

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.13
(396 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
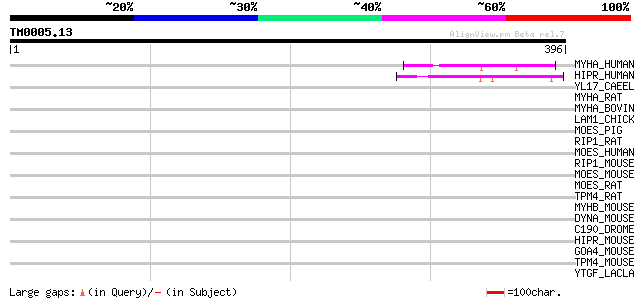
Score E
Sequences producing significant alignments: (bits) Value
MYHA_HUMAN (P35580) Myosin heavy chain, nonmuscle type B (Cellul... 45 4e-04
HIPR_HUMAN (O75146) Huntingtin interacting protein 1 related (Hi... 44 7e-04
YL17_CAEEL (Q11102) Hypothetical protein C02F12.7 in chromosome X 44 0.001
MYHA_RAT (Q9JLT0) Myosin heavy chain, nonmuscle type B (Cellular... 44 0.001
MYHA_BOVIN (Q27991) Myosin heavy chain, nonmuscle type B (Cellul... 44 0.001
LAM1_CHICK (P14731) Lamin B1 43 0.002
MOES_PIG (P26042) Moesin (Membrane-organizing extension spike pr... 42 0.002
RIP1_RAT (Q9JIR0) Peripheral-type benzodiazepine receptor-associ... 42 0.004
MOES_HUMAN (P26038) Moesin (Membrane-organizing extension spike ... 42 0.004
RIP1_MOUSE (Q7TNF8) Peripheral-type benzodiazepine receptor-asso... 40 0.008
MOES_MOUSE (P26041) Moesin (Membrane-organizing extension spike ... 40 0.008
MOES_RAT (O35763) Moesin (Membrane-organizing extension spike pr... 40 0.010
TPM4_RAT (P09495) Tropomyosin alpha 4 chain (Tropomyosin 4) (TM-4) 40 0.014
MYHB_MOUSE (O08638) Myosin heavy chain, smooth muscle isoform (S... 40 0.014
DYNA_MOUSE (O08788) Dynactin 1 (150 kDa dynein-associated polype... 40 0.014
C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 1... 40 0.014
HIPR_MOUSE (Q9JKY5) Huntingtin interacting protein 1 related (Hi... 39 0.018
GOA4_MOUSE (Q91VW5) Golgi autoantigen, golgin subfamily A member... 39 0.018
TPM4_MOUSE (Q6IRU2) Tropomyosin alpha 4 chain (Tropomyosin 4) 39 0.023
YTGF_LACLA (Q9CEE2) Hypothetical UPF0144 protein ytgF 39 0.030
>MYHA_HUMAN (P35580) Myosin heavy chain, nonmuscle type B (Cellular
myosin heavy chain, type B) (Nonmuscle myosin heavy
chain-B) (NMMHC-B)
Length = 1976
Score = 44.7 bits (104), Expect = 4e-04
Identities = 30/114 (26%), Positives = 61/114 (53%), Gaps = 10/114 (8%)
Query: 282 LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK---ISD 338
+ E+ L N+ F++E+K ++ + + +LAE+ +A+ + R+++ ISD
Sbjct: 983 MEEEILLLEDQNSKFIKEKKL----MEDRIAECSSQLAEEEEKAKNLAKIRNKQEVMISD 1038
Query: 339 LEKALGKLRQEAQEESEAKVDL---VASKDQTLSSMETKLEHLRAELAKKDEAL 389
LE+ L K + QE +AK L ++ ++ +++ L+ +LAKK+E L
Sbjct: 1039 LEERLKKEEKTRQELEKAKRKLDGETTDLQDQIAELQAQIDELKLQLAKKEEEL 1092
Score = 39.7 bits (91), Expect = 0.014
Identities = 39/140 (27%), Positives = 63/140 (44%), Gaps = 27/140 (19%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ + L ++ R + + +++K F QL A+E S + AE+ RAE ++
Sbjct: 1424 RLQQELDDLTVDLDHQRQVASNLEKKQKKF-DQLLAEEKSISARYAEERDRAEAEAREKE 1482
Query: 334 RKISDLEKALGKLRQEAQEESEAK--------VDLVASKD-----------------QTL 368
K L +AL + EA+EE E + DL++SKD Q +
Sbjct: 1483 TKALSLARALEE-ALEAKEEFERQNKQLRADMEDLMSSKDDVGKNVHELEKSKRALEQQV 1541
Query: 369 SSMETKLEHLRAELAKKDEA 388
M T+LE L EL ++A
Sbjct: 1542 EEMRTQLEELEDELQATEDA 1561
Score = 38.9 bits (89), Expect = 0.023
Identities = 32/120 (26%), Positives = 58/120 (47%), Gaps = 8/120 (6%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ E R + ++R L A +ERK + +K+ +L + ++ E + + RD
Sbjct: 1583 RDEQNEEKKRLLIKQVRELEAELEDERKQRALAVASKK-KMEIDLKDLEAQIEAANKARD 1641
Query: 334 RKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLS---SMETKLEHLRAELAKKDEALS 390
I L K +++ +E EA+ AS+D+ + E KL+ L AE+ + E L+
Sbjct: 1642 EVIKQLRKLQAQMKDYQRELEEAR----ASRDEIFAQSKESEKKLKSLEAEILQLQEELA 1697
Score = 34.3 bits (77), Expect = 0.57
Identities = 29/111 (26%), Positives = 55/111 (49%), Gaps = 6/111 (5%)
