

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.4
(283 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
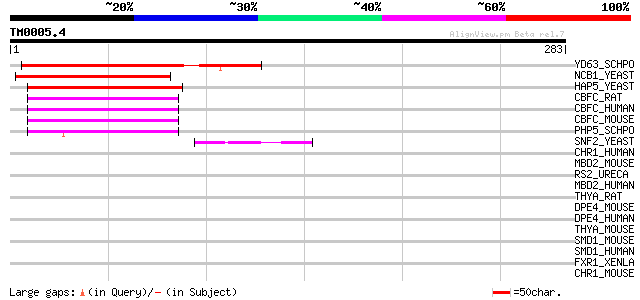
Score E
Sequences producing significant alignments: (bits) Value
YD63_SCHPO (Q10315) Hypothetical protein C17G8.03c in chromosome I 99 9e-21
NCB1_YEAST (P40096) Class 2 transcription repressor (NC2) 67 5e-11
HAP5_YEAST (Q02516) Transcriptional activator HAP5 60 6e-09
CBFC_RAT (Q62725) Nuclear transcription factor Y subunit gamma (... 56 9e-08
CBFC_HUMAN (Q13952) Nuclear transcription factor Y subunit gamma... 56 9e-08
CBFC_MOUSE (P70353) Nuclear transcription factor Y subunit gamma... 52 1e-06
PHP5_SCHPO (P79007) Transcriptional activator php5 50 8e-06
SNF2_YEAST (P22082) Transcription regulatory protein SNF2 (SWI/S... 45 2e-04
CHR1_HUMAN (Q9NRG0) Chromatin accessibility complex protein 1 (C... 43 0.001
MBD2_MOUSE (Q9Z2E1) Methyl-CpG binding protein 2 (Methyl-CpG bin... 42 0.002
RS2_URECA (P49154) 40S ribosomal protein S2 41 0.003
MBD2_HUMAN (Q9UBB5) Methyl-CpG binding protein 2 (Methyl-CpG bin... 41 0.003
THYA_RAT (P06302) Prothymosin alpha 41 0.004
DPE4_MOUSE (Q9CQ36) DNA polymerase epsilon p12 subunit (DNA poly... 41 0.004
DPE4_HUMAN (Q9NR33) DNA polymerase epsilon p12 subunit (DNA poly... 41 0.004
THYA_MOUSE (P26350) Prothymosin alpha 40 0.008
SMD1_MOUSE (P62315) Small nuclear ribonucleoprotein Sm D1 (snRNP... 39 0.011
SMD1_HUMAN (P62314) Small nuclear ribonucleoprotein Sm D1 (snRNP... 39 0.011
FXR1_XENLA (P51115) Fragile X mental retardation syndrome relate... 39 0.011
CHR1_MOUSE (Q9JKP8) Chromatin accessibility complex protein 1 (C... 39 0.014
>YD63_SCHPO (Q10315) Hypothetical protein C17G8.03c in chromosome I
Length = 199
Score = 99.4 bits (246), Expect = 9e-21
Identities = 56/124 (45%), Positives = 76/124 (61%), Gaps = 9/124 (7%)
Query: 7 TRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLH 66
+RFP ARIKKIMQAD+DVGK+A PV++SKALELF+Q + + + T AK + H
Sbjct: 22 SRFPVARIKKIMQADQDVGKVAQVTPVIMSKALELFMQSIIQESCKQTRLHQAKRVTVSH 81
Query: 67 LKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTI--PKRRKAAGDDCNDSDEEV 124
LKH VQS FDFL+D+V KVPD + P +R P+ R+AA + ++
Sbjct: 82 LKHAVQSVEQFDFLQDIVEKVPD-------APPIKAERKTKRPRARRAANEGEHNESVPA 134
Query: 125 KRGK 128
K+ K
Sbjct: 135 KKVK 138
>NCB1_YEAST (P40096) Class 2 transcription repressor (NC2)
Length = 142
Score = 67.0 bits (162), Expect = 5e-11
Identities = 31/79 (39%), Positives = 51/79 (64%)
Query: 4 KLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMN 63
++ T FP A++KKIMQ DED+GK++ A PV+ ++LE F+ L ++ E+ +G K +
