

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0001.6
(381 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
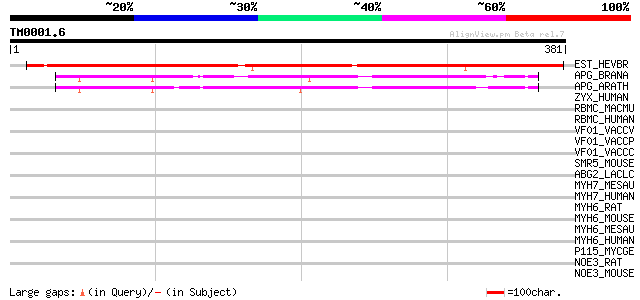
Score E
Sequences producing significant alignments: (bits) Value
EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early nodule... 336 5e-92
APG_BRANA (P40603) Anter-specific proline-rich protein APG (Prot... 87 9e-17
APG_ARATH (P40602) Anter-specific proline-rich protein APG precu... 83 1e-15
ZYX_HUMAN (Q15942) Zyxin (Zyxin 2) 34 0.71
RBMC_MACMU (Q8SQ27) RNA-binding protein 12 (RNA binding motif pr... 33 0.92
RBMC_HUMAN (Q9NTZ6) RNA-binding protein 12 (RNA binding motif pr... 33 0.92
VF01_VACCV (P24356) Protein F1 (Fragment) 32 2.7
VF01_VACCP (P68451) Protein F1 (Fragment) 32 2.7
VF01_VACCC (P68450) Protein F1 32 2.7
SMR5_MOUSE (Q91ZW3) SWI/SNF related matrix associated actin depe... 32 2.7
ABG2_LACLC (Q48726) Abortive phage resistance protein abiGii 32 3.5
MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isof... 31 4.6
MYH7_HUMAN (P12883) Myosin heavy chain, cardiac muscle beta isof... 31 4.6
MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isofo... 31 4.6
MYH6_MOUSE (Q02566) Myosin heavy chain, cardiac muscle alpha iso... 31 4.6
MYH6_MESAU (P13539) Myosin heavy chain, cardiac muscle alpha iso... 31 4.6
MYH6_HUMAN (P13533) Myosin heavy chain, cardiac muscle alpha iso... 31 4.6
P115_MYCGE (P47540) P115 protein homolog 31 6.0
NOE3_RAT (P63057) Noelin 3 precursor (Olfactomedin 3) (Optimedin) 31 6.0
NOE3_MOUSE (P63056) Noelin 3 precursor (Olfactomedin 3) (Optimedin) 31 6.0
>EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early
nodule-specific protein homolog) (Latex allergen Hev b
13)
Length = 391
Score = 336 bits (862), Expect = 5e-92
Identities = 181/376 (48%), Positives = 239/376 (63%), Gaps = 15/376 (3%)
Query: 12 LSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDC 71
LS LC+ S+ S C FPAI+NFGDSNSDTGG +AAF P+ PPYGE+F + + R
Sbjct: 14 LSFLLCMLSLAYA-SETCDFPAIFNFGDSNSDTGGKAAAFYPLNPPYGETFFHRSTGRYS 72
Query: 72 DGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQY-GISPFSL 130
DGRLI+DFIAE NLPYLS YL+SLG+N++HGA+FAT GSTI+ I + G SPF L
Sbjct: 73 DGRLIIDFIAESFNLPYLSPYLSSLGSNFKHGADFATAGSTIKLPTTIIPAHGGFSPFYL 132
Query: 131 DMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEE--FSKALYTFDIGQNDLSVGFRMMN 188
D+Q+ QF+QF R++ + + E VPEE F KALYTFDIGQNDL+ GF +
Sbjct: 133 DVQYSQFRQFIPRSQFIRETGGIFAEL----VPEEYYFEKALYTFDIGQNDLTEGFLNLT 188
Query: 189 FDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYG 248
+++ ++PD+VN ++ VK IY+LG RTFWIHNT PIGCL L Y P D G
Sbjct: 189 VEEVNATVPDLVNSFSANVKKIYDLGARTFWIHNTGPIGCLSFILTY---FPWAEKDSAG 245
Query: 249 CVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKIC 308
C K N +A FN +LK+ V +LR +LP A +VD+Y+ KY L S + GF PL C
