

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0545.3
(2087 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
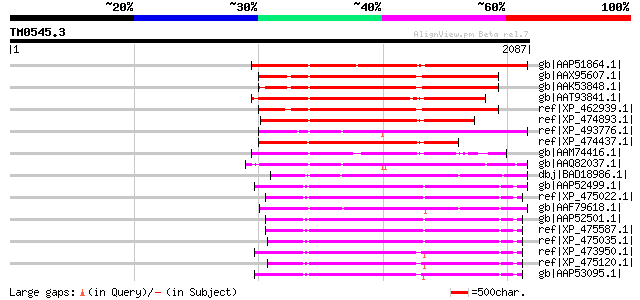
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sati... 1097 0.0
gb|AAX95607.1| RNase H, putative [Oryza sativa (japonica cultiva... 985 0.0
gb|AAK53848.1| Putative retroelement [Oryza sativa] 970 0.0
gb|AAT93841.1| putative polyprotein [Oryza sativa (japonica cult... 969 0.0
ref|XP_462939.1| putative gag-pol protein [Oryza sativa (japonic... 966 0.0
ref|XP_474893.1| OSJNBa0021F22.20 [Oryza sativa (japonica cultiv... 904 0.0
ref|XP_493776.1| unnamed protein product [Oryza sativa (japonica... 862 0.0
ref|XP_474437.1| OSJNBa0070M12.15 [Oryza sativa (japonica cultiv... 859 0.0
gb|AAM74416.1| Putative retroelement [Oryza sativa (japonica cul... 814 0.0
gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum] 791 0.0
dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera] 690 0.0
gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonic... 686 0.0
ref|XP_475022.1| OSJNBb0093G06.3 [Oryza sativa (japonica cultiva... 686 0.0
gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||... 682 0.0
gb|AAP52501.1| putative gag-pol precursor [Oryza sativa (japonic... 682 0.0
ref|XP_475587.1| putative polyprotein [Oryza sativa (japonica cu... 681 0.0
ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultiva... 681 0.0
ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultiv... 681 0.0
ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cu... 676 0.0
gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cul... 676 0.0
>gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37530550|ref|NP_919577.1| putative
retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|18581535|gb|AAK52540.2| Putative
retroelement pol polyprotein [Oryza sativa]
Length = 1580
Score = 1097 bits (2836), Expect = 0.0
Identities = 545/1120 (48%), Positives = 758/1120 (67%), Gaps = 34/1120 (3%)
Query: 973 RLDCIYDDGPLGFEKPVVDVSKKMEA-----QDPLEEVD--LGDGLEKRPTYISALIDPE 1025
+++ + D L E P ME +D ++++D G GL I P
Sbjct: 479 KIEIVPADSQLKMENPSYCFEGVMEGSNVYTKDTVDDLDDKQGQGLMSADDLEEIDISPA 538
Query: 1026 L---KDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLV 1082
+ + +++LLKEF+DCF W+Y EMPG +R +VE +LP+K +P +QLPRR D+L
Sbjct: 539 IGQGRHLLIELLKEFRDCFTWEYYEMPGHSRSIVEHRLPLKPGVRPHQQLPRRCKADMLE 598
Query: 1083 KIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIA 1142
+K EI+RL FIR RY +W++++VPVIKKNGK+RVCIDFR LN ATPKDEY M +A
Sbjct: 599 PVKAEIKRLYDAGFIRPCRYAEWVSSIVPVIKKNGKVRVCIDFRYLNKATPKDEYPMLVA 658
Query: 1143 EMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGL 1202
+ +VD+A+GH+ LS +DG +GYNQIF+A ED+ KT FRCPGA+G +EWVV+ F LK+AG
Sbjct: 659 DQLVDAASGHKILSFMDGNAGYNQIFMAEEDIHKTTFRCPGAIGLFEWVVITFVLKSAG- 717
Query: 1203 KNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNP 1262
ATYQR MN I+HD I ++VYIDD+VVKS + H+ LRK FER RKYGLKMNP
Sbjct: 718 ----ATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEIEDHIADLRKVFERTRKYGLKMNP 773
Query: 1263 LKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITN 1322
KCAFGV AG FLGF+VH++GIE+ + AI PP K +LQ ++GKINF+RRFI+
Sbjct: 774 TKCAFGVSAGQFLGFLVHERGIEVTQRSINAIKKIKPPEDKTELQEMIGKINFVRRFISI 833
Query: 1323 LSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATD 1382
LS K + F+ LLRLK + F W AE QKA D +K+YLS+PPV+IPP KG P +LY+SA D
Sbjct: 834 LSGKLEPFTPLLRLKADQQFAWGAEQQKALDNIKEYLSSPPVLIPPQKGIPFRLYLSAGD 893
Query: 1383 ETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVM 1442
++I S+L QE E ER +FYLSR L AETRY+ +EKLCLCLYFSC +L++Y+ +
Sbjct: 894 KSISSVLIQELER-KERVVFYLSRRLLNAETRYSPVEKLCLCLYFSCTRLRHYLLSNECT 952
Query: 1443 VFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIV 1502
V D++K+MLS PIL RIGKW +LTE+ L Y KAIKGQA+A+F+ D+ + +
Sbjct: 953 VICKADVVKYMLSAPILKGRIGKWIFSLTEFDLRYESPKAIKGQAIANFIVDYR--DDSI 1010
Query: 1503 TYVGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLE 1562
V + PW L+FDGS +G GIG+ I+SP+GA +F + I+ +NN+ EYEA++ GL+
Sbjct: 1011 GSVEVVPWTLFFDGSVCTHGCGIGLVIISPRGACFEFAYTIKPYATNNQAEYEAVLKGLQ 1070
Query: 1563 ILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIE 1622
+L + A + I GDS LV+ QL EY+C ++ L Y K L+ +F + + HV R +
Sbjct: 1071 LLKEVEADTIEIMGDSLLVISQLAGEYECKNDTLIVYNEKCQELMREFRLVTLKHVSREQ 1130
Query: 1623 NQEANELAQIASGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVEYLQNP 1682
N EAN+LAQ ASGY + + + ++I+ + +T++DWR + +YLQ+P
Sbjct: 1131 NIEANDLAQGASGYKL----MIKHVQIEVAV------------ITADDWRYYVYQYLQDP 1174
Query: 1683 VGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAK 1742
S RK++Y+AL Y +L +EL+ + I+G LLKCLS + A +A+ H+G+CG HQ+ K