Query: 285 EVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQ---SSRAEVSLEFRDRKISDLEK 341
EVA L+ + +A + ++ + + +EL+EQ + R + +LE + + K
Sbjct: 1175 EVAELKKALEEETKNHEAQIQDMRQRHATALEELSEQLEQAKRFKANLEKNKQGLETDNK 1234
Query: 342 ALG---KLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEAL 389
L K+ Q+ + ESE K + ++ Q L + ++ + LR ELA+K L
Sbjct: 1235 ELACEVKVLQQVKAESEHKRKKLDAQVQELHAKVSEGDRLRVELAEKASKL 1285
Score = 32.0 bits (71), Expect = 2.8
Identities = 37/155 (23%), Positives = 61/155 (38%), Gaps = 37/155 (23%)
Query: 272 YKRSAQDAEV------------LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELA 319
+KR DA+V LR E+A + ++ L + + K FAK+ A
Sbjct: 1252 HKRKKLDAQVQELHAKVSEGDRLRVELAEKASKLQNELDNVSTLLEEAEKKGIKFAKDAA 1311
Query: 320 EQSSRAEVSLEFRDR----------KISDLEKALGKLRQEAQEESEAK------------ 357
S+ + + E +I LE+ L+++ +EE EA+
Sbjct: 1312 SLESQLQDTQELLQEETRQKLNLSSRIRQLEEEKNSLQEQQEEEEEARKNLEKQVLALQS 1371
Query: 358 --VDLVASKDQTLSSMETKLEHLRAELAKKDEALS 390
D D L ++E+ LE + +L K EALS
Sbjct: 1372 QLADTKKKVDDDLGTIES-LEEAKKKLLKDAEALS 1405
>HIPR_HUMAN (O75146) Huntingtin interacting protein 1 related
(Hip1-related) (Hip 12)
Length = 1068
Score = 43.9 bits (102), Expect = 7e-04
Identities = 34/126 (26%), Positives = 65/126 (50%), Gaps = 14/126 (11%)
Query: 277 QDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDR-- 334
++ E+LRSE+ +++ E + ++AQLK++ + EL EQ + + +L ++
Sbjct: 361 REVEMLRSELEKIKL-------EAQRYIAQLKSQVNALEGELEEQRKQKQKALVDNEQLR 413
Query: 335 -KISDLEKAL--GKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKK--DEAL 389
+++ L A G+ Q +EE+E K ++ L ++L H+ AEL +K D A
Sbjct: 414 HELAQLRAAQLEGERSQGLREEAERKASATEARYNKLKEKHSELVHVHAELLRKNADTAK 473
Query: 390 SMMRTQ 395
+ TQ
Sbjct: 474 QLTVTQ 479
Score = 34.3 bits (77), Expect = 0.57
Identities = 38/138 (27%), Positives = 57/138 (40%), Gaps = 24/138 (17%)
Query: 273 KRSAQDAEVLRSEVARLRAL------NAGFVEERKAFLAQLKAKETSFAKELAEQSSRAE 326
+++ D E LR E+A+LRA + G EE + + A E + K + S
Sbjct: 403 QKALVDNEQLRHELAQLRAAQLEGERSQGLREEAER---KASATEARYNKLKEKHSELVH 459
Query: 327 VSLEFRDRKISDLEKALGKLRQEAQEESEAKVDL--------------VASKDQTLSSME 372
V E RK +D K L +Q +E + K L + K L ++
Sbjct: 460 VHAELL-RKNADTAKQLTVTQQSQEEVARVKEQLAFQVEQVKRESELKLEEKSDQLEKLK 518
Query: 373 TKLEHLRAELAKKDEALS 390
+LE ELA+ EALS
Sbjct: 519 RELEAKAGELARAQEALS 536
>YL17_CAEEL (Q11102) Hypothetical protein C02F12.7 in chromosome X
Length = 1130
Score = 43.5 bits (101), Expect = 0.001
Identities = 32/131 (24%), Positives = 62/131 (46%), Gaps = 11/131 (8%)
Query: 275 SAQDAEVLRSEVARLRALNAGFVEERK----AFLAQLKAKETSFAKELAEQSSRAEVSLE 330
SA+ A++L S A EER+ +F Q+ K+ + AE+ + E L+
Sbjct: 782 SAKTAQLLESNTVESETKLASVTEEREKEIQSFQTQISEKDNEVLTK-AERINELETCLK 840
Query: 331 FRDRKISDLEKALGKLRQEAQEESEAKV------DLVASKDQTLSSMETKLEHLRAELAK 384
R+ +++ + L + Q+ EE+ + + + K+ T++ M +L+ E+AK
Sbjct: 841 EREVELTGMRTKLDDMTQQLNEETTVVLFDNSIQEKIDEKEATINEMNERLKSRENEIAK 900
Query: 385 KDEALSMMRTQ 395
E + M +TQ
Sbjct: 901 LHEEMYMQKTQ 911
Score = 31.6 bits (70), Expect = 3.7
Identities = 35/142 (24%), Positives = 60/142 (41%), Gaps = 23/142 (16%)
Query: 275 SAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRA--------- 325
+ +DAE+L + ++ E+R L + KE KE ++ R
Sbjct: 588 TGKDAEILNLRKQLEKEIS--HTEDRNRLLHENTQKELEAHKETHTETVRVLEAEIDQFK 645
Query: 326 ---EVSLEF---RDRKISDLEKALGKLRQEAQEESEAKVDLVA---SKDQTLSSMETK-- 374
E E+ + KI +LE L E ++ +L A KD L +E+K
Sbjct: 646 SAFENEQEYGKEKSAKIRELEAQNKTLLSEMEKVKHVAENLEAFTSDKDNLLEELESKNK 705
Query: 375 -LEHLRAELAKKDEALSMMRTQ 395