Sbjct: 246 CAKAYNEVAQHFNHKLKEIVAQLRKDLPLATFVHVDIYSVKYSLFSEPEKHGFEFPLITC 305
Query: 309 CGY---HVNDTHIWCG-TIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILN 364
CGY + CG T+ +G + +C PS+ V+WDG HY EAAN + ++I
Sbjct: 306 CGYGGKYNFSVTAPCGDTVTADDGTKIVVGSCACPSVRVNWDGAHYTEAANEYFFDQIST 365
Query: 365 GSFTDPPTLITQACYR 380
G+F+DPP + AC++
Sbjct: 366 GAFSDPPVPLNMACHK 381
>APG_BRANA (P40603) Anter-specific proline-rich protein APG (Protein
CEX) (Fragment)
Length = 449
Score = 86.7 bits (213), Expect = 9e-17
Identities = 90/349 (25%), Positives = 147/349 (41%), Gaps = 48/349 (13%)
Query: 32 PAIYNFGDSNSDTGG---ISAAFEPIPPPYGESFPQK-PSARDCDGRLIVDFIAEKLNLP 87
PA++ FGDS DTG + + PYG FP + R +GR+ D+I++ L +
Sbjct: 124 PAVFFFGDSIFDTGNNNNLDTKLKCNYRPYGMDFPMGVATGRFSNGRVASDYISKYLGVK 183
Query: 88 YL-SAYLNSL--------GTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFK 138
+ AY++ ++ G +FA+GG+ Q ++ LD Q F+
Sbjct: 184 EIVPAYVDKKLQQNNELQQSDLLTGVSFASGGAGYLPQTSESWKVTTM---LD-QLTYFQ 239
Query: 139 QFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPD 198
+K R K+L + KT + SK G NDL + ++ +
Sbjct: 240 DYKKRMKKLVGKKKTK---------KIVSKGAAIVVAGSNDLIYTYFGNGAQHLKNDVDS 290
Query: 199 IVNQLA----SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQN 254
+A S V +Y G R + T PIGC P K + C +D N
Sbjct: 291 FTTMMADSAASFVLQLYGYGARRIGVIGTPPIGCTPSQRVKKKKI---------CNEDLN 341
Query: 255 VMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVN 314
A FN +L + +L LP + I Y D+Y+ ++ + ++ GF + K CC +
Sbjct: 342 YAAQLFNSKLVIILGQLSKTLPNSTIVYGDIYSIFSKMLESPEDYGFEEIKKPCCKIGLT 401
Query: 315 DTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRIL 363
++C N NA S Y+ WDG+H ++ A + ++NR L
Sbjct: 402 KGGVFCKERTLKN----MSNA----SSYLFWDGLHPSQRA-YEISNRKL 441
>APG_ARATH (P40602) Anter-specific proline-rich protein APG
precursor
Length = 534
Score = 82.8 bits (203), Expect = 1e-15
Identities = 82/344 (23%), Positives = 144/344 (41%), Gaps = 34/344 (9%)
Query: 32 PAIYNFGDSNSDTGG---ISAAFEPIPPPYGESFPQK-PSARDCDGRLIVDFIAEKLNLP 87
PA++ FGDS DTG + + PYG F + + R +G + D++A+ + +
Sbjct: 203 PAVFFFGDSVFDTGNNNNLETKIKSNYRPYGMDFKFRVATGRFSNGMVASDYLAKYMGVK 262
Query: 88 YL-SAYLNSL--GTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKART 144
+ AYL+ + G +FA+GG+ N T + + LD Q F+ + +
Sbjct: 263 EIVPAYLDPKIQPNDLLTGVSFASGGAGY---NPTTSEAANAIPMLD-QLTYFQDYIEKV 318
Query: 145 KQLYQEAKTALERSKLPVPEEF-SKALYTFDIGQNDLSVGFRMMNFDQMRESMPD----I 199
+L ++ K+ + + L + SK + G NDL + + +++ + I
Sbjct: 319 NRLVRQEKSQYKLAGLEKTNQLISKGVAIVVGGSNDLIITYFGSGAQRLKNDIDSYTTII 378
Query: 200 VNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVE 259
+ AS V +Y G R + T P+GC+P K + C ++ N +
Sbjct: 379 ADSAASFVLQLYGYGARRIGVIGTPPLGCVPSQRLKKKKI---------CNEELNYASQL 429
Query: 260 FNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIW 319
FN +L + +L LP + Y+D+Y ++ GF + K CC +
Sbjct: 430 FNSKLLLILGQLSKTLPNSTFVYMDIYTIISQMLETPAAYGFEETKKPCCKTGLLSAGAL 489
