

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.6
(566 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
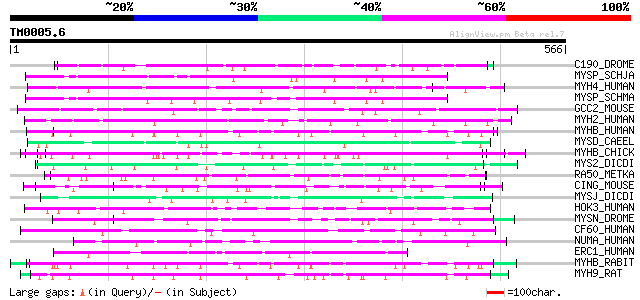
Score E
Sequences producing significant alignments: (bits) Value
C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 1... 63 2e-09
MYSP_SCHJA (Q05870) Paramyosin (Antigen Sj97) 61 7e-09
MYH4_HUMAN (Q9Y623) Myosin heavy chain, skeletal muscle, fetal (... 59 4e-08
MYSP_SCHMA (P06198) Paramyosin 58 8e-08
GCC2_MOUSE (Q8CHG3) GRIP and coiled-coil domain-containing prote... 57 2e-07
MYH2_HUMAN (Q9UKX2) Myosin heavy chain, skeletal muscle, adult 2... 56 2e-07
MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (S... 55 4e-07
MYSD_CAEEL (P02567) Myosin heavy chain D (MHC D) 55 6e-07
MYHB_CHICK (P10587) Myosin heavy chain, gizzard smooth muscle 55 6e-07
MYS2_DICDI (P08799) Myosin II heavy chain, non muscle 54 1e-06
RA50_METKA (Q8TXI4) DNA double-strand break repair rad50 ATPase 53 2e-06
CING_MOUSE (P59242) Cingulin 53 2e-06
MYSJ_DICDI (P54697) Myosin IJ heavy chain 52 3e-06
HOK3_HUMAN (Q86VS8) Hook homolog 3 (hHK3) 52 3e-06
MYSN_DROME (Q99323) Myosin heavy chain, non-muscle (Zipper prote... 52 4e-06
CF60_HUMAN (Q8NB25) Hypothetical protein C6orf60 52 4e-06
NUMA_HUMAN (Q14980) Nuclear mitotic apparatus protein 1 (NuMA pr... 52 5e-06
ERC1_HUMAN (Q8IUD2) ERC protein 1 (ELKS protein) 52 5e-06
MYHB_RABIT (P35748) Myosin heavy chain, smooth muscle isoform (S... 51 7e-06
MYH9_RAT (Q62812) Myosin heavy chain, nonmuscle type A (Cellular... 51 7e-06
>C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 190)
(Microtubule binding protein 190) (d-CLIP-190)
Length = 1690
Score = 62.8 bits (151), Expect = 2e-09
Identities = 99/465 (21%), Positives = 201/465 (42%), Gaps = 58/465 (12%)
Query: 46 ECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRP 105
E + + + AT + + + L AE N + L E + + L +++ I+
Sbjct: 1118 EAIQVANANISATNAELSTVLEVL-QAEKSETNHIFELFEMEADMNSE-RLIEKVTGIKE 1175
Query: 106 VLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFK-KLGELKSH 164
L++ + DER K+F +++ ++ +Q + S +++ Q T K KL E++
Sbjct: 1176 ELKETHLQLDERQKKFEELEEKL----------KQAQQSEQKLQQESQTSKEKLTEIQQS 1225
Query: 165 LQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKT-------------LSEIHPSLGDS 211
LQ+LQ+ +++ V++ + E S+++ K L E L +S
Sbjct: 1226 LQELQDSVKQKEELVQNLEEKVRESSSIIEAQNTKLNESNVQLENKTSCLKETQDQLLES 1285
Query: 212 SKGTPQSISNDTLASLTGYIHSLKQEK---QQRLLKVQELTKFLVDLWDLIETPIDEQKA 268
K Q + A L+G + +++ + L+KV+EL K L + + +D Q+A
Sbjct: 1286 QKKEKQL--QEEAAKLSGELQQVQEANGDIKDSLVKVEELVKVLEEKLQAATSQLDAQQA 1343
Query: 269 FNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYR 328
N L V G L + + E ++Q+L ++K+ + +++N L+E+
Sbjct: 1344 TNK--ELQELLVKSQENEGNLQGESLAVTE-KLQQLEQANG-ELKEALCQKENGLKELQG 1399
Query: 329 GVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFA 388
+ + + +L S +S N ++ D L+ + R +E+ E L ++++
Sbjct: 1400 KL------DESNTVLESQKKSHN-EIQDKLEQAQQKERTLQEETSKLAEQLSQLKQ---- 1448
Query: 389 AEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIV------VSKIPSIVENLTTKVKAW 442
A EE + + + +G + + AE +++ S +++E L +V
Sbjct: 1449 ANEEL---QKSLQQKQLLLEKGNEFDTQLAEYQKVIDEMDDAASVKSALLEQLQNRVA-- 1503
Query: 443 EMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQ 487
E+E + + A LE ++RQL E KSRE L+ Q
Sbjct: 1504 ELETALRQAND-AQKTAYLETKELRRQLESLELEKSREVLSLKAQ 1547
Score = 45.1 bits (105), Expect = 5e-04
Identities = 90/474 (18%), Positives = 194/474 (39%), Gaps = 60/474 (12%)
Query: 49 DIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLE 108
D+ R K K E DA+ + + L ++ E LK ++ + L+
Sbjct: 371 DLLREKQQHVEKLMVERDLDREDAQNQALQLQKNINE----------LKARIVELESALD 420
Query: 109 DLRSKK--------------DERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLT 154
+ R K DE + K +I + +I S + + DL
Sbjct: 421 NERKKTEELQCSIDEAQFCGDELNAQSQVYKEKIHDLESKITKLVSATPSLQSILPPDLP 480
Query: 155 FKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKG 214
G L+ + LQ + ++Q++V+S I+ E + + K L+E +L
Sbjct: 481 SDD-GALQEEIAKLQEKMTIQQKEVESRIAEQLEEEQRLRENV-KYLNEQIATLQSELVS 538
Query: 215 TPQSISNDTLA-----SLTGYIHSLKQEKQQRLLKVQ-ELTKFLVDLWDLIETPIDEQKA 268
+++ +L+ +L + LK+E +++ + Q E T+ L E ++ +
Sbjct: 539 KDEALEKFSLSECGIENLRRELELLKEENEKQAQEAQAEFTR------KLAEKSVEVLRL 592
Query: 269 FNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNE-LEEIY 327
+ + L A+ D + + +D E ++ EV +M+D + N+ L+E+
Sbjct: 593 SSELQNL-KATSDSLESERVNKTDECEILQTEV---------RMRDEQIRELNQQLDEVT 642
Query: 328 RGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQA----QSRREILDRVE 383
+++ +A + L + G + S LL+ + ++ Q+KEQA + ++ ++
Sbjct: 643 TQLNVQKADSSALDDMLRLQKEGTEEKSTLLEKTEKELVQSKEQAAKTLNDKEQLEKQIS 702
Query: 384 KWRFAAEEEKWLDEYERDENRYSAVR----GAHKNLKRAEKARIVVSKIPSIVENLTTKV 439
+ AE+EK + E EN + ++ + L + K S E ++
Sbjct: 703 DLKQLAEQEKLV--REMTENAINQIQLEKESIEQQLALKQNELEDFQKKQSESEVHLQEI 760
Query: 440 KAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQE 493
KA +K L E L+ L++ Q+ L E+ + + E+ + +++ ++++
Sbjct: 761 KAQNTQKDFE-LVESGESLKKLQQQLEQKTLGHEKLQAALEELKKEKETIIKEK 813
Score = 42.4 bits (98), Expect = 0.003
Identities = 102/540 (18%), Positives = 212/540 (38%), Gaps = 80/540 (14%)
Query: 18 LLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELI 77
LL Q+ + E+GE+ + + +E + ++++A ++ + A++ AE
Sbjct: 908 LLEQITKLKSEVGETQAALSSCHTDVESKT-----KQLEAANAALEKVNKEYAESRAEAS 962
Query: 78 NLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAG 137
+L + E + T+ L+ RS + SK +I+ G
Sbjct: 963 DLQDKVKEITD-------------TLHAELQAERSSSSALHTKLSKFSDEIA------TG 1003
Query: 138 YEQPKSSSEEVDQSDLTFKK-LGELKSHLQDLQNE----KILRQQKVKSHISTISELSAV 192
+++ S ++ Q L +K L EL+ LQD Q+ K ++K KS +I L
Sbjct: 1004 HKELTSKADAWSQEMLQKEKELQELRQQLQDSQDSQTKLKAEGERKEKSFEESIKNLQEE 1063
Query: 193 MSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFL 252
++ + L + S GT T+ L + E Q + E + +
Sbjct: 1064 VTKAKTENL--------ELSTGT-----QTTIKDLQERLEITNAELQHKEKMASEDAQKI 1110
Query: 253 VDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAE--------VQRL 304
DL L+E + +S ++ + ++ + E E E ++++
Sbjct: 1111 ADLKTLVEAIQVANANISATNAELSTVLEVLQAEKSETNHIFELFEMEADMNSERLIEKV 1170
Query: 305 NALKASKMKDL---VFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSM 361
+K ++K+ + +RQ + EE+ + SE Q + + ++ LQ +
Sbjct: 1171 TGIK-EELKETHLQLDERQKKFEELEEKLKQAQQSEQKLQQESQTSKEKLTEIQQSLQEL 1229
Query: 362 DDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEK- 420
D ++Q +E Q+ E + R A+ K + + EN+ S ++ L ++K
Sbjct: 1230 QDSVKQKEELVQNLEEKV-RESSSIIEAQNTKLNESNVQLENKTSCLKETQDQLLESQKK 1288
Query: 421 -------ARIVVSKIPSIVE-NLTTKVKAWEMEKGIPFLYEKAPLLQS-----------L 461
A + ++ + E N K ++E+ + L EK S L
Sbjct: 1289 EKQLQEEAAKLSGELQQVQEANGDIKDSLVKVEELVKVLEEKLQAATSQLDAQQATNKEL 1348
Query: 462 EEYHVQRQ-----LREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQSTTANTIV 516
+E V+ Q L+ E + + ++L++ + +EAL + K+ G+ +NT++
Sbjct: 1349 QELLVKSQENEGNLQGESLAVTEKLQQLEQANGELKEALCQKENGLKELQGKLDESNTVL 1408
>MYSP_SCHJA (Q05870) Paramyosin (Antigen Sj97)
Length = 866
Score = 61.2 bits (147), Expect = 7e-09
Identities = 101/486 (20%), Positives = 214/486 (43%), Gaps = 67/486 (13%)
Query: 17 SLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAEL 76
S L++L+ + D++ ++ +N +L E ++ D ++A++ + + A++ L
Sbjct: 170 SQLNRLKTLTDDLQRQLTELNNAKSRLTSENFELLHINQD----YEAQILNY-SKAKSSL 224
Query: 77 INLVSSLGES-SSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEI 135
+ V L S SR + L+ QL +++ ++L++K DE +E S +++Q+S+ +I
Sbjct: 225 ESQVDDLKRSLDDESRNRFNLQAQLTSLQMDYDNLQAKYDEESEEASNLRNQVSKFNADI 284
Query: 136 AGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSI 195
A + K E + +++ + +L + +L E + ++++K+ ++ +L +++
Sbjct: 285 AALKS-KFERELMSKTEEFEEMKRKLTMRITEL--EDVAERERLKA--VSLEKLKTKLTL 339
Query: 196 DFWKTLSEIHPSLGDSSKGTPQSISNDTLAS--------LTGYIHSLKQEKQQRLLKVQE 247
+ SEI ++ + ++ S ++LAS LT +++L + Q +
Sbjct: 340 EIKDLQSEIESLSLENGELIRRAKSAESLASELQRRVDELTIEVNTLTSQNNQLESENMR 399
Query: 248 LTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTH----GCLSS------------ 291
L + DL D E + N + + +S+ + + L S
Sbjct: 400 LKSLVNDLTDKNNALERENRQMNDQVKELKSSLRDANRRLTDLEALRSQLEAERDNLASA 459
Query: 292 ---------DVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRG------------V 330
D+ ++ +A LN LK S+M+ + +R ELE + +
Sbjct: 460 LHDAEEALRDMDQKYQASQAALNHLK-SEMEQRLRERDEELESLRKSTTRTIEELTVTIT 518
Query: 331 HMDVDSEAARQILTSLIESGNIDLSDLLQSMD----DQIRQAKEQAQSRREI---LDRVE 383
M+V ++ L ES DL L + + + +++ K AQ +++ LD
Sbjct: 519 EMEVKYKSELSRLKKRYESSIADLEIQLDATNKANANLMKENKNLAQRVKDLETFLDDER 578
Query: 384 KWRFAAEEEKWLDEYERDE--NRYSAVRGAHKNLKRAEK-ARIVVSKIPSIVENLTTKVK 440
+ R AAE + E++R + N +R A +NL+R K A + + S V LT +V
Sbjct: 579 RLREAAENNLQITEHKRIQLANEVEELRSAMENLERLRKHAETELEETQSRVSELTIQVN 638
Query: 441 AWEMEK 446
+K
Sbjct: 639 TLSNDK 644
>MYH4_HUMAN (Q9Y623) Myosin heavy chain, skeletal muscle, fetal
(Myosin heavy chain IIb) (MyHC-IIb)
Length = 1939
Score = 58.5 bits (140), Expect = 4e-08
Identities = 110/526 (20%), Positives = 221/526 (41%), Gaps = 63/526 (11%)
Query: 19 LSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELIN 78
+ +L+ +E ++ S + L +C D+ R + + ++ KAEL + ++ A +E+
Sbjct: 1316 IEELKRQLEEETKAKSTLAHALQSARHDC-DLLREQYEEEQEAKAELQRGMSKANSEVAQ 1374
Query: 79 L-----VSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICV 133
++ + K L Q+L +E + SK K ++++++ + +
Sbjct: 1375 WRTKYETDAIQRTEELEEAKKKLAQRLQDAEEHVEAVNSKCASLEKTKQRLQNEVEDLMI 1434
Query: 134 EIAGYEQPKSSSEEVDQSDLTFKK-LGELKSHLQDLQNEKILRQQKVKSHISTISELSAV 192
++ E+ ++ +D+ F K L E K ++ Q E Q++ +S + + ++
Sbjct: 1435 DV---ERSNAACIALDKKQRNFDKVLAEWKQKYEETQAELEASQKESRSLSTELFKVKNA 1491
Query: 193 M--SIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTG--YIHSLKQEKQQRLLKVQEL 248
S+D +TL +K Q IS+ T G +IH L++ K+Q + EL
Sbjct: 1492 YEESLDHLETLKR-------ENKNLQQEISDLTEQIAEGGKHIHELEKVKKQLDHEKSEL 1544
Query: 249 TKFLVDL--------WDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAE 300
L + ++ ++ + + + R I+ +E+ L + + VE+
Sbjct: 1545 QTSLEEAEASLEHEEGKILRIQLELNQVKSEIDRKIAEKDEELDQ---LKRNHLRVVESM 1601
Query: 301 VQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQS 360
L+A S+ L K++ E + + ++ + A + L +L + I L D
Sbjct: 1602 QSTLDAEIRSRNDALRIKKKMEGDLNEMEIQLNHANRQAAEALRNLRNTQGI-LKDTQLH 1660
Query: 361 MDDQIR---QAKEQ--------------AQSRREILDRVEKWRFAAEEEKWLDEYERDEN 403
+DD IR KEQ + R L+R E+ R AE+E LD ER +
Sbjct: 1661 LDDAIRGQDDLKEQLAMVERRANLMQAEVEELRASLERTERGRKMAEQE-LLDASERVQL 1719
Query: 404 RYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLEE 463
++ K+ E +S+I +E++ + + E EK + + A + + L++
Sbjct: 1720 LHTQNTSLINTKKKLETD---ISQIQGEMEDIVQEARNAE-EKAKKAITDAAMMAEELKK 1775
Query: 464 -----YHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKK 504
H++R + E+ Q RL E EQ AL G + +K
Sbjct: 1776 EQDTSAHLERMKKNMEQTVKDLQLRLGE---AEQLALKGGKKQIQK 1818
Score = 47.8 bits (112), Expect = 8e-05
Identities = 89/431 (20%), Positives = 174/431 (39%), Gaps = 41/431 (9%)
Query: 19 LSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELIN 78
L LQM D++ + +LEQ+ D+ + + + + +LC L A+ +L
Sbjct: 1011 LDDLQMEEDKVNTLTKAKT----KLEQQVDDL-----EGSLEQEKKLCMDLERAKRKLEG 1061
Query: 79 LVSSLGESSSFSRG-KGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAG 137
+ ES+ + K L ++L + +L+ K ++ +++ +I ++ I
Sbjct: 1062 DLKLAQESTMDTENDKQQLNEKLKKKEFEMSNLQGKIEDEQALAMQLQKKIKELQARIEE 1121
Query: 138 YEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDF 197
E EE++ + K + +S L E R ++ S EL+ +F
Sbjct: 1122 LE------EEIEAERASRAKAEKQRSDLSRELEEISERLEEAGGATSAQIELNKKREAEF 1175
Query: 198 WKTLSEIHPSL--GDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDL 255
K ++ S +++ + D++A L I SL++ KQ+ + EL + DL
Sbjct: 1176 QKMRRDLEESTLQHEATAAALRKKHADSVAELGKQIDSLQRVKQKLEKEKSELKMEINDL 1235
Query: 256 WDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDL 315
+ET + F + R + + E+ T +I ++ A+ RL+ + L
Sbjct: 1236 ASNMETVSKAKANFEKMCRTLEDQLSEIKTKEEEQQRLINELSAQKARLHTESGEFSRQL 1295
Query: 316 VFKR-------------QNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMD 362
K ++EE+ R + + A+ L ++S D DLL+
Sbjct: 1296 DEKDAMVSQLSRGKQAFTQQIEELKRQLE---EETKAKSTLAHALQSARHD-CDLLREQY 1351
Query: 363 DQIRQAKEQAQ-SRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEK- 420
++ ++AK + Q + V +WR E D +R E A + + L+ AE+
Sbjct: 1352 EEEQEAKAELQRGMSKANSEVAQWRTKYE----TDAIQRTEELEEAKKKLAQRLQDAEEH 1407
Query: 421 ARIVVSKIPSI 431
V SK S+
Sbjct: 1408 VEAVNSKCASL 1418
