

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0004a.3
(148 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
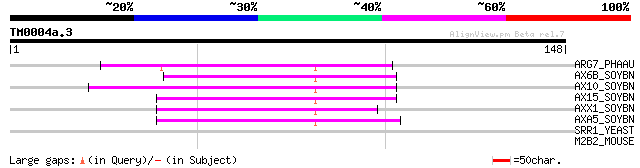
ARG7_PHAAU (P32295) Indole-3-acetic acid induced protein ARG7 60 2e-09
AX6B_SOYBN (P33083) Auxin-induced protein 6B 58 9e-09
AX10_SOYBN (P33080) Auxin-induced protein X10A 57 1e-08
AX15_SOYBN (P33081) Auxin-induced protein 15A 55 6e-08
AXX1_SOYBN (P33082) Auxin-induced protein X15 54 9e-08
AXA5_SOYBN (P33079) Auxin-induced protein 10A5 53 2e-07
SRR1_YEAST (Q06688) SRR1-like protein 28 5.5
M2B2_MOUSE (O54782) Epididymis-specific alpha-mannosidase precur... 28 7.2
>ARG7_PHAAU (P32295) Indole-3-acetic acid induced protein ARG7
Length = 92
Score = 59.7 bits (143), Expect = 2e-09
Identities = 36/80 (45%), Positives = 41/80 (51%), Gaps = 2/80 (2%)
Query: 25 KQISLRRHDSSDPRP-PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RG 82
K +S R SS P G L VYVG +RF IP LN P+F LL + EEEFG
Sbjct: 10 KTLSARNEASSKVLDAPKGYLAVYVGENMKRFVIPVSHLNQPLFQDLLSQAEEEFGYDHP 69
Query: 83 SGGLILPCDVCFFTHIVKRL 102
GGL +PC F HI L
Sbjct: 70 MGGLTIPCSEDLFQHITSCL 89
>AX6B_SOYBN (P33083) Auxin-induced protein 6B
Length = 90
Score = 57.8 bits (138), Expect = 9e-09
Identities = 30/63 (47%), Positives = 37/63 (58%), Gaps = 1/63 (1%)
Query: 42 GCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHIVK 100
G L VYVG + +RF IP +LN P F LL + EEEFG +GGL +PC F HI
Sbjct: 28 GYLAVYVGEKMRRFVIPVSYLNKPSFQDLLSQAEEEFGYHHPNGGLTIPCSEDVFQHITS 87
Query: 101 RLH 103
L+
Sbjct: 88 FLN 90
>AX10_SOYBN (P33080) Auxin-induced protein X10A
Length = 92
Score = 57.4 bits (137), Expect = 1e-08
Identities = 32/83 (38%), Positives = 43/83 (51%), Gaps = 1/83 (1%)
Query: 22 MRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL- 80
+R I+ + S P G L VYVG + +RF IP +LN P F LL++ EEEFG
Sbjct: 8 IRKTSIAANQASSKSVEVPKGYLVVYVGDKMRRFLIPVSYLNQPSFQDLLNQAEEEFGYD 67
Query: 81 RGSGGLILPCDVCFFTHIVKRLH 103
GGL +PC F + L+
Sbjct: 68 HPMGGLTIPCKEDEFLTVTSHLN 90
>AX15_SOYBN (P33081) Auxin-induced protein 15A
Length = 82
Score = 55.1 bits (131), Expect = 6e-08
Identities = 30/65 (46%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG + +RF IP +LN P F LL + EEEFG GGL +PC F I
Sbjct: 18 PKGYLAVYVGEKLKRFVIPVSYLNQPSFQDLLSQAEEEFGYDHPMGGLTIPCSEDVFQCI 77
Query: 99 VKRLH 103
L+
Sbjct: 78 TSCLN 82
>AXX1_SOYBN (P33082) Auxin-induced protein X15
Length = 82
Score = 54.3 bits (129), Expect = 9e-08
Identities = 28/60 (46%), Positives = 34/60 (56%), Gaps = 1/60 (1%)
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG + +RF IP ++N P F LL + EEEFG GGL +PC F I
Sbjct: 18 PKGYLAVYVGEKMKRFVIPVSYMNQPSFQDLLTQAEEEFGYDHPMGGLTIPCSEEVFQRI 77
>AXA5_SOYBN (P33079) Auxin-induced protein 10A5
Length = 93
Score = 53.1 bits (126), Expect = 2e-07
Identities = 28/66 (42%), Positives = 37/66 (55%), Gaps = 1/66 (1%)
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G VYVG + +RF IP +LN P F LL + EEEFG GGL +PC F ++
Sbjct: 27 PKGYAAVYVGDKMRRFTIPVSYLNEPSFQELLSQAEEEFGYDHPMGGLTIPCKEEEFLNV 86
Query: 99 VKRLHK 104
L++
Sbjct: 87 TAHLNE 92
>SRR1_YEAST (Q06688) SRR1-like protein
Length = 274
Score = 28.5 bits (62), Expect = 5.5
Identities = 15/39 (38%), Positives = 18/39 (45%)
Query: 94 FFTHIVKRLHKNENKYGKLTLEQFVITFSDAAANFDSCK 132
F T I KR KN N KL + I + A F SC+
Sbjct: 206 FATFIPKRKRKNRNNSSKLKVTPSDIDYDSIAPKFKSCQ 244
>M2B2_MOUSE (O54782) Epididymis-specific alpha-mannosidase precursor
(EC 3.2.1.24) (Mannosidase alpha class 2B member 2)
Length = 1018
Score = 28.1 bits (61), Expect = 7.2
Identities = 15/43 (34%), Positives = 21/43 (47%), Gaps = 4/43 (9%)
Query: 19 QVMMRW----KQISLRRHDSSDPRPPSGCLYVYVGHERQRFAI 57
Q++ RW K L+ H +S PRPP G + E + F I
Sbjct: 972 QMLQRWHWSTKTDHLKGHPTSPPRPPGGSIITVYPKEIRTFFI 1014
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.141 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,269,315
Number of Sequences: 164201
Number of extensions: 690882
Number of successful extensions: 1454
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 1445
Number of HSP's gapped (non-prelim): 8
length of query: 148
length of database: 59,974,054
effective HSP length: 100
effective length of query: 48
effective length of database: 43,553,954
effective search space: 2090589792
effective search space used: 2090589792
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0004a.3