Query: 320 CGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRIL 363
C K C S Y+ WDGVH + A + N++L
Sbjct: 490 C--------KKSTSKICPNTSSYLFWDGVHPTQRA-YKTINKVL 524
>ZYX_HUMAN (Q15942) Zyxin (Zyxin 2)
Length = 572
Score = 33.9 bits (76), Expect = 0.71
Identities = 14/29 (48%), Positives = 16/29 (54%)
Query: 38 GDSNSDTGGISAAFEPIPPPYGESFPQKP 66
GD + G + AF P PPP ESFP P
Sbjct: 80 GDGDDAEGALGGAFPPPPPPIEESFPPAP 108
>RBMC_MACMU (Q8SQ27) RNA-binding protein 12 (RNA binding motif
protein 12) (SH3/WW domain anchor protein in the
nucleus) (SWAN)
Length = 932
Score = 33.5 bits (75), Expect = 0.92
Identities = 24/58 (41%), Positives = 27/58 (46%), Gaps = 15/58 (25%)
Query: 16 LCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPP------YGESFPQKPS 67
L V S E N PP FP NFG SN A PIPPP +G++ P PS
Sbjct: 708 LTVGSKEANNGPPFNFPG--NFGGSN-------AFGPPIPPPGLGGGAFGDARPGMPS 756
>RBMC_HUMAN (Q9NTZ6) RNA-binding protein 12 (RNA binding motif
protein 12) (SH3/WW domain anchor protein in the
nucleus) (SWAN) (HRIHFB2091)
Length = 932
Score = 33.5 bits (75), Expect = 0.92
Identities = 24/58 (41%), Positives = 27/58 (46%), Gaps = 15/58 (25%)
Query: 16 LCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPP------YGESFPQKPS 67
L V S E N PP FP NFG SN A PIPPP +G++ P PS
Sbjct: 708 LTVGSKEANNGPPFNFPG--NFGGSN-------AFGPPIPPPGLGGGAFGDARPGMPS 756
>VF01_VACCV (P24356) Protein F1 (Fragment)
Length = 207
Score = 32.0 bits (71), Expect = 2.7
Identities = 20/70 (28%), Positives = 32/70 (45%), Gaps = 1/70 (1%)
Query: 229 LPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAA 288
LP N+ Y+ + LD +D +MAV + K R+ L +LP +Y+D+
Sbjct: 41 LPENMVYRFDKSTNILDYLSTERDHVMMAVRYYMS-KQRLDDLYRQLPTKTRSYIDIINI 99
Query: 289 KYGLISNTKN 298
+SN N
Sbjct: 100 YCDKVSNDYN 109
>VF01_VACCP (P68451) Protein F1 (Fragment)
Length = 206
Score = 32.0 bits (71), Expect = 2.7
Identities = 20/70 (28%), Positives = 32/70 (45%), Gaps = 1/70 (1%)
Query: 229 LPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAA 288
LP N+ Y+ + LD +D +MAV + K R+ L +LP +Y+D+
Sbjct: 41 LPENMVYRFDKSTNILDYLSTERDHVMMAVRYYMS-KQRLDDLYRQLPTKTRSYIDIINI 99
Query: 289 KYGLISNTKN 298
+SN N
Sbjct: 100 YCDKVSNDYN 109
>VF01_VACCC (P68450) Protein F1
Length = 226
Score = 32.0 bits (71), Expect = 2.7
Identities = 20/70 (28%), Positives = 32/70 (45%), Gaps = 1/70 (1%)
Query: 229 LPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAA 288
LP N+ Y+ + LD +D +MAV + K R+ L +LP +Y+D+
Sbjct: 41 LPENMVYRFDKSTNILDYLSTERDHVMMAVRYYMS-KQRLDDLYRQLPTKTRSYIDIINI 99
Query: 289 KYGLISNTKN 298
+SN N
Sbjct: 100 YCDKVSNDYN 109
>SMR5_MOUSE (Q91ZW3) SWI/SNF related matrix associated actin
dependent regulator of chromatin, subfamily A member 5
(EC 3.6.1.-) (Sucrose nonfermenting protein 2 homolog)
(mSnf2h)
Length = 1051
Score = 32.0 bits (71), Expect = 2.7