+EHL+ E+A+ +E +S T+
Sbjct: 706 NIEHLKQEIAQLNEKISTKETE 727
>MYHA_RAT (Q9JLT0) Myosin heavy chain, nonmuscle type B (Cellular
myosin heavy chain, type B) (Nonmuscle myosin heavy
chain-B) (NMMHC-B)
Length = 1976
Score = 43.5 bits (101), Expect = 0.001
Identities = 30/114 (26%), Positives = 60/114 (52%), Gaps = 10/114 (8%)
Query: 282 LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK---ISD 338
+ EV L N+ F++E+K ++ + + +LAE+ +A+ + R+++ ISD
Sbjct: 983 MEEEVLLLEDQNSKFIKEKKL----MEDRIAECSSQLAEEEEKAKNLAKIRNKQEVMISD 1038
Query: 339 LEKALGKLRQEAQEESEAKVDL---VASKDQTLSSMETKLEHLRAELAKKDEAL 389
LE+ L K + QE +AK L ++ ++ +++ L+ +L KK+E L
Sbjct: 1039 LEERLKKEEKTRQELEKAKRKLDGETTDLQDQIAELQAQVDELKVQLTKKEEEL 1092
Score = 38.1 bits (87), Expect = 0.040
Identities = 32/120 (26%), Positives = 58/120 (47%), Gaps = 8/120 (6%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ E R + ++R L A +ERK + +K+ +L + ++ E + + RD
Sbjct: 1583 RDEQNEEKKRLLLKQVRELEAELEDERKQRALAVASKK-KMEIDLKDLEAQIEAANKARD 1641
Query: 334 RKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLS---SMETKLEHLRAELAKKDEALS 390
I L K +++ +E EA+ AS+D+ + E KL+ L AE+ + E L+
Sbjct: 1642 EVIKQLRKLQAQMKDYQRELEEAR----ASRDEIFAQSKESEKKLKSLEAEILQLQEELA 1697
Score = 38.1 bits (87), Expect = 0.040
Identities = 38/140 (27%), Positives = 62/140 (44%), Gaps = 27/140 (19%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ + L ++ R + + +++K F QL A+E + AE+ RAE ++
Sbjct: 1424 RLQQELDDLTVDLDHQRQIVSNLEKKQKKF-DQLLAEEKGISARYAEERDRAEAEAREKE 1482
Query: 334 RKISDLEKALGKLRQEAQEESEAK--------VDLVASKD-----------------QTL 368
K L +AL + EA+EE E + DL++SKD Q +
Sbjct: 1483 TKALSLARALEE-ALEAKEEFERQNKQLRADMEDLMSSKDDVGKNVHELEKSKRALEQQV 1541
Query: 369 SSMETKLEHLRAELAKKDEA 388
M T+LE L EL ++A
Sbjct: 1542 EEMRTQLEELEDELQATEDA 1561
Score = 34.7 bits (78), Expect = 0.44
Identities = 29/111 (26%), Positives = 55/111 (49%), Gaps = 6/111 (5%)
Query: 285 EVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQ---SSRAEVSLEFRDRKISDLEK 341
EVA L+ + +A + ++ + + +EL+EQ + R + +LE + + K
Sbjct: 1175 EVAELKKALEDETKNHEAQIQDMRQRHATALEELSEQLEQAKRFKANLEKNKQGLETDNK 1234
Query: 342 ALG---KLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEAL 389
L K+ Q+ + ESE K + ++ Q L + ++ + LR ELA+K L
Sbjct: 1235 ELACEVKVLQQVKAESEHKRKKLDAQVQELHAKVSEGDRLRVELAEKANKL 1285
>MYHA_BOVIN (Q27991) Myosin heavy chain, nonmuscle type B (Cellular
myosin heavy chain, type B) (Nonmuscle myosin heavy
chain-B) (NMMHC-B)
Length = 1976
Score = 43.5 bits (101), Expect = 0.001
Identities = 29/114 (25%), Positives = 61/114 (53%), Gaps = 10/114 (8%)
Query: 282 LRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK---ISD 338
+ E+ L N+ F++E+K ++ + + +LAE+ +A+ + R+++ ISD
Sbjct: 983 MEEEILLLEDQNSKFIKEKKL----MEDRIAECSSQLAEEEEKAKNLAKIRNKQEVMISD 1038
Query: 339 LEKALGKLRQEAQEESEAKVDL---VASKDQTLSSMETKLEHLRAELAKKDEAL 389
LE+ L K + QE +AK L ++ ++ +++ L+ ++AKK+E L
Sbjct: 1039 LEERLKKEEKTRQELEKAKRKLDGETTDLQDQIAELQAQIDELKIQVAKKEEEL 1092
Score = 39.3 bits (90), Expect = 0.018
Identities = 38/140 (27%), Positives = 64/140 (45%), Gaps = 27/140 (19%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ + L ++ R + + +++K F QL A+E + + AE+ RAE ++
Sbjct: 1424 RLQQELDDLLVDLDHQRQIVSNLEKKQKKF-DQLLAEEKNISARYAEERDRAEAEAREKE 1482
Query: 334 RKISDLEKALGKLRQEAQEESEAK--------VDLVASKD-----------------QTL 368
K L +AL + EA+EE+E + DL++SKD Q +
Sbjct: 1483 TKALSLARALEE-ALEAREEAERQNKQLRADMEDLMSSKDDVGKNVHELEKSKRALEQQV 1541
Query: 369 SSMETKLEHLRAELAKKDEA 388
M T+LE L EL ++A