Sbjct: 48 RIKTHFPPAKVKKIMQTDEDIGKVSQATPVIAGRSLEFFIALLVKKSGEMARGQGTKRIT 107
Query: 64 SLHLKHCVQSYNVFDFLRD 82
+ LK + + FDFLR+
Sbjct: 108 AEILKKTILNDEKFDFLRE 126
>HAP5_YEAST (Q02516) Transcriptional activator HAP5
Length = 242
Score = 60.1 bits (144), Expect = 6e-09
Identities = 28/79 (35%), Positives = 48/79 (60%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARI+K+M+ DEDV I+ P++ +KA E+F+ +L R + + + +T+ +
Sbjct: 161 PFARIRKVMKTDEDVKMISAEAPIIFAKACEIFITELTMRAWCVAERNKRRTLQKADIAE 220
Query: 70 CVQSYNVFDFLRDVVSKVP 88
+Q ++FDFL DVV + P
Sbjct: 221 ALQKSDMFDFLIDVVPRRP 239
>CBFC_RAT (Q62725) Nuclear transcription factor Y subunit gamma
(NF-Y protein chain C) (Nuclear factor YC) (NF-YC)
(CCAAT-binding transcription factor subunit C) (CBF-C)
Length = 335
Score = 56.2 bits (134), Expect = 9e-08
Identities = 29/77 (37%), Positives = 45/77 (57%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ DEDV I+ PVL +KA ++F+ +L R + T +T+ +
Sbjct: 44 PLARIKKIMKLDEDVKMISAEAPVLFAKAAQIFITELTLRAWIHTEDNKRRTLQRNDIAM 103
Query: 70 CVQSYNVFDFLRDVVSK 86
+ ++ FDFL D+V +
Sbjct: 104 AITKFDQFDFLIDIVPR 120
>CBFC_HUMAN (Q13952) Nuclear transcription factor Y subunit gamma
(NF-Y protein chain C) (Nuclear factor YC) (NF-YC)
(CCAAT-binding transcription factor subunit C) (CBF-C)
(Transactivator HSM-1/2)
Length = 458
Score = 56.2 bits (134), Expect = 9e-08
Identities = 29/77 (37%), Positives = 45/77 (57%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ DEDV I+ PVL +KA ++F+ +L R + T +T+ +
Sbjct: 44 PLARIKKIMKLDEDVKMISAEAPVLFAKAAQIFITELTLRAWIHTEDNKRRTLQRNDIAM 103
Query: 70 CVQSYNVFDFLRDVVSK 86
+ ++ FDFL D+V +
Sbjct: 104 AITKFDQFDFLIDIVPR 120
>CBFC_MOUSE (P70353) Nuclear transcription factor Y subunit gamma
(NF-Y protein chain C) (Nuclear factor YC) (NF-YC)
(CCAAT-binding transcription factor subunit C) (CBF-C)
Length = 335
Score = 52.4 bits (124), Expect = 1e-06
Identities = 27/77 (35%), Positives = 43/77 (55%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ DEDV I+ PVL +K ++F+ +L R + T + + +
Sbjct: 44 PLARIKKIMKLDEDVKMISAEAPVLFAKGAQIFITELTLRAWIRTEDNKRRPLQRNDIAM 103
Query: 70 CVQSYNVFDFLRDVVSK 86
+ ++ FDFL D+V +
Sbjct: 104 AITKFDQFDFLIDIVPR 120
>PHP5_SCHPO (P79007) Transcriptional activator php5
Length = 415
Score = 49.7 bits (117), Expect = 8e-06
Identities = 26/79 (32%), Positives = 44/79 (54%), Gaps = 2/79 (2%)
Query: 10 PAARIKKIMQADEDVGK--IALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
P ARIKK+M+ D+DV I+ P L +K E+F+ +L R + + +T+ +
Sbjct: 110 PLARIKKVMKTDDDVKNKMISAEAPFLFAKGSEIFIAELTMRAWLHAKKNQRRTLQRSDI 169
Query: 68 KHCVQSYNVFDFLRDVVSK 86
+ V ++DFL D++SK
Sbjct: 170 ANAVSKSEMYDFLIDIISK 188
