

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0002.6
(260 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
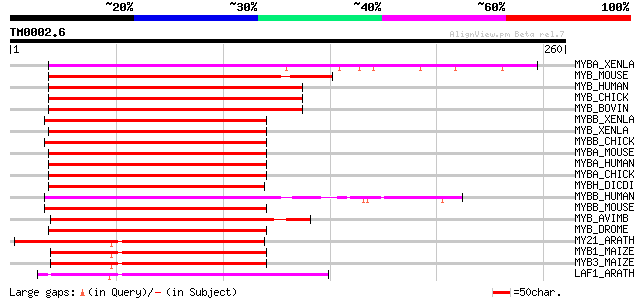
Score E
Sequences producing significant alignments: (bits) Value
MYBA_XENLA (Q05935) Myb-related protein A (A-Myb) (XAMYB) (MYB-r... 136 6e-32
MYB_MOUSE (P06876) Myb proto-oncogene protein (C-myb) 133 4e-31
MYB_HUMAN (P10242) Myb proto-oncogene protein (C-myb) 132 1e-30
MYB_CHICK (P01103) Myb proto-oncogene protein (C-myb) 132 1e-30
MYB_BOVIN (P46200) Myb proto-oncogene protein (C-myb) 132 1e-30
MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related p... 130 3e-30
MYB_XENLA (Q08759) Myb protein 128 2e-29
MYBB_CHICK (Q03237) Myb-related protein B (B-Myb) 128 2e-29
MYBA_MOUSE (P51960) Myb-related protein A (A-Myb) 127 2e-29
MYBA_HUMAN (P10243) Myb-related protein A (A-Myb) 127 2e-29
MYBA_CHICK (P52550) Myb-related protein A (A-Myb) 127 2e-29
MYBH_DICDI (P34127) Myb-like protein (Fragment) 126 5e-29
MYBB_HUMAN (P10244) Myb-related protein B (B-Myb) 126 5e-29
MYBB_MOUSE (P48972) Myb-related protein B (B-Myb) 126 6e-29
MYB_AVIMB (P01104) Transforming protein Myb 124 2e-28
MYB_DROME (P04197) Myb protein 124 3e-28
MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related prot... 111 2e-24
MYB1_MAIZE (P20024) Myb-related protein Zm1 107 2e-23
MYB3_MAIZE (P20025) Myb-related protein Zm38 105 8e-23
LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-re... 104 2e-22
>MYBA_XENLA (Q05935) Myb-related protein A (A-Myb) (XAMYB)
(MYB-related protein 2) (XMYB2)
Length = 728
Score = 136 bits (342), Expect = 6e-32
Identities = 87/256 (33%), Positives = 134/256 (51%), Gaps = 27/256 (10%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
VKG W+ +ED+ +++LV ++GP+ WS+I+ + GR GK CR RW N L+PDV+ +T
Sbjct: 85 VKGPWTKEEDQRVIELVHKYGPKKWSIIAKHLKGRIGKQCRERWHNHLNPDVKKSSWTEE 144
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLP----LASPSP 134
ED II AH GN+WA I++LLPGRTDN+IKNHWNST++R+ + L PS
Sbjct: 145 EDRIIYSAHKRMGNRWAEIAKLLPGRTDNSIKNHWNSTMKRKVEQEGYLQDLMNCDRPSK 204
Query: 135 SPSPSPPLPFSMAKRPHGY----ELLDRYDGL--------MDQGYPF-KRHCVEKDDEES 181
+ S P + + Y + RY L + + F ++ V+ DD E
Sbjct: 205 LQAKSCAAPNHLQAQNQFYIPVQTQIPRYSSLSHDDNCIEIQNSFSFIQQPFVDADDPEK 264
Query: 182 RDGVEEAETM---TEEEVLASSVPTSLSL-----FPMGEKKMVAVAEKEEKKMDMQ--EG 231
++E E + E EV VP+ SL + MGE ++ EE+ D +
Sbjct: 265 EKRIKELELLLMSAENEVRRKRVPSGSSLTWSESYHMGESMSNTMSHLEEQTHDFYSLDE 324
Query: 232 LMRTMMQRMIAEEVRA 247
+ T Q+ + +++ A
Sbjct: 325 IPNTSGQQSVEDKILA 340
Score = 31.6 bits (70), Expect = 2.0
Identities = 16/59 (27%), Positives = 29/59 (49%), Gaps = 1/59 (1%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPF 75
D K SW+ +ED + + G R W+ I+ +PGR+ S + W + + V+ +
Sbjct: 135 DVKKSSWTEEEDRIIYSAHKRMGNR-WAEIAKLLPGRTDNSIKNHWNSTMKRKVEQEGY 192
>MYB_MOUSE (P06876) Myb proto-oncogene protein (C-myb)
Length = 636
Score = 133 bits (335), Expect = 4e-31
Identities = 64/133 (48%), Positives = 88/133 (66%), Gaps = 4/133 (3%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WSVI+ + GR GK CR RW N L+P+V+ +T