Sbjct: 1542 EEMRTQLEELEDELQATEDA 1561
Score = 38.9 bits (89), Expect = 0.023
Identities = 32/120 (26%), Positives = 58/120 (47%), Gaps = 8/120 (6%)
Query: 274 RSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRD 333
R Q+ E R + ++R L A +ERK + +K+ +L + ++ E + + RD
Sbjct: 1583 RDEQNEEKKRLLIKQVRELEAELEDERKQRALAVASKK-KMEIDLKDLEAQIEAANKARD 1641
Query: 334 RKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLS---SMETKLEHLRAELAKKDEALS 390
I L K +++ +E EA+ AS+D+ + E KL+ L AE+ + E L+
Sbjct: 1642 EVIKQLRKLQAQMKDYQRELEEAR----ASRDEIFAQSKESEKKLKSLEAEILQLQEELA 1697
Score = 34.3 bits (77), Expect = 0.57
Identities = 29/111 (26%), Positives = 55/111 (49%), Gaps = 6/111 (5%)
Query: 285 EVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQ---SSRAEVSLEFRDRKISDLEK 341
EVA L+ + +A + ++ + + +EL+EQ + R + +LE + + K
Sbjct: 1175 EVAELKKALEEETKSHEAQIQDMRQRHATALEELSEQLEQAKRFKANLEKNKQGLETDNK 1234
Query: 342 ALG---KLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEAL 389
L K+ Q+ + ESE K + ++ Q L + ++ + LR ELA+K L
Sbjct: 1235 ELACEVKVLQQVKAESEHKRKKLDAQVQELHAKVSEGDRLRVELAEKANKL 1285
>LAM1_CHICK (P14731) Lamin B1
Length = 584
Score = 42.7 bits (99), Expect = 0.002
Identities = 32/88 (36%), Positives = 49/88 (55%), Gaps = 2/88 (2%)
Query: 273 KRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFR 332
K A+ +VL S + LNA V+ R+ F A L AKE + A L ++ S+ E + R
Sbjct: 108 KLRAEHEQVLSSYAKKDSDLNAAQVKLRE-FEAALNAKEAALATALGDKRSQEEELEDLR 166
Query: 333 DRKISDLEKALGKLRQEAQEESEAKVDL 360
D +I+ LE +L ++E +E+ KVDL
Sbjct: 167 D-QIAQLEVSLAAAKKELADETLQKVDL 193
Score = 32.0 bits (71), Expect = 2.8
Identities = 30/123 (24%), Positives = 56/123 (45%), Gaps = 1/123 (0%)
Query: 273 KRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFR 332
+R ++ +V E++ L+ L + + + L + EL + + E L
Sbjct: 61 RRVSEREQVCGREISGLKELFETELADARKTLDDTARERAKLQIELGKLRAEHEQVLSSY 120
Query: 333 DRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMM 392
+K SDL A KLR E + AK +A+ S E +LE LR ++A+ + +L+
Sbjct: 121 AKKDSDLNAAQVKLR-EFEAALNAKEAALATALGDKRSQEEELEDLRDQIAQLEVSLAAA 179
Query: 393 RTQ 395
+ +
Sbjct: 180 KKE 182
>MOES_PIG (P26042) Moesin (Membrane-organizing extension spike
protein)
Length = 576
Score = 42.4 bits (98), Expect = 0.002
Identities = 37/96 (38%), Positives = 49/96 (50%), Gaps = 5/96 (5%)
Query: 298 EERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAK 357
EE L Q++ + +EL EQ+ RA + R R S+ EK L K RQEA+E EA
Sbjct: 344 EELMERLKQIEEQTKKAQQELEEQTRRALALEQERKRAQSEAEK-LAKERQEAEEAKEAL 402
Query: 358 VDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMR 393
L AS+DQ + + LE AEL + L M R
Sbjct: 403 --LKASRDQKKTQEQLALE--MAELTARISQLEMAR 434
>RIP1_RAT (Q9JIR0) Peripheral-type benzodiazepine
receptor-associated protein 1 (PRAX-1) (RIM binding
protein 1) (RIM-BP1)
Length = 1847
Score = 41.6 bits (96), Expect = 0.004
Identities = 49/221 (22%), Positives = 89/221 (40%), Gaps = 29/221 (13%)
Query: 192 PKPNQPGEPSLATPVSSPRAQDDSVDK----PPLSAKGFANREPPYSLSPTPAQVARMDE 247
P P PGE L V SP+ +K L ++ R+ SL + R E
Sbjct: 304 PAPGAPGETVLEDDVESPQVVLGEPEKQLRVQQLESELCKKRKKCESLEQEARKKQRRCE 363
Query: 248 VVKQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQL 307
++ Q R ++ A A + E + E A L+ G +ER + L +
Sbjct: 364 ELELQ--LRAAQNENARLVEENSRLSGKATEKEQVEWENAELKGQLLGVTQERDSALRKS 421
Query: 308 KAKETSF----------------AKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQ 351
+ ++ ++L E+ +A +SL+ + ++ L++A + EA+
Sbjct: 422 QGLQSKLESLEQVLEHMRKVAQRRQQLEEEHEQARLSLQEKQEEVRRLQQA----QAEAK 477
Query: 352 EESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMM 392