>SNF2_YEAST (P22082) Transcription regulatory protein SNF2 (SWI/SNF
complex component SNF2) (Regulatory protein SWI2)
(Regulatory protein GAM1) (Transcription factor TYE3)
Length = 1703
Score = 45.1 bits (105), Expect = 2e-04
Identities = 29/60 (48%), Positives = 31/60 (51%), Gaps = 11/60 (18%)
Query: 95 GHSEPSADDRTIPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGR 154
G + PS D IPK R AG S RG+ GRGRGRGRGRGRGRGR
Sbjct: 1475 GDNSPSEDFMDIPKPR-TAGKTSVKSARTSTRGR----------GRGRGRGRGRGRGRGR 1523
>CHR1_HUMAN (Q9NRG0) Chromatin accessibility complex protein 1
(CHRAC-1) (CHRAC-15) (HuCHRAC15) (DNA polymerase
epsilon subunit p15)
Length = 131
Score = 42.7 bits (99), Expect = 0.001
Identities = 25/77 (32%), Positives = 38/77 (48%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P +RI+ IM++ +V I VL +KA ELF+Q L +Y + K + L +
Sbjct: 20 PLSRIRVIMKSSPEVSSINQEALVLTAKATELFVQCLATYSYRHGSGKEKKVLTYSDLAN 79
Query: 70 CVQSYNVFDFLRDVVSK 86
Q F FL D++ K
Sbjct: 80 TAQQSETFQFLADILPK 96
>MBD2_MOUSE (Q9Z2E1) Methyl-CpG binding protein 2 (Methyl-CpG
binding domain protein 2)
Length = 414
Score = 41.6 bits (96), Expect = 0.002
Identities = 21/37 (56%), Positives = 24/37 (64%), Gaps = 1/37 (2%)
Query: 126 RGKMPELSHTGST-GRGRGRGRGRGRGRGRTAVRETP 161
RG+ + + G GRGRGRGRGRGRGRGR R P
Sbjct: 61 RGRWKQAARGGGVCGRGRGRGRGRGRGRGRGRGRGRP 97
Score = 29.6 bits (65), Expect = 8.7
Identities = 16/30 (53%), Positives = 17/30 (56%), Gaps = 11/30 (36%)
Query: 136 GSTGRGRGRGR-----------GRGRGRGR 154
G+ G GRGRGR GRGRGRGR
Sbjct: 53 GARGGGRGRGRWKQAARGGGVCGRGRGRGR 82
>RS2_URECA (P49154) 40S ribosomal protein S2
Length = 278
Score = 41.2 bits (95), Expect = 0.003
Identities = 18/21 (85%), Positives = 18/21 (85%)
Query: 136 GSTGRGRGRGRGRGRGRGRTA 156
G GRGRGRGRGRGRGRGR A
Sbjct: 19 GGRGRGRGRGRGRGRGRGRGA 39
Score = 39.3 bits (90), Expect = 0.011
Identities = 17/19 (89%), Positives = 17/19 (89%)
Query: 136 GSTGRGRGRGRGRGRGRGR 154
G GRGRGRGRGRGRGRGR
Sbjct: 17 GFGGRGRGRGRGRGRGRGR 35
Score = 30.8 bits (68), Expect = 3.9
Identities = 14/19 (73%), Positives = 14/19 (73%)
Query: 136 GSTGRGRGRGRGRGRGRGR 154
G G GRGRGRGRGRGR
Sbjct: 13 GFRGGFGGRGRGRGRGRGR 31
>MBD2_HUMAN (Q9UBB5) Methyl-CpG binding protein 2 (Methyl-CpG
binding domain protein 2) (Demethylase) (DMTase)
Length = 411
Score = 41.2 bits (95), Expect = 0.003
Identities = 21/37 (56%), Positives = 23/37 (61%), Gaps = 1/37 (2%)
Query: 126 RGKMPELSHTGST-GRGRGRGRGRGRGRGRTAVRETP 161
RG+ + G GRGRGRGRGRGRGRGR R P
Sbjct: 61 RGRWKQAGRGGGVCGRGRGRGRGRGRGRGRGRGRGRP 97
Score = 29.6 bits (65), Expect = 8.7
Identities = 16/30 (53%), Positives = 17/30 (56%), Gaps = 11/30 (36%)