Sbjct: 1175 SQSASRKLRYKALKYILLDDELYYRTIDGVLLKCLSTDQAKVAIGEVHEGICGTHQSAHK 1234
Query: 1743 MKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLI 1802
MKW+L G +WPT+++DC Y +GCQDCQK IQ PAS ++ IIKPWPFRGW ID+I
Sbjct: 1235 MKWLLRCAGYFWPTMLEDCFRYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMI 1294
Query: 1803 GEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQ 1862
G I+P SSK HK+I+VA DYFTKWVEAIPL+ V I+F+Q HI+YR G+P++ITTDQ
Sbjct: 1295 GMINPPSSKGHKFILVATDYFTKWVEAIPLKKVDSGDAIQFVQEHIIYRVGIPQTITTDQ 1354
Query: 1863 GTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHESL 1922
G++FV + FA+S GIKLL S+PYYAQANGQ EA+NK LI LIK+ + P++WH L
Sbjct: 1355 GSIFVSDEFIQFADSMGIKLLNSSPYYAQANGQAEASNKSLIKLIKRKISDYPRQWHTRL 1414
Query: 1923 SQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYWNMMLDEM 1982
++ LW+YR + + P++L Y +A+LP EV + S R + Q ++ ++ Y N+M DE
Sbjct: 1415 AEALWSYRMACHGSIQVPPYKLVYRHKAILPWEVRIGSRRTELQNDLTADEYSNLMADER 1474
Query: 1983 VNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILPMDKKDRVYGKWTPN 2042
+L + RL AL + + K+R+ + YNKKV + F G+ +WK+ILP+ +D +GKW+PN
Sbjct: 1475 EDLVQSRLRALAKVIKDKERVTRHYNKKVVPKDFSEGELIWKLILPIGTRDSKFGKWSPN 1534
Query: 2043 WEGPFTVEKTLPNNAYVIKELGNQRQCVTINGKYLKAYKP 2082
WEGPF + K + AY+++ L + +NGKYLK Y P
Sbjct: 1535 WEGPFQIHKVVSKGAYMLQGLDGEVYDRALNGKYLKKYYP 1574
>gb|AAX95607.1| RNase H, putative [Oryza sativa (japonica cultivar-group)]
Length = 2282
Score = 985 bits (2546), Expect = 0.0
Identities = 493/966 (51%), Positives = 659/966 (68%), Gaps = 36/966 (3%)
Query: 1000 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELK 1059
D LEE+D+G G RPT+I + E + ++++LLKEF+DCFAW+Y EMPGL+R +VE +
Sbjct: 1334 DDLEEIDIGPGDRPRPTFICKNLSTEFRAKLIELLKEFRDCFAWEYYEMPGLSRSIVEHR 1393
Query: 1060 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKM 1119
LPIK +P +Q RR D+L +K EI+RL FIR RY +W++++VPVIKKN
Sbjct: 1394 LPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPCRYAEWVSSIVPVIKKN--- 1450
Query: 1120 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAF 1179
DLN ATPKDEY MP+A+ +VD+A+G++ LS +DG GYNQIF+A ED+ KTAF
Sbjct: 1451 -------DLNKATPKDEYPMPVADQLVDAASGNKILSYMDGNVGYNQIFMAEEDIHKTAF 1503
Query: 1180 RCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1239
RCPGA+G +EWVVM FGLK+AG ATYQR MN I+HD I ++VYIDD+VVKS
Sbjct: 1504 RCPGAIGLFEWVVMTFGLKSAG-----ATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEI 1558
Query: 1240 DGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSP 1299
+ H+ LRK FER RKYGLKMNP KCAFGV G FLGF+VH++GIE+ + AI P
Sbjct: 1559 EDHIADLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIEVTQRSVNAIKKIQP 1618
Query: 1300 PTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYL 1359
P +K +LQ + GKINF+RRFI+ LS + + F+ LLRLK + F W AE QKA D++K+YL
Sbjct: 1619 PENKTELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWGAEQQKALDDIKEYL 1678
Query: 1360 SNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIE 1419
S+PPV+IPP KG +LY+SA +++IGS+L QE DG ER +FYLSR L ETRY+ +E
Sbjct: 1679 SSPPVLIPPQKGISFRLYLSAGEKSIGSVLIQE-LDGKERVVFYLSRRLLDVETRYSPME 1737
Query: 1420 KLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAP 1479
KLCLCLYFSC +L +Y+ + V D+IK+MLS PIL R+GKW +LTE+ L Y
Sbjct: 1738 KLCLCLYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGKWIFSLTEFDLRYES 1797
Query: 1480 LKAIKGQAVADFLADHTVPKEIVTYVGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKF 1539
KAIKGQA+ADF+ +H + + V + PW +FDGS + GIG+ I+SP+GA KF
Sbjct: 1798 PKAIKGQAIADFIVEHC--DDSIGSVEVVPWTSFFDGSVCTHDCGIGLVIISPRGACFKF 1855
Query: 1540 KFRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKY 1599
+ I+ +NN+ EYEA++ GL++L + A + I GDS LV+ QL EY+C S+ L Y
Sbjct: 1856 AYTIKPYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDSLLVISQLAGEYECKSDTLMVY 1915
Query: 1600 YVKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMVDKLKLTELIKIKEKLSPSDLD 1659
K L+ +F + + HV R +N EAN+LAQ ASGY P D
Sbjct: 1916 NEKCQELMQEFRLVTLKHVSREQNIEANDLAQGASGY-----------------KPMIKD 1958
Query: 1660 IMC-IDNLTSNDWRKPIVEYLQNPVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLS 1718
+ I +T+ DWR + +YL NP S RK++Y+AL YT+L +EL+ + I+G LLKCLS
Sbjct: 1959 VKAEIAAITAGDWRYDVHQYLHNPSQSASRKLRYKALKYTLLDDELYYRTIDGVLLKCLS 2018
Query: 1719 ENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQ 1778
+ A +A+ H+G+CG HQ+ KMKW+L R +WPT+++DC +Y +GCQDCQK IQ
Sbjct: 2019 ADQAKVAIGEVHEGICGTHQSAHKMKWLLRRARYFWPTMLEDCFKYYKGCQDCQKFGAIQ 2078
Query: 1779 HVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQE 1838
PAS ++ IIKPWPFRGW ID+IG I SSK HK+I+VA DYFTK VEAIPL+ V
Sbjct: 2079 RAPASAMNPIIKPWPFRGWGIDMIGMISRPSSKGHKFILVATDYFTKLVEAIPLKKVDSG 2138
Query: 1839 TVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEA 1898