Score = 42.7 bits (99), Expect = 0.003
Identities = 103/521 (19%), Positives = 211/521 (39%), Gaps = 59/521 (11%)
Query: 16 ASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAE 75
A++ + + +E+ ++++ R +LE++ + + + K D + +AE LADAE
Sbjct: 854 ANMKEEFEKTKEELAKTEAKRK----ELEEKMVTLMQEKNDLQLQVQAE-ADALADAEER 908
Query: 76 LINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEI 135
L+ + + + + + ++ + +L +KK + E S++K I + + +
Sbjct: 909 CDQLIKTKIQLEAKIK---EVTERAEDEEEINAELTAKKRKLEDECSELKKDIDDLELTL 965
Query: 136 AGYEQPKSSSE--------EVDQSDLTFKKLGELKSHLQDLQNEKI--LRQQKVKSHIST 185
A E+ K ++E E+ D T KL + K LQ+ + + L+ ++ K + T
Sbjct: 966 AKVEKEKHATENKVKNLTEEMAGLDETIAKLTKEKKALQEAHQQTLDDLQMEEDKVNTLT 1025
Query: 186 ISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISND--TLASLTGYIHSLKQEKQQRLL 243
++ +D + E L + + + D T + KQ+ ++L
Sbjct: 1026 KAKTKLEQQVDDLEGSLEQEKKLCMDLERAKRKLEGDLKLAQESTMDTENDKQQLNEKLK 1085
Query: 244 KVQ-ELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGC-------LSSDVIE 295
K + E++ + D + QK + I +E+ SD+
Sbjct: 1086 KKEFEMSNLQGKIEDEQALAMQLQKKIKELQARIEELEEEIEAERASRAKAEKQRSDLSR 1145
Query: 296 QVEAEVQRLNAL--KASKMKDLVFKRQNELEEIYRGV-HMDVDSEAARQILTSLIESGNI 352
++E +RL S +L KR+ E +++ R + + EA L
Sbjct: 1146 ELEEISERLEEAGGATSAQIELNKKREAEFQKMRRDLEESTLQHEATAAALRKKHADSVA 1205
Query: 353 DLSDLLQSMDDQIRQ--AKEQAQSRREILDRVEKWRFAAEEEKWLDEYERD-ENRYSAVR 409
+L + S+ +++Q KE+++ + EI D ++ + ++ R E++ S ++
Sbjct: 1206 ELGKQIDSL-QRVKQKLEKEKSELKMEINDLASNMETVSKAKANFEKMCRTLEDQLSEIK 1264
Query: 410 GAHKNLKR------AEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLEE 463
+ +R A+KAR+ + ++ +G K Q +EE
Sbjct: 1265 TKEEEQQRLINELSAQKARLHTESGEFSRQLDEKDAMVSQLSRG------KQAFTQQIEE 1318
Query: 464 YHVQRQLREEEKRKSREQKRLQ----------EQHAVEQEA 494
++RQL EE K KS LQ EQ+ EQEA
Sbjct: 1319 --LKRQLEEETKAKSTLAHALQSARHDCDLLREQYEEEQEA 1357
Score = 40.4 bits (93), Expect = 0.012
Identities = 104/541 (19%), Positives = 221/541 (40%), Gaps = 78/541 (14%)
Query: 15 CASLLSQLQMIWDEIGES--DSDRDN---MLLQLEQECLDIYRRKVDATRKHKAELCQWL 69
CASL Q + +E+ + D +R N + L +Q D KV A K K E Q
Sbjct: 1415 CASLEKTKQRLQNEVEDLMIDVERSNAACIALDKKQRNFD----KVLAEWKQKYEETQAE 1470
Query: 70 ADAE--------AELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDE---RV 118
+A EL + ++ ES TLK++ ++ + DL + E +
Sbjct: 1471 LEASQKESRSLSTELFKVKNAYEESLDHLE---TLKRENKNLQQEISDLTEQIAEGGKHI 1527
Query: 119 KEFSKIKSQISQICVEI-AGYEQPKSSSEEVDQSDLTFK-KLGELKSHLQDLQNEKILRQ 176
E K+K Q+ E+ E+ ++S E + L + +L ++KS + EK
Sbjct: 1528 HELEKVKKQLDHEKSELQTSLEEAEASLEHEEGKILRIQLELNQVKSEIDRKIAEKDEEL 1587
Query: 177 QKVK-SHISTISELSAVMSIDFWKT-------------LSEIHPSLGDSSKGTPQSISND 222
++K +H+ + + + + + L+E+ L +++ +++ N
Sbjct: 1588 DQLKRNHLRVVESMQSTLDAEIRSRNDALRIKKKMEGDLNEMEIQLNHANRQAAEALRN- 1646
Query: 223 TLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLW---DLIETPIDEQKAFNHVTR----- 274
L + G + + + +L + L + +L++ ++E +A T
Sbjct: 1647 -LRNTQGILKDTQLHLDDAIRGQDDLKEQLAMVERRANLMQAEVEELRASLERTERGRKM 1705
Query: 275 ----LISAS--VDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYR 328
L+ AS V + T + +++E ++ ++ +M+D+V + +N E+ +
Sbjct: 1706 AEQELLDASERVQLLHTQNTSLINTKKKLETDISQIQG----EMEDIVQEARNAEEKAKK 1761
Query: 329 GVH---MDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKW 385
+ M + Q ++ +E ++ ++ + ++ +A++ A + +++K
Sbjct: 1762 AITDAAMMAEELKKEQDTSAHLERMKKNMEQTVKDLQLRLGEAEQLALKGGK--KQIQKL 1819
Query: 386 RFAAEEEKWLDEYERDENRYSAVRGAHKNLKRA-------EKARIVVSKIPSIVENLTTK 438
E + E E+ N AV+G K+ +R E+ R + ++ +V+ L TK
Sbjct: 1820 EARVRELESEVESEQKHN-VEAVKGLRKHERRVKELTYQTEEDRKNILRLQDLVDKLQTK 1878
Query: 439 VKAWEM------EKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQ 492
VKA++ E+ L + L LEE + + E + K R + R + +
Sbjct: 1879 VKAYKRQAEEAEEQSNVNLAKFRKLQHELEEAEERADIAESQVNKLRVKSREVHTKVISE 1938
Query: 493 E 493
E
Sbjct: 1939 E 1939
>MYSP_SCHMA (P06198) Paramyosin
Length = 866
Score = 57.8 bits (138), Expect = 8e-08
Identities = 106/491 (21%), Positives = 219/491 (44%), Gaps = 77/491 (15%)
Query: 17 SLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAEL 76
S L++L+ + D++ ++ +N +L E ++ D ++A++ + + A++ L
Sbjct: 170 SQLNRLKSLTDDLQRQLTELNNAKSRLTSENFELLHINQD----YEAQILNY-SKAKSSL 224
Query: 77 INLVSSLGES-SSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEI 135
+ V L S ++ + L+ QL +++ ++L++K DE +E S ++SQ+S+ +I
Sbjct: 225 ESQVDDLKRSLDDEAKNRFNLQAQLTSLQMDYDNLQAKYDEESEEASNLRSQVSKFNADI 284
Query: 136 AGYE-----QPKSSSEEVDQSDLTF-KKLGELKS----------HLQDLQNEKILRQQKV 179
A + + S +EE ++ F ++ EL+ L+ L+ + L + +
Sbjct: 285 AALKSKFERELMSKTEEFEEMKRKFTMRITELEDTAERERLKAVSLEKLKTKLTLEIKDL 344
Query: 180 KSHISTIS----------ELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISND-----TL 224
+S I ++S + + ++ D + + E+ + + Q S + +
Sbjct: 345 QSEIESLSLENSELIRRAKAAESLASDLQRRVDELTIEVNTLTSQNSQLESENLRLKSLV 404
Query: 225 ASLTGYIHSLKQEKQQRLLKVQELTKFLVD----LWDL--IETPIDEQKAFNHVTRLISA 278
LT + L++E +Q +V+EL L D L DL + + ++ ++ L SA
Sbjct: 405 NDLTDKNNLLERENRQMNDQVKELKSSLRDANRRLTDLEALRSQLEAER-----DNLASA 459
Query: 279 SVD-EVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRG-------- 329
D E + H D+ ++ +A LN LK S+M+ + +R ELE + +
Sbjct: 460 LHDAEEALH-----DMDQKYQASQAALNHLK-SEMEQRLRERDEELESLRKSTTRTIEEL 513
Query: 330 ----VHMDVDSEAARQILTSLIESGNIDLSDLLQSMD----DQIRQAKEQAQSRREI--- 378
M+V ++ L ES DL L + + + +++ K +Q +++
Sbjct: 514 TVTITEMEVKYKSELSRLKKRYESNIADLEIQLDTANKANANLMKENKNLSQRVKDLETF 573
Query: 379 LDRVEKWRFAAEEEKWLDEYERDE--NRYSAVRGAHKNLKRAEK-ARIVVSKIPSIVENL 435
LD + R AAE + E++R + N +R +NL+R K A + + S V L
Sbjct: 574 LDEERRLREAAENNLQITEHKRLQLANEIEEIRSTLENLERLRKHAETELEEAQSRVSEL 633
Query: 436 TTKVKAWEMEK 446
T +V +K
Sbjct: 634 TIQVNTLTNDK 644
>GCC2_MOUSE (Q8CHG3) GRIP and coiled-coil domain-containing protein 2
(Golgi coiled coil protein GCC185)
Length = 1679
Score = 56.6 bits (135), Expect = 2e-07
Identities = 100/536 (18%), Positives = 221/536 (40%), Gaps = 56/536 (10%)
Query: 9 SPSRTTCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQW 68
SPS SL+ +L+ + + + D+D + +++ + ++++D+ RK L +
Sbjct: 927 SPSVKDQLSLVKELEEKIESLEKESKDKDEKISKIKLVAVKA-KKELDSNRKEAQTLREE 985
Query: 69 LADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSK-IKSQ 127
L +E L +S+ E F +G + K L E L +K ER F + I+
Sbjct: 986 LESVRSEKDRLSASMKE---FLQGAESYKSLLLEYDKQSEQLDVEK-ERAHNFERHIEDL 1041
Query: 128 ISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKSHL------------QDLQNEKILR 175
Q+ YE+ S +E++ T + +L ++L E++ +
Sbjct: 1042 TKQLRNSTCQYERLTSDNEDLLARIETLQANAKLLEAQILEVQKAKGVVEKELDAEELQK 1101
Query: 176 QQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLK 235
+QK+K H+ST++EL + + F K ++ ++ + + + D + + +++
Sbjct: 1102 EQKIKEHVSTVNELEE-LQLQFQKEKKQLQKTMQEL-----ELVKKDAQQTT---LMNME 1152
Query: 236 QEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIE 295
+RL+K EL + L + IE E K + + + + ++
Sbjct: 1153 IADYERLMK--ELNQKLTNKNSTIEDLEQEMKIQKEKQETLQEEITSLQSS-------VQ 1203
Query: 296 QVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLS 355
E + ++ L K+L +Q E + + + + EA++Q +E I L+
Sbjct: 1204 HYEEKNTKIKQLLVKTKKELADAKQAETDHLLLQASLKGELEASQQ----QVEVYKIQLA 1259
Query: 356 DLL---QSMDDQIRQAKEQ------AQSRREILDRVEKWRFAAEEEKWLDEYERDENRYS 406
++ + + ++ + EQ A +R + + E AE+ E+E + R
Sbjct: 1260 EMTSEKHKIHEHLKTSAEQHQRTLSAYQQRVVALQEESRAAKAEQAAVTSEFESYKVRVH 1319
Query: 407 AVRGAHKNLK----RAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLE 462
V KN E A+ + +++ L K+K + + + + LQ+
Sbjct: 1320 NVLKQQKNKSVSQVETEGAKQEREHLEMLIDQL--KIKLQDSQNSLQISVSEYQTLQAEH 1377
Query: 463 EYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQSTTANTIVGT 518
+ ++R R ++ ++E + ++ +V+ E +S + + Q T+ N + T
Sbjct: 1378 DTLLERHNRMLQETVTKEAELREKLCSVQSENTM-MKSEHSQTMCQLTSQNEALRT 1432
Score = 32.3 bits (72), Expect = 3.4
Identities = 87/450 (19%), Positives = 174/450 (38%), Gaps = 77/450 (17%)
Query: 94 GTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEE------ 147
GT K +L T+ EDL +++ K K++ +++ E+ E+ KS +
Sbjct: 15 GTPKSKLETLPR--EDLIKFAKKQMMLLQKAKARCTELDKEV---EELKSKPVDGGTDDI 69
Query: 148 ----VDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSE 203
++ D + E + L+ E + +Q+V+ ++ + E + E
Sbjct: 70 IKVLTERLDALLLEKAETEQQCLCLKKENVKMKQEVEDSVTKLEETHKEFEQSHRNYVKE 129
Query: 204 IHPSLGDSSKGTPQSISNDTLASLTGYIHS--------LKQEKQQRLLKVQELTKFLVDL 255
I +S N+ +A +HS L++E ++ + K EL + L
Sbjct: 130 I------------ESCKNELMA-----VHSEHSKETAILQKELEEAVHKQVELREQLKSQ 172
Query: 256 WDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDL 315
D + Q+ ++T + + +SD +Q +Q++ KA +
Sbjct: 173 SDSEDNVRKLQEEIQNITAAFEEQISCLEKKLEATSDEKQQEIIHLQKVIEDKAQHYQKD 232
Query: 316 VFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAK---EQA 372
+ Q E+ ++ R H + +E QI TS E +++ L ++ Q ++ E+
Sbjct: 233 INTFQAEILQL-RATHKEEVTELMSQIETSAKEH-EAEINKLKENRVTQCEASENIPEKY 290
Query: 373 QSRREILDRV--------EKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIV 424
Q E L+ V + A +E+ ++ D+ VR +LK E +
Sbjct: 291 QCESENLNEVASDASPESQNCSVALQEDPSAEQTVCDK-----VRQLEDSLKELESQHSI 345
Query: 425 VSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPL--------LQSLEEYHVQRQLREEEK- 475
+ + + NL K++ F +E+ L L E+ +V +L+ E +
Sbjct: 346 LKDEVTYMNNLKLKLEMDAQHIKDEFFHEREDLEFKINELLLAKEEQGYVVEKLKYERED 405
Query: 476 ----------RKSREQKRLQEQHAVEQEAL 495
+ ++E +RLQE H E L
Sbjct: 406 LNRQLCCAVEQHNKEIQRLQEHHQKEVSEL 435
>MYH2_HUMAN (Q9UKX2) Myosin heavy chain, skeletal muscle, adult 2
(Myosin heavy chain IIa) (MyHC-IIa)
Length = 1941
Score = 56.2 bits (134), Expect = 2e-07
Identities = 114/547 (20%), Positives = 228/547 (40%), Gaps = 81/547 (14%)
Query: 16 ASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAE 75
A++ + Q I DE+ +S++ R +LE++ + + + K D + +AE + LADAE
Sbjct: 856 ATMKEEFQKIKDELAKSEAKRK----ELEEKMVTLLKEKNDLQLQVQAE-AEGLADAEER 910
Query: 76 LINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEI 135
L+ + + + + + ++ + +L +KK + E S++K I + + +
Sbjct: 911 CDQLIKTKIQLEAKIK---EVTERAEDEEEINAELTAKKRKLEDECSELKKDIDDLELTL 967
Query: 136 AGYEQPKSSSE--------EVDQSDLTFKKLGELKSHLQDLQNEKI--LRQQKVKSHIST 185
A E+ K ++E E+ D T KL + K LQ+ + + L+ ++ K + T
Sbjct: 968 AKVEKEKHATENKVKNLTEEMAGLDETIAKLTKEKKALQEAHQQTLDDLQAEEDKVNTLT 1027
Query: 186 ISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKV 245
+++ +D + E L + + + D L I ++ EKQQ K+
Sbjct: 1028 KAKIKLEQQVDDLEGSLEQEKKLRMDLERAKRKLEGD-LKLAQESIMDIENEKQQLDEKL 1086
Query: 246 QELTKFLVDLWDLIETPIDEQKAFNHVTRLI---SASVDEVSTHGCLSSDVIEQVEAEVQ 302
++ + +L IE DEQ + + I A ++E+ E++EAE
Sbjct: 1087 KKKEFEISNLQSKIE---DEQALGIQLQKKIKELQARIEELE----------EEIEAE-- 1131
Query: 303 RLNALKASKMKDLVFKRQNELEE--------IYRGVHMDVDSEAARQILTSLIESGNID- 353
R + KA K + + + E+ E + M+ EA Q + +E +
Sbjct: 1132 RASRAKAEKQRSDLSRELEEISERLEEAGGATSAQIEMNKKREAEFQKMRRDLEEATLQH 1191
Query: 354 ---LSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRG 410
+ L + D + + EQ + + + ++EK + +E + +D+ + S +G
Sbjct: 1192 EATAATLRKKHADSVAELGEQIDNLQRVKQKLEKEK--SEMKMEIDDLASNVETVSKAKG 1249
Query: 411 AHKNLKRAEKARIVVSKIPS-----IVENLTTKVKAWEMEKG--IPFLYEKAPLLQSLEE 463
+ + R + ++ K ++ +LT + + E G L EK L+ L
Sbjct: 1250 NLEKMCRTLEDQLSELKSKEEEQQRLINDLTAQRGRLQTESGEFSRQLDEKEALVSQLSR 1309
Query: 464 ---------YHVQRQLREEEKRKSREQKRLQ----------EQHAVEQEALFGSRSATKK 504
++RQL EE K K+ LQ EQ+ EQE S++ ++
Sbjct: 1310 GKQAFTQQIEELKRQLEEEIKAKNALAHALQSSRHDCDLLREQYEEEQE----SKAELQR 1365
Query: 505 PLGQSTT 511
L ++ T
Sbjct: 1366 ALSKANT 1372
Score = 38.5 bits (88), Expect = 0.047
Identities = 69/355 (19%), Positives = 149/355 (41%), Gaps = 40/355 (11%)
Query: 54 KVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSK 113
++D ++ K +L + ++ + E+ +L S++ + S+ KG L++ T+ L +L+SK
Sbjct: 1212 QIDNLQRVKQKLEKEKSEMKMEIDDLASNV---ETVSKAKGNLEKMCRTLEDQLSELKSK 1268
Query: 114 KDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELK-SHLQDLQNEK 172
++E+ + + + +Q ++ E + S ++D+ + +L K + Q ++ K
Sbjct: 1269 EEEQQRLINDLTAQRGRLQTESGEF------SRQLDEKEALVSQLSRGKQAFTQQIEELK 1322
Query: 173 ILRQQKVKSHISTISEL-SAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYI 231
++++K+ + L S+ D + E SK Q + + +
Sbjct: 1323 RQLEEEIKAKNALAHALQSSRHDCDLLREQYEEE----QESKAELQRALSKANTEVAQWR 1378
Query: 232 HSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSS 291
+ + QR +++E K L Q A HV + + T L +
Sbjct: 1379 TKYETDAIQRTEELEEAKKKLAQRL---------QAAEEHVEAVNAKCASLEKTKQRLQN 1429
Query: 292 DVIEQVEAEVQRLNAL------KASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTS 345
+V E + +V+R NA K ++ + + + EE + ++ + AR + T