E E V L+ S TL SM+ ++ L + + E S++
Sbjct: 478 REHEGAVQLLES---TLDSMQARVRELEGQCRSQTERFSLL 515
>MOES_HUMAN (P26038) Moesin (Membrane-organizing extension spike
protein)
Length = 576
Score = 41.6 bits (96), Expect = 0.004
Identities = 37/96 (38%), Positives = 49/96 (50%), Gaps = 5/96 (5%)
Query: 298 EERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAK 357
EE L Q++ + +EL EQ+ RA + R R S+ EK L K RQEA+E EA
Sbjct: 344 EELMERLKQIEEQTKKAQQELEEQTRRALELEQERKRAQSEAEK-LAKERQEAEEAKEAL 402
Query: 358 VDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMR 393
L AS+DQ + + LE AEL + L M R
Sbjct: 403 --LQASRDQKKTQEQLALE--MAELTARISQLEMAR 434
>RIP1_MOUSE (Q7TNF8) Peripheral-type benzodiazepine
receptor-associated protein 1 (PRAX-1) (Peripheral
benzodiazepine receptor interacting protein) (PBR-IP)
(RIM binding protein 1) (RIM-BP1)
Length = 1846
Score = 40.4 bits (93), Expect = 0.008
Identities = 53/237 (22%), Positives = 94/237 (39%), Gaps = 26/237 (10%)
Query: 177 RAPVISAAFCGPEE-----TPKPNQPGEPSLATPVSSP----RAQDDSVDKPPLSAKGFA 227
R P+ A GP P P PGE L V SP R + L ++
Sbjct: 283 REPLRPARSPGPTAPSRVGAPAPGAPGEAVLQDDVESPQVVLREPEKQQRVQQLESELCK 342
Query: 228 NREPPYSLSPTPAQVARMDEVVKQQGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVA 287
R+ SL + R E ++ Q R ++ A A + E + E +
Sbjct: 343 KRKKCESLEQEARKKQRRCEELELQ--LRAAQNENARLVEENSRLSGRATEKEQVEWENS 400
Query: 288 RLRALNAGFVEERKAFLAQLKAKETSF---------AKELAEQSSRAEVSLEFRDRKISD 338
L+ G +ER + L + + ++ +E+A++ + EV E + +
Sbjct: 401 ELKGQLLGVTQERDSALLKSQGLQSKLESLEQVLKHMREVAQRRQQLEVEHEQARLSLQE 460
Query: 339 LEKALGKLRQ---EAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALSMM 392
++ + +L+Q EA+ E E V L+ S TL SM+ ++ L + + E S++
Sbjct: 461 KQEEVRRLQQAQAEAKREHEGAVQLLES---TLDSMQARVRELEGQCRSQTERFSLL 514
>MOES_MOUSE (P26041) Moesin (Membrane-organizing extension spike
protein)
Length = 576
Score = 40.4 bits (93), Expect = 0.008
Identities = 29/99 (29%), Positives = 50/99 (50%), Gaps = 1/99 (1%)
Query: 298 EERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAK 357
EE L Q++ + +EL EQ+ RA + R R S+ EK L K RQEA+E EA
Sbjct: 344 EELMEKLKQIEEQTKKAQQELEEQTRRALELEQERKRAQSEAEK-LAKERQEAEEAKEAL 402
Query: 358 VDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQA 396
+ + +T + +++ L A +++ + A ++A
Sbjct: 403 LQASRDQKKTQEQLASEMAELTARISQLEMARKKKESEA 441
>MOES_RAT (O35763) Moesin (Membrane-organizing extension spike
protein)
Length = 576
Score = 40.0 bits (92), Expect = 0.010
Identities = 29/99 (29%), Positives = 50/99 (50%), Gaps = 1/99 (1%)
Query: 298 EERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAK 357
EE L Q++ + +EL EQ+ RA + R R S+ EK L K RQEA+E EA
Sbjct: 344 EELMEKLKQIEEQTKKAQQELEEQTRRALELEQERKRAQSEAEK-LAKERQEAEEAKEAL 402
Query: 358 VDLVASKDQTLSSMETKLEHLRAELAKKDEALSMMRTQA 396
+ + +T + +++ L A +++ + A ++A
Sbjct: 403 LQASRDQKKTQEQLASEMAELTARVSQLEMARKKKESEA 441
>TPM4_RAT (P09495) Tropomyosin alpha 4 chain (Tropomyosin 4) (TM-4)
Length = 248
Score = 39.7 bits (91), Expect = 0.014
Identities = 30/102 (29%), Positives = 50/102 (48%), Gaps = 15/102 (14%)
Query: 283 RSEVARLRALNAGFVEERKAFLAQLKA----------KETSFAKE---LAEQSSRAEVSL 329
R+EV+ L++ + EE K LK+ KE + +E L+++ AE
Sbjct: 146 RAEVSELKS--SDLEEELKNVTNNLKSLEAASEKYSEKEDKYEEEIKLLSDKLKEAETRA 203
Query: 330 EFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSM 371
EF +R +S LEK + L ++ + E V L + DQTL+ +
Sbjct: 204 EFAERTVSKLEKTIDDLEEKLAQAKEENVGLHQTLDQTLNEL 245
>MYHB_MOUSE (O08638) Myosin heavy chain, smooth muscle isoform (SMMHC)