I+F+Q HI+YRFG+P++ITTDQG++FV + F +S GIKLL S+PYYAQANGQ EA
Sbjct: 2139 DAIQFVQEHIIYRFGIPQTITTDQGSIFVSDEFVQFVDSMGIKLLNSSPYYAQANGQAEA 2198
Query: 1899 ANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYL 1958
+NK LI LIK+ P++WH L + LW+YR + + P++L YG EA+LP EV +
Sbjct: 2199 SNKSLIKLIKRKNSDYPRQWHTRLDEALWSYRMACHGSIQVPPYKLVYGHEAILPWEVRI 2258
Query: 1959 QSCRIQ 1964
S R +
Sbjct: 2259 GSRRTE 2264
>gb|AAK53848.1| Putative retroelement [Oryza sativa]
Length = 2014
Score = 970 bits (2507), Expect = 0.0
Identities = 489/966 (50%), Positives = 653/966 (66%), Gaps = 51/966 (5%)
Query: 1000 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELK 1059
D LEE+D+G PE + ++++LLKEF+DCFAW+Y EMPGL+R +VE +
Sbjct: 1081 DDLEEIDIG---------------PEFRAKLIELLKEFRDCFAWEYYEMPGLSRSIVEHR 1125
Query: 1060 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKM 1119
LPIK +P +Q RR D+L +K EI+RL FIR RY +W++++VPVIKKN
Sbjct: 1126 LPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPCRYAEWVSSIVPVIKKN--- 1182
Query: 1120 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAF 1179
DLN ATPKDEY MP+A+ +VD+A+G++ LS +DG GYNQIF+A ED+ KTAF
Sbjct: 1183 -------DLNKATPKDEYPMPVADQLVDAASGNKILSYMDGNVGYNQIFMAEEDIHKTAF 1235
Query: 1180 RCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1239
RCPGA+G +EWVVM FGLK+AG ATYQR MN I+HD I ++VYIDD+VVKS
Sbjct: 1236 RCPGAIGLFEWVVMTFGLKSAG-----ATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEI 1290
Query: 1240 DGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSP 1299
+ H+ LRK FER RKYGLKMNP KCAFGV G FLGF+VH++GIE+ + AI P
Sbjct: 1291 EDHIADLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIEVTQRSVNAIKKIQP 1350
Query: 1300 PTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYL 1359
P +K +LQ + GKINF+RRFI+ LS + + F+ LLRLK + F W AE QKA D++K+YL
Sbjct: 1351 PENKTELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWGAEQQKALDDIKEYL 1410
Query: 1360 SNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIE 1419
S+PPV+IPP KG +LY+SA +++IGS+L QE DG ER +FYLSR L ETRY+ +E
Sbjct: 1411 SSPPVLIPPQKGISFRLYLSAGEKSIGSVLIQE-LDGKERVVFYLSRRLLDVETRYSPME 1469
Query: 1420 KLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAP 1479
KLCLCLYFSC +L +Y+ + V D+IK+MLS PIL R+GKW +LTE+ L Y
Sbjct: 1470 KLCLCLYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGKWIFSLTEFDLRYES 1529
Query: 1480 LKAIKGQAVADFLADHTVPKEIVTYVGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKF 1539
KAIKGQA+ADF+ +H + + V + PW +FDGS + GIG+ I+SP+GA KF
Sbjct: 1530 PKAIKGQAIADFIVEHC--DDSIGSVEVVPWTSFFDGSVCTHDCGIGLVIISPRGACFKF 1587
Query: 1540 KFRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKY 1599
+ I+ +NN+ EYEA++ GL++L + A + I GDS LV+ QL EY+C S+ L Y
Sbjct: 1588 AYTIKPYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDSLLVISQLAGEYECKSDTLMVY 1647
Query: 1600 YVKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMVDKLKLTELIKIKEKLSPSDLD 1659
K L+ +F + + HV R +N EAN+LAQ ASGY P D
Sbjct: 1648 NEKCQELMQEFRLVTLKHVSREQNIEANDLAQGASGY-----------------KPMIKD 1690
Query: 1660 IMC-IDNLTSNDWRKPIVEYLQNPVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLS 1718
+ I +T+ DWR + +YL NP S RK++Y+AL YT+L +EL+ + I+G LLKCLS
Sbjct: 1691 VKAEIAAITAGDWRYDVHQYLHNPSQSASRKLRYKALKYTLLDDELYYRTIDGVLLKCLS 1750
Query: 1719 ENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQ 1778
+ A +A+ H+G+CG HQ+ KMKW+L R +WPT+++DC +Y +GCQDCQK IQ
Sbjct: 1751 ADQAKVAIGEVHEGICGTHQSAHKMKWLLRRARYFWPTMLEDCFKYYKGCQDCQKFGAIQ 1810
Query: 1779 HVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQE 1838
PAS ++ IIKPWPFRGW ID+IG I SSK HK+I+VA DYFTK VEAIPL+ V
Sbjct: 1811 RAPASAMNPIIKPWPFRGWGIDMIGMISRPSSKGHKFILVATDYFTKLVEAIPLKKVDSG 1870
Query: 1839 TVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEA 1898
I+F+Q HI+YRFG+P++ITTDQG++FV + F +S GIKLL S+PYYAQANGQ EA
Sbjct: 1871 DAIQFVQEHIIYRFGIPQTITTDQGSIFVSDEFVQFVDSMGIKLLNSSPYYAQANGQAEA 1930
Query: 1899 ANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYL 1958
+NK LI LIK+ P++WH L + LW+YR + + P++L YG EA+LP EV +
Sbjct: 1931 SNKSLIKLIKRKNSDYPRQWHTRLDEALWSYRMACHGSIQVPPYKLVYGHEAILPWEVRI 1990
Query: 1959 QSCRIQ 1964
S R +
Sbjct: 1991 GSRRTE 1996
>gb|AAT93841.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1739
Score = 969 bits (2505), Expect = 0.0
Identities = 486/938 (51%), Positives = 653/938 (68%), Gaps = 44/938 (4%)
Query: 974 LDCIYDDGPLGFEKPVVDVSKKMEAQDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKL 1033
+D +YD GF M A D LEE+D+G G RPT++S + PE + ++++L
Sbjct: 844 VDDLYDKQGQGF----------MSADD-LEEIDIGPGDRPRPTFVSKNLSPEFRTKLIEL 892
Query: 1034 LKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLK 1093
LKEF+DCFAW+Y EMPGL+R +VE +LPIK +P +Q RR D+L +K EI+RL