Query: 136 GSTGRGRGRGR-----------GRGRGRGR 154
G+ G GRGRGR GRGRGRGR
Sbjct: 53 GARGGGRGRGRWKQAGRGGGVCGRGRGRGR 82
>THYA_RAT (P06302) Prothymosin alpha
Length = 111
Score = 40.8 bits (94), Expect = 0.004
Identities = 31/103 (30%), Positives = 47/103 (45%), Gaps = 8/103 (7%)
Query: 180 DTNTDMTIHDSSETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPP 239
DT++++T D E KE+ +E AE+ + NA EN +++A +E E
Sbjct: 6 DTSSEITTKDLKEKKEVVEE-----AENGRDAPANGNAQNEENGEQEADNEVDEEEEEGG 60
Query: 240 TESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEE 282
E + EE D D+ D A G + ED+ED D E
Sbjct: 61 EEEEEEEEGDGEEEDGDE---DEEAEAPTGKRVAEDDEDDDVE 100
>DPE4_MOUSE (Q9CQ36) DNA polymerase epsilon p12 subunit (DNA
polymerase epsilon subunit 4)
Length = 118
Score = 40.8 bits (94), Expect = 0.004
Identities = 21/74 (28%), Positives = 39/74 (52%)
Query: 7 TRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLH 66
+R P AR+K +++AD DV ++++A ELF++ + Y Q KT+
Sbjct: 40 SRLPLARVKALVKADPDVTLAGQEAIFILARAAELFVETIAKDAYCCAQQGKRKTLQRRD 99
Query: 67 LKHCVQSYNVFDFL 80
L + +++ + F FL
Sbjct: 100 LDNAIEAVDEFAFL 113
>DPE4_HUMAN (Q9NR33) DNA polymerase epsilon p12 subunit (DNA
polymerase epsilon subunit 4)
Length = 117
Score = 40.8 bits (94), Expect = 0.004
Identities = 21/74 (28%), Positives = 39/74 (52%)
Query: 7 TRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLH 66
+R P AR+K +++AD DV ++++A ELF++ + Y Q KT+
Sbjct: 39 SRLPLARVKALVKADPDVTLAGQEAIFILARAAELFVETIAKDAYCCAQQGKRKTLQRRD 98
Query: 67 LKHCVQSYNVFDFL 80
L + +++ + F FL
Sbjct: 99 LDNAIEAVDEFAFL 112
>THYA_MOUSE (P26350) Prothymosin alpha
Length = 110
Score = 39.7 bits (91), Expect = 0.008
Identities = 30/101 (29%), Positives = 46/101 (44%), Gaps = 8/101 (7%)
Query: 180 DTNTDMTIHDSSETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPP 239
DT++++T D E KE+ +E AE+ + NA EN +++A +E E
Sbjct: 6 DTSSEITTKDLKEKKEVVEE-----AENGRDAPANGNAQNEENGEQEADNEVDEEEEEGG 60
Query: 240 TESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYD 280
E + EE D D+ D A G + ED+ED D
Sbjct: 61 EEEEEEEEGDGEEEDGDE---DEEAEAPTGKRVAEDDEDDD 98
>SMD1_MOUSE (P62315) Small nuclear ribonucleoprotein Sm D1 (snRNP
core protein D1) (Sm-D1) (Sm-D autoantigen)
Length = 119
Score = 39.3 bits (90), Expect = 0.011
Identities = 21/40 (52%), Positives = 25/40 (62%), Gaps = 3/40 (7%)
Query: 119 DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVR 158
D + +VK K ++ GRGRGRGRGRGRGRGR R
Sbjct: 82 DVEPKVKSKKREAVA---GRGRGRGRGRGRGRGRGRGGPR 118
Score = 35.4 bits (80), Expect = 0.16
Identities = 16/23 (69%), Positives = 17/23 (73%)
Query: 141 GRGRGRGRGRGRGRTAVRETPHQ 163
GRGRGRGRGRGRGR R P +