Length = 1972
Score = 39.7 bits (91), Expect = 0.014
Identities = 36/125 (28%), Positives = 58/125 (45%), Gaps = 15/125 (12%)
Query: 276 AQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRK 335
+++ ++L V+ L N EE+ L +LK+K S EL EV L+ ++
Sbjct: 998 SKERKLLEERVSDLTT-NLAEEEEKAKNLTKLKSKHESMISEL-------EVRLKKEEKS 1049
Query: 336 ISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRA-------ELAKKDEA 388
+LEK KL +A + E DL A + + K E L+A E+A+K+ A
Sbjct: 1050 RQELEKLKRKLEGDASDFHEQIADLQAQIAELKMQLAKKEEELQAALARLDEEIAQKNNA 1109
Query: 389 LSMMR 393
L +R
Sbjct: 1110 LKKIR 1114
Score = 31.2 bits (69), Expect = 4.8
Identities = 28/119 (23%), Positives = 57/119 (47%), Gaps = 29/119 (24%)
Query: 297 VEERKAFLAQLKAKETSFAKELAEQSS----------RAEVSLEFRDRKISDLEKALGKL 346
++ ++ ++ L+ K+ F + LAE+ + RAE ++ K L +AL +
Sbjct: 1436 LDNQRQLVSNLEKKQKKFDQLLAEEKNISSKYADERDRAEAEAREKETKALSLARALEE- 1494
Query: 347 RQEAQEESE--------AKVDLVASKD----------QTLSSMETKLEHLRAELAKKDE 387
EA+EE E DLV+SKD ++ ++ET++E ++ +L + ++
Sbjct: 1495 ALEAKEELERTNKMLKAEMEDLVSSKDDVGKNVHELEKSKRALETQMEEMKTQLEESED 1553
Score = 30.4 bits (67), Expect = 8.2
Identities = 33/120 (27%), Positives = 53/120 (43%), Gaps = 9/120 (7%)
Query: 275 SAQDAEVLRSEVARLRALNAGFVEERKAFLAQ---LKAKETSFAKELAEQSSRAEVSLEF 331
+A E+ + L EE ++ AQ ++ K T +EL EQ + + +
Sbjct: 1162 TATQQELRAKREQEVTVLKKALDEETRSHEAQVQEMRQKHTQAVEELTEQLEQFKRAKAN 1221
Query: 332 RDRKISDLEK----ALGKLR--QEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKK 385
D+ LEK G+LR +A++E E K + + Q L S + E RAEL+ K
Sbjct: 1222 LDKSKQTLEKENADLAGELRVLGQAKQEVEHKKKKLEVQLQDLQSKCSDGERARAELSDK 1281
>DYNA_MOUSE (O08788) Dynactin 1 (150 kDa dynein-associated
polypeptide) (DP-150) (DAP-150) (p150-glued)
Length = 1281
Score = 39.7 bits (91), Expect = 0.014
Identities = 61/299 (20%), Positives = 116/299 (38%), Gaps = 51/299 (17%)
Query: 102 LYWTQEVRQVIDPPEDSLTPE--DKTVISFLARLPVLDCARVIEKIPQVSEGLLTVFEKK 159
++ Q QV + D+ +PE D + L R D A K+ + K
Sbjct: 87 IFVRQSQIQVFEDGADTTSPETPDSSASKVLKREGA-DAAAKTSKLRGLKPKKAPTARKT 145
Query: 160 GNVSPHP-------VAATSSSDGDRAPVISAAFCGPEETPKPNQPGEPSLATPVSSPRAQ 212
P P VA SSS G +A G + +P+ P + LA P+ A
Sbjct: 146 TTRRPKPTRPASTGVAGPSSSLGPSG----SASAGELSSSEPSTPAQTPLAAPIIPTPAL 201
Query: 213 DDSVDKPPLSAKGFANREPPYSLSPTPAQVARMDEVVKQQGLDRVSEGAFATAFHAFHHY 272
PPL + P AQV ++E ++ L
Sbjct: 202 TSPGAAPPLPS-------PSKEEEGLRAQVRDLEEKLETLRL------------------ 236
Query: 273 KRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFR 332
KRS A++ E +++ +E+ + + ++++ ++ + L E A+ +LE +
Sbjct: 237 KRSEDKAKLKELEKHKIQ------LEQVQEWKSKMQEQQADLQRRLKEARKEAKEALEAK 290
Query: 333 DRKISDLEKALGKL------RQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKK 385
+R + ++ + ++ A+E +E+ V + + + + T LE L+AE+ +K
Sbjct: 291 ERYMEEMADTADAIEMATLDKEMAEERAESLQQEVEALKERVDELTTDLEILKAEIEEK 349
Score = 36.2 bits (82), Expect = 0.15
Identities = 50/207 (24%), Positives = 91/207 (43%), Gaps = 37/207 (17%)
Query: 192 PKPNQPGEPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVARMDEVVKQ 251
PKP +P +A P SS + SA ++ EP TPAQ ++
Sbjct: 150 PKPTRPASTGVAGPSSSLGPSGSA------SAGELSSSEPS-----TPAQTPLAAPIIPT 198
Query: 252 QGLDRVSEGAFATAFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKE 311
L T+ A +++ E LR++V L EE+ L ++++
Sbjct: 199 PAL---------TSPGAAPPLPSPSKEEEGLRAQVRDL--------EEKLETLRLKRSED 241
Query: 312 TSFAKELAEQSSRAEVSLEFRDR---KISDLEKALGKLRQEAQEESEAKVDLVASKDQT- 367
+ KEL + + E E++ + + +DL++ L + R+EA+E EAK + T
Sbjct: 242 KAKLKELEKHKIQLEQVQEWKSKMQEQQADLQRRLKEARKEAKEALEAKERYMEEMADTA 301