Sbjct: 893 LKEFRDCFAWEYYEMPGLSRSIVEHRLPIKPGVRPRQQPLRRCKADMLEPVKAEIKRLYD 952
Query: 1094 CKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHE 1153
FIR RY W++++VPVIKKNGK+RVCIDFRDLN ATPKDEY MP+A+ +VD+A+G++
Sbjct: 953 AGFIRPWRYAKWVSSIVPVIKKNGKVRVCIDFRDLNKATPKDEYPMPVADQLVDAASGNK 1012
Query: 1154 YLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVM 1213
LS +DG +GYNQIF+A ED+ KTAFRCP A+G +EWVVM FGLK+AG ATYQR M
Sbjct: 1013 ILSFMDGNAGYNQIFMAEEDIHKTAFRCPCAIGLFEWVVMTFGLKSAG-----ATYQRAM 1067
Query: 1214 NTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGD 1273
N I+HD I + VYIDD+VVKS + + LRK FER RKYGLKMNP KCAFGV AG
Sbjct: 1068 NYIYHDLIGWLVGVYIDDVVVKSKEIEDQIADLRKVFERTRKYGLKMNPTKCAFGVSAGQ 1127
Query: 1274 FLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSL 1333
FLGF+VH +GIE+ + AI PP +K +LQ ++GKI+F+RRFI+NLS + + F+ L
Sbjct: 1128 FLGFLVHDRGIEVTQRSVNAIKKIQPPENKTELQEMIGKIHFVRRFISNLSGRLEPFTPL 1187
Query: 1334 LRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQED 1393
LRLK + F AE QKA D++K+YLS+PPV+IPP KG P +LY+SA +++IGS+L QE
Sbjct: 1188 LRLKADQQFTCGAEQQKALDDIKEYLSSPPVLIPPHKGIPFRLYLSAGEKSIGSVLIQEL 1247
Query: 1394 EDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHM 1453
E G ER +FYLSR L AETRY+ ++KLCLCLYFSC +L++Y+ + V D++K+M
Sbjct: 1248 E-GKERVVFYLSRRLLNAETRYSPVKKLCLCLYFSCTRLRHYLLSNECTVICKADVVKYM 1306
Query: 1454 LSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTYVGIQPWKLY 1513
LS PIL R+GKW +LTE+ Y KAIKGQA+ADF+ +H + + V I PW L+
Sbjct: 1307 LSAPILKGRVGKWIFSLTEFDHRYESPKAIKGQAIADFIVEHR--DDSIGSVEIVPWTLF 1364
Query: 1514 FDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGAKNVV 1573
FDGS +G GIG+ I+SP+GA +F + I+ +NN+ EYEA++ GL++L + A +
Sbjct: 1365 FDGSVCTHGCGIGLVIISPRGACFEFAYTIKPYATNNQAEYEAVLKGLQLLKKVEADTIE 1424
Query: 1574 IKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANELAQIA 1633
I GDS LV+ QL EY+C ++ L Y K L+ +F R++N EAN+LAQ A
Sbjct: 1425 IMGDSLLVISQLAGEYECKNDTLMVYNEKCQELMKEF---------RLQNIEANDLAQGA 1475
Query: 1634 SGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVEYLQNPVGSTDRKVKYR 1693
SGY + + +K++ I +T++DWR + YL NP S RK++Y+
Sbjct: 1476 SGYK----PMIKDVKVE------------IAAMTADDWRYDVHRYLSNPSQSASRKLRYK 1519
Query: 1694 ALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLY 1753
AL YT+L +EL+ + I+G LLKCLS + A +A+ H+G+CG HQ+ KMKW+L R G +
Sbjct: 1520 ALKYTLLDDELYYRTIDGVLLKCLSADQAKVAIGEVHEGICGTHQSAHKMKWLLRRAGYF 1579
Query: 1754 WPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQH 1813
W T+++DC Y +GCQDCQK IQ PAS ++ IIKPWPFRGW ID+IG I+P SSK H
Sbjct: 1580 WSTMLEDCFRYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMINPPSSKGH 1639
Query: 1814 KYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAA 1873
K+I+VA DYFTKWVEAIPL+ V I+F+Q +I+YRFG P++ITTDQG++FV +
Sbjct: 1640 KFILVATDYFTKWVEAIPLKKVDSGDAIQFVQEYIIYRFGTPQTITTDQGSIFVSDEFVQ 1699
Query: 1874 FAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHV 1911
FA+S IKLL S+PYYAQANGQ EA+NK LI LIK+ +
Sbjct: 1700 FADSMSIKLLNSSPYYAQANGQAEASNKSLIKLIKRKI 1737
>ref|XP_462939.1| putative gag-pol protein [Oryza sativa (japonica cultivar-group)]
Length = 2273
Score = 966 bits (2498), Expect = 0.0
Identities = 486/966 (50%), Positives = 650/966 (66%), Gaps = 45/966 (4%)
Query: 1000 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELK 1059
D LEE+D+G G RPT+I + E + ++++LLKEF+DCFAW+Y EMPGL+R +VE +
Sbjct: 1334 DDLEEIDIGPGDRPRPTFICKNLSTEFRAKLIELLKEFRDCFAWEYYEMPGLSRSIVEHR 1393
Query: 1060 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKM 1119
LPIK +P +Q RR D+L +K EI+RL FIR RY +W++N
Sbjct: 1394 LPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPCRYAEWVSN----------- 1442
Query: 1120 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAF 1179
LN ATPKDEY MP+A+ +VD+A+G++ LS +DG GYNQIF+A ED+ KTAF
Sbjct: 1443 --------LNKATPKDEYPMPVADQLVDAASGNKILSYMDGNVGYNQIFMAEEDIHKTAF 1494
Query: 1180 RCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1239
RCPGA+G +EWVVM FGLK+AG ATYQR MN I+HD I ++VYIDD+VVKS
Sbjct: 1495 RCPGAIGLFEWVVMTFGLKSAG-----ATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEI 1549
Query: 1240 DGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSP 1299
+ H+ LRK FER RKYGLKMNP KCAFGV G FLGF+VH++GIE+ + AI P
Sbjct: 1550 EDHIADLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIEVTQRSVNAIKKIQP 1609
Query: 1300 PTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYL 1359
P +K +LQ + GKINF+RRFI+ LS + + F+ LLRLK + F W AE QKA D++K+YL
Sbjct: 1610 PENKTELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWGAEQQKALDDIKEYL 1669
Query: 1360 SNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIE 1419