Sbjct: 1430 EV-EDLMLDVERTNAACAALDKKQRNFDKILAEWKQKCEETH--AELEASQKEARSLGTE 1486
Query: 346 LIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYER 400
L + N +S+D +E ++EI D E+ AE K + E E+
Sbjct: 1487 LFKIKNA----YEESLDQLETLKRENKNLQQEISDLTEQ---IAEGGKRIHELEK 1534
>MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (SMMHC)
Length = 1972
Score = 55.5 bits (132), Expect = 4e-07
Identities = 112/545 (20%), Positives = 233/545 (42%), Gaps = 89/545 (16%)
Query: 18 LLSQLQMIWDEIGESDSDRDNMLLQ----------LEQECL--DIYRRKVDATRK----H 61
+ Q+ + +++ E ++ R + L+ LE E L D K+ RK
Sbjct: 948 MAQQMLDLEEQLEEEEAARQKLQLEKVTAEAKIKKLEDEILVMDDQNNKLSKERKLLEER 1007
Query: 62 KAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEF 121
++L LA+ E + NL + S ++ R LE L+ K + +F
Sbjct: 1008 ISDLTTNLAEEEEKAKNLTKLKNKHESMISELEVRLKKEEKSRQELEKLKRKLEGDASDF 1067
Query: 122 SK----IKSQISQICVEIAGYEQPKSSS-----EEVDQSDLTFKKLGELKSHLQDLQ--- 169
+ +++QI+++ +++A E+ ++ +E+ Q + KK+ EL+ H+ DLQ
Sbjct: 1068 HEQIADLQAQIAELKMQLAKKEEELQAALARLDDEIAQKNNALKKIRELEGHISDLQEDL 1127
Query: 170 -NEKILRQQKVKSHISTISELSAVM-----SIDFWKTLSEIHPSLGDSSKGTPQSISNDT 223
+E+ R + K EL A+ ++D T E+ +++ +T
Sbjct: 1128 DSERAARNKAEKQKRDLGEELEALKTELEDTLDSTATQQELRAKREQEVTVLKKALDEET 1187
Query: 224 LASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEV 283
+ +++ +Q+ V+ELT+ L + + + +D+ K + + + E+
Sbjct: 1188 ----RSHEAQVQEMRQKHAQAVEELTEQL-EQFKRAKANLDKNK--QTLEKENADLAGEL 1240
Query: 284 STHGCLSSDV---IEQVEAEVQRLNA------LKASKMKDLVFKRQNELEEIYRGVHMDV 334
G +V +++EA+VQ L + +++ D V K QNE+E + G+ +
Sbjct: 1241 RVLGQAKQEVEHKKKKLEAQVQELQSKCSDGERARAELNDKVHKLQNEVESV-TGMLNEA 1299
Query: 335 DSEAARQIL-TSLIESGNIDLSDLLQ-------SMDDQIRQAKEQAQSRREILDRVEKWR 386
+ +A + + + S D +LLQ ++ ++RQ +E+ S ++ LD + +
Sbjct: 1300 EGKAIKLAKDVASLSSQLQDTQELLQEETRQKLNVSTKLRQLEEERNSLQDQLDEEMEAK 1359
Query: 387 FAAEEE------KWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVK 440
E + D ++ ++ S V + KR +K +I ++ + K
Sbjct: 1360 QNLERHISTLNIQLSDSKKKLQDFASTVEALEEGKKRFQK------EIENLTQQYEEKAA 1413
Query: 441 AWE-MEKGIPFLYEKAPLLQSLEEYHV----QRQLREEEKRKSR-------EQKRLQEQH 488
A++ +EK K L Q L++ V QRQL ++K R E+K + ++
Sbjct: 1414 AYDKLEK------TKNRLQQELDDLVVDLDNQRQLVSNLEKKQRKFDQLLAEEKNISSKY 1467
Query: 489 AVEQE 493
A E++
Sbjct: 1468 ADERD 1472
Score = 45.8 bits (107), Expect = 3e-04
Identities = 96/480 (20%), Positives = 196/480 (40%), Gaps = 56/480 (11%)
Query: 45 QECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIR 104
QE LD R + K K +L + L + EL + + S + +Q++ ++
Sbjct: 1124 QEDLDSERAARNKAEKQKRDLGEELEALKTELEDTLDSTATQQEL---RAKREQEVTVLK 1180
Query: 105 PVLEDLRSKKDERVKEFSKIKSQ-ISQICVEIAGYEQPKSSSE------EVDQSDLT--F 155
L++ + +V+E + +Q + ++ ++ +++ K++ + E + +DL
Sbjct: 1181 KALDEETRSHEAQVQEMRQKHAQAVEELTEQLEQFKRAKANLDKNKQTLEKENADLAGEL 1240
Query: 156 KKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLG--DSSK 213
+ LG+ K ++ + + + Q+++S S A ++ K +E+ G + ++
Sbjct: 1241 RVLGQAKQEVEHKKKKLEAQVQELQSKCSDGERARAELNDKVHKLQNEVESVTGMLNEAE 1300
Query: 214 GTPQSISNDTLASLTGYIHS---LKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQ-KAF 269
G ++ D +ASL+ + L QE+ ++ L V + L + + ++ +DE+ +A
Sbjct: 1301 GKAIKLAKD-VASLSSQLQDTQELLQEETRQKLNVSTKLRQLEEERNSLQDQLDEEMEAK 1359
Query: 270 NHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRG 329
++ R +ST SD ++++ + AL+ K + Q E+E + +
Sbjct: 1360 QNLER-------HISTLNIQLSDSKKKLQDFASTVEALEEGKKRF-----QKEIENLTQ- 1406
Query: 330 VHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAA 389
E L ++ N L Q +DD + Q Q + + K+
Sbjct: 1407 -----QYEEKAAAYDKLEKTKN----RLQQELDDLVVDLDNQRQLVSNLEKKQRKFDQLL 1457
Query: 390 EEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVE-NLTTKVKAWEME--- 445
EEK + DE + K K AR + + + E T K+ EME
Sbjct: 1458 AEEKNISSKYADERDRAEAEAREKETKALSLARALEEALEAKEELERTNKMLKAEMEDLV 1517
Query: 446 -------KGIPFLYE-KAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFG 497
K + L + K L +EE Q + E+E + + + K E V +AL G
Sbjct: 1518 SSKDDVGKNVHELEKSKRALETQMEEMKTQLEELEDELQATEDAKLRLE---VNMQALKG 1574
Score = 42.0 bits (97), Expect = 0.004
Identities = 104/505 (20%), Positives = 207/505 (40%), Gaps = 70/505 (13%)
Query: 42 QLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLA 101
QL E +I + D + +AE + E + ++L +L E+ K L++
Sbjct: 1455 QLLAEEKNISSKYADERDRAEAEA----REKETKALSLARALEEALE---AKEELERTNK 1507
Query: 102 TIRPVLEDLRSKKDE---RVKEFSKIK----SQISQICVEIAGYEQPKSSSEEVD-QSDL 153
++ +EDL S KD+ V E K K +Q+ ++ ++ E ++E+ + ++
Sbjct: 1508 MLKAEMEDLVSSKDDVGKNVHELEKSKRALETQMEEMKTQLEELEDELQATEDAKLRLEV 1567
Query: 154 TFKKL-GELKSHLQ--DLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGD 210
+ L G+ + LQ D QNE+ RQ + + H E L D
Sbjct: 1568 NMQALKGQFERDLQARDEQNEEKRRQLQRQLH--------------------EYETELED 1607
Query: 211 SSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFN 270
K ++++ L G + L+ + + +E K L L + K F
Sbjct: 1608 ERK--QRALAAAAKKKLEGDLKDLELQADSAIKGREEAIKQLRKLQA-------QMKDFQ 1658
Query: 271 HVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLN-----ALKASKMKDLVFKR-QNELE 324
AS DE+ + + +EA++ +L A +A K DL + EL
Sbjct: 1659 RELEDARASRDEIFATAKENEKKAKSLEADLMQLQEDLAAAERARKQADLEKEELAEELA 1718
Query: 325 EIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEK 384
G + D + + + +E + +++M D++R+A +QA+ L +
Sbjct: 1719 SSLSGRNALQDEKRRLEARIAQLEEELEEEQGNMEAMSDRVRKATQQAEQLSNEL--ATE 1776
Query: 385 WRFAAEEEKWLDEYERD----ENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVK 440
A + E + ER ++ + GA K+ ++ A + +KI + E + + +
Sbjct: 1777 RSTAQKNESARQQLERQNKELRSKLHEMEGAVKSKFKSTIAAL-EAKIAQLEEQV--EQE 1833
Query: 441 AWEMEKGIPFLYEKAP-----LLQSLEEYHVQRQLREEEKR---KSREQKRLQEQHAVEQ 492
A E + L +K LLQ +E + Q +E+ ++ + ++ KR E+ E
Sbjct: 1834 AREKQAATKSLKQKDKKLKEILLQVEDERKMAEQYKEQAEKGNARVKQLKRQLEEAEEES 1893
Query: 493 EALFGSRSATKKPLGQSTTANTIVG 517
+ + +R ++ L ++T +N +G
Sbjct: 1894 QRINANRRKLQRELDEATESNEAMG 1918
Score = 32.0 bits (71), Expect = 4.4
Identities = 58/292 (19%), Positives = 104/292 (34%), Gaps = 56/292 (19%)
Query: 27 DEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELI--NLVSSLG 84
DEI + + + LE + + + A R K AD E E + L SSL
Sbjct: 1669 DEIFATAKENEKKAKSLEADLMQLQEDLAAAERARKQ------ADLEKEELAEELASSLS 1722
Query: 85 ESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSS 144
++ K L+ ++A + LE+ + + K Q Q+ E+A
Sbjct: 1723 GRNALQDEKRRLEARIAQLEEELEEEQGNMEAMSDRVRKATQQAEQLSNELATERSTAQK 1782
Query: 145 SEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHI-STISELSAVMSIDFWKTLSE 203
+E Q ++ EL+S L +++ VKS STI+ L A
Sbjct: 1783 NESARQQ--LERQNKELRSKLHEMEG-------AVKSKFKSTIAALEA------------ 1821
Query: 204 IHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPI 263
+A L + +EKQ +++ K L ++ +E
Sbjct: 1822 -------------------KIAQLEEQVEQEAREKQAATKSLKQKDKKLKEILLQVEDER 1862
Query: 264 DEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDL 315
+ + +A V ++ +E+ E E QR+NA + ++L
Sbjct: 1863 KMAEQYKEQAEKGNARVKQLKRQ-------LEEAEEESQRINANRRKLQREL 1907
>MYSD_CAEEL (P02567) Myosin heavy chain D (MHC D)
Length = 1938
Score = 54.7 bits (130), Expect = 6e-07
Identities = 108/538 (20%), Positives = 209/538 (38%), Gaps = 82/538 (15%)
Query: 19 LSQLQMIWDE----IGESDSDR-----DNMLLQLEQECLDIYRRKVD-ATRKHKAELCQW 68
+++LQ DE IGE S + DN L + E L+I+ ++ A ++L +
Sbjct: 1263 VAELQQKVDEQSRQIGEYTSTKGRLSNDNSDLARQVEELEIHLATINRAKTAFSSQLVEA 1322
Query: 69 LADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQI 128
AE EL E F L+ +L +LE+ + KD+ ++ S+I S+I
Sbjct: 1323 KKAAEDEL-------HERQEFHAACKNLEHELDQCHELLEEQINGKDDIQRQLSRINSEI 1375
Query: 129 SQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKS------- 181
SQ G E S E + ++ +L+ L QN K++ +K K
Sbjct: 1376 SQWKARYEG-EGLVGSEELEELKRKQMNRVMDLQEALSAAQN-KVISLEKAKGKLLAETE 1433
Query: 182 --------HISTISELSAVMS-----IDFWK-----TLSEIHPSLGDSSKGTPQSIS-ND 222
H++ I+ L +D WK EI + DS + +
Sbjct: 1434 DARSDVDRHLTVIASLEKKQRAFDKIVDDWKRKVDDIQKEIDATTRDSRNTSTEVFKLRS 1493
Query: 223 TLASLTGYIHSLKQEKQQRLLKVQELTKFLV-------DLWDLIETPIDEQKAFNHVTRL 275
++ +L+ I +L++E + +++++ + + ++ + E+ H
Sbjct: 1494 SMDNLSEQIETLRRENKIFSQEIRDINEQITQGGRTYQEVHKSVRRLEQEKDELQHALDE 1553
Query: 276 ISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVD 335
A+++ + ++Q+ +E+++ K + ++ Q LE I + +
Sbjct: 1554 AEAALEAEESKVLRLQIEVQQIRSEIEKRIQEKEEEFENTRKNHQRALESIQASLETEAK 1613
Query: 336 SEA--AR------------QILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDR 381
S+A AR +I N+D L+ + DQ+++ + Q + +
Sbjct: 1614 SKAELARAKKKLETDINQLEIALDHANKANVDAQKNLKKLFDQVKELQGQVDDEQRRREE 1673
Query: 382 VEKWRFAAEEEK--WLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKV 439
+ + AAE+ L E E +R A HK E+A + S I N
Sbjct: 1674 IRENYLAAEKRLAIALSESEDLAHRIEA-SDKHKKQLEIEQAELKSSNTELIGNNAALSA 1732
Query: 440 KAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKS-------REQKRLQEQHAV 490
++E + + L+EY + + EE RK+ E+ R +++HAV
Sbjct: 1733 MKRKVENEVQIARNE------LDEYLNELKASEERARKAAADADRLAEEVRQEQEHAV 1784
Score = 38.9 bits (89), Expect = 0.036
Identities = 90/488 (18%), Positives = 192/488 (38%), Gaps = 69/488 (14%)
Query: 44 EQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVS-SLGESSSFSRGKGTLKQQLAT 102
EQ+ L+ +K ++ RK + E L+ ++L+ + + G S++ L
Sbjct: 862 EQKVLEAEAKKAESARKSQEEAYAKLSAERSKLLEALELTQGGSAAIEEKLTRLNSARQE 921
Query: 103 IRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELK 162
+ L D + E ++ + ++ Q + E+ E K S E VD G L
Sbjct: 922 VEKSLNDANDRLSEHEEKNADLEKQRRKAQQEV---ENLKKSIEAVD---------GNLA 969
Query: 163 SHLQDL---QNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDS--SKGTPQ 217
L++ +N+ Q ++ S TI +++ K L E + L D ++ Q
Sbjct: 970 KSLEEKAAKENQIHSLQDEMNSQDETIGKINKEK-----KLLEENNRQLVDDLQAEEAKQ 1024
Query: 218 SISNDTLASLTGYIHSLKQ--EKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRL 275
+ +N L + +++ E+++R+ E +K V+ E K
Sbjct: 1025 AQANRLRGKLEQTLDEMEEAVEREKRIRAETEKSKRKVE---------GELKGAQE---- 1071
Query: 276 ISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVD 335
++DE+S + +++ EA++ L + + E R + ++ +
Sbjct: 1072 ---TIDELSAIKLETDASLKKKEADIHALGVRIEDEQALANRLTRQSKENAQRIIEIEDE 1128
Query: 336 SEAARQILTSLIESG---NIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEE 392
E RQ + + +L +L + +D+Q +Q + Q + ++ + K+R
Sbjct: 1129 LEHERQSRSKADRARAELQRELDELNERLDEQNKQLEIQQDNNKKKDSEIIKFR------ 1182
Query: 393 KWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLY 452
+ LDE KN+ ++ ++ K + LT + A + K
Sbjct: 1183 RDLDE---------------KNMANEDQMAMIRRKNNDQISALTNTLDALQKSKA-KIEK 1226
Query: 453 EKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQSTTA 512
EK L + L++ + Q ++E + EQ+RL +Q+ ++ L + +G+ T+
Sbjct: 1227 EKGVLQKELDDINAQ---VDQETKSRVEQERLAKQYEIQVAELQQKVDEQSRQIGEYTST 1283
Query: 513 NTIVGTPN 520
+ N
Sbjct: 1284 KGRLSNDN 1291
>MYHB_CHICK (P10587) Myosin heavy chain, gizzard smooth muscle
Length = 1978
Score = 54.7 bits (130), Expect = 6e-07
Identities = 102/481 (21%), Positives = 203/481 (41%), Gaps = 57/481 (11%)
Query: 37 DNMLLQLEQECLDIYRRKVDATRKHK------AELCQWLADAEAELINLV-------SSL 83
D + ++E + L + + T++ K ++L LA+ E + NL S +
Sbjct: 982 DGKIKKMEDDILIMEDQNNKLTKERKLLEERVSDLTTNLAEEEEKAKNLTKLKNKHESMI 1041
Query: 84 GESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKS 143
E + + +Q+L I+ LE S E++ E +++QI+++ ++A E+
Sbjct: 1042 SELEVRLKKEEKSRQELEKIKRKLEGESSDLHEQIAE---LQAQIAELKAQLAKKEEELQ 1098
Query: 144 SS-----EEVDQSDLTFKKLGELKSHLQDLQ----NEKILRQQKVKSHISTISELSAVMS 194
++ +E Q + KK+ EL+SH+ DLQ +EK R + K EL A+
Sbjct: 1099 AALARLEDETSQKNNALKKIRELESHISDLQEDLESEKAARNKAEKQKRDLSEELEALK- 1157
Query: 195 IDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVD 254
+E+ +L + T Q + +T +L++E + +VQE+ +
Sbjct: 1158 -------TELEDTL--DTTATQQELRAKREQEVTVLKRALEEETRTHEAQVQEMRQKHTQ 1208
Query: 255 LWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKD 314
+ + +++ K ++++ + + Q + +V+ +++D
Sbjct: 1209 AVEELTEQLEQFKRAKANLDKTKQTLEKDNADLANEIRSLSQAKQDVEHKKKKLEVQLQD 1268
Query: 315 LVFKR------QNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQA 368
L K + EL E + ++V++ + L + ES NI L+ + ++ Q++
Sbjct: 1269 LQSKYSDGERVRTELNEKVHKLQIEVENVTS---LLNEAESKNIKLTKDVATLGSQLQDT 1325
Query: 369 KEQAQSR-REILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKR-AEKARIVVS 426
+E Q R+ L+ K R +++ L E +E A +NL+R I +S
Sbjct: 1326 QELLQEETRQKLNVTTKLRQLEDDKNSLQEQLDEEVE------AKQNLERHISTLTIQLS 1379
Query: 427 KIPSIVENLTTKVKAWEMEKGIPFLY-EKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQ 485
++ T V+ ME+G L E L Q EE + EK K+R Q+ L
