTLQP+N + KD L +++G+
Sbjct: 61 VDAHIPNYPSLHPQLICQLHNVTMHADVETDEVYAQMTLQPLNAQEQKDPYLPAELGVP- 119
Query: 119 NRQPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSF 178
+RQPT +FCKTLTASDTSTHGGFSVPRRAAEK+FPPLDYS QPPAQEL+A+DLHD W F
Sbjct: 120 SRQPTNYFCKTLTASDTSTHGGFSVPRRAAEKVFPPLDYSQQPPAQELMARDLHDNEWKF 179
Query: 179 RHIYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSV 238
RHI+RGQPKRHLLTTGWSVFVS KRL AGDSVLFI ++K QLLLGIRRANR Q + SSV
Sbjct: 180 RHIFRGQPKRHLLTTGWSVFVSAKRLVAGDSVLFIWNDKNQLLLGIRRANRPQTVMPSSV 239
Query: 239 ISSDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMY-TQVSLGMRFR 297
+SSDSMH+G+LAAAAHAAA NS FTIFYNPRASPSEFVIPLAK+ KA+Y T+VS+GMRFR
Sbjct: 240 LSSDSMHLGLLAAAAHAAATNSRFTIFYNPRASPSEFVIPLAKYVKAVYHTRVSVGMRFR 299
Query: 298 MMFETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPV 357
M+FETEES VRR+MGTITGI DLD RW +S WR+++VGWDESTAGER RVS+WEIEP+
Sbjct: 300 MLFETEESSVRRYMGTITGICDLDPTRWANSHWRSVKVGWDESTAGERQPRVSLWEIEPL 359
Query: 358 VTPFYICPPPF-FRPKFPRQPGMPD----DESDIENAFKRAMPWLGDDFGMKDASGSIFP 412
T F + P PF R K P PG+P E D+ + + W D G++ + F
Sbjct: 360 TT-FPMYPSPFPLRLKRPWPPGLPSFHGLKEDDMGMSMSSPLMW---DRGLQSLN---FQ 412
Query: 413 GLSLVQWMSMQQNNQFSAAQSGCFPPMLSPNTLHSNLGTDDPSKLLSFQAPALSAPSLQF 472
G+ + WM + + ++ L G DP+K + ++P
Sbjct: 413 GMGVNPWMQPRLDTSGLLGMQNDVYQAMAAAALQDMRGI-DPAKAAASLLQFQNSPGFSM 471
Query: 473 NKPNLPNQINQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQL 532
P+L Q Q QQQ QQQQQL Q QQQQQQL QQQQQQ QQ
Sbjct: 472 QSPSL---------------VQPQMLQQQLSQQQQQLS-QQQQQQQQLSQQQQQQLSQQQ 515
Query: 533 QQQQQQQLQQQQSQQQQLQSLLQIPMNQLQQPRQQQLPEPKNLPLPQQQLPQQLGQQSQK 592
QQQ QQ QQQ SQQQQ Q+ L +P + QP+ Q Q Q L QQ Q+
Sbjct: 516 QQQLSQQQQQQLSQQQQQQAYLGVP--ETHQPQSQ----------AQSQSNNHLSQQQQQ 563
Query: 593 AIMSNGVVASNQITNQLVLQQQQLLSGGISPQQSIHS 629
+ ++ AS+ + Q SP QS+ S
Sbjct: 564 VVDNHNPSASSAAVVSAMSQFGSASQPNTSPLQSMTS 600
Score = 162 bits (411), Expect = 4e-39
Identities = 149/481 (30%), Positives = 215/481 (43%), Gaps = 51/481 (10%)
Query: 652 QQASLLQRQQLPQQTQLQQSSLHLLQQNQPQRAPQQPPVATMSQQTPSEQQLHLQLLQKL 711
Q SL+Q Q L QQ QQ L QQ Q Q + QQ + QQ QQL Q Q+L
Sbjct: 472 QSPSLVQPQMLQQQLSQQQQQLSQQQQQQQQLSQQQQQQLSQQQQ----QQLSQQQQQQL 527
Query: 712 QQHQQHQQLLSTSSPLLQPQLLQQQSTPQQNQQLTQLPASQHQTQQLGNNAFSTEKVLNS 771
Q QQ Q L QPQ Q+ Q N L+Q Q Q N + S+ V+++
Sbjct: 528 SQQQQQQAYLGVPETH-QPQ---SQAQSQSNNHLSQ---QQQQVVDNHNPSASSAAVVSA 580
Query: 772 NSFSSSLMQPHQLPVNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQISPSNLLK 831
S S QP+ P+ + SL ++ S G+ P ISP + L
Sbjct: 581 MSQFGSASQPNTSPLQ----SMTSLCHQQSFSDTNGGNNP------------ISPLHTLL 624
Query: 832 RNQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFD---QLKYKGTITDQME 888
N + +S L+ ++ S M S D Q G Q
Sbjct: 625 SN----------FSQDESSQLLHLTRTNSAMTSSGWPSKRPAVDSSFQHSGAGNNNTQSV 674
Query: 889 ASSSGTSYCLDPGNVQQSLPLSNFCMEGDVQSHSRSNLPFDSNLDGLTPDPVLMRGYDSQ 948
G S+ + SLP E ++ ++ P L G+ D + +
Sbjct: 675 LEQLGQSHTSNVPPNAVSLPPFPGGRECSIEQEGSASDPHSHLLFGVNIDSSSLLMPNGM 734
Query: 949 KDLQNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGVLNNGLR- 1007
+L+++ G + ++++ ++ G MT C I+++G L +
Sbjct: 735 SNLRSIGIEGGDSTT---LPFTSSNFNNDFSGNLAMTTPSSC-----IDESGFLQSSENL 786
Query: 1008 ANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVY 1067
++ + T+ KV K GS GR +D++++ Y ELR++LARMFG+EGQLEDP R+ W+LV+
Sbjct: 787 GSENPQSNTFVKVYKSGSFGRSLDISKFSSYHELRSELARMFGLEGQLEDPVRSGWQLVF 846
Query: 1068 VDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDG--DLGNVPIPNQACSGTDSG 1125
VD END+LL+GDDPW EFVS V IKILS EVQQM G L + P N +G
Sbjct: 847 VDRENDVLLLGDDPWPEFVSSVWCIKILSPQEVQQMGKRGLELLNSAPSSNNVDKLPSNG 906
Query: 1126 N 1126
N
Sbjct: 907 N 907
>ARFH_ARATH (Q9FGV1) Auxin response factor 8
Length = 811
Score = 561 bits (1447), Expect = e-159
Identities = 327/637 (51%), Positives = 411/637 (64%), Gaps = 38/637 (5%)
Query: 14 EGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETD-FIPSYPNLPS 72
EGE K +NSELWHACAGPLVSLP GS VVYFPQGHSEQVAA+ KE D IP+YP+LP
Sbjct: 14 EGE-KCLNSELWHACAGPLVSLPSSGSRVVYFPQGHSEQVAATTNKEVDGHIPNYPSLPP 72
Query: 73 KLICMLHNVALHADPETDEVYAQMTLQPVN-KYDKDAILASDMGLKQNRQPTEFFCKTLT 131
+LIC LHNV +HAD ETDEVYAQMTLQP+ + K+ + ++G+ ++QP+ +FCKTLT
Sbjct: 73 QLICQLHNVTMHADVETDEVYAQMTLQPLTPEEQKETFVPIELGI-PSKQPSNYFCKTLT 131
Query: 132 ASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLL 191
ASDTSTHGGFSVPRRAAEK+FPPLDY++QPPAQEL+A+DLHD W FRHI+RGQPKRHLL
Sbjct: 132 ASDTSTHGGFSVPRRAAEKVFPPLDYTLQPPAQELIARDLHDVEWKFRHIFRGQPKRHLL 191
Query: 192 TTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAA 251
TTGWSVFVS KRL AGDSV+FIR+EK QL LGIR A R Q + SSV+SSDSMHIG+LAA
Sbjct: 192 TTGWSVFVSAKRLVAGDSVIFIRNEKNQLFLGIRHATRPQTIVPSSVLSSDSMHIGLLAA 251
Query: 252 AAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMY-TQVSLGMRFRMMFETEESGVRRH 310
AAHA+A NS FT+F++PRAS SEFVI L+K+ KA++ T++S+GMRFRM+FETEES VRR+
Sbjct: 252 AAHASATNSCFTVFFHPRASQSEFVIQLSKYIKAVFHTRISVGMRFRMLFETEESSVRRY 311
Query: 311 MGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVT-PFY--ICPPP 367
MGTITGISDLDS RW +S WR+++VGWDESTAGER RVS+WEIEP+ T P Y + P
Sbjct: 312 MGTITGISDLDSVRWPNSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSLFPLR 371
Query: 368 FFRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFGMKDASGSIFPGLSLVQWMSMQQN-N 426
RP +PD D+ + G+ G+ + +P + L WM + + +
Sbjct: 372 LKRPWHAGTSSLPDGRGDLGSGLTWLRGGGGEQQGLLPLN---YPSVGLFPWMQQRLDLS 428