Sbjct: 91 IKGPWTKEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEE 150
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPSP 138
ED II QAH GN+WA I++LLPGRTDNAIKNHWNST+RR+ + L PS +
Sbjct: 151 EDRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRKVEQEGYL----QEPSKAS 206
Query: 139 SPPLPFSMAKRPH 151
P+ S K H
Sbjct: 207 QTPVATSFQKNNH 219
>MYB_HUMAN (P10242) Myb proto-oncogene protein (C-myb)
Length = 640
Score = 132 bits (331), Expect = 1e-30
Identities = 60/119 (50%), Positives = 85/119 (71%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WSVI+ + GR GK CR RW N L+P+V+ +T
Sbjct: 91 IKGPWTKEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEE 150
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPS 137
ED II QAH GN+WA I++LLPGRTDNAIKNHWNST+RR+ + L +S + P+
Sbjct: 151 EDRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRKVEQEGYLQESSKASQPA 209
>MYB_CHICK (P01103) Myb proto-oncogene protein (C-myb)
Length = 641
Score = 132 bits (331), Expect = 1e-30
Identities = 61/119 (51%), Positives = 85/119 (71%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WSVI+ + GR GK CR RW N L+P+V+ +T
Sbjct: 91 IKGPWTKEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEE 150
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPS 137
ED II QAH GN+WA I++LLPGRTDNAIKNHWNST+RR+ + L +S + PS
Sbjct: 151 EDRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRKVEQEGYLQESSKAGLPS 209
>MYB_BOVIN (P46200) Myb proto-oncogene protein (C-myb)
Length = 640
Score = 132 bits (331), Expect = 1e-30
Identities = 60/119 (50%), Positives = 85/119 (71%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WSVI+ + GR GK CR RW N L+P+V+ +T
Sbjct: 91 IKGPWTKEEDQRVIELVQKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEE 150
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPS 137
ED II QAH GN+WA I++LLPGRTDNAIKNHWNST+RR+ + L +S + P+
Sbjct: 151 EDRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRKVEQEGYLQESSKASQPA 209
>MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related
protein 1) (XMYB1)
Length = 743
Score = 130 bits (327), Expect = 3e-30
Identities = 56/104 (53%), Positives = 80/104 (76%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D VKG W+ +EDE +++LV ++G ++W++I+ + GR GK CR RW N L+P+V+ +T
Sbjct: 80 DLVKGPWTKEEDEKVIELVKKYGTKHWTLIAKQLRGRMGKQCRERWHNHLNPEVKKSSWT 139
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II QAH V GN+WA I++LLPGRTDNA+KNHWNST++R+
Sbjct: 140 EEEDRIICQAHKVLGNRWAEIAKLLPGRTDNAVKNHWNSTIKRK 183
>MYB_XENLA (Q08759) Myb protein
Length = 624
Score = 128 bits (321), Expect = 2e-29
Identities = 55/102 (53%), Positives = 77/102 (74%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WSVI+ + GR GK CR RW N L+P+V+ +T
Sbjct: 88 IKGPWTKEEDQRVIELVHKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKKSSWTEE 147
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED I +AH GN+WA I++LLPGRTDNAIKNHWNST+RR+
Sbjct: 148 EDRTIYEAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRK 189
>MYBB_CHICK (Q03237) Myb-related protein B (B-Myb)