Sbjct: 1380 DSKKKLQEFTATVET--MEEGKKKLQREIESLTQQFEEKAASYD--KLEKTKNRLQQELD 1435
Query: 486 E 486
+
Sbjct: 1436 D 1436
Score = 53.5 bits (127), Expect = 1e-06
Identities = 97/550 (17%), Positives = 244/550 (43%), Gaps = 50/550 (9%)
Query: 12 RTTCASLLSQLQMIWDEIG---ESDSDRDNMLL----QLEQECLDIYRR-KVDATRKHKA 63
R A+ +L+ I E+ E + +R L +++Q+ LD+ + + + + K
Sbjct: 915 RVRLAAKKQELEEILHEMEARIEEEEERSQQLQAEKKKMQQQMLDLEEQLEEEEAARQKL 974
Query: 64 ELCQWLADAEAELI--NLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEF 121
+L + AD + + + +++ +++ ++ + L+++++ + L + K K
Sbjct: 975 QLEKVTADGKIKKMEDDILIMEDQNNKLTKERKLLEERVSDLTTNLAEEEEKAKNLTKLK 1034
Query: 122 SKIKSQISQICVEIAGYEQPKSSSEEV------DQSDLTFKKLGELKSHLQDLQNEKILR 175
+K +S IS++ V + E+ + E++ + SDL +++ EL++ + +L+ + +
Sbjct: 1035 NKHESMISELEVRLKKEEKSRQELEKIKRKLEGESSDL-HEQIAELQAQIAELKAQLAKK 1093
Query: 176 QQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLK 235
++++++ ++ + + ++ + + K + E+ + D + + ++ A K
Sbjct: 1094 EEELQAALARLEDETSQKN-NALKKIRELESHISDLQ----EDLESEKAARN-------K 1141
Query: 236 QEKQQRLLKVQELTKFLVDLWDLIETPIDEQ----KAFNHVTRLISASVDEVSTHGCLSS 291
EKQ+R L +EL +L D ++T +Q K VT L A +E TH
Sbjct: 1142 AEKQKRDLS-EELEALKTELEDTLDTTATQQELRAKREQEVTVLKRALEEETRTHEAQVQ 1200
Query: 292 DV-------IEQVEAEVQRLNALKAS--KMKDLVFKRQNELEEIYRGVHM---DVDSEAA 339
++ +E++ ++++ KA+ K K + K +L R + DV+ +
Sbjct: 1201 EMRQKHTQAVEELTEQLEQFKRAKANLDKTKQTLEKDNADLANEIRSLSQAKQDVEHKKK 1260
Query: 340 R-QILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEY 398
+ ++ ++S D + +++++ + + + ++ +L+ E ++
Sbjct: 1261 KLEVQLQDLQSKYSDGERVRTELNEKVHKLQIEVENVTSLLNEAESKNIKLTKDVATLGS 1320
Query: 399 ERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWE-MEKGIPFL-YEKAP 456
+ + + + L K R + S+ E L +V+A + +E+ I L + +
Sbjct: 1321 QLQDTQELLQEETRQKLNVTTKLRQLEDDKNSLQEQLDEEVEAKQNLERHISTLTIQLSD 1380
Query: 457 LLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQSTTANTIV 516
+ L+E+ + EE K+K + + Q E+ A + TK L Q + +V
Sbjct: 1381 SKKKLQEFTATVETMEEGKKKLQREIESLTQQFEEKAASYDKLEKTKNRL-QQELDDLVV 1439
Query: 517 GTPNGRRMLT 526
N R++++
Sbjct: 1440 DLDNQRQLVS 1449
Score = 46.2 bits (108), Expect = 2e-04
Identities = 121/542 (22%), Positives = 213/542 (38%), Gaps = 89/542 (16%)
Query: 17 SLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAEL 76
S++S+L++ + +S + + + +LE E D++ + + + AEL LA E EL
Sbjct: 1039 SMISELEVRLKKEEKSRQELEKIKRKLEGESSDLHEQ-IAELQAQIAELKAQLAKKEEEL 1097
Query: 77 INLVSSL-GESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEI 135
++ L E+S + +++ + I + EDL S+K R K K K +S+ +E
Sbjct: 1098 QAALARLEDETSQKNNALKKIRELESHISDLQEDLESEKAARNKA-EKQKRDLSEE-LEA 1155
Query: 136 AGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKS----HISTISELSA 191
E + Q +L K+ E+ + L+ E + +V+ H + EL+
Sbjct: 1156 LKTELEDTLDTTATQQELRAKREQEVTVLKRALEEETRTHEAQVQEMRQKHTQAVEELT- 1214
Query: 192 VMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKF 251
+ L + + + K T Q++ D A L I SL Q KQ K ++L
Sbjct: 1215 -------EQLEQFKRAKANLDK-TKQTLEKDN-ADLANEIRSLSQAKQDVEHKKKKLEVQ 1265
Query: 252 LVDLW------DLIETPIDE-----QKAFNHVTRLISAS-------VDEVSTHGCLSSDV 293
L DL + + T ++E Q +VT L++ + +V+T G D
Sbjct: 1266 LQDLQSKYSDGERVRTELNEKVHKLQIEVENVTSLLNEAESKNIKLTKDVATLGSQLQDT 1325
Query: 294 IEQVEAEV-QRLNAL-KASKMKDLVFKRQNELEEIYRGV-----HMDV------DSEAAR 340
E ++ E Q+LN K +++D Q +L+E H+ DS+
Sbjct: 1326 QELLQEETRQKLNVTTKLRQLEDDKNSLQEQLDEEVEAKQNLERHISTLTIQLSDSKKKL 1385
Query: 341 QILTSLIES---GNIDL-------------------------SDLLQSMDDQIRQAKEQA 372
Q T+ +E+ G L + L Q +DD + Q
Sbjct: 1386 QEFTATVETMEEGKKKLQREIESLTQQFEEKAASYDKLEKTKNRLQQELDDLVVDLDNQR 1445
Query: 373 QSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIV 432
Q + + +K+ EEK + DE + K K AR + + +
Sbjct: 1446 QLVSNLEKKQKKFDQMLAEEKNISSKYADERDRAEAEAREKETKALSLARALEEALEAKE 1505
Query: 433 E-NLTTKVKAWEME----------KGIPFLYE-KAPLLQSLEEYHVQRQLREEEKRKSRE 480
E T K+ EME K + L + K L Q +EE Q + E+E + + +
Sbjct: 1506 ELERTNKMLKAEMEDLVSSKDDVGKNVHELEKSKRTLEQQVEEMKTQLEELEDELQAAED 1565
Query: 481 QK 482
K
Sbjct: 1566 AK 1567
Score = 45.8 bits (107), Expect = 3e-04
Identities = 93/490 (18%), Positives = 191/490 (38%), Gaps = 65/490 (13%)
Query: 29 IGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSS 88
+ ++ D ++ +LE + D+ + D R + EL + + + E+ N+ S L E+ S
Sbjct: 1248 LSQAKQDVEHKKKKLEVQLQDLQSKYSDGERV-RTELNEKVHKLQIEVENVTSLLNEAES 1306
Query: 89 FSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEV 148
+ L + +AT+ L+D + E ++ + +++ Q+ + ++ E
Sbjct: 1307 KNI---KLTKDVATLGSQLQDTQELLQEETRQKLNVTTKLRQLEDDKNSLQEQLDEEVEA 1363
Query: 149 DQ------SDLTF------KKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSID 196
Q S LT KKL E + ++ ++ K Q++++S E +A
Sbjct: 1364 KQNLERHISTLTIQLSDSKKKLQEFTATVETMEEGKKKLQREIESLTQQFEEKAASYD-K 1422
Query: 197 FWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLW 256
KT + + L D + D L + +++ Q L + + ++ D
Sbjct: 1423 LEKTKNRLQQELDDLV------VDLDNQRQLVSNLEKKQKKFDQMLAEEKNISSKYADER 1476
Query: 257 DLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEA--EVQRLNALKASKMKD 314
D E E++ T+ +S L+ + E +EA E++R N + ++M+D
Sbjct: 1477 DRAEAEAREKE-----TKALS-----------LARALEEALEAKEELERTNKMLKAEMED 1520
Query: 315 LVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQS 374
LV + +++ + VH +E L ++ M Q+ + +++ Q+
Sbjct: 1521 LVSSK----DDVGKNVHE--------------LEKSKRTLEQQVEEMKTQLEELEDELQA 1562
Query: 375 RREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVEN 434
+ R+E A + + D RDE R K L E K ++
Sbjct: 1563 AEDAKLRLEVNMQAMKSQFERDLQARDEQNEEKRRQLLKQLHEHETELEDERKQRALAAA 1622
Query: 435 LTTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEA 494
K +E + L + E + +QLR+ + + Q+ L + A +E
Sbjct: 1623 AKKK-----LEVDVKDLESQVDSANKAREEAI-KQLRKLQAQMKDYQRDLDDARAAREEI 1676
Query: 495 LFGSRSATKK 504
+R KK
Sbjct: 1677 FATARENEKK 1686
>MYS2_DICDI (P08799) Myosin II heavy chain, non muscle
Length = 2116
Score = 53.5 bits (127), Expect = 1e-06
Identities = 104/523 (19%), Positives = 211/523 (39%), Gaps = 76/523 (14%)
Query: 29 IGESDSDRDNMLLQLEQECLDIYRRKVDA--TRKHKAELCQWLADAEAELINLVSSLGES 86
I + D++ D++ +L++E + D TRK A+L + +A+ E++ +
Sbjct: 1535 IRKKDAEIDDLRARLDRETESRIKSDEDKKNTRKQFADLEAKVEEAQREVVTI------- 1587
Query: 87 SSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSE 146
R K L+ + + L+ ++ R+K K K ++ Q E E+ S +
Sbjct: 1588 ---DRLKKKLESDIIDLSTQLD---TETKSRIK-IEKSKKKLEQTLAERRAAEEGSSKAA 1640
Query: 147 EVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWK------- 199
+ + ++++ EL++ L + ++K+KS ++ + E+ + +
Sbjct: 1641 DEEIRKQVWQEVDELRAQLDSERAALNASEKKIKSLVAEVDEVKEQLEDEILAKDKLVKA 1700
Query: 200 ------TLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQE---KQQRLLKVQELTK 250
L E+ L + +S D+ LT + +K++ + ++ K+ E K
Sbjct: 1701 KRALEVELEEVRDQLEEEEDS--RSELEDSKRRLTTEVEDIKKKYDAEVEQNTKLDEAKK 1758
Query: 251 FLVDLWDLIETPI-DEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKA 309
L D D ++ + DE+K N R E + D + +++AEV
Sbjct: 1759 KLTDDVDTLKKQLEDEKKKLNESERAKKRLESE-------NEDFLAKLDAEV-------- 1803
Query: 310 SKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAK 369
K + K + + E+ + ++ EAA + T + + D D L+S +Q +
Sbjct: 1804 -KNRSRAEKDRKKYEKDLKDTKYKLNDEAATKTQTEIGAAKLEDQIDELRSKLEQEQAKA 1862
Query: 370 EQAQSRREILD--------RVE---KWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRA 418
QA ++ L+ ++E K + E+EK E E +E R + +
Sbjct: 1863 TQADKSKKTLEGEIDNLRAQIEDEGKIKMRLEKEKRALEGELEELRETVEEAEDSKSEAE 1922
Query: 419 EKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREE---EK 475
+ R+V ++ NL ++ A E+ + K+ L + + E + +L EE
Sbjct: 1923 QSKRLVELELEDARRNLQKEIDAKEIAED-----AKSNLQREIVE--AKGRLEEESIART 1975
Query: 476 RKSREQKRLQEQHAVEQEALFGSRSATKKPLGQSTTANTIVGT 518
R +KRL+ E +AL A +K Q N + T
Sbjct: 1976 NSDRSRKRLE----AEIDALTAQVDAEQKAKNQQIKENKKIET 2014
Score = 50.8 bits (120), Expect = 9e-06
Identities = 106/498 (21%), Positives = 196/498 (39%), Gaps = 78/498 (15%)
Query: 27 DEIGESDSDRDNMLLQL------EQECLDIYRRKVDATRKHKAELCQWLADAEAEL---- 76
D++ +S D ++ +L L E+E L DA K EL + D E+EL
Sbjct: 852 DKLEKSLKDTESNVLDLQRQLKAEKETLKAMYDSKDALEAQKRELEIRVEDMESELDEKK 911
Query: 77 INLVSSLGESSSFSRGKGTLKQQLAT---IRPVLEDLRSKKDERVKEFSKIK-------S 126
+ L + + S L+++L +R LE L+ K +E ++E ++ S
Sbjct: 912 LALENLQNQKRSVEEKVRDLEEELQEEQKLRNTLEKLKKKYEEELEEMKRVNDGQSDTIS 971
Query: 127 QISQICVEIAGY--EQPKSSSEEVDQSDLTFKKLGELKSHLQDL----------QNEKIL 174
++ +I E+ E +S SEE + K L+S L DL ++E +
Sbjct: 972 RLEKIKDELQKEVEELTESFSEESKDKGVLEKTRVRLQSELDDLTVRLDSETKDKSELLR 1031
Query: 175 RQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSL 234
+++K++ + + E A +++ Q +N L ++
Sbjct: 1032 QKKKLEEELKQVQEALAA-----------------ETAAKLAQEAANKKLQGEYTELNEK 1074
Query: 235 KQEKQQRLLKVQELTKFLVDLWDLIETPIDEQK----AFNHVTRLISASVDEVSTHGCLS 290
+ V++ K L + +DE+K A + + A ++E+
Sbjct: 1075 FNSEVTARSNVEKSKKTLESQLVAVNNELDEEKKNRDALEKKKKALDAMLEEMK------ 1128
Query: 291 SDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESG 350
D +E E + L LK + D+ R N++ E+ + A + + S +E
Sbjct: 1129 -DQLESTGGEKKSLYDLKVKQESDMEALR-NQISELQSTI-------AKLEKIKSTLEGE 1179
Query: 351 NIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEE---EKWLDEYERD-ENRYS 406
L L++ +Q+ ++ + Q ++ LD +K AEE ++ LD+ ++ E S
Sbjct: 1180 VARLQGELEA--EQLAKSNVEKQKKKVELDLEDKSAQLAEETAAKQALDKLKKKLEQELS 1237
Query: 407 AVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKA-WEMEKGIPFLYEKAPLLQSLEEYH 465
V+ L A + +E +K E E+ EK L E H
Sbjct: 1238 EVQ---TQLSEANNKNVNSDSTNKHLETSFNNLKLELEAEQKAKQALEKKRLGLESELKH 1294
Query: 466 VQRQLREEEKRKSREQKR 483
V QL EE+K+K +KR
Sbjct: 1295 VNEQLEEEKKQKESNEKR 1312
Score = 38.5 bits (88), Expect = 0.047
Identities = 100/460 (21%), Positives = 188/460 (40%), Gaps = 59/460 (12%)
Query: 75 ELINLVSSLGES-SSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICV 133
EL V L ES S S+ KG L++ ++ L+DL + D K+ S++ Q ++
Sbjct: 979 ELQKEVEELTESFSEESKDKGVLEKTRVRLQSELDDLTVRLDSETKDKSELLRQKKKLEE 1038
Query: 134 EIAGYEQPKSS--SEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSA 191
E+ ++ ++ + ++ Q K GE + +E R KS + S+L A
Sbjct: 1039 ELKQVQEALAAETAAKLAQEAANKKLQGEYTELNEKFNSEVTARSNVEKSKKTLESQLVA 1098
Query: 192 VMS-IDFWKTLSEIHPSLGDSSKGTPQSISN--DTLASLTGYIHSL-----KQEKQQRLL 243
V + +D K + +L K + D L S G SL KQE L
Sbjct: 1099 VNNELDEEKKNRD---ALEKKKKALDAMLEEMKDQLESTGGEKKSLYDLKVKQESDMEAL 1155
Query: 244 K--VQELTKFLVDLWDL---IETPIDEQKAFNHVTRLISASVDEVSTHGCL-----SSDV 293
+ + EL + L + +E + + +L ++V++ L S+ +
Sbjct: 1156 RNQISELQSTIAKLEKIKSTLEGEVARLQGELEAEQLAKSNVEKQKKKVELDLEDKSAQL 1215
Query: 294 IEQVEAEVQRLNALKASKMKDLVFKRQNELEEIY-RGVHMDVDSE-------------AA 339
E+ A+ Q L+ LK K++ + + Q +L E + V+ D ++ A
Sbjct: 1216 AEETAAK-QALDKLK-KKLEQELSEVQTQLSEANNKNVNSDSTNKHLETSFNNLKLELEA 1273
Query: 340 RQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYE 399
Q +E + L L+ +++Q+ + K+Q +S + +EK E + D+ E
Sbjct: 1274 EQKAKQALEKKRLGLESELKHVNEQLEEEKKQKESNEKRKVDLEK-----EVSELKDQIE 1328
Query: 400 RDENRYSAVRGAHKNLKRAE-------KARIVVSKIPSIVENLTTKVKAWEME------K 446
+ AV A KN K +E A +V S+ S+ + T + K E+ +
Sbjct: 1329 EEVASKKAVTEA-KNKKESELDEIKRQYADVVSSRDKSVEQLKTLQAKNEELRNTAEEAE 1387
Query: 447 GIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQE 486
G E++ + + L EE +K + +K +++
Sbjct: 1388 GQLDRAERSKKKAEFDLEEAVKNLEEETAKKVKAEKAMKK 1427
Score = 37.4 bits (85), Expect = 0.11
Identities = 81/398 (20%), Positives = 167/398 (41%), Gaps = 66/398 (16%)
Query: 104 RPVLEDLRSKKD--ERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGEL 161
RP+L+ +K+ E+ +E ++KS ++ + Q D K L +
Sbjct: 818 RPLLKRRNFEKEIKEKEREILELKSNLT----------------DSTTQKDKLEKSLKDT 861
Query: 162 KSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISN 221
+S++ DLQ + ++ +K+ + L A + + ++ L D K +++ N
Sbjct: 862 ESNVLDLQRQLKAEKETLKAMYDSKDALEA-QKRELEIRVEDMESEL-DEKKLALENLQN 919
Query: 222 DTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVD 281
S+ + L++E Q+ Q+L L L E ++E K N D
Sbjct: 920 QK-RSVEEKVRDLEEELQEE----QKLRNTLEKLKKKYEEELEEMKRVN------DGQSD 968
Query: 282 EVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQ 341
+S + ++++ EV+ L + + KD + LE+ + ++D
Sbjct: 969 TISR----LEKIKDELQKEVEELTESFSEESKD-----KGVLEKTRVRLQSELDD----- 1014
Query: 342 ILTSLIESGNIDLSDLL---QSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEY 398
LT ++S D S+LL + ++++++Q +E + + K A +K EY
Sbjct: 1015 -LTVRLDSETKDKSELLRQKKKLEEELKQVQEALAA-----ETAAKLAQEAANKKLQGEY 1068
Query: 399 ERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWE-MEKGIPFLYEKAPL 457
++++ A N+++++K + S++ ++ L + K + +EK +K L