S+PPV+IPP KG +LY+SA +++IGS+L QE DG ER +FYLSR L ETRY+ +E
Sbjct: 1670 SSPPVLIPPQKGISFRLYLSAGEKSIGSVLIQE-LDGKERVVFYLSRRLLDVETRYSPME 1728
Query: 1420 KLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAP 1479
KLCLCLYFSC +L +Y+ + V D+IK+MLS PIL R+GKW +LTE+ L Y
Sbjct: 1729 KLCLCLYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGKWIFSLTEFDLRYES 1788
Query: 1480 LKAIKGQAVADFLADHTVPKEIVTYVGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKF 1539
KAIKGQA+ADF+ +H + + V + PW +FDGS + GIG+ I+SP+GA KF
Sbjct: 1789 PKAIKGQAIADFIVEHC--DDSIGSVEVVPWTSFFDGSVCTHDCGIGLVIISPRGACFKF 1846
Query: 1540 KFRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKY 1599
+ I+ +NN+ EYEA++ GL++L + A + I GDS LV+ QL EY+C S+ L Y
Sbjct: 1847 AYTIKPYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDSLLVISQLAGEYECKSDTLMVY 1906
Query: 1600 YVKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMVDKLKLTELIKIKEKLSPSDLD 1659
K L+ +F + + HV R +N EAN+LAQ ASGY P D
Sbjct: 1907 NEKCQELMQEFRLVTLKHVSREQNIEANDLAQGASGY-----------------KPMIKD 1949
Query: 1660 IMC-IDNLTSNDWRKPIVEYLQNPVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLS 1718
+ I +T+ DWR + +YL NP S RK++Y+AL YT+L +EL+ + I+G LLKCLS
Sbjct: 1950 VKAEIAAITAGDWRYDVHQYLHNPSQSASRKLRYKALKYTLLDDELYYRTIDGVLLKCLS 2009
Query: 1719 ENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQ 1778
+ A +A+ H+G+CG HQ+ KMKW+L R +WPT+++DC +Y +GCQDCQK IQ
Sbjct: 2010 ADQAKVAIGEVHEGICGTHQSAHKMKWLLRRARYFWPTMLEDCFKYYKGCQDCQKFGAIQ 2069
Query: 1779 HVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQE 1838
PAS ++ IIKPWPFRGW ID+IG I SSK HK+I+VA DYFTK VEAIPL+ V
Sbjct: 2070 RAPASAMNPIIKPWPFRGWGIDMIGMISRPSSKGHKFILVATDYFTKLVEAIPLKKVDSG 2129
Query: 1839 TVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEA 1898
I+F+Q HI+YRFG+P++ITTDQG++FV + F +S GIKLL S+PYYAQANGQ EA
Sbjct: 2130 DAIQFVQEHIIYRFGIPQTITTDQGSIFVSDEFVQFVDSMGIKLLNSSPYYAQANGQAEA 2189
Query: 1899 ANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYL 1958
+NK LI LIK+ P++WH L + LW+YR + + P++L YG EA+LP EV +
Sbjct: 2190 SNKSLIKLIKRKNSDYPRQWHTRLDEALWSYRMACHGSIQVPPYKLVYGHEAILPWEVRI 2249
Query: 1959 QSCRIQ 1964
S R +
Sbjct: 2250 GSRRTE 2255
>ref|XP_474893.1| OSJNBa0021F22.20 [Oryza sativa (japonica cultivar-group)]
gi|21743195|emb|CAD40050.1| OSJNBa0085C10.2 [Oryza sativa
(japonica cultivar-group)] gi|38346487|emb|CAE03726.2|
OSJNBa0021F22.20 [Oryza sativa (japonica cultivar-group)]
Length = 861
Score = 904 bits (2336), Expect = 0.0
Identities = 443/858 (51%), Positives = 605/858 (69%), Gaps = 24/858 (2%)
Query: 1010 GLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPV 1069
G + RPT+IS + E + ++++LLKEF+DCFAW+Y EMPGL+R +VE +LPIK +P
Sbjct: 26 GDQPRPTFISKNLSSEFRTKLIELLKEFRDCFAWEYYEMPGLSRSIVEHRLPIKPGVRPR 85
Query: 1070 KQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLN 1129
+Q PRR D+L +K EI+RL FI RY +W++++VPVIKKNGK+RVCIDFRDLN
Sbjct: 86 QQPPRRCKADMLEPVKAEIKRLYDAGFIHPYRYAEWVSSIVPVIKKNGKVRVCIDFRDLN 145
Query: 1130 AATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYE 1189
ATPKDEY MP+A+ + D+A+G++ LS +D +GYNQIF+A ED+ KTAFRCP A+G +E
Sbjct: 146 KATPKDEYPMPVADQLFDAASGNKILSFMDRNTGYNQIFMAEEDIHKTAFRCPSAIGLFE 205
Query: 1190 WVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKS 1249
WVVM F LK+AG ATYQR MN I+HD I + ++V IDD+VVKS D H+ LRK
Sbjct: 206 WVVMTFDLKSAG-----ATYQRAMNYIYHDLIGSLVEVDIDDVVVKSKEVDDHIADLRKV 260
Query: 1250 FERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSL 1309
FER RKYGLKMNP KCAFGV A FLGF+VH++GIE+ + AI PP +K +LQ +
Sbjct: 261 FERTRKYGLKMNPTKCAFGVSARQFLGFLVHERGIEVTQRSVNAIQKIKPPENKTELQEM 320
Query: 1310 LGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPI 1369
+GKINF+RRFI+NLS + + F+ LLRLK + F W AE QKA D +K+YLS+PPV+IPP
Sbjct: 321 IGKINFVRRFISNLSGRLEPFTPLLRLKADQQFTWGAEQQKALDNIKEYLSSPPVLIPPQ 380
Query: 1370 KGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFSC 1429
KG P +LY+SA +++ GS+L QE E G ER +FY+SR L AETRY+ +EKLCLCLYF C
Sbjct: 381 KGIPFRLYLSAGEKSNGSVLIQELE-GKERVVFYISRRLLDAETRYSPMEKLCLCLYFLC 439
Query: 1430 VKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVA 1489
+L++Y+ + V D++++MLS PIL +IG W +LTE+ L Y KAIKGQA+A
Sbjct: 440 TRLRHYLLSNECTVICKADVVRYMLSAPILKGQIGNWIFSLTEFDLRYESPKAIKGQAIA 499
Query: 1490 DFLADHTVPKEIVTYVGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSN 1549
DF+ +H + + V I PW L+FDGS +G GIG+ I+SP+GA +F + I+ +N
Sbjct: 500 DFIVEHR--DDSIGSVEIVPWTLFFDGSVCTHGCGIGLVIISPRGACFEFAYTIKSYATN 557
Query: 1550 NEVEYEALISGLEILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAK 1609
N+ EYEA++ GL++L + A + I GDS LV+ QL EY+C S+ Y K L+ +
Sbjct: 558 NQAEYEAVLKGLQLLKEVEANIIEIMGDSLLVISQLAGEYECKSDTFIVYNEKCLELMKE 617