Length = 686
Score = 128 bits (321), Expect = 2e-29
Identities = 54/104 (51%), Positives = 79/104 (75%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D VKG W+ +ED+ +++LV ++G + W++I+ + GR GK CR RW N L+P+V+ +T
Sbjct: 80 DLVKGPWTKEEDQKVIELVKKYGTKQWTLIAKHLKGRLGKQCRERWHNHLNPEVKKSSWT 139
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II +AH V GN+WA I++LLPGRTDNA+KNHWNST++R+
Sbjct: 140 EEEDRIIFEAHKVLGNRWAEIAKLLPGRTDNAVKNHWNSTIKRK 183
>MYBA_MOUSE (P51960) Myb-related protein A (A-Myb)
Length = 751
Score = 127 bits (320), Expect = 2e-29
Identities = 54/102 (52%), Positives = 78/102 (75%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WS+I+ + GR GK CR RW N L+P+V+ +T
Sbjct: 86 IKGPWTKEEDQRVIELVQKYGPKRWSLIAKHLKGRIGKQCRERWHNHLNPEVKKSSWTEE 145
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II +AH GN+WA I++LLPGRTDN+IKNHWNST+RR+
Sbjct: 146 EDRIIYEAHKRLGNRWAEIAKLLPGRTDNSIKNHWNSTMRRK 187
Score = 30.8 bits (68), Expect = 3.4
Identities = 15/56 (26%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPF 75
K SW+ +ED + + + G R W+ I+ +PGR+ S + W + + V+ +
Sbjct: 139 KSSWTEEEDRIIYEAHKRLGNR-WAEIAKLLPGRTDNSIKNHWNSTMRRKVEQEGY 193
>MYBA_HUMAN (P10243) Myb-related protein A (A-Myb)
Length = 752
Score = 127 bits (320), Expect = 2e-29
Identities = 54/102 (52%), Positives = 78/102 (75%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WS+I+ + GR GK CR RW N L+P+V+ +T
Sbjct: 86 IKGPWTKEEDQRVIELVQKYGPKRWSLIAKHLKGRIGKQCRERWHNHLNPEVKKSSWTEE 145
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II +AH GN+WA I++LLPGRTDN+IKNHWNST+RR+
Sbjct: 146 EDRIIYEAHKRLGNRWAEIAKLLPGRTDNSIKNHWNSTMRRK 187
Score = 30.8 bits (68), Expect = 3.4
Identities = 15/56 (26%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPF 75
K SW+ +ED + + + G R W+ I+ +PGR+ S + W + + V+ +
Sbjct: 139 KSSWTEEEDRIIYEAHKRLGNR-WAEIAKLLPGRTDNSIKNHWNSTMRRKVEQEGY 193
>MYBA_CHICK (P52550) Myb-related protein A (A-Myb)
Length = 757
Score = 127 bits (320), Expect = 2e-29
Identities = 53/102 (51%), Positives = 78/102 (75%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ +ED+ +++LV ++GP+ WS+I+ + GR GK CR RW N L+P+V+ +T
Sbjct: 86 IKGPWTKEEDQRVIELVQKYGPKRWSLIAKHLKGRIGKQCRERWHNHLNPEVKKSSWTEA 145
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED +I +AH GN+WA I++LLPGRTDN+IKNHWNST+RR+
Sbjct: 146 EDRVIYEAHKRLGNRWAEIAKLLPGRTDNSIKNHWNSTMRRK 187
Score = 30.0 bits (66), Expect = 5.9
Identities = 15/56 (26%), Positives = 28/56 (49%), Gaps = 1/56 (1%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPF 75
K SW+ ED + + + G R W+ I+ +PGR+ S + W + + V+ +
Sbjct: 139 KSSWTEAEDRVIYEAHKRLGNR-WAEIAKLLPGRTDNSIKNHWNSTMRRKVEQEGY 193
>MYBH_DICDI (P34127) Myb-like protein (Fragment)
Length = 451
Score = 126 bits (317), Expect = 5e-29
Identities = 53/101 (52%), Positives = 75/101 (73%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
VKG+W+ ED+ +++LV +GP+ WS I+ + GR GK CR RW N L+P+++ ++
Sbjct: 200 VKGAWTKDEDDKVIELVKTYGPKKWSDIALHLKGRMGKQCRERWHNHLNPNIKKEAWSDE 259
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRR 119