Query: 427 QFSAAQSGCFPPMLSPNTLHSNLGTDDPSKLLSFQAPALSAPSLQFNKPNLPNQINQLHQ 486
Q + + ML+ N+G DP L Q L P Q+ L Q
Sbjct: 429 QMGTDNNQQYQAMLAAGL--QNIGGGDP---LRQQFVQLQEPHHQY-----------LQQ 472
Query: 487 SPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQLQQQQSQ 546
S QQQQQQQ + + Q Q + L QQ +Q+ QQQQL QQ
Sbjct: 473 SASHNSDLMLQQQQQQQASRHLMHAQTQIMSENLPQQNMRQEVSNQPAGQQQQL--QQPD 530
Query: 547 QQQLQSLLQIPMNQLQQPRQQ-QLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGVVASNQI 605
Q + ++ LQQ +QQ ++P P + + + A +G + + I
Sbjct: 531 QNAYLNAFKMQNGHLQQWQQQSEMPSPSFMKSDFTDSSNKFATTASPA-SGDGNLLNFSI 589
Query: 606 TNQLVLQQQQLLSGGISPQQSIHSASKTTFPLTSLPQ 642
T Q VL +QL + G SP+ AS T SLPQ
Sbjct: 590 TGQSVL-PEQLTTEGWSPK-----ASNTFSEPLSLPQ 620
Score = 130 bits (326), Expect = 3e-29
Identities = 130/458 (28%), Positives = 186/458 (40%), Gaps = 100/458 (21%)
Query: 670 QSSLHLLQQNQPQRAPQQPPVATMSQQTPSEQQLHLQLLQKLQQHQQH-QQLLSTSSPLL 728
Q L L Q Q +A Q L Q +Q + H Q+ QQ S +S L+
Sbjct: 422 QQRLDLSQMGTDNNQQYQAMLAAGLQNIGGGDPLRQQFVQLQEPHHQYLQQSASHNSDLM 481
Query: 729 QPQLLQQQST---------------PQQN--QQLTQLPASQHQT-QQLGNNAFSTEKVLN 770
Q QQQ++ PQQN Q+++ PA Q Q QQ NA+ LN
Sbjct: 482 LQQQQQQQASRHLMHAQTQIMSENLPQQNMRQEVSNQPAGQQQQLQQPDQNAY-----LN 536
Query: 771 SNSFSSSLMQPHQLPVNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQISPSNLL 830
+ + +Q Q P +PS + S + +T + NLL
Sbjct: 537 AFKMQNGHLQQWQQQSEMP-----------SPSFMKSDFTDSSNKFATTASPASGDGNLL 585
Query: 831 KRNQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTITDQMEAS 890
+ + L L E S + N FS Q +
Sbjct: 586 NFSITGQSVLPEQLTTEGWSP-----------KASNTFSEPLSLPQ-------------A 621
Query: 891 SSGTSYCLDPGNVQQSLPLSNFCMEGDVQSHSRSNLPFDSNLDGLTPDPVLMRGYDSQKD 950
G S L+PGN P + +L G+ PD L
Sbjct: 622 YPGKSLALEPGN------------------------PQNPSLFGVDPDSGLF-------- 649
Query: 951 LQNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGVLNNGLRANQ 1010
L + + + + D E + +S G N ++ C D +L+ + N
Sbjct: 650 LPSTVPRFASSSGDAEA----SPMSLTDSGFQNSLYS--CMQDTTHE---LLHGAGQINS 700
Query: 1011 TQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDH 1070
+ + + + KV K GSVGR +D++R+ Y ELR +L +MF IEG LEDP R+ W+LV+VD
Sbjct: 701 SNQTKNFVKVYKSGSVGRSLDISRFSSYHELREELGKMFAIEGLLEDPLRSGWQLVFVDK 760
Query: 1071 ENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGD 1108
ENDILL+GDDPWE FV+ V IKILS +V QM G+
Sbjct: 761 ENDILLLGDDPWESFVNNVWYIKILSPEDVHQMGDHGE 798
>ARFE_ARATH (P93024) Auxin response factor 5 (Transcription factor
MONOPTEROS) (Auxin-responsive protein IAA24)
Length = 902
Score = 509 bits (1311), Expect = e-143
Identities = 311/689 (45%), Positives = 399/689 (57%), Gaps = 46/689 (6%)
Query: 15 GERK-TINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKE-TDFIPSYPNLPS 72
G RK INSELWHACAGPLV LP VGSLV YF QGHSEQVA S ++ T +P+YPNLPS
Sbjct: 45 GTRKPVINSELWHACAGPLVCLPQVGSLVYYFSQGHSEQVAVSTRRSATTQVPNYPNLPS 104
Query: 73 KLICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMG-LKQNRQPTEFFCKTLT 131
+L+C +HNV LHAD ++DE+YAQM+LQPV+ ++D D G L+ ++ PTEFFCKTLT
Sbjct: 105 QLMCQVHNVTLHADKDSDEIYAQMSLQPVHS-ERDVFPVPDFGMLRGSKHPTEFFCKTLT 163
Query: 132 ASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLL 191
ASDTSTHGGFSVPRRAAEK+FPPLDYS QPP QELV +DLH+ TW+FRHIYRGQPKRHLL
Sbjct: 164 ASDTSTHGGFSVPRRAAEKLFPPLDYSAQPPTQELVVRDLHENTWTFRHIYRGQPKRHLL 223
Query: 192 TTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAA 251
TTGWS+FV +KRL AGDSVLFIRDEK QL++G+RRANRQQ AL SSV+S+DSMHIG+LAA
Sbjct: 224 TTGWSLFVGSKRLRAGDSVLFIRDEKSQLMVGVRRANRQQTALPSSVLSADSMHIGVLAA 283
Query: 252 AAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMY-TQVSLGMRFRMMFETEESGVRRH 310
AAHA AN +PF IFYNPRA P+EFVIPLAK+ KA+ +Q+S+GMRF MMFETE+SG RR+
Sbjct: 284 AAHATANRTPFLIFYNPRACPAEFVIPLAKYRKAICGSQLSVGMRFGMMFETEDSGKRRY 343
Query: 311 MGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTP--FYICPPPF 368
MGTI GISDLD RW S+WRNLQV WDE ++P+RVS W+IE TP +I P
Sbjct: 344 MGTIVGISDLDPLRWPGSKWRNLQVEWDEPGCNDKPTRVSPWDIE---TPESLFIFPSLT 400
Query: 369 FRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFG--MKDASGSIFPGLSLVQWMSMQQNN 426
K P E++ + KR + + D M AS L++ M NN
Sbjct: 401 SGLKRQLHPSYFAGETEWGSLIKRPLIRVPDSANGIMPYASFPSMASEQLMKMMMRPHNN 460
Query: 427 QFSAAQSGCFPPMLSPNTLHSNLGTDDPSKLLSFQAPALSAPSLQFNKPNLPNQINQLHQ 486
Q F + N + N G K + P + K + N+L
Sbjct: 461 Q----NVPSFMSEMQQNIVMGNGGLLGDMK--------MQQPLMMNQKSEMVQPQNKLTV 508
Query: 487 SPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQLQQQQS- 545
+P Q+Q Q + + + L + Q L+Q +Q Q S
Sbjct: 509 NP-----SASNTSGQEQNLSQSMSAPAKPENSTLSGCSSGRVQHGLEQSMEQASQVTTST 563
Query: 546 --QQQQLQSLLQIPMNQLQQPRQQQLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGVVASN 603
++++ LLQ P Q L + Q+ Q I + ++
Sbjct: 564 VCNEEKVNQLLQKPGASSPVQADQCL-----------DITHQIYQPQSDPINGFSFLETD 612
Query: 604 QITNQLVLQQQQLLSGGISPQQSIHSASKTTFPLTSLPQESQFLQQIDQQASLLQRQQL- 662
++T+Q + Q L+G + S + L F D Q + L+ Q
Sbjct: 613 ELTSQ--VSSFQSLAGSYKQPFILSSQDSSAVVLPDSTNSPLFHDVWDTQLNGLKFDQFS 670
Query: 663 PQQTQLQQSSLHLLQQNQPQRAPQQPPVA 691
P Q +S ++ N PP++
Sbjct: 671 PLMQQDLYASQNICMSNSTTSNILDPPLS 699
Score = 150 bits (378), Expect = 2e-35
Identities = 161/578 (27%), Positives = 237/578 (40%), Gaps = 155/578 (26%)
Query: 556 IPMNQLQQPRQQQLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGVVASNQITNQLVLQQQQ 615
+P +QL + P Q +P + + Q +M NG + + Q ++ Q+
Sbjct: 437 MPYASFPSMASEQLMKMMMRPHNNQNVPSFMSEMQQNIVMGNGGLLGDMKMQQPLMMNQK 496