Query: 1610 FNEIGIGHVPRIENQEANELAQIASGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSN 1669
F + + HV R +N EAN+LAQ ASGY + + +K++ I ++++
Sbjct: 618 FRLVTLKHVSREQNLEANDLAQGASGYK----PMIKDVKVE------------IAAMSAD 661
Query: 1670 DWRKPIVEYLQNPVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAA 1729
DWR + +YL NP S RK++Y+AL YT+LG+EL+ + I+G LLKCLS + A +A+
Sbjct: 662 DWRYDVHQYLSNPSQSASRKLRYKALKYTLLGDELYYRVIDGVLLKCLSADQAKVAIGEV 721
Query: 1730 HDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSII 1789
H+G+CG HQ+ KMKW+L R G +WPT+++ Y +GCQDCQK IQ PAS ++ II
Sbjct: 722 HEGICGTHQSAHKMKWLLRRTGYFWPTMLEVYFRYYKGCQDCQKFGAIQRAPASAMNPII 781
Query: 1790 KPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIV 1849
KPWPFRGW ID+IG I+P SSK HK+I+VA DYFTKWVEAIPL+ V I+F+Q HI+
Sbjct: 782 KPWPFRGWEIDMIGMINPPSSKGHKFILVATDYFTKWVEAIPLKKVDSGDAIQFVQEHII 841
Query: 1850 YRFGLPESITTDQGTVFV 1867
Y+FGLP++IT DQG++F+
Sbjct: 842 YQFGLPQTITMDQGSIFM 859
>ref|XP_493776.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
gi|9927274|dbj|BAB08213.2| Similar to Arabidopsis
thaliana chromosome II BAC F26H6; putative retroelement
pol polyprotein (AC006920) [Oryza sativa (japonica
cultivar-group)]
Length = 2876
Score = 862 bits (2228), Expect = 0.0
Identities = 459/1115 (41%), Positives = 666/1115 (59%), Gaps = 50/1115 (4%)
Query: 1000 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELK 1059
D L E++LG + RP ++S ++ E ++ L EF+DCFAW Y EMPGL+ + K
Sbjct: 1776 DELVEMNLGTEDDPRPIFVSGMLTEEEREDYRSFLMEFRDCFAWTYKEMPGLDSRVATHK 1835
Query: 1060 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKM 1119
L I +PVKQ PRR P+ ++ E++RL+ FI+ +Y WLAN+VPV KKNG++
Sbjct: 1836 LAIDPQFRPVKQPPRRLRPEFQDQVIAEVDRLINVGFIKEIQYPRWLANIVPVEKKNGQV 1895
Query: 1120 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAF 1179
RVC+DFRDLN A PKD++ +PI EM+VDS G+ LS GYNQI + D TAF
Sbjct: 1896 RVCVDFRDLNRACPKDDFPLPITEMVVDSTTGYGALS------GYNQIKMDLLDAFDTAF 1949
Query: 1180 RCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1239
R P G + + VMPFGLKNAG ATYQR M + D I ++ Y+DD+VVK+
Sbjct: 1950 RTPK--GNFYYTVMPFGLKNAG-----ATYQRAMQFVLDDLIHHSVECYVDDMVVKTKDH 2002
Query: 1240 DGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSP 1299
+ H LR FER+R++ LKMNPLKCAF V +G FLGFV+ +GIEI K KAIL+ P
Sbjct: 2003 EHHQEDLRIVFERLRRHQLKMNPLKCAFAVQSGVFLGFVIRHRGIEIEPKKIKAILNMPP 2062
Query: 1300 PTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYL 1359
P K L+ L GK+ ++RRFI+NLS + + FS L+ KK F W+ E Q FD +K+YL
Sbjct: 2063 PQELKDLRKLQGKLAYIRRFISNLSGRIQPFSKLM--KKGTPFVWDEECQNGFDSIKRYL 2120
Query: 1360 SNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIE 1419
NPPV+ P+KGRP+ LYI+ +IG++LAQ +++G E A +YLSR + GAE Y+ IE
Sbjct: 2121 LNPPVLAAPVKGRPLILYIATQPASIGALLAQHNDEGKEVACYYLSRTMVGAEQNYSPIE 2180
Query: 1420 KLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAP 1479
KLCL L F+ KL++Y+ + + + D I+++LS+P+L R+GKWAL + EY +T+ P
Sbjct: 2181 KLCLALIFALKKLRHYMLAHQIQLIARADPIRYVLSQPVLTGRLGKWALLMMEYDITFVP 2240
Query: 1480 LKAIKGQAVADFLADHTVP-----------KEIVTYVGIQPWKLYFDGSSHKN----GT- 1523
KAIKGQA+A+FLA H +P +EI T + W+LYFDG+S K+ GT
Sbjct: 2241 QKAIKGQALAEFLATHPMPDDSPLIANLPDEEIFTAELQEQWELYFDGASRKDINPDGTP 2300
Query: 1524 ----GIGMFIMSPQGAPTKFKFRI-EGNCSNNEVEYEALISGLEILLALGAKNVVIKGDS 1578
G G+ +PQG F + + CSNNE EYEALI GL + L++ +++ GDS
Sbjct: 2301 RRRAGAGLVFKTPQGGVIYHSFSLLKEECSNNEAEYEALIFGLLLALSMEVRSLRAHGDS 2360
Query: 1579 ELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMV 1638
L+++Q+ Y+ L YY A L+ KF I + HVPR +N A+ LA++A+ +
Sbjct: 2361 RLIIRQINNIYEVRKPELVPYYTVARRLMDKFEHIEVIHVPRSKNAPADALAKLAAALVF 2420
Query: 1639 DKLKLTELIKIKEK--------LSPSDLDIMCIDNLTSNDWRKPIVEYLQN---PVGSTD 1687
+++ ++E+ L P +++I+ ++ DWR+P ++Y ++ P +
Sbjct: 2421 QGDNPAQIV-VEERWLLPAVLELIPEEVNIIITNSAEEEDWRQPFLDYFKHGSLPEDPVE 2479
Query: 1688 RKVKYRAL-SYTILGNELFKKNING-TLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMKW 1745
R+ R L SY L+K++ LL+C+ ++A + H G+CG HQ+G KM
Sbjct: 2480 RRQLQRRLPSYIYKAGVLYKRSYGQEVLLRCVDRSEANRVLQEVHHGVCGGHQSGPKMYH 2539
Query: 1746 ILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGEI 1805
+ G YWP IM DC++ A+ C CQ H +H P + LH + WPF W ID+IG I
Sbjct: 2540 SIRLVGYYWPGIMADCLKTAKTCHGCQIHDNFKHQPPAPLHPTVPSWPFDAWGIDVIGLI 2599
Query: 1806 HPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGTV 1865
+P SS+ H++I+ A DYF+KW EA+PL+ V VI F++ HI+YRFG+P IT+D
Sbjct: 2600 NPPSSRGHRFILTATDYFSKWAEAVPLREVKSSDVINFLERHIIYRFGVPHRITSDNAKA 2659
Query: 1866 FVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHESLSQV 1925
F +K+ F E + IK ST YY QANG EA NK L ++KK V + + WH+ L +
Sbjct: 2660 FKSQKIYRFMEKYKIKWNYSTGYYPQANGMAEAFNKTLGKILKKTVDKHRRDWHDRLYEA 2719