Sbjct: 1069 TELNEKFNSEVTARSNVEKSKKT--LESQLVAVNNELDEEKKNRDALEK------KKKAL 1120
Query: 458 LQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEAL 495
LEE Q + EK+ + K QE + EAL
Sbjct: 1121 DAMLEEMKDQLESTGGEKKSLYDLKVKQES---DMEAL 1155
>RA50_METKA (Q8TXI4) DNA double-strand break repair rad50 ATPase
Length = 876
Score = 53.1 bits (126), Expect = 2e-06
Identities = 101/505 (20%), Positives = 221/505 (43%), Gaps = 71/505 (14%)
Query: 36 RDNMLLQLEQECLDIYRRKVDAT------RKHKAELCQWLADAEAELI-------NLVSS 82
R + +L + + +R VD T +K + + + L AEA+L +L S
Sbjct: 138 RQGEIAKLVEATREERKRIVDRTLGLAEFKKAREQAHELLRVAEAKLETFRERVRDLKGS 197
Query: 83 LGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDE---RVKEFSKIKSQISQICVEIAGYE 139
E R LK+++ + P +E+L+ + +E +EF +++ ++ + +I +
Sbjct: 198 KKELKRVERELEELKREVKELEPEVEELKERLNELREAKREFERLEGELRLLENKIESLK 257
Query: 140 QPKSSS----EEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISEL------ 189
+ EE +++ ++LG++ S +++L+NE+ +++++ + + +L
Sbjct: 258 GRRDDLRKLVEEGKEAERELQRLGDVPSKVRELENEEAELRRRIEELRNLLDDLRSLRNR 317
Query: 190 --SAVMSIDFWKTLSEIHPSLGDSSKGTPQSI--SNDTLASLTGYIHSLKQEKQQRLLKV 245
SA ++ K E L D + P+ + D + + + L++E++ + K+
Sbjct: 318 LESAEEELEGVKRELE---ELKDEAGVDPERLVEFKDKIVEASERLRDLRREEELK-RKL 373
Query: 246 QELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQR-- 303
++++ L +L D ET E + I + E+ +++E++E+ +
Sbjct: 374 EKVSDELSELGDREETLQSEYEELQERLDEIQGELKEIRVK---EKELLERIESLREAEG 430
Query: 304 -----LNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLL 358
L L + + L+ + ELE + +G D+ E R+ L +ES +L
Sbjct: 431 ECPVCLRKLPRERAEKLLRDAEKELERL-QGREEDLRKE--RRELKDRLESVRRELEGTK 487
Query: 359 QSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRG-------- 410
+ M ++R+ +E+ + E ++ +++ E ++E E R AVR
Sbjct: 488 ERM-WRLRERREELERELEEIEELKEELADLSRELGVEEDRLPELRDLAVRAESLLRDLE 546
Query: 411 --------AHKNLKRA-EKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQ-S 460
K L+R ++ V+ + PS VE++ +++ E E+ + +K +
Sbjct: 547 RRRGDVLRLEKELERTLDRCEKVIGRTPSGVEDVEEELRRLEEER--DHVGQKLREAEGE 604
Query: 461 LEEYHVQRQLREEEKRKSREQKRLQ 485
LE YH L E+ KR +K L+
Sbjct: 605 LERYH---NLEEKVKRAREARKELK 626
Score = 47.0 bits (110), Expect = 1e-04
Identities = 100/473 (21%), Positives = 196/473 (41%), Gaps = 62/473 (13%)
Query: 42 QLEQECLDIYRRKVDATRKHKAELCQWLAD---AEAELINLVSSLGESSSFSRGKGTLKQ 98
+LE E L + K+++ + + +L + + + AE EL L + + L++
Sbjct: 241 RLEGE-LRLLENKIESLKGRRDDLRKLVEEGKEAERELQRLGDVPSKVRELENEEAELRR 299
Query: 99 QLATIRPVLEDLRS---KKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTF 155
++ +R +L+DLRS + + +E +K ++ ++ E AG + P+ E D
Sbjct: 300 RIEELRNLLDDLRSLRNRLESAEEELEGVKRELEELKDE-AGVD-PERLVEFKD------ 351
Query: 156 KKLGELKSHLQDLQNEKILRQ--QKVKSHISTISELSAVMSIDF---WKTLSEIHPSLGD 210
K+ E L+DL+ E+ L++ +KV +S + + + ++ + L EI L +
Sbjct: 352 -KIVEASERLRDLRREEELKRKLEKVSDELSELGDREETLQSEYEELQERLDEIQGELKE 410
Query: 211 SSKGTPQSISN-DTLASLTG----YIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDE 265
+ + ++L G + L +E+ ++LL+ E K L L E E
Sbjct: 411 IRVKEKELLERIESLREAEGECPVCLRKLPRERAEKLLRDAE--KELERLQGREEDLRKE 468
Query: 266 QKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEE 325
++ + ++ + E++E E++ + LK EL +
Sbjct: 469 RRELKDRLESVRRELEGTKERMWRLRERREELERELEEIEELK------------EELAD 516
Query: 326 IYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSR---REILDRV 382
+ R + ++ D + L ES L DL + D +R KE ++ +++ R
Sbjct: 517 LSRELGVEEDRLPELRDLAVRAES---LLRDLERRRGDVLRLEKELERTLDRCEKVIGRT 573
Query: 383 EKWRFAAEEEKWLDEYERD---------ENRYSAVRGAHKNLKRAEKARIVVSKIPSIVE 433
EEE E ERD E + +KRA +AR + +I +E
Sbjct: 574 PSGVEDVEEELRRLEEERDHVGQKLREAEGELERYHNLEEKVKRAREARKELKRIERDLE 633
Query: 434 NLTTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQE 486
+ K + ++E+ + L E+ LEE +L EK+ R + +L E
Sbjct: 634 D--AKGRLEQVERNLEGLRERYGSEDRLEE-----ELESVEKKYERVRDKLSE 679
>CING_MOUSE (P59242) Cingulin
Length = 1191
Score = 53.1 bits (126), Expect = 2e-06
Identities = 106/494 (21%), Positives = 207/494 (41%), Gaps = 74/494 (14%)
Query: 15 CASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEA 74
C L L+ E+ +S + NM L L QE +H EA
Sbjct: 391 CHRLQELLERRKGEVQQSSKELQNMKLLLGQE----------EGLRH---------GLEA 431
Query: 75 ELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVE 134
++ L L S S GK +L + L R +LE+L K +RV+E +++ + ++
Sbjct: 432 QVKELQLKLKHSQSPDSGKESLLKDLLDTRELLEELLEGK-QRVEEQLRLRER-ELTALK 489
Query: 135 IAGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMS 194
A E+ S +EV+ L +++ D + + Q + H + +E + S
Sbjct: 490 GALKEEVASHDQEVEHVRLQYQR---------DTEQLRRSMQDATQDHAALEAERQKMSS 540
Query: 195 IDFWKTLSEIHPSLGDSSKGTP--QSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFL 252
+ + E+ L ++S+ T QS+ L + KQE Q ++ +E+ + L
Sbjct: 541 L-----VRELQRELEETSEETGHWQSMFQKNKEEL----RATKQELLQLRMEKEEMEEEL 591
Query: 253 VDLWDLIETPIDEQKAFNHVTRLISASVDEV-STHGCLSSDVIEQVEAEV---------- 301
+ ++++ +++ +A T + E+ T G L EQ EV
Sbjct: 592 GEKMEVLQRDLEQARASTRDTHQVEELKKELRRTQGELKELQAEQQNQEVTGRHRNQVLE 651
Query: 302 QRLNALKASKMKDLVFKRQN-ELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQS 360
++L AL+ + ++QN +L++ + + D + + ++ + E+ + L +
Sbjct: 652 KQLAALREEADRGRELEQQNLQLQKTLQQLRQDCEEASKAKVAS---ETEAMMLGQRRAT 708
Query: 361 MDDQIRQAKEQ-AQSRREILDRVEKWRFA--------AEEEKWLDEYERDENRYSAVRGA 411
++ +R+ +E+ + RR IL ++ + A A E + D+ R E + A
Sbjct: 709 VETTLRETQEENDEFRRRILGLEQQLKEARGLAEGGEAVEARLRDKVHRLEVEKQQLEEA 768
Query: 412 HKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFL-YEKAPLLQSLEEYHVQRQL 470
N + E+ + +K +V+ E ++G+ L E+ L ++LEE QR+
Sbjct: 769 -LNAAQEEEGNLAAAK-------RALEVRLDEAQRGLARLGQEQQALNRALEEEGKQREA 820
Query: 471 REEEKRKSREQKRL 484
K + EQKRL
Sbjct: 821 LRRSKAELEEQKRL 834
Score = 52.8 bits (125), Expect = 2e-06
Identities = 105/520 (20%), Positives = 208/520 (39%), Gaps = 105/520 (20%)
Query: 27 DEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKA------ELCQWLADAEAELINLV 80
+E+ D + +++ LQ +++ + R DAT+ H A ++ + + + EL
Sbjct: 494 EEVASHDQEVEHVRLQYQRDTEQLRRSMQDATQDHAALEAERQKMSSLVRELQRELEETS 553
Query: 81 SSLGE-SSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYE 139
G S F + K++L + L LR +K+E +E + K ++ Q E
Sbjct: 554 EETGHWQSMFQKN----KEELRATKQELLQLRMEKEEMEEELGE-KMEVLQ-----RDLE 603
Query: 140 QPKSSSEEVDQSDLTFKKL----GELKSHLQDLQNEKIL---RQQKVKSHISTISELS-- 190
Q ++S+ + Q + K+L GELK + QN+++ R Q ++ ++ + E +
Sbjct: 604 QARASTRDTHQVEELKKELRRTQGELKELQAEQQNQEVTGRHRNQVLEKQLAALREEADR 663
Query: 191 ----AVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGY----IHSLKQEKQQ-- 240
++ KTL ++ ++SK ++++T A + G + + +E Q+
Sbjct: 664 GRELEQQNLQLQKTLQQLRQDCEEASKA---KVASETEAMMLGQRRATVETTLRETQEEN 720
Query: 241 -----RLL-------------------------KVQELTKFLVDLWDLIETPIDEQKAFN 270
R+L KV L L + + +E+
Sbjct: 721 DEFRRRILGLEQQLKEARGLAEGGEAVEARLRDKVHRLEVEKQQLEEALNAAQEEEGNLA 780
Query: 271 HVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNAL--KASKMKDLVFKRQNELEEIYR 328
R + +DE + ++ E Q LN + K ++ + + + ELEE R
Sbjct: 781 AAKRALEVRLDEAQRG-------LARLGQEQQALNRALEEEGKQREALRRSKAELEEQKR 833
Query: 329 GVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFA 388
++ VD L +E D LQ + Q+ KE+A R+E+ D
Sbjct: 834 LLNRTVDR------LNKELEQIGDDSKLALQQLQAQMEDYKEKA--RKEVADA------Q 879
Query: 389 AEEEKWLDEYERDENRYSAVRGAHKNLKRA--------EKARIVVSKIPSIVENLTTKVK 440
+ + W E E++ S ++ + L++A + AR+ + ++ L
Sbjct: 880 RQAKDWASEAEKNSGGLSRLQDELQRLRQALQTSQAERDTARLDKELLAQRLQGLEQ--- 936
Query: 441 AWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSRE 480
E E F +KA L+SLEE + + +E++ + E
Sbjct: 937 --EAENKKRFQDDKARQLKSLEEKVSRLEAELDEEKNTVE 974
Score = 47.0 bits (110), Expect = 1e-04
Identities = 88/402 (21%), Positives = 172/402 (41%), Gaps = 55/402 (13%)
Query: 107 LEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQ-SDLTFKKLGELKSHL 165
+E+L+ K DE VK+ K++ S++ +E Q + +EE + +L ++ GE++
Sbjct: 356 MEELQKKLDEEVKKRQKLEP--SRVGLE----RQLEEKAEECHRLQELLERRKGEVQQSS 409
Query: 166 QDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLA 225
++LQN K+L Q+ +++ + L H DS K + DT
Sbjct: 410 KELQNMKLLLGQEEGLRHGLEAQVKELQ-------LKLKHSQSPDSGKESLLKDLLDTRE 462
Query: 226 SLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVST 285
L + ++ ++Q L+ +ELT L + + + E + HV +++
Sbjct: 463 LLEELLEGKQRVEEQLRLRERELTALKGALKEEVASHDQEVE---HVRLQYQRDTEQLRR 519
Query: 286 HGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTS 345
++ +EAE Q KM LV + Q ELEE SE S
Sbjct: 520 SMQDATQDHAALEAERQ--------KMSSLVRELQRELEET---------SEETGH-WQS 561
Query: 346 LIESGNIDLSDLLQSMDDQIRQAKEQAQS----RREILDR-VEKWRFAAEEEKWLDEYER 400
+ + +L Q + Q+R KE+ + + E+L R +E+ R + + ++E ++
Sbjct: 562 MFQKNKEELRATKQELL-QLRMEKEEMEEELGEKMEVLQRDLEQARASTRDTHQVEELKK 620
Query: 401 DENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQS 460
+ R +G K L+ ++ + V + + V +EK + L E+A +
Sbjct: 621 ELRR---TQGELKELQAEQQNQEVTGRHRNQV-----------LEKQLAALREEADRGRE 666
Query: 461 LEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSAT 502
LE+ ++Q Q ++ R+ E+ + + + + G R AT
Sbjct: 667 LEQQNLQLQKTLQQLRQDCEEASKAKVASETEAMMLGQRRAT 708
Score = 36.6 bits (83), Expect = 0.18
Identities = 91/490 (18%), Positives = 189/490 (38%), Gaps = 47/490 (9%)
Query: 28 EIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESS 87
E+ + + L QL Q+C + + KV A+ L Q A E L + E+
Sbjct: 666 ELEQQNLQLQKTLQQLRQDCEEASKAKV-ASETEAMMLGQRRATVET---TLRETQEEND 721
Query: 88 SFSRGKGTLKQQLATIRPVLED-------LRSKKDERVKEFSKIKSQISQICVEIAGYEQ 140
F R L+QQL R + E LR K E +++ ++ E
Sbjct: 722 EFRRRILGLEQQLKEARGLAEGGEAVEARLRDKVHRLEVEKQQLEEALNAAQEEEGNLAA 781
Query: 141 PKSSSE-EVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWK 199
K + E +D++ +LG+ + L E+ +++ ++ + + E +++ +
Sbjct: 782 AKRALEVRLDEAQRGLARLGQEQQALNRALEEEGKQREALRRSKAELEEQKRLLNRTVDR 841
Query: 200 TLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLI 259
E+ +GD SK Q + A + Y ++E + ++ +
Sbjct: 842 LNKELE-QIGDDSKLALQQLQ----AQMEDYKEKARKEVADAQRQAKDWASEAEKNSGGL 896
Query: 260 ETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKR 319
DE + + A D L + ++ +E E + + K + L
Sbjct: 897 SRLQDELQRLRQALQTSQAERDTARLDKELLAQRLQGLEQEAENKKRFQDDKARQL---- 952
Query: 320 QNELEEIYRGVHMDVDSEAAR-QILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREI 378
LEE + ++D E ++LT + G D D L++ Q R A++ + +
Sbjct: 953 -KSLEEKVSRLEAELDEEKNTVELLTDRVNRGR-DQVDQLRTELMQERSARQDLECDKIS 1010
Query: 379 LDRVEK---WRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENL 435
L+R K R A+ E +++ S + ++ L+ +A + +++++
Sbjct: 1011 LERQNKDLKTRLASSEG-----FQKPSASLSQLESQNQLLQERLQAE---EREKTVLQST 1062
Query: 436 TTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQ-----RQLREEEKR-------KSREQKR 483
K++ E I E+ + ++ ++ RQ+ E E+ + + Q+
Sbjct: 1063 NRKLERRVKELSIQIDDERQHVNDQKDQLTLRVKALKRQVDEAEEEIERLDSLRKKAQRE 1122
Query: 484 LQEQHAVEQE 493
L+EQH V ++
Sbjct: 1123 LEEQHEVNEQ 1132
>MYSJ_DICDI (P54697) Myosin IJ heavy chain
Length = 2245
Score = 52.4 bits (124), Expect = 3e-06
Identities = 96/511 (18%), Positives = 204/511 (39%), Gaps = 93/511 (18%)
Query: 32 SDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSR 91
S+ + + QLE E D HK++L L E + G+ +
Sbjct: 1229 SEEEAKKQINQLELELTD-----------HKSKLQIQLQLTEQSNEKIKKLKGKLEEYQD 1277
Query: 92 GKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQP----KSSSEE 147
K L+Q+L I+ + + +K+ + + + +K + +Q+ ++ ++ KS+ EE
Sbjct: 1278 EKKQLQQELERIKQSKQSVEDEKNSLITQLTTVKFESTQVSTNVSHQKEKITTLKSTIEE 1337
Query: 148 VDQSDLTFKKLGELKS-------HLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKT 200
++ K +G+L++ ++ +Q E ++Q+ S+L + SID K+
Sbjct: 1338 LN------KSIGKLQAEQKNKDDEIRKIQFELNDQKQQFTRQTKEFSDLQSQQSIDRPKS 1391
Query: 201 LSEIH------PSLGDSSKGTPQSIS---------NDTLASLTGYIHSLKQEKQ------ 239
IH +L + QS+ DT+ L + L Q K+
Sbjct: 1392 EITIHSLERTNETLKSDFERVQQSLKQQERDCQQYKDTINRLENEVKQLTQLKERFENEF 1451
Query: 240 -----------QRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGC 288
Q + ++E+T + IE ++E+K H+TR I DE+
Sbjct: 1452 FVAKEQNSNQTQESVYLKEVTTQMQQNQSRIERELEEKK--QHITR-IDDERDELKKQ-- 1506
Query: 289 LSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIE 348
+ Q++ + ++ + +L R+ EL+ RG + + SL
Sbjct: 1507 -----LTQLQQQHEQSSTQLLLAQNELERLRKKELKYKERGHETSKQQDQFNMEIQSLRI 1561
Query: 349 SGNIDLSDL------LQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEE-EKWLDEYERD 401
+ N L L + + D++ +K++AQ +RE + +++ A ++ +W+
Sbjct: 1562 TNNDQLKSLQDYEQEKKKLKDKLSSSKQEAQQQRESIIKMDAELSAIKQHSQWV------ 1615