Sbjct: 97 GRGRGRGRGRGRGRGRGRGGPRR 119
>SMD1_HUMAN (P62314) Small nuclear ribonucleoprotein Sm D1 (snRNP
core protein D1) (Sm-D1) (Sm-D autoantigen)
Length = 119
Score = 39.3 bits (90), Expect = 0.011
Identities = 21/40 (52%), Positives = 25/40 (62%), Gaps = 3/40 (7%)
Query: 119 DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVR 158
D + +VK K ++ GRGRGRGRGRGRGRGR R
Sbjct: 82 DVEPKVKSKKREAVA---GRGRGRGRGRGRGRGRGRGGPR 118
Score = 35.4 bits (80), Expect = 0.16
Identities = 16/23 (69%), Positives = 17/23 (73%)
Query: 141 GRGRGRGRGRGRGRTAVRETPHQ 163
GRGRGRGRGRGRGR R P +
Sbjct: 97 GRGRGRGRGRGRGRGRGRGGPRR 119
>FXR1_XENLA (P51115) Fragile X mental retardation syndrome related
protein 1
Length = 648
Score = 39.3 bits (90), Expect = 0.011
Identities = 36/181 (19%), Positives = 68/181 (36%), Gaps = 9/181 (4%)
Query: 88 PDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRG 147
P+Y+ G+G + ++ RK D + + E+ + + S GRGR G
Sbjct: 422 PNYTSGYGTNSELSNPSETESERKEELSDWSLAGEDERESRQQRDSRRRPGGRGRSGSAG 481
Query: 148 RGRGRGRTAVRETPHQAVEPEP-----CASVQQGIQHDTNTDMTIHDSSETKELPKENAA 202
RGRG R +P+ + + DT+ + H+++ + +
Sbjct: 482 RGRGGSRGGKSSISSVLKDPDSNPYSLLDNTESDQTADTDASESHHNTNRRRRSRRRRTD 541
Query: 203 VPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDS 262
+ + NA+ NEN T IS ++S+ +P +L+ K +
Sbjct: 542 EDSSLMDGMTELDNASVNEN----GLVTVADYISRAESQSRQRNLPKETLAKGKKEKVKD 597
Query: 263 V 263
V
Sbjct: 598 V 598
>CHR1_MOUSE (Q9JKP8) Chromatin accessibility complex protein 1
(CHRAC-1) (DNA polymerase epsilon subunit p15) (YC-like
protein 1) (YCL1) (NF-YC-like protein)
Length = 129
Score = 38.9 bits (89), Expect = 0.014
Identities = 23/77 (29%), Positives = 36/77 (45%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P +RI+ IM++ +V I VL +KA ELF+Q L +Y + K + L
Sbjct: 20 PLSRIRVIMKSSPEVSSINQEALVLTAKATELFVQYLATCSYRHGSGKAKKALTYSDLAS 79
Query: 70 CVQSYNVFDFLRDVVSK 86
+ FL D++ K
Sbjct: 80 TAEDSETLQFLADILPK 96
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.310 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,845,916
Number of Sequences: 164201
Number of extensions: 1583989
Number of successful extensions: 7837
Number of sequences better than 10.0: 174
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 114
Number of HSP's that attempted gapping in prelim test: 7239
Number of HSP's gapped (non-prelim): 389
length of query: 283
length of database: 59,974,054
effective HSP length: 109
effective length of query: 174
effective length of database: 42,076,145
effective search space: 7321249230
effective search space used: 7321249230
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0005.4