ED II HA+HGNKWA I++ LPGRTDNAIKNHWNS+++R
Sbjct: 260 EDQIIRDQHAIHGNKWAEIAKFLPGRTDNAIKNHWNSSMKR 300
>MYBB_HUMAN (P10244) Myb-related protein B (B-Myb)
Length = 700
Score = 126 bits (317), Expect = 5e-29
Identities = 79/212 (37%), Positives = 118/212 (55%), Gaps = 30/212 (14%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D VKG W+ +ED+ +++LV ++G + W++I+ + GR GK CR RW N L+P+V+ +T
Sbjct: 80 DLVKGPWTKEEDQKVIELVKKYGTKQWTLIAKHLKGRLGKQCRERWHNHLNPEVKKSCWT 139
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSP 136
ED II +AH V GN+WA I+++LPGRTDNA+KNHWNST++R+ L S S
Sbjct: 140 EEEDRIICEAHKVLGNRWAEIAKMLPGRTDNAVKNHWNSTIKRKVDTGGFL-----SESK 194
Query: 137 SPSPPLPFSMAKRPHGYELLDRYDGLMD------QG-----YPFKRHCVEKDDEESRDGV 185
PP+ + EL D+ DGL QG +P + K++E S + +
Sbjct: 195 DCKPPVYLLL-------ELEDK-DGLQSAQPTEGQGSLLTNWPSVPPTI-KEEENSEEEL 245
Query: 186 EEAETMTEEEVLASSV-----PTSLSLFPMGE 212
A T E+E + + + P L FP E
Sbjct: 246 AAATTSKEQEPIGTDLDAVRTPEPLEEFPKRE 277
>MYBB_MOUSE (P48972) Myb-related protein B (B-Myb)
Length = 704
Score = 126 bits (316), Expect = 6e-29
Identities = 53/104 (50%), Positives = 79/104 (75%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D VKG W+ +ED+ +++LV ++G + W++I+ + GR GK CR RW N L+P+V+ +T
Sbjct: 80 DLVKGPWTKEEDQKVIELVKKYGTKQWTLIAKHLKGRLGKQCRERWHNHLNPEVKKSCWT 139
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II +AH V GN+WA I+++LPGRTDNA+KNHWNST++R+
Sbjct: 140 EEEDRIICEAHKVLGNRWAEIAKMLPGRTDNAVKNHWNSTIKRK 183
>MYB_AVIMB (P01104) Transforming protein Myb
Length = 382
Score = 124 bits (312), Expect = 2e-28
Identities = 58/122 (47%), Positives = 80/122 (65%), Gaps = 5/122 (4%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
KG W+ +ED+ +++ V ++GP+ WS I+ + GR GK CR RW N L+P+V+ +T E
Sbjct: 21 KGPWTKEEDQRVIEHVQKYGPKRWSDIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTEEE 80
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPSPS 139
D II QAH GN+WA I++LLPGRTDNA+KNHWNST+RR+ + P S
Sbjct: 81 DRIIYQAHKRLGNRWAEIAKLLPGRTDNAVKNHWNSTMRRKVEQE-----GYPQESSKAG 135
Query: 140 PP 141
PP
Sbjct: 136 PP 137
>MYB_DROME (P04197) Myb protein
Length = 657
Score = 124 bits (310), Expect = 3e-28
Identities = 54/102 (52%), Positives = 74/102 (71%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ ED+ ++KLV GP+ W++I+ + GR GK CR RW N L+P+++ +T
Sbjct: 135 IKGPWTRDEDDMVIKLVRNFGPKKWTLIARYLNGRIGKQCRERWHNHLNPNIKKTAWTEK 194
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II QAH GN+WA I++ LPGRTDNAIKNHWNST+RR+
Sbjct: 195 EDEIIYQAHLELGNQWAKIAKRLPGRTDNAIKNHWNSTMRRK 236
>MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related protein
21) (AtMYB21) (Myb homolog 3) (AtMyb3)
Length = 226
Score = 111 bits (277), Expect = 2e-24
Identities = 53/119 (44%), Positives = 74/119 (61%), Gaps = 3/119 (2%)
Query: 3 GGGDSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRL 60
GGG S G + + KG W+ +ED L+ + HG W+ + SAG+ R+GKSCRL
Sbjct: 5 GGGSSGGSGSSAEAEVRKGPWTMEEDLILINYIANHGDGVWNSLAKSAGLK-RTGKSCRL 63
Query: 61 RWCNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRR 119