Query: 402 ENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSL 461
EN ++ ++ +N + E + + ++ + + +K E E + L Q L
Sbjct: 1616 ENSFTDMK--QRNQELIESSALYKQQLLQQTSTIDSTIKEKE--------NEISKLQQQL 1665
Query: 462 EEYHVQRQLREEEKRKSREQKRLQEQHAVEQ 492
E + Q +EE ++ +L+ +Q
Sbjct: 1666 ETSNQQLHQLKEELNSMKQSNQLESTEQSKQ 1696
>HOK3_HUMAN (Q86VS8) Hook homolog 3 (hHK3)
Length = 718
Score = 52.4 bits (124), Expect = 3e-06
Identities = 108/525 (20%), Positives = 219/525 (41%), Gaps = 122/525 (23%)
Query: 16 ASLLSQLQMIWDEIGESDSDRD-------------NMLLQLEQECL------DIYRRKVD 56
+SLL++ Q++ + + +SDS D L QL++E D YR + +
Sbjct: 214 SSLLAENQVLMERLNQSDSIEDPNSPAGRRHLQLQTQLEQLQEETFRLEAAKDDYRIRCE 273
Query: 57 ATRKHKAELCQW------LADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDL 110
K +EL Q LAD L + + L SS L+ Q+ + + LEDL
Sbjct: 274 ELEKEISELRQQNDELTTLADEAQSLKDEIDVLRHSSD---KVSKLEGQVESYKKKLEDL 330
Query: 111 RSKK--------------------DERVKEFSKIKSQISQICVEIAGYEQPKSS-SEEVD 149
+ +E +++ + +SQ+ ++ + S S++ D
Sbjct: 331 GDLRRQVKLLEEKNTMYMQNTVSLEEELRKANAARSQLETYKRQVVELQNRLSEESKKAD 390
Query: 150 QSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLG 209
+ D +K+L K + LQ EK + + S TI EL V + + T + P LG
Sbjct: 391 KLDFEYKRL---KEKVDSLQKEKDRLRTERDSLKETIEELRCVQAQEGQLTTQGLMP-LG 446
Query: 210 DSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAF 269
S+D+LA+ + E +++L+++Q K L ++++ +
Sbjct: 447 SQE-------SSDSLAA-----EIVTPEIREKLIRLQHENKML---------KLNQEGSD 485
Query: 270 NHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRG 329
N L+ + +D+ + + E++ N L ++ ++ Q+++EE+ +
Sbjct: 486 NEKIALLQSLLDDANLR-----------KNELETENRLVNQRLLEV----QSQVEELQKS 530
Query: 330 VHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRF-- 387
+ D S+A +L L L+ +++ +A + Q +R I++ +E RF
Sbjct: 531 LQ-DQGSKAEDSVL----------LKKKLEEHLEKLHEANNELQKKRAIIEDLEP-RFNN 578
Query: 388 -AAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKI-PSIVENLTTKVKAWEME 445
+ + E+ + + E + +K K EKA+ V+ + P + +++A + +
Sbjct: 579 SSLKIEELQEALRKKEEEMKQMEERYK--KYLEKAKSVIRTLDPKQNQGAAPEIQALKNQ 636
Query: 446 KGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAV 490
L E+ L SLE+ E K++ Q+ ++E++ V
Sbjct: 637 -----LQERDRLFHSLEK----------EYEKTKSQREMEEKYIV 666
>MYSN_DROME (Q99323) Myosin heavy chain, non-muscle (Zipper protein)
(Myosin II) (Non-muscle MHC)
Length = 2057
Score = 52.0 bits (123), Expect = 4e-06
Identities = 98/425 (23%), Positives = 167/425 (39%), Gaps = 51/425 (12%)
Query: 107 LEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKSHLQ 166
++DL + +E K++ + Q+ +I YE+ + +++ +Q L KKL L+
Sbjct: 1036 IQDLEEQLEEEEAARQKLQLEKVQLDAKIKKYEEDLALTDDQNQKLLKEKKL--LEERAN 1093
Query: 167 DLQNEKILRQQKVK-------SHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSI 219
DL ++K K H +TI+EL + D + D SK ++
Sbjct: 1094 DLSQTLAEEEEKAKHLAKLKAKHEATITELEERLHKD------QQQRQESDRSKRKIETE 1147
Query: 220 SNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISAS 279
D L + + + Q + +ELT+ L+ + + T QKA R + +
Sbjct: 1148 VADLKEQLNERRVQVDEMQAQLAKREEELTQTLLRIDEESATKATAQKA----QRELESQ 1203
Query: 280 VDEVSTHGCLSSDVIEQVEAEVQRLNALKASKM-KDLVFKRQNELEEIYRGVHMDVDSEA 338
+ E+ Q + E ++ KA K+ +DL ELE + + +D+ A
Sbjct: 1204 LAEI------------QEDLEAEKAARAKAEKVRRDL----SEELEALKNELLDSLDTTA 1247
Query: 339 ARQILTSLIESGNIDLSDLLQSMDDQ--------IRQAKEQAQSRREILDRVEKWRFAAE 390
A+Q L S E +L+ L +S++++ + +Q I D++E R A
Sbjct: 1248 AQQELRSKREQ---ELATLKKSLEEETVNHEGVLADMRHKHSQELNSINDQLENLRKAKT 1304
Query: 391 EEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPF 450
+ EN A N R E R I E +VK E+E+
Sbjct: 1305 VLEKAKGTLEAENADLATELRSVNSSRQENDRRRKQAESQIAE---LQVKLAEIERARSE 1361
Query: 451 LYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQST 510
L EK LQ E ++ QL E E + S K + EA T++ LG S+
Sbjct: 1362 LQEKCTKLQQ-EAENITNQLEEAELKASAAVKSASNMESQLTEAQQLLEEETRQKLGLSS 1420
Query: 511 TANTI 515
I
Sbjct: 1421 KLRQI 1425
Score = 47.0 bits (110), Expect = 1e-04
Identities = 95/475 (20%), Positives = 198/475 (41%), Gaps = 68/475 (14%)
Query: 44 EQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGE----SSSFSRGKGTLKQQ 99
EQE + + + T H+ L EL ++ L + + KGTL+ +
Sbjct: 1257 EQELATLKKSLEEETVNHEGVLADMRHKHSQELNSINDQLENLRKAKTVLEKAKGTLEAE 1316
Query: 100 LATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLG 159
A + L + S + E + + +SQI+++ V++A E+ +S +E K
Sbjct: 1317 NADLATELRSVNSSRQENDRRRKQAESQIAELQVKLAEIERARSELQE---------KCT 1367
Query: 160 ELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSI 219
+L+ +++ N+ + K + + + S + + ++ E LG SSK + I
Sbjct: 1368 KLQQEAENITNQLEEAELKASAAVKSASNMESQLTEAQQLLEEETRQKLGLSSK--LRQI 1425
Query: 220 SNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISAS 279
++ A L + + K+ K+ E+T + ++ E D K + ++
Sbjct: 1426 ESEKEA-LQEQLEEDDEAKRNYERKLAEVTTQMQEIKKKAEEDADLAKELEEGKKRLNKD 1484
Query: 280 VDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAA 339
++ + ++++ A+ RL+ K K Q+ELE+ ++ EA
Sbjct: 1485 IEALERQ-------VKELIAQNDRLDKSKK--------KIQSELED------ATIELEAQ 1523
Query: 340 RQILTSLIESGNIDLSDLL---QSMDDQIRQAKEQAQSRREILDRVEKWRFAAEE-EKWL 395
R + L E + +L +++ +QI Q ++ A+ RE ++ K + E ++
Sbjct: 1524 RTKVLEL-EKKQKNFDKILAEEKAISEQIAQERDTAE--REAREKETKVLSVSRELDEAF 1580
Query: 396 DEYERDENRYSAVRG-----------AHKNLKRAEKA-RIVVSKIPSIVENLTTKVKAWE 443
D+ E EN+ ++ A KN+ EKA R + S++ + K + E
Sbjct: 1581 DKIEDLENKRKTLQNELDDLANTQGTADKNVHELEKAKRALESQLAEL------KAQNEE 1634
Query: 444 MEKGIPFLYEKAPL-----LQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQE 493
+E + L E A L +Q+L + L +EE + + + +++ +E E
Sbjct: 1635 LEDDLQ-LTEDAKLRLEVNMQALRSQFERDLLAKEEGAEEKRRGLVKQLRDLETE 1688
Score = 42.4 bits (98), Expect = 0.003
Identities = 89/460 (19%), Positives = 199/460 (42%), Gaps = 51/460 (11%)
Query: 12 RTTCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLAD 71
R S +++LQ+ EI + S+ +L+QE +I + +A K A + + ++
Sbjct: 1338 RKQAESQIAELQVKLAEIERARSELQEKCTKLQQEAENITNQLEEAELKASAAV-KSASN 1396
Query: 72 AEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQI 131
E++L L E + G + +Q+ + + L++ + DE + + + ++++
Sbjct: 1397 MESQLTEAQQLLEEETRQKLGLSSKLRQIESEKEALQEQLEEDDEAKRNYERKLAEVTTQ 1456
Query: 132 CVEI-AGYEQPKSSSEEVDQSDLTFKK-LGELKSHLQDL--QNEKILR-QQKVKSHISTI 186
EI E+ ++E+++ K + L+ +++L QN+++ + ++K++S +
Sbjct: 1457 MQEIKKKAEEDADLAKELEEGKKRLNKDIEALERQVKELIAQNDRLDKSKKKIQSELEDA 1516
Query: 187 S---ELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLL 243
+ E ++ K L + K + I+ + + ++E +++
Sbjct: 1517 TIELEAQRTKVLELEKKQKNFDKILAE-EKAISEQIAQER--------DTAEREAREKET 1567
Query: 244 KVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQR 303
KV +++ L + +D IE +++K L + D +T G +V E+++
Sbjct: 1568 KVLSVSRELDEAFDKIEDLENKRKT------LQNELDDLANTQGTADKNV-----HELEK 1616
Query: 304 LNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQI-LTSLIESGNIDLSDLLQSMD 362
S++ +L K QN EE+ + + D++ ++ + +L DL + +
Sbjct: 1617 AKRALESQLAEL--KAQN--EELEDDLQLTEDAKLRLEVNMQALRSQFERDLLAKEEGAE 1672
Query: 363 DQIRQAKEQAQSRREILDRVEKWRFAAEEEKW-----LDEYERDENRYSAVR-GAHKNLK 416
++ R +Q + LD K R AA K L E E ++ V+ A K+ K
Sbjct: 1673 EKRRGLVKQLRDLETELDEERKQRTAAVASKKKLEGDLKEIETTMEMHNKVKEDALKHAK 1732
Query: 417 R-----------AEKARIVVSKIPSIVENLTTKVKAWEME 445
+ AE+A+ ++ ++ + KVKA E E
Sbjct: 1733 KLQAQVKDALRDAEEAKAAKEELQALSKEADGKVKALEAE 1772
Score = 39.7 bits (91), Expect = 0.021
Identities = 59/295 (20%), Positives = 119/295 (40%), Gaps = 51/295 (17%)
Query: 27 DEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGES 86
+E+ + D + LE E L + + R +A + D AE + ++ +
Sbjct: 1753 EELQALSKEADGKVKALEAEVLQLTEDLASSERARRAAETE--RDELAE--EIANNANKG 1808
Query: 87 SSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSE 146
S K L+ ++AT+ LE+ +S + + K + QI Q+ E+A + +S +
Sbjct: 1809 SLMIDEKRRLEARIATLEEELEEEQSNSEVLLDRSRKAQLQIEQLTTELA--NEKSNSQK 1866
Query: 147 EVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHP 206
+ L ++ ELK+ L +++ ++ KVK+ I+T+ ++ +
Sbjct: 1867 NENGRALLERQNKELKAKLAEIET---AQRTKVKATIATLE-----------AKIANLEE 1912
Query: 207 SLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLL--KVQELTKFLVDLWDLIETPID 264
L + K L Q+K R + K++ELT + D ++ +
Sbjct: 1913 QLENEGK------------------ERLLQQKANRKMDKKIKELTMNIEDERRHVDQHKE 1954
Query: 265 EQKAFNHVTRLISASVDEVS-----------THGCLSSDVIEQVEAEVQRLNALK 308
+ N +L+ ++DE + D+IE EA + +N+LK
Sbjct: 1955 QMDKLNSRIKLLKRNLDETEEELQKEKTQKRKYQRECEDMIESQEAMNREINSLK 2009
>CF60_HUMAN (Q8NB25) Hypothetical protein C6orf60
Length = 1020
Score = 52.0 bits (123), Expect = 4e-06
Identities = 99/527 (18%), Positives = 216/527 (40%), Gaps = 84/527 (15%)
Query: 12 RTTCASLLSQLQMIWDEIGE--SDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWL 69
R T L ++LQM+ DE G + L + + + + ++++D ++ + L
Sbjct: 179 RKTIGKLKTELQMVQDEAGSLLDKCQKLQTALAIAENNVQVLQKQLDDAKEGEMALLSKH 238
Query: 70 ADAEAELINLVSSLGESSSFSRGK----GTLKQQLATIRPVLEDLRSKKDERVKEFSKIK 125
+ E+EL L + +S K G L+ T ++DL S+K S++
Sbjct: 239 KEVESELAAARERLQQQASDLVLKASHIGMLQATQMTQEVTIKDLESEK-------SRVN 291
Query: 126 SQISQICVEIAGYEQPKSSSEEVDQSDLTF--KKLGELKSHLQDLQNEKILR-QQKVKSH 182
++SQ+ E A S +E + + KK+ E K Q+ ++ Q +++
Sbjct: 292 ERLSQLEEERAFLRSKTQSLDEEQKQQILELEKKVNEAKRTQQEYYERELKNLQSRLEEE 351
Query: 183 ISTISELSAVMSIDFWKTLSEI----HPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEK 238
++ ++E + KTL E+ H ++ +++ ++ + L+++
Sbjct: 352 VTQLNEAHS-------KTLEELAWKHHMAI--------EAVHSNAIRDKKKLQMDLEEQH 396
Query: 239 QQRLLKVQE----LTKFLVDLWDLIETPIDE-QKAFNHVTRLISASVDEVSTHGCLSSDV 293
+ L ++E L + L +L +++E ++ + H+ ++ S + + L + +
Sbjct: 397 NKDKLNLEEDKNQLQQELENLKEVLEDKLNTANQEIGHLQDMVRKSEQGLGSAEGLIASL 456
Query: 294 IEQVEAEVQRLNALKAS--KMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGN 351
+ E L+ K S + KD + + ELE+ + + + + ++ E
Sbjct: 457 QDSQERLQNELDLTKDSLKETKDALLNVEGELEQ---------ERQQHEETIAAMKEEEK 507
Query: 352 IDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGA 411
+ + + ++ K R+E E+ R EE+K + ++ ++
Sbjct: 508 LKVDKMAHDLE-----IKWTENLRQECSKLREELRLQHEEDK-----KSAMSQLLQLKDR 557
Query: 412 HKNLKR---AEKARIVVSKIPSIVENLTTKVKAWE---MEKGIPFLYEKAPLLQSLEEYH 465
KN R +K ++++I + +NL ++ + + F E+ L Q LEE
Sbjct: 558 EKNAARDSWQKKVEDLLNQISLLKQNLEIQLSQSQTSLQQLQAQFTQERQRLTQELEELE 617
Query: 466 VQRQLR---------------EEEKRKSRE--QKRLQEQHAVEQEAL 495
Q Q R EEEK K + + LQ++H+ E ++L
Sbjct: 618 EQHQQRHKSLKEAHVLAFQTMEEEKEKEQRALENHLQQKHSAELQSL 664
>NUMA_HUMAN (Q14980) Nuclear mitotic apparatus protein 1 (NuMA
protein) (SP-H antigen)
Length = 2115
Score = 51.6 bits (122), Expect = 5e-06
Identities = 94/469 (20%), Positives = 189/469 (40%), Gaps = 77/469 (16%)
Query: 66 CQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIK 125
CQ L ++++ ++ L E + G L +L L+ L+ +E +E SK
Sbjct: 299 CQDLKTEKSQMDRKINQLSEEN------GDLSFKLREFASHLQQLQDALNELTEEHSKAT 352
Query: 126 SQI----SQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKS 181
+ +Q+ E++ Q K EE ++++ KL +L+ HL LQ+ + +V
Sbjct: 353 QEWLEKQAQLEKELSAALQDKKCLEE--KNEILQGKLSQLEEHLSQLQDNPPQEKGEVLG 410
Query: 182 HI---STISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEK 238
+ T+ + +A ++ + T + + ++ +G + A L ++EK
Sbjct: 411 DVLQLETLKQEAATLAAN--NTQLQARVEMLETERGQQE-------AKLLAERGHFEEEK 461
Query: 239 QQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVE 298
QQ L+ + DL Q + +++++ HG + + +
Sbjct: 462 QQ-------LSSLITDL----------QSSISNLSQAKEELEQASQAHGARLTAQVASLT 504
Query: 299 AEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLL 358
+E+ LNA + ++L +Q E+ + E A Q L +E LS L
Sbjct: 505 SELTTLNATIQQQDQELAGLKQQAKEKQAQLAQTLQQQEQASQGLRHQVEQ----LSSSL 560
Query: 359 QSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRA 418
+ + Q+++ E+ ++ R+ D ++ AAEE + ERD A K L+
Sbjct: 561 KQKEQQLKEVAEKQEATRQ--DHAQQLATAAEERE-ASLRERD--------AALKQLEAL 609
Query: 419 EKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLEEYH------------- 465
EK + +I + + + EKA L + +EE
Sbjct: 610 EKEKAAKLEILQQQLQVANEARDSAQTSVTQAQREKAELSRKVEELQACVETARQEQHEA 669
Query: 466 ------VQRQLREEEKRKSREQKRLQEQHAVEQ--EALFGSRSATKKPL 506
++ QLR E+++ + +++ QE+ +++ +AL S TK L
Sbjct: 670 QAQVAELELQLRSEQQKATEKERVAQEKDQLQEQLQALKESLKVTKGSL 718
Score = 38.5 bits (88), Expect = 0.047
Identities = 98/498 (19%), Positives = 181/498 (35%), Gaps = 119/498 (23%)
Query: 99 QLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQ------------------ 140
Q+ ++ L D RS +DE E ++ + +++ +IA +Q
Sbjct: 215 QMRRLKKQLADERSNRDELELELAENRKLLTEKDAQIAMMQQRIDRLALLNEKQAASPLE 274
Query: 141 PKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKT 200
PK E D+++ +L E QDL+ EK +K+ +LS + +F