Query: 616 LLSGGISPQQSIH---SASKTTFPLTSLPQESQFLQQIDQQASLLQRQQLPQQTQLQQSS 672
S + PQ + SAS T+ QE Q + A P+ + L S
Sbjct: 497 --SEMVQPQNKLTVNPSASNTS------GQEQNLSQSMSAPAK-------PENSTLSGCS 541
Query: 673 LHLLQQNQPQRAPQQPPVATMSQQTPSEQQLHLQLLQKLQQHQQHQQLLSTSSPLLQPQL 732
+Q Q Q V T T ++ QLLQK
Sbjct: 542 SGRVQHGLEQSMEQASQVTT---STVCNEEKVNQLLQK---------------------- 576
Query: 733 LQQQSTPQQNQQLTQLPASQHQTQQLGNNAFS-------TEKVLNSNSFSSSLMQPHQLP 785
S+P Q Q + +Q Q N FS T +V + S + S QP L
Sbjct: 577 -PGASSPVQADQCLDITHQIYQPQSDPINGFSFLETDELTSQVSSFQSLAGSYKQPFIL- 634
Query: 786 VNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQISPSNLLKRNQQVPATLGGSLV 845
++Q S A+ P + SP ++ + N LK +Q P
Sbjct: 635 -----SSQDSSAVV----------LPDSTNSPLFHDVWDTQLNGLKFDQFSPL------- 672
Query: 846 VESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTITDQMEASSSGTSYCLDPGNVQQ 905
+ Q+L + ++ + N S TS LDP
Sbjct: 673 ------MQQDLYASQNICMSN-------------------------STTSNILDP----- 696
Query: 906 SLPLSN-----FCM--EGDVQSHSRSNLPFDSNLDGLTPDPVLMRGYDSQKDLQNMLSNY 958
PLSN FC + D Q+H L ++N
Sbjct: 697 --PLSNTVLDDFCAIKDTDFQNHPSGCLVGNNNTS------------------------- 729
Query: 959 GGAPRDIETELSTADIS-SQSFGVPNMTFNPGCSG----DVGINDTGVLNNGLRAN---- 1009
+D+++++++A + SQ+F + N G +G +V +D + N ++
Sbjct: 730 --FAQDVQSQITSASFADSQAFSRQDFPDNSGGTGTSSSNVDFDDCSLRQNSKGSSWQKI 787
Query: 1010 QTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVD 1069
T R+RTYTKVQK GSVGR IDVT +K Y+EL++ + MFG+EG L PQ + WKLVYVD
Sbjct: 788 ATPRVRTYTKVQKTGSVGRSIDVTSFKDYEELKSAIECMFGLEGLLTHPQSSGWKLVYVD 847
Query: 1070 HENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDG 1107
+E+D+LLVGDDPWEEFV CV+ I+ILS TEVQQMS +G
Sbjct: 848 YESDVLLVGDDPWEEFVGCVRCIRILSPTEVQQMSEEG 885
>ARFB_ARATH (Q94JM3) Auxin response factor 2 (ARF1-binding protein)
(ARF1-BP)
Length = 859
Score = 387 bits (994), Expect = e-107
Identities = 195/362 (53%), Positives = 248/362 (67%), Gaps = 3/362 (0%)
Query: 23 ELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSKLICMLHNVA 82
ELWHACAGPLV++P V YFPQGH EQV AS + + +LPSKL+C + NV
Sbjct: 61 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKLLCRVINVD 120
Query: 83 LHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTASDTSTHGGFS 142
L A+ +TDEVYAQ+TL P D++AI R FCKTLTASDTSTHGGFS
Sbjct: 121 LKAEADTDEVYAQITLLPEANQDENAIEKEAPLPPPPRFQVHSFCKTLTASDTSTHGGFS 180
Query: 143 VPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTTGWSVFVSTK 202
V RR A++ PPLD S QPP QELVAKDLH W FRHI+RGQP+RHLL +GWSVFVS+K
Sbjct: 181 VLRRHADECLPPLDMSRQPPTQELVAKDLHANEWRFRHIFRGQPRRHLLQSGWSVFVSSK 240
Query: 203 RLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNSPF 262
RL AGD+ +F+R E +L +G+RRA RQQ + SSVISS SMH+G+LA A HA + + F
Sbjct: 241 RLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTMF 300
Query: 263 TIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDLDS 322
T++Y PR SPSEF++P ++ +++ S+GMRF+M FE EE+ +R GTI GI + D
Sbjct: 301 TVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEESDP 360
Query: 323 ARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPPPFFRPKFPRQ---PGM 379
RW S+WR+L+V WDE+++ RP RVS W++EP + P + P P RPK PR P
Sbjct: 361 TRWPKSKWRSLKVRWDETSSIPRPDRVSPWKVEPALAPPALSPVPMPRPKRPRSNIAPSS 420
Query: 380 PD 381
PD
Sbjct: 421 PD 422
Score = 90.1 bits (222), Expect = 3e-17
Identities = 55/164 (33%), Positives = 93/164 (56%), Gaps = 7/164 (4%)
Query: 942 MRGYDSQKDLQNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGV 1001
M G DS +N L++ G + ++ D+S QS G + + N+
Sbjct: 665 MNGTDSTMSQRNNLNDAAGLTQIASPKVQ--DLSDQSKGSKSTNDHREQGRPFQTNNPHP 722
Query: 1002 LNNGLRANQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQR 1060
+ + N + R+ TKV K+G ++GR +D+++++ Y+EL +L R+F G+L P++
Sbjct: 723 KDAQTKTNSS---RSCTKVHKQGIALGRSVDLSKFQNYEELVAELDRLFEFNGELMAPKK 779
Query: 1061 TDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMS 1104
DW +VY D END++LVGDDPW+EF V+ I I + EV++M+
Sbjct: 780 -DWLIVYTDEENDMMLVGDDPWQEFCCMVRKIFIYTKEEVRKMN 822
>ARFA_ARATH (Q8L7G0) Auxin response factor 1
Length = 665
Score = 380 bits (976), Expect = e-104
Identities = 197/363 (54%), Positives = 259/363 (71%), Gaps = 9/363 (2%)
Query: 23 ELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDF-IPSYPNLPSKLICMLHNV 81
ELWHACAGPLV+LP G V YFP+GH EQ+ ASM + + +PS+ NLPSK++C + N+
Sbjct: 22 ELWHACAGPLVTLPREGERVYYFPEGHMEQLEASMHQGLEQQMPSF-NLPSKILCKVINI 80
Query: 82 ALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPT-EFFCKTLTASDTSTHGG 140
A+PETDEVYAQ+TL P + D+ + D +++ + T FCKTLTASDTSTHGG
Sbjct: 81 QRRAEPETDEVYAQITLLP--ELDQSEPTSPDAPVQEPEKCTVHSFCKTLTASDTSTHGG 138
Query: 141 FSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTTGWSVFVS 200
FSV RR A+ PPLD S QPP QELVA DLH++ W FRHI+RGQP+RHLLTTGWSVFVS
Sbjct: 139 FSVLRRHADDCLPPLDMSQQPPWQELVATDLHNSEWHFRHIFRGQPRRHLLTTGWSVFVS 198
Query: 201 TKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNS 260
+K+L AGD+ +F+R E ++L +G+RR RQQ + SSVISS SMHIG+LA AAHA +
Sbjct: 199 SKKLVAGDAFIFLRGENEELRVGVRRHMRQQTNIPSSVISSHSMHIGVLATAAHAITTGT 258
Query: 261 PFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDL 320
F++FY PR S SEF++ + ++ +A ++S+GMRF+M FE EE+ +R GTI G+ +
Sbjct: 259 IFSVFYKPRTSRSEFIVSVNRYLEAKTQKLSVGMRFKMRFEGEEAPEKRFSGTIVGVQEN 318
Query: 321 DSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVV---TPFYICPPPFFRPKFPRQP 377
S+ W S+WR+L+V WDE ++ RP RVS WE+EP+V TP PP R K PR P
Sbjct: 319 KSSVWHDSEWRSLKVQWDEPSSVFRPERVSPWELEPLVANSTPSSQPQPP-QRNKRPRPP 377