Query: 1926 LWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYWNMMLDEMVNL 1985
LWAYR + T TP+ L YG EAVLP E+ L S R+ +E+ + + E+ +
Sbjct: 2720 LWAYRVTVRTPTQATPYSLVYGNEAVLPLEIQLPSLRVAIHDELTKDEQIRLRFQELDAV 2779
Query: 1986 DEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILPMDKKDRVYGKWTPNWEG 2045
+EERL AL L + + +AY+K V+ R F G+ V + P+ ++ GK+ P WEG
Sbjct: 2780 EEERLGALQNLELYRQNMVRAYDKLVKQRVFRKGELVLVLRRPIVVTHKMKGKFEPKWEG 2839
Query: 2046 PFTVEKTLPNNAYVIKELGNQRQCVTINGKYLKAY 2080
P+ +E+ AY + + + ING++LK Y
Sbjct: 2840 PYVIEQAYDGGAYQLIDHQGSQPMPPINGRFLKKY 2874
>ref|XP_474437.1| OSJNBa0070M12.15 [Oryza sativa (japonica cultivar-group)]
gi|38345832|emb|CAE01862.2| OSJNBa0070M12.15 [Oryza
sativa (japonica cultivar-group)]
Length = 1685
Score = 859 bits (2219), Expect = 0.0
Identities = 423/804 (52%), Positives = 567/804 (69%), Gaps = 24/804 (2%)
Query: 1000 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELK 1059
D L+EVD+G G RPT+IS + E + ++++LLKEF+DCFAW+Y EMPGL+R +VE +
Sbjct: 904 DDLDEVDIGPGDRPRPTFISKNLSSEFRTKLMELLKEFRDCFAWEYYEMPGLSRSIVEHR 963
Query: 1060 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKM 1119
LP+K +P +Q PRR D+L +K EI+RL FI RY +W++++VPVIKKNGK+
Sbjct: 964 LPLKPGIRPHQQPPRRCKADMLEPVKAEIKRLYDAGFIHPCRYAEWVSSIVPVIKKNGKV 1023
Query: 1120 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAF 1179
RVCIDFRDLN ATPKDEY MP+A+ +VD+A+GH+ LS +DG +GYNQIF+ ED+ KT F
Sbjct: 1024 RVCIDFRDLNKATPKDEYPMPVADQLVDAASGHKILSFMDGNAGYNQIFMTEEDIHKTVF 1083
Query: 1180 RCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1239
RCPGA+G +EWVVM F GLK+AGATYQR MN I+HD I ++VYIDD+VVKS
Sbjct: 1084 RCPGAIGLFEWVVMTF-----GLKSAGATYQRAMNYIYHDLIGWLVEVYIDDVVVKSREI 1138
Query: 1240 DGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSP 1299
+ H+ LRK FER RKYGLKMNP KCAFGV AG FLGF+VH++GIEI + AI P
Sbjct: 1139 EDHIADLRKVFERTRKYGLKMNPTKCAFGVSAGQFLGFLVHERGIEIPQRSINAIKKIKP 1198
Query: 1300 PTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYL 1359
P +K +LQ ++GKINF+RRFI+NLS K + F+ LLRL+ + F W AE QKA D +K+YL
Sbjct: 1199 PGNKTELQEMIGKINFVRRFISNLSGKLEPFTPLLRLRADQQFTWGAEQQKALDNIKEYL 1258
Query: 1360 SNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIE 1419
S+PPV+IPP KG P LY+SA D++IGS+L QE E G ER +FYLSR L AETRY+ +E
Sbjct: 1259 SSPPVLIPPQKGIPFWLYLSAGDKSIGSVLIQELE-GKERVVFYLSRRLLDAETRYSPVE 1317
Query: 1420 KLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAP 1479
KLCLCLYFSC KL++Y+ + V D+IK+MLS PIL R+GKW +LTE+ L +
Sbjct: 1318 KLCLCLYFSCTKLRHYLLSNECTVVCKADVIKYMLSAPILKVRVGKWIFSLTEFDLRHES 1377
Query: 1480 LKAIKGQAVADFLADHTVPKEIVTYVGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKF 1539
KAIKGQAVADF+ H + V I PW L+FDGS +G GIG+ I++P GA +F
Sbjct: 1378 PKAIKGQAVADFIVGHR--DGSIGLVDIMPWVLFFDGSVCSHGYGIGLVIIAPWGASFEF 1435
Query: 1540 KFRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKY 1599
+ I + +NN+ EYEA++ GL++L + A + I GDS LV+ QL EY+C + L Y
Sbjct: 1436 AYAITHHVTNNQAEYEAVLKGLQLLQEVAADAIEIMGDSLLVISQLAGEYECKDDTLMVY 1495
Query: 1600 YVKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMVDKLKLTELIKIKEKLSPSDLD 1659
Y K L+++F + + H+ R +N EAN+L+Q ASGY +T+ I+I+
Sbjct: 1496 YEKCRTLISEFRLVTLRHISREQNIEANDLSQGASGYK----PMTKDIEIE--------- 1542
Query: 1660 IMCIDNLTSNDWRKPIVEYLQNPVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLSE 1719
I +T++DWR + +YLQNP S RK++Y+AL YT+L ++L+ + I+G LLKCLS
Sbjct: 1543 ---IATITADDWRYDVFQYLQNPSQSALRKLRYKALKYTLLDDDLYYRTIDGVLLKCLSA 1599
Query: 1720 NDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQH 1779
+ A +A+ H+G+CG HQ+ KMKW+L R G +WPT+++DC Y +GCQDCQK IQ
Sbjct: 1600 DQAKVAIGEIHEGICGTHQSAHKMKWLLRRAGYFWPTMLEDCFRYYKGCQDCQKFGAIQR 1659
Query: 1780 VPASELHSIIKPWPFRGWAIDLIG 1803
PAS ++ II+PWPFRGW ID+IG
Sbjct: 1660 APASAMNPIIQPWPFRGWGIDMIG 1683
>gb|AAM74416.1| Putative retroelement [Oryza sativa (japonica cultivar-group)]
Length = 1781
Score = 814 bits (2103), Expect = 0.0
Identities = 440/1036 (42%), Positives = 621/1036 (59%), Gaps = 126/1036 (12%)
Query: 974 LDCIYDDGPLG----FEKPVVDVSKKMEAQ-----DPLEEVDLGDGLEKRPTYISALIDP 1024
+DC+ G + + K VD + Q D LEE+D+G G RPT+IS + P
Sbjct: 860 IDCLSGKGVVEGSNVYTKDTVDDLDDKQGQGFMSADDLEEIDIGPGDRPRPTFISKNLSP 919
Query: 1025 ELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKI 1084
E + ++++LLKE++DCFAW+Y EMPGL+R +VE +LPIK +P +Q PRR D+L +
Sbjct: 920 EFRTKLIELLKEYRDCFAWEYYEMPGLSRSIVEHRLPIKTGVRPYQQPPRRCKADMLEAV 979
Query: 1085 KEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEM 1144
K E+ RL FIR RY +W++N+VPVIKKNGK+RVCIDFRDLN AT KDEY M +A+
Sbjct: 980 KAEVARLYDADFIRPCRYAEWVSNMVPVIKKNGKVRVCIDFRDLNKATLKDEYPMSVADQ 1039
Query: 1145 MVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKN 1204
+VD+A+GH+ L+ +DG +GYNQIF+ ED+ KTAFRCPGA+G +EWVVM FGLK AG