Sbjct: 275 PKELEELRDKNESLTMRLHETLKQCQDLKTEKSQMDRKINQLSEENGDLSFKLR-EFASH 333
Query: 201 LSEIHPSLG----DSSKGTPQ--------------------------SISNDTLASLTGY 230
L ++ +L + SK T + I L+ L +
Sbjct: 334 LQQLQDALNELTEEHSKATQEWLEKQAQLEKELSAALQDKKCLEEKNEILQGKLSQLEEH 393
Query: 231 IHSLK----QEKQQRLLKVQELTKF-------------LVDLWDLIETPIDEQKA----- 268
+ L+ QEK + L V +L L +++ET +Q+A
Sbjct: 394 LSQLQDNPPQEKGEVLGDVLQLETLKQEAATLAANNTQLQARVEMLETERGQQEAKLLAE 453
Query: 269 ---FNHVTRLISASVDEVST------------------HGCLSSDVIEQVEAEVQRLNAL 307
F + +S+ + ++ + HG + + + +E+ LNA
Sbjct: 454 RGHFEEEKQQLSSLITDLQSSISNLSQAKEELEQASQAHGARLTAQVASLTSELTTLNAT 513
Query: 308 KASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQ 367
+ ++L +Q E+ + E A Q L +E LS L+ + Q+++
Sbjct: 514 IQQQDQELAGLKQQAKEKQAQLAQTLQQQEQASQGLRHQVE----QLSSSLKQKEQQLKE 569
Query: 368 AKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSK 427
E+ ++ R+ D ++ AAEE + ERD A K L+ EK + +
Sbjct: 570 VAEKQEATRQ--DHAQQLATAAEERE-ASLRERD--------AALKQLEALEKEKAAKLE 618
Query: 428 IPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLEE------------YHVQRQLREEEK 475
I + + + EKA L + +EE + Q Q+ E E
Sbjct: 619 ILQQQLQVANEARDSAQTSVTQAQREKAELSRKVEELQACVETARQEQHEAQAQVAELEL 678
Query: 476 RKSREQKRLQEQHAVEQE 493
+ EQ++ E+ V QE
Sbjct: 679 QLRSEQQKATEKERVAQE 696
Score = 37.7 bits (86), Expect = 0.080
Identities = 103/476 (21%), Positives = 189/476 (39%), Gaps = 62/476 (13%)
Query: 42 QLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLA 101
Q +QE D R ++A R +AE L + +L LG S S + +++LA
Sbjct: 1137 QKQQEQADSLERSLEAERASRAERDSALETLQGQLEEKAQELGHSQS---ALASAQRELA 1193
Query: 102 TIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGEL 161
R ++D +DE + ++ + + + I+ E+ S +K GE
Sbjct: 1194 AFRTKVQDHSKAEDEWKAQVARGRQEAERKNSLISSLEEEVSILNR-----QVLEKEGES 1248
Query: 162 KSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISN 221
K L+ L + + QK++ + + +A S +E +L + + +
Sbjct: 1249 K-ELKRLVMAESEKSQKLEERLRLLQAETASNS----ARAAERSSALREEVQSLREEAEK 1303
Query: 222 DTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVD 281
+AS +L+QE + + +EL + L W ++K F L + ++
Sbjct: 1304 QRVAS-----ENLRQELTSQAERAEELGQEL-KAW--------QEKFFQKEQALSTLQLE 1349
Query: 282 EVSTHGCLSSDV----------IEQVEAEVQRLNALKASK-----MKDLVFKRQNELEEI 326
ST +S + EQ AE + L+ SK ++ + + Q EL E+
Sbjct: 1350 HTSTQALVSELLPAKHLCQQLQAEQAAAEKRHREELEQSKQAAGGLRAELLRAQRELGEL 1409
Query: 327 YRGVHMDVDSEAARQILTSLIESGNIDLSDLLQS---MDDQIRQAKEQAQSRREILDRVE 383
+ E Q L + S LS L ++ + ++ R E+A R+ L+ VE
Sbjct: 1410 IPLRQKVAEQERTAQQLRAEKASYAEQLSMLKKAHGLLAEENRGLGERANLGRQFLE-VE 1468
Query: 384 KWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTK---VK 440
EK++ E +AVR A + AE R S + E +T K K
Sbjct: 1469 ---LDQAREKYVQE-------LAAVR-ADAETRLAEVQREAQSTAREL-EVMTAKYEGAK 1516
Query: 441 AWEMEKGIPFLYEKAPLLQSLEEYHV-QRQLREEEKRKSREQKRLQEQHAVEQEAL 495
+E+ F E+ L +E+ V QR+ ++ + S++ + V+Q+ L
Sbjct: 1517 VKVLEERQRFQEERQKLTAQVEQLEVFQREQTKQVEELSKKLADSDQASKVQQQKL 1572
Score = 36.2 bits (82), Expect = 0.23
Identities = 78/378 (20%), Positives = 156/378 (40%), Gaps = 36/378 (9%)
Query: 17 SLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAEL 76
SL+ Q + E+ E + R + +L+Q + ++ + + R+ AE AE+E
Sbjct: 745 SLVEQHKRERKELEEERAGRKGLEARLQQ-LGEAHQAETEVLRRELAEAMAAQHTAESEC 803
Query: 77 INLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIA 136
LV + ++ R + + +++ E L + K+E K + ++ + ++A
Sbjct: 804 EQLVKEV--AAWRERYEDSQQEEAQYGAMFQEQLMTLKEE----CEKARQELQEAKEKVA 857
Query: 137 GYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSID 196
G E Q++L + L LQ +Q EK +R QK+ +ST+ E A S +
Sbjct: 858 GIESHSELQISRQQNELA-ELHANLARALQQVQ-EKEVRAQKLADDLSTLQEKMAATSKE 915
Query: 197 FWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLW 256
+ L + G+ + + + + + ++Q L+ Q+ +F
Sbjct: 916 VAR-LETLVRKAGEQQETASRELVKEPARA---------GDRQPEWLEEQQGRQFCSTQA 965
Query: 257 DLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLN----------A 306
L + ++ N + RL +A ++ + Q E EV RL A
Sbjct: 966 ALQAMEREAEQMGNELERLRAALMESQGQ----QQEERGQQEREVARLTQERGRAQADLA 1021
Query: 307 LKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIR 366
L+ + +L + QN L E + V EA LT E + +L+ L QI+
Sbjct: 1022 LEKAARAELEMRLQNALNE--QRVEFATLQEALAHALTEK-EGKDQELAKLRGLEAAQIK 1078
Query: 367 QAKEQAQSRREILDRVEK 384
+ +E Q+ +++ +++ K
Sbjct: 1079 ELEELRQTVKQLKEQLAK 1096
>ERC1_HUMAN (Q8IUD2) ERC protein 1 (ELKS protein)
Length = 1116
Score = 51.6 bits (122), Expect = 5e-06
Identities = 74/377 (19%), Positives = 163/377 (42%), Gaps = 36/377 (9%)
Query: 45 QECLDIYRRKVDATRKHKAELCQWLADAEAELINL---VSSL-GESSSFSRGKGTLKQQL 100
++ LD+ RKV+ +K L + L D E ++ +L V SL ++++ TL++ L
Sbjct: 563 KDMLDVKERKVNVLQKKIENLQEQLRDKEKQMSSLKERVKSLQADTTNTDTALTTLEEAL 622
Query: 101 ATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQP-KSSSEEVD--QSDLTFKK 157
A +E L+ ++D +E + EI Y++ K E+V Q DL+ K+
Sbjct: 623 AEKERTIERLKEQRDRDEREKQE----------EIDNYKKDLKDLKEKVSLLQGDLSEKE 672
Query: 158 --LGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGT 215
L +LK H L + + + ++K+ + E + L + H + + ++ +
Sbjct: 673 ASLLDLKEHASSLASSGLKKDSRLKT-LEIALEQKKEECLKMESQLKKAHEAALE-ARAS 730
Query: 216 PQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRL 275
P+ +D + L I K E + +V L + L ++ + D+ K + R
Sbjct: 731 PE--MSDRIQHLEREITRYKDESSKAQAEVDRLLEILKEVEN---EKNDKDKKIAELERQ 785
Query: 276 ISASVDEVS-------THGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYR 328
+ +V+ S+ ++E+ LN + +++D + K+ + +EE+
Sbjct: 786 VKDQNKKVANLKHKEQVEKKKSAQMLEEARRREDNLND-SSQQLQDSLRKKDDRIEELEE 844
Query: 329 GVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFA 388
+ V A R+++ + ES + ++ + + + K++ +S + L + +
Sbjct: 845 ALRESVQITAEREMVLAQEESARTNAEKQVEELLMAMEKVKQELESMKAKLSSTQ--QSL 902
Query: 389 AEEEKWLDEYERDENRY 405
AE+E L + ++
Sbjct: 903 AEKETHLTNLRAERRKH 919
Score = 38.1 bits (87), Expect = 0.062
Identities = 99/507 (19%), Positives = 195/507 (37%), Gaps = 67/507 (13%)
Query: 13 TTCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADA 72
TT L++ + + + E DRD + +QE +D Y++ + ++ + L L++
Sbjct: 616 TTLEEALAEKERTIERLKEQ-RDRDE---REKQEEIDNYKKDLKDLKEKVSLLQGDLSEK 671
Query: 73 EAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQIC 132
EA L++L +S K + +L T+ LE KK+E +K S++K
Sbjct: 672 EASLLDLKEHASSLASSGLKKDS---RLKTLEIALEQ---KKEECLKMESQLKKAHEAAL 725
Query: 133 VEIAGYEQP-------KSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHIST 185
A E + + D+S ++ L L++++NEK + +K
Sbjct: 726 EARASPEMSDRIQHLEREITRYKDESSKAQAEVDRLLEILKEVENEKNDKDKK------- 778
Query: 186 ISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSIS----NDTLASLTGYIHSLKQEKQQR 241
I+EL + K + H + K D L + + ++K R
Sbjct: 779 IAELERQVKDQNKKVANLKHKEQVEKKKSAQMLEEARRREDNLNDSSQQLQDSLRKKDDR 838
Query: 242 LLKVQELTKFLVDLWDLIETPIDEQK-----AFNHVTRLISASV---DEVSTHGCLSSDV 293
+ +++E + V + E + +++ A V L+ A E+ + S
Sbjct: 839 IEELEEALRESVQITAEREMVLAQEESARTNAEKQVEELLMAMEKVKQELESMKAKLSST 898
Query: 294 IEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNID 353
+ + + L L+A + K L + + E + + + D+ A L+S + +
Sbjct: 899 QQSLAEKETHLTNLRAERRKHLEEVLEMKQEALLAAIS-EKDANIALLELSSSKKKTQEE 957
Query: 354 LSDLLQSMDDQIRQAKEQAQSRREIL-DRVEKWRFAAEEEKW-------------LDEYE 399
++ L + D ++Q K+Q Q+R +++ D E F + L E +
Sbjct: 958 VAALKREKDRLVQQLKQQTQNRMKLMADNYEDDHFKSSHSNQTNHKPSPDQIIQPLLELD 1017
Query: 400 RDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQ 459
++ ++ G L I+ P NL AWE E LQ
Sbjct: 1018 QNRSKLKLYIGHLTTLCHDRDPLILRGLTPPASYNLDDDQAAWENE------------LQ 1065
Query: 460 SLEEYHVQRQLREEEKRKSREQKRLQE 486
+ + QL++E ++ R+ LQE
Sbjct: 1066 KM----TRGQLQDELEKGERDNAELQE 1088
Score = 33.5 bits (75), Expect = 1.5
Identities = 92/492 (18%), Positives = 199/492 (39%), Gaps = 70/492 (14%)
Query: 36 RDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSF-----S 90
RDN ++ L+ + ++ R D RK D E + L SS+ +F
Sbjct: 147 RDNTIMDLQTQLKEVLREN-DLLRK----------DVEVKESKLSSSMNSIKTFWSPELK 195
Query: 91 RGKGTLKQQLATIRPVLEDLRSKKDER---------VKEFSKIKSQISQICVEIAGYEQP 141
+ + K + + I E R ++E +++ +I+ ++Q+ ++Q
Sbjct: 196 KERALRKDEASKITIWKEQYRVVQEENQHMQMTIQALQDELRIQRDLNQL------FQQD 249
Query: 142 KSS-SEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSI--DFW 198
SS + E ++LT + L + + E L ++ ++ I ++ +
Sbjct: 250 SSSRTGEPCVAELTEENFQRLHAEHERQAKELFLLRKTLEEMELRIETQKQTLNARDESI 309
Query: 199 KTLSEIHPSLGDSSKGTPQSISNDT-LASLTGYIHSLK-----QEKQQRLLKVQELTKF- 251
K L E+ S G S+K T + LA ++H L+ +EK+ +L+ + +F
Sbjct: 310 KKLLEMLQSKGLSAKATEEDHERTRRLAEAEMHVHHLESLLEQKEKENSMLREEMHRRFE 369
Query: 252 -------LVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQV--EAEVQ 302
L +IE + + R + + + ++G LS++ E+ + EV
Sbjct: 370 NAPDSAKTKALQTVIEMKDSKISSMERGLRDLEEEIQMLKSNGALSTEEREEEMKQMEVY 429
Query: 303 RLNAL----KASKMKDLVFKRQNELEEIYR-GVHMDVDSEAARQILTSLIESGNIDLSDL 357
R ++ K ++K+ + ++ + EE+ + + + +Q L S ++ + L
Sbjct: 430 RSHSKFMKNKVEQLKEELSSKEAQWEELKKKAAGLQAEIGQVKQEL-SRKDTELLALQTK 488
Query: 358 LQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKR 417
L+++ +Q +K+ + +E L E+ + E +E + +
Sbjct: 489 LETLTNQFSDSKQHIEVLKESLTAKEQRAAILQTEVDALRLRLEEKETMLNKKTKQIQDM 548
Query: 418 AEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRK 477
AE+ +I + + L K + K +LQ E ++Q QLR++EK+
Sbjct: 549 AEEKGTQAGEIHDLKDMLDVKER-------------KVNVLQKKIE-NLQEQLRDKEKQM 594
Query: 478 SREQKRLQEQHA 489
S ++R++ A
Sbjct: 595 SSLKERVKSLQA 606
>MYHB_RABIT (P35748) Myosin heavy chain, smooth muscle isoform (SMMHC)
Length = 1972
Score = 51.2 bits (121), Expect = 7e-06
Identities = 107/547 (19%), Positives = 226/547 (40%), Gaps = 93/547 (17%)
Query: 18 LLSQLQMIWDEIGESDSDRDNMLLQ----------LEQECL--DIYRRKVDATRK----H 61
+ Q+ + +++ E ++ R + L+ LE + L D K+ RK
Sbjct: 948 MAQQMLDLEEQLEEEEAARQKLQLEKVTAEAKIKKLEDDILVMDDQNNKLSKERKLLEER 1007
Query: 62 KAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERV--- 118
++L LA+ E + NL + S ++ R LE L+ K D
Sbjct: 1008 ISDLTTNLAEEEEKAKNLTKLKNKHESMISELEVRLKKEEKSRQELEKLKRKMDGEASDL 1067
Query: 119 -KEFSKIKSQISQICVEIAGYEQPKSSS-----EEVDQSDLTFKKLGELKSHLQDLQ--- 169
++ + +++QI+++ +++A E+ ++ +E Q + KK+ EL+ H+ DLQ
Sbjct: 1068 HEQIADLQAQIAELKMQLAKKEEELQAALARLEDETSQKNNALKKIRELEGHISDLQEDL 1127
Query: 170 -NEKILRQQKVKSHISTISELSAVM-----SIDFWKTLSEIHPSLGDSSKGTPQSISNDT 223
+E+ R + K EL A+ ++D T E+ +++ +T
Sbjct: 1128 DSERAARNKAEKQKRDLGEELEALKTELEDTLDTTATQQELRAKREQEVTVLKKALDEET 1187
Query: 224 LASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEV 283
+ +++ +Q+ V+ELT+ L + + + +D+ K + + + E+
Sbjct: 1188 ----RSHEAQVQEMRQKHTQVVEELTEQL-EQFKRAKANLDKTK--QTLEKENADLAGEL 1240
Query: 284 STHGCLSSDV---IEQVEAEVQRLNA------LKASKMKDLVFKRQNELEEIYRGVHMDV 334
G +V +++E ++Q L + +++ D V K QNE+E + G+ +
Sbjct: 1241 RVLGQAKQEVEHKKKKLEVQLQELQSKCSDGERARAELNDKVHKLQNEVESV-TGMLSEA 1299
Query: 335 DSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEE-- 392
+ +A + L + S L D + + ++ RQ + R++ D + +EE
Sbjct: 1300 EGKAIK--LAKEVASLGSQLQDTQELLQEETRQKLNVSTKLRQLEDERNSLQEQLDEEME 1357
Query: 393 --------------KWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTK 438
+ D ++ ++ S V + KR +K +I S+ + K
Sbjct: 1358 AKQNLERHISTLNIQLSDSKKKLQDFASTVESLEEGKKRFQK------EIESLTQQYEEK 1411
Query: 439 VKAWE-MEKGIPFLYEKAPLLQSLEEYHV----QRQLREEEKRKSR-------EQKRLQE 486
A++ +EK K L Q L++ V QRQL ++K + E+K +
Sbjct: 1412 AAAYDKLEK------TKNRLQQELDDLVVDLDNQRQLVSNLEKKQKKFDQLLAEEKNISS 1465
Query: 487 QHAVEQE 493
++A E++
Sbjct: 1466 KYADERD 1472
Score = 47.0 bits (110), Expect = 1e-04
Identities = 99/499 (19%), Positives = 200/499 (39%), Gaps = 65/499 (13%)
Query: 21 QLQMIWDEIGESDSDRDNMLLQLE---------QECLDIYRRKVDATRKHKAELCQWLAD 71
+LQ + + S ++N L ++ QE LD R + K K +L + L
Sbjct: 1091 ELQAALARLEDETSQKNNALKKIRELEGHISDLQEDLDSERAARNKAEKQKRDLGEEL-- 1148
Query: 72 AEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQI 131
EA L +L +++ + +Q++ ++ L++ + +V+E + +Q+ +
Sbjct: 1149 -EALKTELEDTLDTTATQQELRAKREQEVTVLKKALDEETRSHEAQVQEMRQKHTQVVEE 1207
Query: 132 CVEIAGYEQPKSSSEEVDQSDLTFKK-----------LGELKSHLQDLQNEKILRQQKVK 180
E EQ K + +D++ T +K LG+ K ++ + + ++ Q+++
Sbjct: 1208 LTE--QLEQFKRAKANLDKTKQTLEKENADLAGELRVLGQAKQEVEHKKKKLEVQLQELQ 1265
Query: 181 SHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSIS-NDTLASLTGYIHS---LKQ 236
S S A ++ K +E+ G S+ ++I +ASL + L Q
Sbjct: 1266 SKCSDGERARAELNDKVHKLQNEVESVTGMLSEAEGKAIKLAKEVASLGSQLQDTQELLQ 1325
Query: 237 EKQQRLLKVQELTKFLVDLWDLIETPIDEQ-KAFNHVTRLISASVDEVSTHGCLSSDVIE 295
E+ ++ L V + L D + ++ +DE+ +A ++ R +ST SD +