Query: 378 GMP 380
G+P
Sbjct: 378 GLP 380
Score = 82.8 bits (203), Expect = 5e-15
Identities = 42/97 (43%), Positives = 68/97 (69%), Gaps = 2/97 (2%)
Query: 1009 NQTQRMRTYTKVQKRGS-VGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVY 1067
+Q++++R+ TKV +GS VGR ID+TR + Y++L L MF I+G+L + + W++VY
Sbjct: 536 SQSRQIRSCTKVHMQGSAVGRAIDLTRSECYEDLFKKLEEMFDIKGELLESTKK-WQVVY 594
Query: 1068 VDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMS 1104
D E+D+++VGDDPW EF V+ I I + EV+++S
Sbjct: 595 TDDEDDMMMVGDDPWNEFCGMVRKIFIYTPEEVKKLS 631
>ARFD_ARATH (Q9ZTX9) Auxin response factor 4
Length = 788
Score = 353 bits (905), Expect = 2e-96
Identities = 187/374 (50%), Positives = 243/374 (64%), Gaps = 12/374 (3%)
Query: 11 NSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNL 70
+S G +I SELWHACAGPL LP G++VVYFPQGH EQ A IP + +L
Sbjct: 53 SSSTGSASSIYSELWHACAGPLTCLPKKGNVVVYFPQGHLEQDAMVSYSSPLEIPKF-DL 111
Query: 71 PSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDK---DAILASDMGLKQNRQPTE--- 124
+++C + NV L A+ +TDEVY Q+TL P+ ++ + ++G ++ R +
Sbjct: 112 NPQIVCRVVNVQLLANKDTDEVYTQVTLLPLQEFSMLNGEGKEVKELGGEEERNGSSSVK 171
Query: 125 ----FFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRH 180
FCKTLTASDTSTHGGFSVPRRAAE F PLDY Q P+QEL+AKDLH W FRH
Sbjct: 172 RTPHMFCKTLTASDTSTHGGFSVPRRAAEDCFAPLDYKQQRPSQELIAKDLHGVEWKFRH 231
Query: 181 IYRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVIS 240
IYRGQP+RHLLTTGWS+FVS K L +GD+VLF+RDE +L LGIRRA R + L S+I
Sbjct: 232 IYRGQPRRHLLTTGWSIFVSQKNLVSGDAVLFLRDEGGELRLGIRRAARPRNGLPDSIIE 291
Query: 241 SDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMF 300
+S IL+ A+A + S F +FY+PRA+ +EFVIP K+ ++ + V +G RFRM F
Sbjct: 292 KNSCS-NILSLVANAVSTKSMFHVFYSPRATHAEFVIPYEKYITSIRSPVCIGTRFRMRF 350
Query: 301 ETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTP 360
E ++S RR G +TG+ DLD RW +S+WR L V WDES + RVS WEI+P V+
Sbjct: 351 EMDDSPERRCAGVVTGVCDLDPYRWPNSKWRCLLVRWDESFVSDHQERVSPWEIDPSVSL 410
Query: 361 FYICPPPFFRPKFP 374
++ RPK P
Sbjct: 411 PHLSIQSSPRPKRP 424
Score = 92.4 bits (228), Expect = 6e-18
Identities = 55/153 (35%), Positives = 85/153 (54%), Gaps = 7/153 (4%)
Query: 957 NYGGAPRDIETELS-TADISSQSFGVP-NMTFNPGCSGDVGINDTGVLNNGLRANQTQRM 1014
N GG P ++ +L D+ + G NM + GC + + Q+
Sbjct: 608 NEGGLPNNVTADLPFKIDMMGKQKGSELNMNASSGCKL---FGFSLPVETPASKPQSSSK 664
Query: 1015 RTYTKVQKRGS-VGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDHEND 1073
R TKV K+GS VGR ID++R GYD+L +L R+F +EG L DP++ W+++Y D END
Sbjct: 665 RICTKVHKQGSQVGRAIDLSRLNGYDDLLMELERLFNMEGLLRDPEK-GWRILYTDSEND 723
Query: 1074 ILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLD 1106
+++VGDDPW +F + V I + + EV+ + D
Sbjct: 724 MMVVGDDPWHDFCNVVWKIHLYTKEEVENANDD 756
>ARFI_ARATH (Q9XED8) Auxin response factor 9
Length = 638
Score = 341 bits (874), Expect = 8e-93
Identities = 176/355 (49%), Positives = 233/355 (65%), Gaps = 4/355 (1%)
Query: 23 ELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQK-ETDFIPSYPNLPSKLICMLHNV 81
ELW CAGPLV +P V YFPQGH EQ+ AS Q+ + + + LP K++C + NV
Sbjct: 12 ELWKLCAGPLVDVPQAQERVYYFPQGHMEQLEASTQQVDLNTMKPLFVLPPKILCNVMNV 71
Query: 82 ALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTASDTSTHGGF 141
+L A+ +TDEVYAQ+TL PV + + + R F K LTASDTSTHGGF
Sbjct: 72 SLQAEKDTDEVYAQITLIPVGTEVDEPMSPDPSPPELQRPKVHSFSKVLTASDTSTHGGF 131
Query: 142 SVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTTGWSVFVST 201
SV R+ A + PPLD + Q P QELVA+D+H W F+HI+RGQP+RHLLTTGWS FV++
Sbjct: 132 SVLRKHATECLPPLDMTQQTPTQELVAEDVHGYQWKFKHIFRGQPRRHLLTTGWSTFVTS 191
Query: 202 KRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNSP 261
KRL AGD+ +F+R E +L +G+RRAN QQ ++ SSVISS SMH+G+LA A HA +
Sbjct: 192 KRLVAGDTFVFLRGENGELRVGVRRANLQQSSMPSSVISSHSMHLGVLATARHATQTKTM 251
Query: 262 FTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDLD 321
F ++Y PR S+F+I L K+ +AM + S+GMRF+M FE E+S RR+ GT+ G+ D
Sbjct: 252 FIVYYKPRT--SQFIISLNKYLEAMSNKFSVGMRFKMRFEGEDSPERRYSGTVIGVKDC- 308
Query: 322 SARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPPPFFRPKFPRQ 376
S WK S+WR L+V WDE + RP++VS WEIEP V + + K PRQ
Sbjct: 309 SPHWKDSKWRCLEVHWDEPASISRPNKVSPWEIEPFVNSENVPKSVMLKNKRPRQ 363
Score = 87.4 bits (215), Expect = 2e-16
Identities = 43/100 (43%), Positives = 68/100 (68%), Gaps = 3/100 (3%)
Query: 1006 LRANQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWK 1064
+++ Q+ R+ TKVQ +G VGR +D+ KGY+EL +D+ ++F I+G+L R W+
Sbjct: 514 VQSKQSSSTRSRTKVQMQGVPVGRAVDLNALKGYNELIDDIEKLFDIKGELRS--RNQWE 571
Query: 1065 LVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMS 1104
+V+ D E D++LVGDDPW EF + V+ I I S EV++M+
Sbjct: 572 IVFTDDEGDMMLVGDDPWPEFCNMVKRIFIWSKEEVKKMT 611
>ARFC_ARATH (O23661) Auxin response factor 3 (ETTIN protein)
Length = 608
Score = 337 bits (864), Expect = 1e-91
Identities = 177/344 (51%), Positives = 227/344 (65%), Gaps = 18/344 (5%)
Query: 23 ELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSKLICMLHNVA 82
ELWHACAGPL+SLP GSLV+YFPQGH EQ DF + LP + C + +V
Sbjct: 54 ELWHACAGPLISLPKRGSLVLYFPQGHLEQAP-------DFSAAIYGLPPHVFCRILDVK 106
Query: 83 LHADPETDEVYAQMTLQP----VNKYDKDAILASDMG------LKQNRQPTEFFCKTLTA 132
LHA+ TDEVYAQ++L P + + ++ I+ D G LK++ P FCKTLTA
Sbjct: 107 LHAETTTDEVYAQVSLLPESEDIERKVREGIIDVDGGEEDYEVLKRSNTP-HMFCKTLTA 165
Query: 133 SDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLT 192
SDTSTHGGFSVPRRAAE FPPLDYS P+QEL+A+DLH W FRHIYRGQP+RHLLT