RW N L PDV+ TP E II++ HA GN+W+ I++ LPGRTDN IKN W + +++
Sbjct: 64 RWLNYLRPDVRRGNITPEEQLIIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQK 122
Score = 31.2 bits (69), Expect = 2.6
Identities = 16/59 (27%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+G+ +P+E +M+L + G R WS I+ +PGR+ + W ++ ++ T T
Sbjct: 75 RGNITPEEQLIIMELHAKWGNR-WSKIAKHLPGRTDNEIKNFWRTRIQKYIKQSDVTTT 132
>MYB1_MAIZE (P20024) Myb-related protein Zm1
Length = 340
Score = 107 bits (268), Expect = 2e-23
Identities = 51/103 (49%), Positives = 69/103 (66%), Gaps = 3/103 (2%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
+GSW+PQED L+ + +HG NW + AG+ R GKSCRLRW N L PD++ FT
Sbjct: 16 RGSWTPQEDMRLIAYIQKHGHTNWRALPKQAGLL-RCGKSCRLRWINYLRPDLKRGNFTD 74
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ II+ H + GNKW+ I+ LPGRTDN IKN WN+ L+++
Sbjct: 75 EEEEAIIRLHGLLGNKWSKIAACLPGRTDNEIKNVWNTHLKKK 117
Score = 35.0 bits (79), Expect = 0.18
Identities = 18/57 (31%), Positives = 32/57 (55%), Gaps = 1/57 (1%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHR 73
D +G+++ +E+E +++L G G + WS I+A +PGR+ + W L V R
Sbjct: 66 DLKRGNFTDEEEEAIIRLHGLLGNK-WSKIAACLPGRTDNEIKNVWNTHLKKKVAQR 121
>MYB3_MAIZE (P20025) Myb-related protein Zm38
Length = 255
Score = 105 bits (263), Expect = 8e-23
Identities = 49/103 (47%), Positives = 71/103 (68%), Gaps = 3/103 (2%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
+G+W+ +EDE L+ + HG W + +AG+ R GKSCRLRW N L PD++ FT
Sbjct: 14 RGAWTKEEDERLVAYIRAHGEGCWRSLPKAAGLL-RCGKSCRLRWINYLRPDLKRGNFTA 72
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED +I++ H++ GNKW+ I+ LPGRTDN IKN+WN+ +RR+
Sbjct: 73 DEDDLIVKLHSLLGNKWSLIAARLPGRTDNEIKNYWNTHVRRK 115
>LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-red
light protein 1) (Myb-related protein 18) (AtMYB18)
Length = 283
Score = 104 bits (259), Expect = 2e-22
Identities = 58/138 (42%), Positives = 81/138 (58%), Gaps = 4/138 (2%)
Query: 14 GDRDRVKGSWSPQEDETLMKLVGQHGPRNWSV--ISAGIPGRSGKSCRLRWCNQLSPDVQ 71
G+R R KG WSP+EDE L + +G W+ I AG+ R+GKSCRLRW N L P ++
Sbjct: 7 GERHR-KGLWSPEEDEKLRSFILSYGHSCWTTVPIKAGLQ-RNGKSCRLRWINYLRPGLK 64
Query: 72 HRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLAS 131
+ E+ I+ H+ GNKW+ I++ LPGRTDN IKN+W+S L+++ +S L A
Sbjct: 65 RDMISAEEEETILTFHSSLGNKWSQIAKFLPGRTDNEIKNYWHSHLKKKWLKSQSLQDAK 124
Query: 132 PSPSPSPSPPLPFSMAKR 149
PS S + KR
Sbjct: 125 SISPPSSSSSSLVACGKR 142
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,963,469
Number of Sequences: 164201
Number of extensions: 1684193
Number of successful extensions: 10856
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 66
Number of HSP's successfully gapped in prelim test: 64
Number of HSP's that attempted gapping in prelim test: 10202
Number of HSP's gapped (non-prelim): 503
length of query: 260
length of database: 59,974,054
effective HSP length: 108
effective length of query: 152
effective length of database: 42,240,346
effective search space: 6420532592
effective search space used: 6420532592
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0002.6