Identities = 15/27 (55%), Positives = 17/27 (62%)
Query: 47 ISAAFEPIPPPYGESFPQKPSARDCDG 73
+S+A EP PPP ES P KPSA G
Sbjct: 1 MSSAVEPPPPPPPESAPSKPSAAGAGG 27
>ABG2_LACLC (Q48726) Abortive phage resistance protein abiGii
Length = 397
Score = 31.6 bits (70), Expect = 3.5
Identities = 19/58 (32%), Positives = 29/58 (49%), Gaps = 2/58 (3%)
Query: 250 VKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGL-ISNTKNEGFVDPLK 306
+KD N++ + F + DRVVK ++E P T+ D K I+ KN G P +
Sbjct: 260 IKDVNIIKILFESYINDRVVKFKSE-PSLKFTFDDSSNFKESFPITGKKNMGIFVPFQ 316
>MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1934
Score = 31.2 bits (69), Expect = 4.6
Identities = 25/88 (28%), Positives = 43/88 (48%), Gaps = 5/88 (5%)
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSK---LPVPEEFSKALYTFDIGQNDLSVGFR 185
+LD + F + A KQ Y+E+++ LE S+ + E K ++ L F+
Sbjct: 1440 ALDKKQRNFDKILAEWKQKYEESQSELESSQKEARSLSTELFKLKNAYEESLEHLET-FK 1498
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYEL 213
N ++E + D+ QL S K+I+EL
Sbjct: 1499 REN-KNLQEEISDLTEQLGSTGKSIHEL 1525
>MYH7_HUMAN (P12883) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1935
Score = 31.2 bits (69), Expect = 4.6
Identities = 25/88 (28%), Positives = 43/88 (48%), Gaps = 5/88 (5%)
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSK---LPVPEEFSKALYTFDIGQNDLSVGFR 185
+LD + F + A KQ Y+E+++ LE S+ + E K ++ L F+
Sbjct: 1441 ALDKKQRNFDKILAEWKQKYEESQSELESSQKEARSLSTELFKLKNAYEESLEHLET-FK 1499
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYEL 213
N ++E + D+ QL S+ K I+EL
Sbjct: 1500 REN-KNLQEEISDLTEQLGSSGKTIHEL 1526
>MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1938
Score = 31.2 bits (69), Expect = 4.6
Identities = 24/88 (27%), Positives = 42/88 (47%), Gaps = 5/88 (5%)
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSK---LPVPEEFSKALYTFDIGQNDLSVGFR 185
+LD + F + A KQ Y+E+++ LE S+ + E K ++ L F+
Sbjct: 1442 ALDKKQRNFDKILAEWKQKYEESQSELESSQKEARSLSTELFKLKNAYEESLEHLET-FK 1500
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYEL 213
N ++E + D+ QL KN++EL
Sbjct: 1501 REN-KNLQEEISDLTEQLGEGGKNVHEL 1527
>MYH6_MOUSE (Q02566) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1938
Score = 31.2 bits (69), Expect = 4.6
Identities = 24/88 (27%), Positives = 42/88 (47%), Gaps = 5/88 (5%)
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSK---LPVPEEFSKALYTFDIGQNDLSVGFR 185
+LD + F + A KQ Y+E+++ LE S+ + E K ++ L F+
Sbjct: 1443 ALDKKQRNFDKILAEWKQKYEESQSELESSQKEARSLSTELFKLKNAYEESLEHLET-FK 1501
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYEL 213
N ++E + D+ QL KN++EL
Sbjct: 1502 REN-KNLQEEISDLTEQLGEGGKNVHEL 1528
>MYH6_MESAU (P13539) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1939
Score = 31.2 bits (69), Expect = 4.6
Identities = 24/88 (27%), Positives = 42/88 (47%), Gaps = 5/88 (5%)