Sbjct: 166 SDTSTHGGFSVPRRAAEDCFPPLDYSQPRPSQELLARDLHGLEWRFRHIYRGQPRRHLLT 225
Query: 193 TGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAA 252
TGWS FV+ K+L +GD+VLF+R + +L LG+RRA++ + + S + +M+ +
Sbjct: 226 TGWSAFVNKKKLVSGDAVLFLRGDDGKLRLGVRRASQIEGTAALSAQYNQNMNHNNFSEV 285
Query: 253 AHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMG 312
AHA + +S F+I YNP+AS S F+IP KF K + +GMRF+ E+E++ RR G
Sbjct: 286 AHAISTHSVFSISYNPKASWSNFIIPAPKFLKVVDYPFCIGMRFKARVESEDASERRSPG 345
Query: 313 TITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEP 356
I+GISDLD RW S+WR L V WD+ A RVS WEIEP
Sbjct: 346 IISGISDLDPIRWPGSKWRCLLVRWDDIVANGHQQRVSPWEIEP 389
>ARFK_ARATH (Q9ZPY6) Auxin response factor 11
Length = 601
Score = 333 bits (855), Expect = 1e-90
Identities = 179/349 (51%), Positives = 236/349 (67%), Gaps = 10/349 (2%)
Query: 22 SELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKET--DFIPSYPNLPSKLICMLH 79
+ELW ACAGPLV +P G V YFPQGH EQ+ AS + IP + NLP K++C +
Sbjct: 20 TELWKACAGPLVEVPRYGERVFYFPQGHMEQLVASTNQGVVDQEIPVF-NLPPKILCRVL 78
Query: 80 NVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPT-EFFCKTLTASDTSTH 138
+V L A+ ETDEVYAQ+TLQP + D+ + D L + +PT + F K LTASDTSTH
Sbjct: 79 SVTLKAEHETDEVYAQITLQP--EEDQSEPTSLDPPLVEPAKPTVDSFVKILTASDTSTH 136
Query: 139 GGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTTGWSVF 198
GGFSV R+ A + P LD + P QELVA+DLH W F+HI+RGQP+RHLLTTGWS F
Sbjct: 137 GGFSVLRKHATECLPSLDMTQPTPTQELVARDLHGYEWRFKHIFRGQPRRHLLTTGWSTF 196
Query: 199 VSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAAN 258
V++KRL AGD+ +F+R E L +G+RR +QQ + +SVISS SM +G+LA A+HA
Sbjct: 197 VTSKRLVAGDAFVFLRGETGDLRVGVRRLAKQQSTMPASVISSQSMRLGVLATASHAVTT 256
Query: 259 NSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGIS 318
+ F +FY PR S+F+I + K+ AM SLGMR+RM FE EES R GTI G
Sbjct: 257 TTIFVVFYKPRI--SQFIISVNKYMMAMKNGFSLGMRYRMRFEGEESPERIFTGTIIGSG 314
Query: 319 DLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPPP 367
DL S++W +S+WR+LQ+ WDE ++ +RP++VS WEIEP +P + P P
Sbjct: 315 DL-SSQWPASKWRSLQIQWDEPSSIQRPNKVSPWEIEP-FSPSALTPTP 361
Score = 81.3 bits (199), Expect = 1e-14
Identities = 47/119 (39%), Positives = 66/119 (54%), Gaps = 6/119 (5%)
Query: 998 DTGVLNNGLRANQTQRMRTYTKVQKRGS-VGRCIDVTRYKGYDELRNDLARMFGIEGQLE 1056
D N+ Q R+ KVQ +G+ VGR +D+T + YDEL +L +MF IEG+L
Sbjct: 473 DPNSSNSPKEQKQQTSTRSRIKVQMQGTAVGRAVDLTLLRSYDELIKELEKMFEIEGELS 532
Query: 1057 DPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQM---SLDGDLGNV 1112
+ W +V+ D E D +LVGDDPW EF + + I EV++M SL GD G +
Sbjct: 533 PKDK--WAIVFTDDEGDRMLVGDDPWNEFCKMAKKLFIYPSDEVKKMRSKSLLGDKGTI 589
>ARFR_ARATH (Q9C5W9) Auxin response factor 18
Length = 602
Score = 332 bits (851), Expect = 4e-90
Identities = 204/508 (40%), Positives = 289/508 (56%), Gaps = 29/508 (5%)
Query: 11 NSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQK--ETDFIPSYP 68
+S + + +ELW CAGPLV +P V YFPQGH EQ+ AS + ++ IP +
Sbjct: 13 SSSRSYQDQLYTELWKVCAGPLVEVPRAQERVFYFPQGHMEQLVASTNQGINSEEIPVF- 71
Query: 69 NLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCK 128
+LP K++C + +V L A+ ETDEVYAQ+TLQP + L + + +Q F K
Sbjct: 72 DLPPKILCRVLDVTLKAEHETDEVYAQITLQPEEDQSEPTSLDPPI-VGPTKQEFHSFVK 130
Query: 129 TLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKR 188
LTASDTSTHGGFSV R+ A + P LD + P QELV +DLH W F+HI+RGQP+R
Sbjct: 131 ILTASDTSTHGGFSVLRKHATECLPSLDMTQATPTQELVTRDLHGFEWRFKHIFRGQPRR 190
Query: 189 HLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGI 248
HLLTTGWS FVS+KRL AGD+ +F+R E L +G+RR R Q + +SVISS SMH+G+
Sbjct: 191 HLLTTGWSTFVSSKRLVAGDAFVFLRGENGDLRVGVRRLARHQSTMPTSVISSQSMHLGV 250
Query: 249 LAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVR 308
LA A+HA + F +FY PR S+F++ + K+ +A+ SLG RFRM FE EES R
Sbjct: 251 LATASHAVRTTTIFVVFYKPRI--SQFIVGVNKYMEAIKHGFSLGTRFRMRFEGEESPER 308
Query: 309 RHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPP-- 366
GTI G DL S++W +S+WR+LQV WDE T +RP +VS WEIEP + I P
Sbjct: 309 IFTGTIVGSGDL-SSQWPASKWRSLQVQWDEPTTVQRPDKVSPWEIEPFLATSPISTPAQ 367
Query: 367 -PFFRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFGMKDASGSIFPGLSLVQWMSMQQ- 424
P + K R P P ++ +F ++P + SI L L Q S+++
Sbjct: 368 QPQSKCKRSR-PIEPSVKTPAPPSFLYSLP---------QSQDSINASLKLFQDPSLERI 417
Query: 425 NNQFSAAQSGCFPPMLSPNTLHSNLGTDDPSKLLSFQAPALSAPSLQFNKPNLPNQINQL 484
+ +S+ S F P P T+ +L F + S + +K +
Sbjct: 418 SGGYSSNNS--FKPETPPPP------TNCSYRLFGFDLTSNSPAPIPQDKQPMDTCGAAK 469
Query: 485 HQSPVSWPQQQQQQQQQQQQQQQQLQLQ 512
Q P++ +Q++QQ + + ++Q+Q
Sbjct: 470 CQEPITPTSMSEQKKQQTSRSRTKVQMQ 497
Score = 82.0 bits (201), Expect = 8e-15
Identities = 44/94 (46%), Positives = 60/94 (63%), Gaps = 3/94 (3%)
Query: 1012 QRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDH 1070
Q R+ TKVQ +G +VGR +D+T K YDEL ++L MF I+GQL R W +V+ D
Sbjct: 486 QTSRSRTKVQMQGIAVGRAVDLTLLKSYDELIDELEEMFEIQGQLL--ARDKWIVVFTDD 543
Query: 1071 ENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMS 1104
E D++L GDDPW EF + I I S EV++M+
Sbjct: 544 EGDMMLAGDDPWNEFCKMAKKIFIYSSDEVKKMT 577
>ARFU_ARATH (Q9C8N9) Putative auxin response factor 21
Length = 606
Score = 298 bits (763), Expect = 6e-80
Identities = 152/345 (44%), Positives = 217/345 (62%), Gaps = 7/345 (2%)
Query: 14 EGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSK 73
+G + + +LW CAGPL +P +G V YFPQG+ E V AS ++E + + +LPSK
Sbjct: 18 DGSKSYMYEQLWKLCAGPLCDIPKLGENVYYFPQGNIELVQASTREELNELQPICDLPSK 77