Query: 368 ----LSSMETKLEHLRAE-LAKKDEAL 389
+++++ ++ RAE L ++ EAL
Sbjct: 302 DAIEMATLDKEMAEERAESLQQEVEAL 328
Score = 33.1 bits (74), Expect = 1.3
Identities = 23/112 (20%), Positives = 51/112 (45%), Gaps = 1/112 (0%)
Query: 279 AEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISD 338
A LR+E+ L +E+R+ + +LK +EL+E + R + + D D
Sbjct: 928 AAALRAEITDAEGLGLK-LEDRETVIKELKKSLKIKGEELSEANVRXSLLEKKLDSAAKD 986
Query: 339 LEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEALS 390
++ + K++ E ++T+ +++ ++ L AE A+ + L+
Sbjct: 987 ADERIEKVQTRLDETQTLLRKKEKDFEETMDALQADIDQLEAEKAELKQRLN 1038
>C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 190)
(Microtubule binding protein 190) (d-CLIP-190)
Length = 1690
Score = 39.7 bits (91), Expect = 0.014
Identities = 35/135 (25%), Positives = 62/135 (45%), Gaps = 3/135 (2%)
Query: 259 EGAFATAFHAFHHYKRSAQDAEVLRSEVARLR-ALNAGFVEERKAFLAQLKAKETSFAKE 317
E A A Y S +A L+ +V + L+A ER + A L K + F+ E
Sbjct: 941 EAANAALEKVNKEYAESRAEASDLQDKVKEITDTLHAELQAERSSSSA-LHTKLSKFSDE 999
Query: 318 LAEQSSRAEVSLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEH 377
+A + +++ EK L +LRQ+ Q+ +++ L A ++ S E +++
Sbjct: 1000 IATGHKELTSKADAWSQEMLQKEKELQELRQQLQDSQDSQTKLKAEGERKEKSFEESIKN 1059
Query: 378 LRAELAK-KDEALSM 391
L+ E+ K K E L +
Sbjct: 1060 LQEEVTKAKTENLEL 1074
Score = 33.1 bits (74), Expect = 1.3
Identities = 19/84 (22%), Positives = 40/84 (47%), Gaps = 1/84 (1%)
Query: 308 KAKETSFAKELAEQSSRAEVSLEFR-DRKISDLEKALGKLRQEAQEESEAKVDLVASKDQ 366
K KE ++ + S ++ L+ +RK E+++ L++E + ++L
Sbjct: 1021 KEKELQELRQQLQDSQDSQTKLKAEGERKEKSFEESIKNLQEEVTKAKTENLELSTGTQT 1080
Query: 367 TLSSMETKLEHLRAELAKKDEALS 390
T+ ++ +LE AEL K++ S
Sbjct: 1081 TIKDLQERLEITNAELQHKEKMAS 1104
>HIPR_MOUSE (Q9JKY5) Huntingtin interacting protein 1 related
(Hip1-related)
Length = 1068
Score = 39.3 bits (90), Expect = 0.018
Identities = 36/123 (29%), Positives = 57/123 (46%), Gaps = 7/123 (5%)
Query: 280 EVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFRDRKISDL 339
E L+ EV LRA E + +++QLK + EL EQ + + +L ++ +L
Sbjct: 357 ENLKREVETLRAELEKIKMEAQRYISQLKGQVNGLEAELEEQRKQKQKALVDNEQLRHEL 416
Query: 340 E--KAL---GKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKK--DEALSMM 392
KAL G Q +EE+E K ++ L ++L + AEL +K D A +
Sbjct: 417 AQLKALQLEGARNQGLREEAERKASATEARYSKLKEKHSELINTHAELLRKNADTAKQLT 476
Query: 393 RTQ 395
TQ
Sbjct: 477 VTQ 479
Score = 35.4 bits (80), Expect = 0.26
Identities = 57/242 (23%), Positives = 92/242 (37%), Gaps = 50/242 (20%)
Query: 188 PEETPKPNQPG---EPSLATPVSSPRAQDDSVDKPPLSAKGFANREPPYSLSPTPAQVAR 244
PEE P+ +P E S A P P D D+ G + + +V
Sbjct: 306 PEEAPEEEEPENLIEISSAPPAGEPVVVADLFDQTFGPPNGSMKDDRDLQIENLKREVET 365
Query: 245 MD---EVVKQQGLDRVSE------GAFATAFHAFHHYKRSAQDAEVLRSEVARLRAL--- 292
+ E +K + +S+ G A +++ D E LR E+A+L+AL
Sbjct: 366 LRAELEKIKMEAQRYISQLKGQVNGLEAELEEQRKQKQKALVDNEQLRHELAQLKALQLE 425
Query: 293 ---NAGFVEE--RKAFL-------------------AQLKAKETSFAKELAEQSSRAEVS 328
N G EE RKA A+L K AK+L E
Sbjct: 426 GARNQGLREEAERKASATEARYSKLKEKHSELINTHAELLRKNADTAKQLTVTQQSQE-- 483
Query: 329 LEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLEHLRAELAKKDEA 388
+++ +++ L ++A+ ESE K++ + L ++ +L ELA+ EA
Sbjct: 484 ------EVARVKEQLAFQMEQAKRESEMKME---EQSDQLEKLKRELAARAGELARAQEA 534
Query: 389 LS 390
LS
Sbjct: 535 LS 536
>GOA4_MOUSE (Q91VW5) Golgi autoantigen, golgin subfamily A member 4
(tGolgin-1)
Length = 2238
Score = 39.3 bits (90), Expect = 0.018
Identities = 35/136 (25%), Positives = 66/136 (47%), Gaps = 9/136 (6%)