Sbjct: 1326 EETRQKLNVSTKLRQLEDERNSLQEQLDEEMEAKQNLER-------HISTLNIQLSDSKK 1378
Query: 296 QVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLS 355
+++ + +L+ K + Q E+E + + E L ++ N
Sbjct: 1379 KLQDFASTVESLEEGKKRF-----QKEIESLTQ------QYEEKAAAYDKLEKTKN---- 1423
Query: 356 DLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNL 415
L Q +DD + Q Q + + +K+ EEK + DE + K
Sbjct: 1424 RLQQELDDLVVDLDNQRQLVSNLEKKQKKFDQLLAEEKNISSKYADERDRAEAEAREKET 1483
Query: 416 KRAEKARIVVSKIPSIVE-NLTTKVKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEE 474
K AR + + + E T K+ EME L+ S ++ V + + E E
Sbjct: 1484 KALSLARALEEALEAKEELERTNKMLKAEME----------DLVSSKDD--VGKNVHELE 1531
Query: 475 KRKSREQKRLQEQHAVEQE 493
K K + +++E +E
Sbjct: 1532 KSKRALETQMEEMKTQLEE 1550
Score = 45.4 bits (106), Expect = 4e-04
Identities = 106/560 (18%), Positives = 222/560 (38%), Gaps = 69/560 (12%)
Query: 2 AATPPSFSPSRTTCASLLSQLQMIWDEIGESDSDRDNMLLQLEQEC------LDIYRRKV 55
A+T S + + L ++E + + +L+QE LD R+ V
Sbjct: 1384 ASTVESLEEGKKRFQKEIESLTQQYEEKAAAYDKLEKTKNRLQQELDDLVVDLDNQRQLV 1443
Query: 56 DATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKD 115
K + + Q LA+ + + + +R K T LA LE+ K+
Sbjct: 1444 SNLEKKQKKFDQLLAEEKNISSKYADERDRAEAEAREKETKALSLAR---ALEEALEAKE 1500
Query: 116 ERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFK-KLGELKSHLQDLQNE-KI 173
E + +K+++ + ++ + + E+++S + ++ E+K+ L++L++E +
Sbjct: 1501 ELERTNKMLKAEMEDL---VSSKDDVGKNVHELEKSKRALETQMEEMKTQLEELEDELQA 1557
Query: 174 LRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHS 233
K++ ++ + + F + L + + + + Y
Sbjct: 1558 TEDAKLRLEVNM-----QALKVQFERDLQARDEQNEEKRRQLQRQLHE--------YETE 1604
Query: 234 LKQEKQQRLLKVQELTKFLVDLWDLIETPID------------------EQKAFNHVTRL 275
L+ E++QR L K DL DL E D + K F
Sbjct: 1605 LEDERKQRALAAAAKKKLEGDLKDL-ELQADSAIKGREEAIKQLLKLQAQMKDFQRELED 1663
Query: 276 ISASVDEVSTHGCLSSDVIEQVEAEVQRLN-----ALKASKMKDLVFKR-QNELEEIYRG 329
AS DE+ + + +EA++ +L A +A K DL + EL G
Sbjct: 1664 ARASRDEIFATAKENEKKAKSLEADLMQLQEDLAAAERARKQADLEKEELAEELASSLSG 1723
Query: 330 VHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAA 389
+ D + + + +E + +++M D++R+A +QA+ L + A
Sbjct: 1724 RNALQDEKRRLEARIAQLEEELEEEQGNMEAMSDRVRKATQQAEQLSNEL--ATERSTAQ 1781
Query: 390 EEEKWLDEYERD----ENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEME 445
+ E + ER +++ + GA K+ ++ A + +KI + E + + +A E +
Sbjct: 1782 KNESARQQLERQNKELKSKLQEMEGAVKSKFKSTIAAL-EAKIAQLEEQV--EQEAREKQ 1838
Query: 446 KGIPFLYE-----KAPLLQSLEEYHVQRQLREEEKR---KSREQKRLQEQHAVEQEALFG 497
L + K LLQ +E + Q +E+ ++ K ++ KR E+ E + +
Sbjct: 1839 AAAKALKQRDKKLKEMLLQVEDERKMAEQYKEQAEKGNAKVKQLKRQLEEAEEESQRINA 1898
Query: 498 SRSATKKPLGQSTTANTIVG 517
+R ++ L ++T +N +G
Sbjct: 1899 NRRKLQRELDEATESNEAMG 1918
Score = 37.4 bits (85), Expect = 0.11
Identities = 63/316 (19%), Positives = 131/316 (40%), Gaps = 48/316 (15%)
Query: 96 LKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTF 155
L+ ++ ++ +L + K + KE + + SQ+ E + + +
Sbjct: 1285 LQNEVESVTGMLSEAEGKAIKLAKEVASLGSQLQDT------QELLQEETRQKLNVSTKL 1338
Query: 156 KKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGT 215
++L + ++ LQ+ +E++ +Q ++ HIST++ + D K L + ++ +G
Sbjct: 1339 RQLEDERNSLQEQLDEEMEAKQNLERHISTLN----IQLSDSKKKLQDFASTVESLEEGK 1394
Query: 216 PQSISNDTLASLTGYIHSL-----KQEKQQRLLKVQELTKFLVDLWD---LIETPIDEQK 267
+ + SLT K EK + L+ QEL +VDL + L+ +QK
Sbjct: 1395 KRF--QKEIESLTQQYEEKAAAYDKLEKTKNRLQ-QELDDLVVDLDNQRQLVSNLEKKQK 1451
Query: 268 AFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIY 327
F+ + A +S+ D E EA + AL ++ + + + ELE
Sbjct: 1452 KFDQLL----AEEKNISSKYADERDRAE-AEAREKETKALSLARALEEALEAKEELERTN 1506
Query: 328 RGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRF 387
+ + +++ DL+ S DD + E +S+R + ++E+ +
Sbjct: 1507 KMLKAEME--------------------DLVSSKDDVGKNVHELEKSKRALETQMEEMKT 1546
Query: 388 AAEEEKWLDEYERDEN 403
EE + DE + E+
Sbjct: 1547 QLEELE--DELQATED 1560
>MYH9_RAT (Q62812) Myosin heavy chain, nonmuscle type A (Cellular
myosin heavy chain, type A) (Nonmuscle myosin heavy
chain-A) (NMMHC-A)
Length = 1961
Score = 51.2 bits (121), Expect = 7e-06
Identities = 106/533 (19%), Positives = 227/533 (41%), Gaps = 84/533 (15%)
Query: 22 LQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATR----KHKAELCQWLADAEAELI 77
+Q + +++ E +S R LQLE+ + +K++ + +L + E +
Sbjct: 945 IQELEEQLEEEESARQK--LQLEKVTTEAKLKKLEEDQIIMEDQNCKLAKEKKLLEDRVA 1002
Query: 78 NLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSK-------------- 123
+ L E S+ LK + + LE+ +++++ +E K
Sbjct: 1003 EFTTDLMEEEEKSKSLAKLKNKHEAMITDLEERLRREEKQRQELEKTRRKLEGDSTDLSD 1062
Query: 124 ----IKSQISQICVEIAGYEQPKSSS-----EEVDQSDLTFKKLGELKSHL----QDLQN 170
+++QI+++ +++A E+ ++ EE Q ++ KK+ EL++ + +DL++
Sbjct: 1063 QIAELQAQIAELKMQLAKKEEELQAALARVEEEAAQKNMALKKIRELETQISELQEDLES 1122
Query: 171 EKILRQQKVKSHISTISELSAVM-----SIDFWKTLSEIHPSLGDSSKGTPQSISNDTLA 225
E+ R + K EL A+ ++D E+ S + + D
Sbjct: 1123 ERACRNKAEKQKRDLGEELEALKTELEDTLDSTAAQQELR-SKREQEVSILKKTLEDEAK 1181
Query: 226 SLTGYIHSLKQEKQQ---RLLKVQELTKFLVDLWDLIETPIDEQKA--FNHVTRLISASV 280
+ I ++Q+ Q L + E TK + + + ++ ++ N V L+
Sbjct: 1182 THEAQIQEMRQKHSQAVEELAEQLEQTKRVKATLEKAKQTLENERGELANEVKALLQGKG 1241
Query: 281 DEVSTHGCLSSDVIEQVEAEVQRLNALKA------SKMKDLVFKRQNELEEIYRGVHMDV 334
D S ++VEA++Q L + +++ D V K Q EL+ + G+
Sbjct: 1242 D--------SEHKRKKVEAQLQELQVKFSEGERVRTELADKVSKLQVELDSV-TGLLNQS 1292
Query: 335 DSEAARQILT---SLIESGNIDLSDLLQ-------SMDDQIRQAKEQAQSRREILDRVEK 384
DS++++ LT S +ES D +LLQ S+ +++Q +++ S RE L+ E+
Sbjct: 1293 DSKSSK--LTKDFSALESQLQDTQELLQEENRQKLSLSTKLKQMEDEKNSFREQLEEEEE 1350
Query: 385 WRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSK-IPSIVENLTTKVKAWE 443
E++ + + + + L+ AE+A+ + K + + + L KV A++
Sbjct: 1351 EAKRNLEKQIATLHAQVTDMKKKMEDGVGCLETAEEAKRRLQKDLEGLSQRLEEKVAAYD 1410
Query: 444 -MEKGIPFLYEKAPLLQSLEEY-----HVQRQLREEEKRKSREQKRLQEQHAV 490
+EK K L Q L++ H ++ + EK++ + + L E+ +
Sbjct: 1411 KLEK------TKTRLQQELDDLLVDLDHQRQSVSNLEKKQKKFDQLLAEEKTI 1457
Score = 49.3 bits (116), Expect = 3e-05
Identities = 117/570 (20%), Positives = 221/570 (38%), Gaps = 91/570 (15%)
Query: 12 RTTCASLLSQLQMIWDEI----GESDSDRDNMLLQ---LEQECLDIYRRKVDATRKHKAE 64
RT A +S+LQ+ D + +SDS + LE + D + R+ K
Sbjct: 1268 RTELADKVSKLQVELDSVTGLLNQSDSKSSKLTKDFSALESQLQDTQELLQEENRQ-KLS 1326
Query: 65 LCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKI 124
L L E E + L E K L++Q+AT+ + D++ K ++ V
Sbjct: 1327 LSTKLKQMEDEKNSFREQLEEEEE--EAKRNLEKQIATLHAQVTDMKKKMEDGVGCLETA 1384
Query: 125 KSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHIS 184
+ ++ ++ G Q E+V D K L+ L DL + ++Q V +
Sbjct: 1385 EEAKRRLQKDLEGLSQ--RLEEKVAAYDKLEKTKTRLQQELDDLLVDLDHQRQSVSNLEK 1442
Query: 185 TISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLK 244
+ +++ + KT+S + D ++ + L+ +++Q+ + L
Sbjct: 1443 KQKKFDQLLAEE--KTISAKYAEERDRAEAEAREKETKALSLARALEEAMEQKAELERLN 1500
Query: 245 VQELTKFLVDLWDLIETPIDEQKAFNHV---TRLISASVDEVSTHGCLSSDVIEQVEAEV 301
Q F ++ DL+ + D K+ + + R + V+E+ T +E++E E+
Sbjct: 1501 KQ----FRTEMEDLMSSKDDVGKSVHELEKSNRALEQQVEEMKTQ-------LEELEDEL 1549
Query: 302 Q-----------RLNALKASKMKDLVFK------RQNELEEIYRGVHMDVDSEAA-RQIL 343
Q L A+KA +DL + ++ +L R + +++ E R I
Sbjct: 1550 QATEDAKLRLEVNLQAMKAQFERDLQGRDEQSEEKKKQLVRQVREMEAELEDERKQRSIA 1609
Query: 344 TSLIESGNIDLSDL--------------------LQS-MDDQIRQAKEQAQSRREILDR- 381
+ + +DL DL LQ+ M D +R + SR EIL +
Sbjct: 1610 MAARKKLEMDLKDLEAHIDTANKNREEAIKQLRKLQAQMKDCMRDVDDTRASREEILAQA 1669
Query: 382 ----------------VEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVV 425
+++ AAE K + ERDE + K E+ R +
Sbjct: 1670 KENEKKLKSMEAEMIQLQEELAAAERAKRQAQQERDELADEIANSSGKGALALEEKRRLE 1729
Query: 426 SKIPSIVENLTTKVKAWEM----EKGIPFLYEKAPLLQSLEEYHVQR--QLREEEKRKSR 479
+ I + E L + E+ K ++ +LE H Q+ R++ +R+++
Sbjct: 1730 ALIALLEEELEEEQGNTELINDRLKKANLQIDQINTDLNLERSHAQKNENARQQLERQNK 1789
Query: 480 EQK-RLQEQHAVEQEALFGSRSATKKPLGQ 508
E K +LQE + + S +A + + Q
Sbjct: 1790 ELKAKLQEMESAVKSKYKASIAALEAKIAQ 1819
Score = 43.5 bits (101), Expect = 0.001
Identities = 72/394 (18%), Positives = 159/394 (40%), Gaps = 63/394 (15%)
Query: 42 QLEQECLDIYRRKVDATRKHKAEL---CQWLADAEAELINLVSSLGESSSFSRGKGTLKQ 98
Q + ++ +++ T++ KA L Q L + EL N V +L +GKG +
Sbjct: 1192 QKHSQAVEELAEQLEQTKRVKATLEKAKQTLENERGELANEVKAL------LQGKGDSEH 1245
Query: 99 QLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKL 158
+ + L++L+ K E + +++ ++S++ VE+ + S+ +S K
Sbjct: 1246 KRKKVEAQLQELQVKFSEGERVRTELADKVSKLQVELDSVTGLLNQSDS--KSSKLTKDF 1303
Query: 159 GELKSHLQDLQNEKILRQQKVKSHIST---------------ISELSAVMSIDFWKTLSE 203
L+S LQD Q E + + + K +ST + E + K ++
Sbjct: 1304 SALESQLQDTQ-ELLQEENRQKLSLSTKLKQMEDEKNSFREQLEEEEEEAKRNLEKQIAT 1362
Query: 204 IHPSLGDSSKGTPQSI-----SNDTLASLTGYIHSLKQEKQQRLLKVQEL----TKFLVD 254
+H + D K + + + L + L Q ++++ +L T+ +
Sbjct: 1363 LHAQVTDMKKKMEDGVGCLETAEEAKRRLQKDLEGLSQRLEEKVAAYDKLEKTKTRLQQE 1422
Query: 255 LWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQ-----VEAEVQRLNALKA 309
L DL+ ++++ +++ + ++ +S+ E+ EA + AL
Sbjct: 1423 LDDLLVDLDHQRQSVSNLEKKQKKFDQLLAEEKTISAKYAEERDRAEAEAREKETKALSL 1482
Query: 310 SKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAK 369
++ + +++ ELE + + +++ DL+ S DD +
Sbjct: 1483 ARALEEAMEQKAELERLNKQFRTEME--------------------DLMSSKDDVGKSVH 1522
Query: 370 EQAQSRREILDRVEKWRFAAEEEKWLDEYERDEN 403
E +S R + +VE+ + EE + DE + E+
Sbjct: 1523 ELEKSNRALEQQVEEMKTQLEELE--DELQATED 1554
Score = 40.8 bits (94), Expect = 0.010
Identities = 83/457 (18%), Positives = 182/457 (39%), Gaps = 37/457 (8%)
Query: 45 QECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLATIR 104
QE L+ R + K K +L + L + EL + + S + +Q+++ ++
Sbjct: 1117 QEDLESERACRNKAEKQKRDLGEELEALKTELEDTLDSTAAQQEL---RSKREQEVSILK 1173
Query: 105 PVLEDLRSKKDERVKEFSKIKSQ-ISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKS 163
LED + +++E + SQ + ++ ++ ++ K++ E+ Q+ + GEL +
Sbjct: 1174 KTLEDEAKTHEAQIQEMRQKHSQAVEELAEQLEQTKRVKATLEKAKQT--LENERGELAN 1231
Query: 164 HLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDT 223
++ L K + K K + + EL S + L D S
Sbjct: 1232 EVKALLQGKGDSEHKRKKVEAQLQELQVKFSEG-----ERVRTELADKV-----SKLQVE 1281
Query: 224 LASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEV 283
L S+TG ++ + + L L D +L++ +E + ++ + DE
Sbjct: 1282 LDSVTGLLNQSDSKSSKLTKDFSALESQLQDTQELLQ---EENRQKLSLSTKLKQMEDEK 1338
Query: 284 STHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNEL------EEIYRGVHMDVDSE 337
++ + E+ + +++ A +++ D+ K ++ + EE R + D++
Sbjct: 1339 NSFREQLEEEEEEAKRNLEKQIATLHAQVTDMKKKMEDGVGCLETAEEAKRRLQKDLEGL 1398
Query: 338 AAR-QILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLD 396
+ R + + + + L Q +DD + Q QS + + +K+ EEK +
Sbjct: 1399 SQRLEEKVAAYDKLEKTKTRLQQELDDLLVDLDHQRQSVSNLEKKQKKFDQLLAEEKTIS 1458
Query: 397 EYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAP 456
+E + K K AR ++ E + K + + K F E
Sbjct: 1459 AKYAEERDRAEAEAREKETKALSLAR-------ALEEAMEQKAELERLNK--QFRTEMED 1509
Query: 457 LLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQE 493
L+ S ++ V + + E EK +++++E +E
Sbjct: 1510 LMSSKDD--VGKSVHELEKSNRALEQQVEEMKTQLEE 1544
Score = 31.6 bits (70), Expect = 5.8
Identities = 23/133 (17%), Positives = 66/133 (49%), Gaps = 6/133 (4%)
Query: 41 LQLEQECLDIYRRKVDATRKHKA--ELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQ 98
LQ++Q D+ + A + A +L + + +A+L + S++ S + L+
Sbjct: 1758 LQIDQINTDLNLERSHAQKNENARQQLERQNKELKAKLQEMESAV--KSKYKASIAALEA 1815
Query: 99 QLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKL 158
++A + L++ ++ K+ + + ++ + +++ ++ +++ + DQ+D +L
Sbjct: 1816 KIAQLEEQLDNETKERQAASKQVRRAEKKLKDVLLQVE--DERRNAEQFKDQADKASTRL 1873
Query: 159 GELKSHLQDLQNE 171
+LK L++ + E
Sbjct: 1874 KQLKRQLEEAEEE 1886
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.128 0.347
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,431,027
Number of Sequences: 164201
Number of extensions: 2441370
Number of successful extensions: 12830
Number of sequences better than 10.0: 539
Number of HSP's better than 10.0 without gapping: 37
Number of HSP's successfully gapped in prelim test: 511
Number of HSP's that attempted gapping in prelim test: 11327
Number of HSP's gapped (non-prelim): 1414
length of query: 566
length of database: 59,974,054
effective HSP length: 116
effective length of query: 450
effective length of database: 40,926,738
effective search space: 18417032100
effective search space used: 18417032100
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0005.6