Query: 74 LICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTAS 133
L C + + L + +DE+YA++TL P D ++ + R F K LTAS
Sbjct: 78 LQCRVIAIHLKVENNSDEIYAEITLMP----DTTQVVIPTQSENRFRPLVNSFTKVLTAS 133
Query: 134 DTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTT 193
DTS +GGFSVP++ A + PPLD S PAQE++A DLHD W FRH YRG P+RH LTT
Sbjct: 134 DTSAYGGFSVPKKHAIECLPPLDMSQPLPAQEILAIDLHDNQWRFRHNYRGTPQRHSLTT 193
Query: 194 GWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAA 253
GW+ F+++K+L GD ++F+R E +L +GIRRA QQ + SS++S D M G++A+A
Sbjct: 194 GWNEFITSKKLVKGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVIASAK 253
Query: 254 HAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGT 313
HA N F + Y PR+ S+F++ KF A+ + ++G RF M FE ++ RR+ GT
Sbjct: 254 HAFDNQCIFIVVYKPRS--SQFIVSYDKFLDAVNNKFNVGSRFTMRFEGDDFSERRYFGT 311
Query: 314 ITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVV 358
I G+SD S WK S+WR+L+V WDE + RP++VS WEIE +V
Sbjct: 312 IIGVSDF-SPHWKCSEWRSLEVQWDEFASFSRPNKVSPWEIEHLV 355
Score = 79.3 bits (194), Expect = 5e-14
Identities = 41/107 (38%), Positives = 64/107 (59%), Gaps = 3/107 (2%)
Query: 1006 LRANQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWK 1064
+++ Q RT TKVQ +G ++GR +D++ GYD+L +L ++F I+GQL+ R WK
Sbjct: 502 IQSKQFSSSRTCTKVQMQGVTIGRAVDLSVLNGYDQLILELEKLFDIKGQLQT--RNQWK 559
Query: 1065 LVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLGN 1111
+ + D + +LVGDDPW EF V+ I I S EV+ + L +
Sbjct: 560 IAFTDSDGYEMLVGDDPWPEFCKMVKKILIYSKEEVKNLKSSKSLSS 606
>ARFT_ARATH (Q9C7I9) Putative auxin response factor 20
Length = 606
Score = 295 bits (755), Expect = 5e-79
Identities = 152/346 (43%), Positives = 214/346 (60%), Gaps = 7/346 (2%)
Query: 14 EGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSK 73
+G + + +LW CAGPL +P +G V YFPQG+ E V AS ++E + + +LPSK
Sbjct: 18 DGSKSYMYEQLWKLCAGPLCDIPKLGENVYYFPQGNIELVDASTREELNELQPICDLPSK 77
Query: 74 LICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTAS 133
L C + + L + +DE YA++TL P D ++ Q R F K LTAS
Sbjct: 78 LQCRVIAIHLKVENNSDETYAEITLMP----DTTQVVIPTQSENQFRPLVNSFTKVLTAS 133
Query: 134 DTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTT 193
DTS +GGF VP++ A + PPLD S PAQEL+AKDLH W FRH YRG P+RH LTT
Sbjct: 134 DTSAYGGFFVPKKHAIECLPPLDMSQPLPAQELLAKDLHGNQWRFRHSYRGTPQRHSLTT 193
Query: 194 GWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAA 253
GW+ F ++K+L GD ++F+R E +L +GIRRA QQ + SS++S D M G++A+A
Sbjct: 194 GWNEFTTSKKLVKGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVIASAK 253
Query: 254 HAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGT 313
HA N F + Y PR+ S+F++ KF AM + +G RF M FE ++ RR+ GT
Sbjct: 254 HALDNQCIFIVVYKPRS--SQFIVSYDKFLDAMNNKFIVGSRFTMRFEGDDFSERRYFGT 311
Query: 314 ITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVT 359
I G++D S WK S+WR+L+V WDE + RP++VS WEIE +++
Sbjct: 312 IIGVNDF-SPHWKCSEWRSLEVQWDEFASFSRPNKVSPWEIEHLMS 356
Score = 77.8 bits (190), Expect = 2e-13
Identities = 40/107 (37%), Positives = 64/107 (59%), Gaps = 3/107 (2%)
Query: 1006 LRANQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWK 1064
+++ Q RT TKVQ +G ++GR +D++ GYD+L +L ++F ++GQL+ R WK
Sbjct: 502 IQSKQFGSTRTCTKVQMQGVTIGRAVDLSVLNGYDQLILELEKLFDLKGQLQT--RNQWK 559
Query: 1065 LVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLGN 1111
+ + D + +LVGDDPW EF V+ I I S EV+ + L +
Sbjct: 560 IAFTDSDGYEMLVGDDPWPEFCKMVKKILIYSKEEVKNLKSSKSLSS 606
>ARFV_ARATH (Q9C8N7) Putative auxin response factor 22
Length = 598
Score = 294 bits (753), Expect = 8e-79
Identities = 155/363 (42%), Positives = 219/363 (59%), Gaps = 9/363 (2%)
Query: 14 EGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSK 73
+G + + +LW CAGPL +P +G + YFPQG+ E V AS ++E + + +LPSK
Sbjct: 18 DGSKSYMYEQLWKLCAGPLCDIPKLGEKIYYFPQGNIELVEASTREELNELKPICDLPSK 77
Query: 74 LICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTAS 133
L C + + L + +DE YA++TL P D ++ Q R F K LTAS
Sbjct: 78 LQCRVIAIQLKVENNSDETYAEITLMP----DTTQVVIPTQNENQFRPLVNSFTKVLTAS 133
Query: 134 DTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTT 193
DTS GGF VP++ A + PPLD S P QEL+A DLH W F H YRG P+RHLLTT
Sbjct: 134 DTS--GGFFVPKKHAIECLPPLDMSQPLPTQELLATDLHGNQWRFNHNYRGTPQRHLLTT 191
Query: 194 GWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAA 253
GW+ F ++K+L AGD ++F+R E +L +GIRRA QQ + SS+IS +SM G++A+A
Sbjct: 192 GWNAFTTSKKLVAGDVIVFVRGETGELRVGIRRAGHQQGNIPSSIISIESMRHGVIASAK 251
Query: 254 HAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGT 313
HA N F + Y PR+ S+F++ KF A+ + ++G RF M FE ++ RR+ GT
Sbjct: 252 HAFDNQCMFIVVYKPRS--SQFIVSYDKFLDAVNNKFNVGSRFTMRFEGDDFSERRYFGT 309
Query: 314 ITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPPPFFRPKF 373
I G+SD S WK S+WRNL+V WDE + RP++VS WEIE ++ + P + K
Sbjct: 310 IIGVSDF-SPHWKCSEWRNLEVQWDEFASFSRPNKVSPWEIEHLMPALNVPRPSLLKNKR 368
Query: 374 PRQ 376
R+
Sbjct: 369 LRE 371
Score = 72.4 bits (176), Expect = 7e-12
Identities = 35/90 (38%), Positives = 58/90 (63%), Gaps = 3/90 (3%)
Query: 1006 LRANQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWK 1064
+++ Q RT TKVQ +G ++ R +D++ GYD+L +L +F ++GQL+ R W+
Sbjct: 500 IQSKQFSSTRTCTKVQMQGVTIERAVDLSVLNGYDQLILELEELFDLKGQLQT--RNQWE 557
Query: 1065 LVYVDHENDILLVGDDPWEEFVSCVQSIKI 1094
+ + D ++D +LVGDDPW EF + V+ I I
Sbjct: 558 IAFTDSDDDKMLVGDDPWPEFCNMVKKILI 587
>ARFN_ARATH (Q9LQE8) Putative auxin response factor 14
Length = 605
Score = 294 bits (753), Expect = 8e-79