Query: 265 AFHAFHHYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSR 324
A H + ++++ E+ R AR R L E+ + L + +++ +E +Q S
Sbjct: 500 ALQKLHAEELASKEQELSRRLEARERELQ----EQMRIALEKSRSEYLKLTQEKEQQESL 555
Query: 325 AEVSLEFRDRKI-SDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSMETKLE----HLR 379
A LE + + I ++ E L +L QEA+ ++L S +++L +T+ E HL
Sbjct: 556 ALEELELQKKAILTESENKLQELGQEAEAYRTRILELETSLEKSLQESKTQSEHLAVHLE 615
Query: 380 AELAKKDEALSMMRTQ 395
AE K ++ L+ + Q
Sbjct: 616 AEKNKHNKELTALAEQ 631
Score = 36.6 bits (83), Expect = 0.11
Identities = 35/118 (29%), Positives = 57/118 (47%), Gaps = 11/118 (9%)
Query: 273 KRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLKAKETSFAKELAEQSSRAEVSLEFR 332
+ S ++ L+ E++R+R A ++ Q+ A + A+ELA + LE R
Sbjct: 466 RASEEERLRLQHELSRVRQEAASMAKKNSE--EQVAALQKLHAEELASKEQELSRRLEAR 523
Query: 333 DRKISD-----LEKALG---KLRQE-AQEESEAKVDLVASKDQTLSSMETKLEHLRAE 381
+R++ + LEK+ KL QE Q+ES A +L K L+ E KL+ L E
Sbjct: 524 ERELQEQMRIALEKSRSEYLKLTQEKEQQESLALEELELQKKAILTESENKLQELGQE 581
Score = 34.7 bits (78), Expect = 0.44
Identities = 33/133 (24%), Positives = 64/133 (47%), Gaps = 22/133 (16%)
Query: 271 HYKRSAQDAEVLRSEVARLRALNAGFVEERKAFLAQLK---AKETSFAKELAEQSSRAEV 327
H K ++ + R++V +L + +EE+ LAQL+ K ++ KE A QS
Sbjct: 868 HKKHVCEELDAQRAQVQQLERQRSE-LEEKVRSLAQLQDSQLKNSTVEKEQARQS----- 921
Query: 328 SLEFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSM----ETKLEHLRAELA 383
+ + E + ++R+E +E E ++SK++++S + ETK ++ +
Sbjct: 922 --------LMEKENIILQMREEQAKEIEILKQTLSSKEESISILHEEYETKFKNQEKRME 973
Query: 384 K-KDEALSMMRTQ 395
K K +A M T+
Sbjct: 974 KIKQKAKEMQETK 986
>TPM4_MOUSE (Q6IRU2) Tropomyosin alpha 4 chain (Tropomyosin 4)
Length = 248
Score = 38.9 bits (89), Expect = 0.023
Identities = 30/102 (29%), Positives = 48/102 (46%), Gaps = 15/102 (14%)
Query: 283 RSEVARLRALNAGFVEERKAFLAQLKA----------KETSFAKE---LAEQSSRAEVSL 329
R+EV+ L+ EE K LK+ KE + +E L+++ AE
Sbjct: 146 RAEVSELKC--GDLEEELKNVTNNLKSLEAASEKYSEKEDKYEEEIKLLSDKLKEAETRA 203
Query: 330 EFRDRKISDLEKALGKLRQEAQEESEAKVDLVASKDQTLSSM 371
EF +R +S LEK + L ++ + E V L + DQTL+ +
Sbjct: 204 EFAERTVSKLEKTIDDLEEKLAQAKEENVGLHQTLDQTLNEL 245
>YTGF_LACLA (Q9CEE2) Hypothetical UPF0144 protein ytgF
Length = 531
Score = 38.5 bits (88), Expect = 0.030
Identities = 41/148 (27%), Positives = 65/148 (43%), Gaps = 18/148 (12%)
Query: 263 ATAFHAFHHYKRSAQDAEVLRSEV---ARLRALNAGFVEERKAFLAQLKAKETSFAKELA 319
A F+A + K++ QDAE L +E A NA E A+ KE +
Sbjct: 18 AFIFYA-QNIKKNKQDAESLFNEAENKANEVMANAKREAESLKMEAEAFKKEARYTLREE 76
Query: 320 EQSSRAEVSLEFRDRK--ISDLEKALGKLRQEA-----------QEESEAKVDLVASKDQ 366
EQ R E+ EF+ + + + EK L K R+E +E ++K + + K
Sbjct: 77 EQKQRREIEDEFKQERQELKETEKRL-KQREEILDRKDDTLTKKEENLDSKEENLVRKTD 135
Query: 367 TLSSMETKLEHLRAELAKKDEALSMMRT 394
TLS E +L H+ + + E +S + T
Sbjct: 136 TLSKREEQLAHIEEKKRLELERISNLST 163
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.130 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,962,525
Number of Sequences: 164201
Number of extensions: 1959171
Number of successful extensions: 9748
Number of sequences better than 10.0: 416
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 365
Number of HSP's that attempted gapping in prelim test: 8969
Number of HSP's gapped (non-prelim): 1070
length of query: 396
length of database: 59,974,054
effective HSP length: 112
effective length of query: 284
effective length of database: 41,583,542
effective search space: 11809725928
effective search space used: 11809725928
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0005.13