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSK---LPVPEEFSKALYTFDIGQNDLSVGFR 185
+LD + F + A KQ Y+E+++ LE S+ + E K ++ L F+
Sbjct: 1443 ALDKKQRNFDKILAEWKQKYEESQSELESSQKEARSLSTELFKLKNAYEESLEHLET-FK 1501
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYEL 213
N ++E + D+ QL KN++EL
Sbjct: 1502 REN-KNLQEEISDLTEQLGEGGKNVHEL 1528
>MYH6_HUMAN (P13533) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1939
Score = 31.2 bits (69), Expect = 4.6
Identities = 24/88 (27%), Positives = 42/88 (47%), Gaps = 5/88 (5%)
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSK---LPVPEEFSKALYTFDIGQNDLSVGFR 185
+LD + F + A KQ Y+E+++ LE S+ + E K ++ L F+
Sbjct: 1443 ALDKKQRNFDKILAEWKQKYEESQSELESSQKEARSLSTELFKLKNAYEESLEHLET-FK 1501
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYEL 213
N ++E + D+ QL KN++EL
Sbjct: 1502 REN-KNLQEEISDLTEQLGEGGKNVHEL 1528
>P115_MYCGE (P47540) P115 protein homolog
Length = 982
Score = 30.8 bits (68), Expect = 6.0
Identities = 20/60 (33%), Positives = 31/60 (51%), Gaps = 3/60 (5%)
Query: 234 FYKHNLPAGYLDPYGCVKDQNVMAVEFN-KQLKDRVVKLRTELPE--AAITYVDLYAAKY 290
F K NL GYL +QN+ +E N ++LK + +L +L E + Y +L AK+
Sbjct: 563 FEKTNLSDGYLSSASLDNEQNINKLENNERELKKELTELEVKLDEMNRKLKYEELLQAKF 622
>NOE3_RAT (P63057) Noelin 3 precursor (Olfactomedin 3) (Optimedin)
Length = 478
Score = 30.8 bits (68), Expect = 6.0
Identities = 27/104 (25%), Positives = 44/104 (41%), Gaps = 6/104 (5%)
Query: 67 SARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGIS 126
SA+D DGR I +A + NL A L N + + + + FQY
Sbjct: 57 SAQDPDGRCICTVVAPEQNLCSRDAKSRQLRQLLEKVQNMSQSIEVLNLRTQRDFQY--- 113
Query: 127 PFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKAL 170
L M+ Q K KA+ +Q+ + KT + + + E+ + L
Sbjct: 114 --VLKME-TQMKGLKAKFRQIEDDRKTLMTKHFQELKEKMDELL 154
>NOE3_MOUSE (P63056) Noelin 3 precursor (Olfactomedin 3) (Optimedin)
Length = 478
Score = 30.8 bits (68), Expect = 6.0
Identities = 27/104 (25%), Positives = 44/104 (41%), Gaps = 6/104 (5%)
Query: 67 SARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGIS 126
SA+D DGR I +A + NL A L N + + + + FQY
Sbjct: 57 SAQDPDGRCICTVVAPEQNLCSRDAKSRQLRQLLEKVQNMSQSIEVLNLRTQRDFQY--- 113
Query: 127 PFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKAL 170
L M+ Q K KA+ +Q+ + KT + + + E+ + L
Sbjct: 114 --VLKME-TQMKGLKAKFRQIEDDRKTLMTKHFQELKEKMDELL 154
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,268,911
Number of Sequences: 164201
Number of extensions: 2127973
Number of successful extensions: 4720
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 4687
Number of HSP's gapped (non-prelim): 37
length of query: 381
length of database: 59,974,054
effective HSP length: 112
effective length of query: 269
effective length of database: 41,583,542
effective search space: 11185972798
effective search space used: 11185972798
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0001.6