Identities = 153/363 (42%), Positives = 216/363 (59%), Gaps = 7/363 (1%)
Query: 14 EGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSK 73
+G + + +LW CAGPL +P +G V YFPQGH E V AS ++E + + + PSK
Sbjct: 18 DGSKSYMYEQLWKLCAGPLCDIPKLGEKVYYFPQGHIELVEASTREELNELQPICDFPSK 77
Query: 74 LICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTAS 133
L C + + L + +DE YA++TL P D ++ Q R F K LTAS
Sbjct: 78 LQCRVIAIQLKVENNSDETYAEITLMP----DTTQVVIPTQNQNQFRPLVNSFTKVLTAS 133
Query: 134 DTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTT 193
DTS HGGFSVP++ A + PPLD S P QE++A DLH W FRHIYRG +RHLLT
Sbjct: 134 DTSVHGGFSVPKKHAIECLPPLDMSQPLPTQEILAIDLHGNQWRFRHIYRGTAQRHLLTI 193
Query: 194 GWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAA 253
GW+ F ++K+L GD ++F+R E +L +GIRRA QQ + SS++S +SM GI+A+A
Sbjct: 194 GWNAFTTSKKLVEGDVIVFVRGETGELRVGIRRAGHQQGNIPSSIVSIESMRHGIIASAK 253
Query: 254 HAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGT 313
HA N F + Y PR+ S+F++ KF + + ++G RF M FE ++ RR GT
Sbjct: 254 HAFDNQCMFIVVYKPRS--SQFIVSYDKFLDVVNNKFNVGSRFTMRFEGDDFSERRSFGT 311
Query: 314 ITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPPPFFRPKF 373
I G+SD S WK S+WR+L+V WDE + RP++VS W+IE + + F + K
Sbjct: 312 IIGVSDF-SPHWKCSEWRSLEVQWDEFASFPRPNQVSPWDIEHLTPWSNVSRSSFLKNKR 370
Query: 374 PRQ 376
R+
Sbjct: 371 SRE 373
Score = 80.5 bits (197), Expect = 2e-14
Identities = 72/262 (27%), Positives = 127/262 (47%), Gaps = 33/262 (12%)
Query: 861 DMQIKNEFSNVKGFDQLKYKGTITDQMEASSSGTSYCLDPGNVQ-QSLPLSNFCMEGDVQ 919
D++ +SNV LK K + ++ S +S+ L P Q Q + + ++
Sbjct: 350 DIEHLTPWSNVSRSSFLKNKRS--REVNEIGSSSSHLLPPTLTQGQEIGQQSMATPMNIS 407
Query: 920 SHSRSNLPFDSNLDGLTPDPVLMRGYDSQKDLQNMLS-NYGGAPRDIETELSTADISS-Q 977
R D D +TP +LM +Q M NY IE ++T ++S +
Sbjct: 408 LRYR-----DITEDAMTPSRLLM-----SYPVQPMAKLNYNNVVTPIEENITTNAVASFR 457
Query: 978 SFGV----PNMTFNP---------GCSGDVGINDTGVLNNG--LRANQTQRMRTYTKVQK 1022
FGV P++ +P + + + +L + +++ Q RT TKVQ
Sbjct: 458 LFGVSLATPSVIKDPVEQIGLEISRLTQEKKFGQSQILRSPTEIQSKQFSSTRTCTKVQM 517
Query: 1023 RG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDHENDILLVGDDP 1081
+G ++GR +D++ GYD+L +L ++F ++GQL+ R W++ + ++E D +LVG+DP
Sbjct: 518 QGVTIGRAVDLSVLNGYDQLILELEKLFDLKGQLQ--ARNQWEIAFTNNEEDKMLVGEDP 575
Query: 1082 WEEFVSCVQSIKILSYTEVQQM 1103
W EF + V+ I I S EV+ +
Sbjct: 576 WPEFCNMVKKIFIYSKEEVKNL 597
>ARFL_ARATH (Q9XID4) Putative auxin response factor 12
Length = 593
Score = 291 bits (746), Expect = 5e-78
Identities = 151/342 (44%), Positives = 208/342 (60%), Gaps = 7/342 (2%)
Query: 14 EGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSK 73
+G + + +LW CAGPL +P +G V YFPQGH E V S ++E + + +LPSK
Sbjct: 18 DGSKSYVYEQLWKLCAGPLCDIPKLGEKVYYFPQGHIELVETSTREELNELQPICDLPSK 77
Query: 74 LICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTAS 133
L C + + L + +DE YA++TL P D ++ Q R F K LTAS
Sbjct: 78 LQCRVIAIHLKVENNSDETYAEITLMP----DTTQVVIPTQNENQFRPLVNSFTKVLTAS 133
Query: 134 DTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTT 193
DTS HGGF VP++ A + P LD S PAQEL+A DLH W F H YRG P+RHLLTT
Sbjct: 134 DTSAHGGFFVPKKHAIECLPSLDMSQPLPAQELLAIDLHGNQWRFNHNYRGTPQRHLLTT 193
Query: 194 GWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAA 253
GW+ F ++K+L AGD ++F+R E +L +GIRRA QQ + SS++S D M G++A+A
Sbjct: 194 GWNAFTTSKKLVAGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVVASAK 253
Query: 254 HAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGT 313
HA N FT+ Y PR+ S+F++ KF A+ + ++G RF M E ++ RR GT
Sbjct: 254 HAFDNQCMFTVVYKPRS--SKFIVSYDKFLDAVNNKFNVGSRFTMRLEGDDFSERRCFGT 311
Query: 314 ITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIE 355
I G+SD S WK S+WR+L+V WDE T+ P +VS W+IE
Sbjct: 312 IIGVSDF-SPHWKCSEWRSLEVQWDEFTSFPGPKKVSPWDIE 352
Score = 75.5 bits (184), Expect = 8e-13
Identities = 39/101 (38%), Positives = 62/101 (60%), Gaps = 3/101 (2%)
Query: 995 GINDTGVLNNGLRANQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEG 1053
G++ T ++ Q RT TKVQ +G ++GR +D++ GYD+L +L ++F I+G
Sbjct: 491 GLSQTLRSPTEIQNKQFSSSRTCTKVQMQGVTIGRAVDLSVLNGYDQLILELEKLFDIKG 550
Query: 1054 QLEDPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKI 1094
QL+ R W++ + D + D +LVGDDPW EF + V+ I I
Sbjct: 551 QLQT--RNQWEIAFTDSDEDKMLVGDDPWPEFCNMVKKIFI 589
>ARFO_ARATH (Q9LQE3) Putative auxin response factor 15
Length = 593
Score = 288 bits (737), Expect = 6e-77
Identities = 153/354 (43%), Positives = 215/354 (60%), Gaps = 7/354 (1%)
Query: 23 ELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSKLICMLHNVA 82
+LW CAGPL +P +G V YFPQG+ E V AS ++E + + +LPSKL C + +
Sbjct: 27 QLWKLCAGPLCDIPKLGEKVYYFPQGNIELVEASTREELNELQPICDLPSKLQCRVIAIH 86
Query: 83 LHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTASDTSTHGGFS 142
L + +DE YA++TL P D ++ Q R F K LTASD S +G FS
Sbjct: 87 LKVENNSDETYAKITLMP----DTTQVVIPTQNENQFRPLVNSFTKVLTASDISANGVFS 142
Query: 143 VPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTTGWSVFVSTK 202
VP++ A + PPLD S PAQEL+A DLH WSFRH YRG P+RHLLTTGW+ F ++K
Sbjct: 143 VPKKHAIECLPPLDMSQPLPAQELLAIDLHGNQWSFRHSYRGTPQRHLLTTGWNEFTTSK 202
Query: 203 RLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNSPF 262
+L GD ++F+R E +L +GIRRA QQ + SS++S D M G++A+A HA N F
Sbjct: 203 KLVKGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVIASAKHAFDNQCMF 262
Query: 263 TIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDLDS 322
+ Y PR+ S+F++ KF A+ + ++G RF M FE ++ RR+ GTI G+S+ S
Sbjct: 263 IVVYKPRS--SQFIVSYDKFLDAVNNKFNVGSRFTMRFEGDDLSERRYFGTIIGVSNF-S 319
Query: 323 ARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPPPFFRPKFPRQ 376
WK S WR+L+V WDE + RP++VS WEIE ++ + F + K R+
Sbjct: 320 PHWKCSDWRSLEVQWDEFASFLRPNKVSPWEIEHLMPALNVPRSSFLKNKRLRE 373
Score = 75.5 bits (184), Expect = 8e-13
Identities = 36/90 (40%), Positives = 59/90 (65%), Gaps = 3/90 (3%)
Query: 1006 LRANQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWK 1064
+++ Q RT TKVQ +G ++GR +D++ GYD+L +L ++F ++GQL+ R WK
Sbjct: 502 IQSKQFSSTRTCTKVQMQGVTIGRAVDLSVLNGYDQLILELEKLFDLKGQLQT--RNQWK 559
Query: 1065 LVYVDHENDILLVGDDPWEEFVSCVQSIKI 1094
+++ + D +LVGDDPW EF + V+ I I
Sbjct: 560 IIFTGSDEDEMLVGDDPWPEFCNMVKRIYI 589
>ARFM_ARATH (Q9FX25) Putative auxin response factor 13
Length = 623
Score = 276 bits (706), Expect = 2e-73
Identities = 149/338 (44%), Positives = 203/338 (59%), Gaps = 9/338 (2%)
Query: 23 ELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSKLICMLHNVA 82
+LW+ CAGPL LP G V YFPQGH E + S + E D I +LPSKL C + +
Sbjct: 27 KLWNICAGPLCVLPKPGEKVYYFPQGHIELIENSTRDELDHIRPIFDLPSKLRCRVVAID 86
Query: 83 LHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTASDTSTHGGFS 142
D TDEVYAQ++L P D ++ + + R FF K LTASD S GG
Sbjct: 87 RKVDKNTDEVYAQISLMP----DTTEVMTHNTTMDTRRPIVYFFSKILTASDVSLSGGLI 142
Query: 143 VPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRHLLTT--GWSVFVS 200
+P++ A + FPPLD S Q LVAKDL+ WSF+H++RG P+RH+ T+ GWSVF +
Sbjct: 143 IPKQYAIECFPPLDMSQPISTQNLVAKDLYGQEWSFKHVFRGTPQRHMFTSGGGWSVFAT 202
Query: 201 TKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNS 260
TKRL GD + +R E +L GIRRA QQ + SSVIS++ M G++A+ +A
Sbjct: 203 TKRLIVGDIFVLLRGENGELRFGIRRAKHQQGHIPSSVISANCMQHGVIASVVNAFKTKC 262
Query: 261 PFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDL 320
F + Y P S S+FVI KF AM +G RFRM FE ++ +R+ GTI G++D+
Sbjct: 263 MFNVVYKP--SSSQFVISYDKFVDAMNNNYIVGSRFRMQFEGKDFSEKRYDGTIIGVNDM 320
Query: 321 DSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVV 358
S WK S+WR+L+V WDE + RP++VS W+IE ++
Sbjct: 321 -SPHWKDSEWRSLKVQWDELSPFLRPNQVSPWDIEHLI 357
Score = 63.9 bits (154), Expect = 2e-09
Identities = 32/92 (34%), Positives = 55/92 (59%), Gaps = 3/92 (3%)
Query: 1015 RTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDHEND 1073
R+ KV +G ++ R +D+T GY++L L +F ++ +L R W++V+ ++E
Sbjct: 510 RSRIKVHMQGVAISRAVDLTAMHGYNQLIQKLEELFDLKDELRT--RNQWEIVFTNNEGA 567
Query: 1074 ILLVGDDPWEEFVSCVQSIKILSYTEVQQMSL 1105
+LVGDDPW EF + + I I S E+++M L
Sbjct: 568 EMLVGDDPWPEFCNMAKRIFICSKEEIKKMKL 599
>ARFJ_ARATH (Q9SKN5) Auxin response factor 10
Length = 693
Score = 272 bits (695), Expect = 4e-72
Identities = 164/407 (40%), Positives = 214/407 (52%), Gaps = 51/407 (12%)
Query: 16 ERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSKLI 75
+ K+++ +LWHACAG +V +P + S V YF QGH+E A DF P +P ++
Sbjct: 3 QEKSLDPQLWHACAGSMVQIPSLNSTVFYFAQGHTEHAHAP----PDF--HAPRVPPLIL 56
Query: 76 CMLHNVALHADPETDEVYAQMTLQPVNKYD----KDAILA----SDMGLKQNRQPTEFFC 127
C + +V AD ETDEV+A++TL P+ D DA+L S G ++ F
Sbjct: 57 CRVVSVKFLADAETDEVFAKITLLPLPGNDLDLENDAVLGLTPPSSDGNGNGKEKPASFA 116
Query: 128 KTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPK 187
KTLT SD + GGFSVPR AE IFP LDYS +PP Q ++AKD+H TW FRHIYRG P+
Sbjct: 117 KTLTQSDANNGGGFSVPRYCAETIFPRLDYSAEPPVQTVIAKDIHGETWKFRHIYRGTPR 176
Query: 188 RHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANR-----------QQPALSS 236
RHLLTTGWS FV+ K+L AGDS++F+R E L +GIRRA R P S
Sbjct: 177 RHLLTTGWSTFVNQKKLIAGDSIVFLRSESGDLCVGIRRAKRGGLGSNAGSDNPYPGFSG 236
Query: 237 SVISSDS-------------------------MHIGILAAAAHAAANNSPFTIFYNPRAS 271
+ +S + + +A A AA F + Y PRAS
Sbjct: 237 FLRDDESTTTTSKLMMMKRNGNNDGNAAATGRVRVEAVAEAVARAACGQAFEVVYYPRAS 296
Query: 272 PSEFVIPLAKFNKAMYTQVSLGMRFRMMFETEESG-VRRHMGTITGISDLDSARWKSSQW 330
EF + A AM + GMRF+M FETE+S + MGT++ + D RW +S W
Sbjct: 297 TPEFCVKAADVRSAMRIRWCSGMRFKMAFETEDSSRISWFMGTVSAVQVADPIRWPNSPW 356
Query: 331 RNLQVGWDESTAGERPSRVSIWEIEPVVTPFYICPPPFFRPKFPRQP 377
R LQV WDE + RVS W +E V I PF K R P
Sbjct: 357 RLLQVAWDEPDLLQNVKRVSPWLVELVSNMPTIHLSPFSPRKKIRIP 403
Score = 38.9 bits (89), Expect = 0.080
Identities = 30/108 (27%), Positives = 51/108 (46%), Gaps = 18/108 (16%)
Query: 1003 NNGLRANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTD 1062
N L+ +T + + + + VGR +D++ Y EL LA MF IE +R+D
Sbjct: 572 NYSLQGLETGHCKVFMESE---DVGRTLDLSVIGSYQELYRKLAEMFHIE------ERSD 622
Query: 1063 --WKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGD 1108
+VY D I +GD+P+ +F+ + + I +M + GD
Sbjct: 623 LLTHVVYRDANGVIKRIGDEPFSDFMKATKRLTI-------KMDIGGD 663
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.128 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 135,108,335
Number of Sequences: 164201
Number of extensions: 6056895
Number of successful extensions: 121793
Number of sequences better than 10.0: 1624
Number of HSP's better than 10.0 without gapping: 1004
Number of HSP's successfully gapped in prelim test: 655
Number of HSP's that attempted gapping in prelim test: 27640
Number of HSP's gapped (non-prelim): 16413
length of query: 1142
length of database: 59,974,054
effective HSP length: 121
effective length of query: 1021
effective length of database: 40,105,733
effective search space: 40947953393
effective search space used: 40947953393
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0006.8