Sbjct: 1040 LVDAASGHKILNFMDGNAGYNQIFMVEEDIHKTAFRCPGAIGLFEWVVMTFGLKRAG--- 1096
Query: 1205 AGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLK 1264
ATYQR MN IFHD I ++VY+DD+VVKS + H+ +LRK FER RKYGLKMNP K
Sbjct: 1097 --ATYQRAMNYIFHDLIGWLVEVYVDDVVVKSKEIENHIANLRKVFERTRKYGLKMNPTK 1154
Query: 1265 CAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLS 1324
CAFGV A FLGF+VH++GIE+ + AI PP KK+LQ L+GK+NF+RRF +NLS
Sbjct: 1155 CAFGVSARQFLGFLVHERGIEVTQRSINAIKKIQPPEDKKKLQELIGKVNFVRRFNSNLS 1214
Query: 1325 EKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDET 1384
+ + F+ LLRLK + F W AE QKA D +K+YLS+PPV+IPP KG P +LY+SA
Sbjct: 1215 GRLEPFTLLLRLKADQEFTWGAEQQKALDNIKEYLSSPPVLIPPQKGIPFRLYLSA---- 1270
Query: 1385 IGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVF 1444
ETRY+ +EKLCLCLYFSC KL++Y + +
Sbjct: 1271 ---------------------------ETRYSPVEKLCLCLYFSCTKLRHYPLFNECTMI 1303
Query: 1445 SHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTY 1504
D++K+ML PIL +GKW ALTE+ L Y K IKGQA+ADF+ DH + +
Sbjct: 1304 CKADVVKYMLPAPILKGIVGKWIFALTEFDLRYESPKVIKGQAIADFIVDHR--DDSIGS 1361
Query: 1505 VGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEIL 1564
V I PW L+FDGS +G GIG+ I+SP+GA +F + I+ +NN+ EYEA++ GL++L
Sbjct: 1362 VDIVPWTLFFDGSVCTHGCGIGLVIISPRGASFEFAYTIKPYATNNQAEYEAVLKGLQLL 1421
Query: 1565 LALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQ 1624
+ A V I GDS LV+ QL +Y+C ++ L Y K L+ F + + HV R +N
Sbjct: 1422 KEVEADAVEIMGDSLLVISQLAGQYECKNDTLMVYNEKCRELMDGFRLVTLKHVSREQNI 1481
Query: 1625 EANELAQIASGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVEYLQNPVG 1684
EAN+LAQ ASGY +T+ IK++ I ++++DWR I +YLQNP
Sbjct: 1482 EANDLAQGASGYK----PMTKDIKVE------------IATISADDWRYDIFQYLQNPSQ 1525
Query: 1685 STDRKVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMK 1744
ST RK+ Y+AL YT+L +EL + I+G LLKCLS + A +A+ ++G+CG HQ+ KMK
Sbjct: 1526 STSRKLLYKALKYTLLDDELCYRTIDGVLLKCLSADQAKVAIGEVNEGICGTHQSAHKMK 1585
Query: 1745 WILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGE 1804
W+L + G +WPTI++DC Y +GCQDCQK IQ PAS ++SIIKP +
Sbjct: 1586 WLLRQAGYFWPTILEDCFRYYKGCQDCQKFGAIQRAPASAMNSIIKPMAIQ-------RP 1638
Query: 1805 IHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGT 1864
+ P ++ D F ++ + + I+ + + Y ++ ++++
Sbjct: 1639 LQPIRARSS-----CTDEFVQFADIMG---------IKLLNSSPYYAQANGQAESSNKSL 1684
Query: 1865 V-FVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHESLS 1923
+ + RK++ + W +L +
Sbjct: 1685 IKLIKRKISDYPRQWHTRL----------------------------------------A 1704
Query: 1924 QVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYWNMMLDEMV 1983
+ LW+YR + + P++L YG EAVLP EV + R + Q E+ ++ Y+N+M DE
Sbjct: 1705 EALWSYRMACHGSIQVPPYKLLYGHEAVLPWEVRIGLRRTELQNELTADEYYNLMADERE 1764
Query: 1984 NLDEERLLA-LDVLTR 1998
+L + RL L ++TR
Sbjct: 1765 DLVQSRLKGLLSIITR 1780
>gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 791 bits (2043), Expect = 0.0
Identities = 433/1164 (37%), Positives = 668/1164 (57%), Gaps = 59/1164 (5%)
Query: 950 DNTCIEENEQSVDECEFEDLRNLRLDCIYDDGPLGFEKPVVDVSKKMEAQDPLEEVDLGD 1009
DN + E+S ++CE +RL L E+ V+ Q+ LE V+LG
Sbjct: 1123 DNPIYQAEEESEEDCELP-AELVRL--------LKQEERVIQPH-----QEELEVVNLGT 1168
Query: 1010 GLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPV 1069
R I A ++ +K R++++L+E+ + FAW Y +MPGL+ ++V +LP++E V
Sbjct: 1169 EDATREIKIGAALEDSVKRRLIEMLREYVEIFAWSYQDMPGLDTDIVVHRLPLREGCPSV 1228
Query: 1070 KQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLN 1129
KQ RR PD+ KIKEE+++ F+ Y W+AN+VPV KK+GK+R+C+D+RDLN
Sbjct: 1229 KQKLRRTSPDMATKIKEEVQKQWDAGFLAVTSYPPWMANIVPVPKKDGKVRMCVDYRDLN 1288
Query: 1130 AATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYE 1189
A+PKD++ +P +++VD+ A S +DG+SGYNQI +A ED+ KT F P GT+
Sbjct: 1289 RASPKDDFPLPHIDVLVDNTAQSSVFSFMDGFSGYNQIKMAPEDMEKTTFITPW--GTFC 1346
Query: 1190 WVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKS 1249
+ VMPFGLKNAG ATYQR M T+FHD + ++VY+DD++ KS + + HL++L+K
Sbjct: 1347 YKVMPFGLKNAG-----ATYQRAMTTLFHDMMHKEIEVYVDDMIAKSQTEEEHLVNLQKL 1401
Query: 1250 FERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSL 1309
F+R+RK+ L++NP KC FGV +G LGF+V +KGIE++ K KAI + P ++KQ++
Sbjct: 1402 FDRLRKFKLRLNPNKCTFGVRSGKLLGFIVSEKGIEVDPAKVKAIQEMPEPKTEKQVRGF 1461
Query: 1310 LGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPI 1369
LG++N++ RFI++L+ + LLR K +W + QKAFD++K+YL PP++IPP+
Sbjct: 1462 LGRLNYIARFISHLTATCEPIFKLLR--KNQAIKWNDDCQKAFDKIKEYLQKPPILIPPV 1519
Query: 1370 KGRPMKLYISATDETIGSMLAQEDEDGN-ERAIFYLSRVLNGAETRYTMIEKLCLCLYFS 1428
GRP+ +Y+S T+ ++G +L + DE G E AI+YLS+ ETRY+++EK C L ++
Sbjct: 1520 PGRPLIMYLSVTENSMGCVLGRHDESGRKEHAIYYLSKKFTDCETRYSLLEKTCCALAWA 1579
Query: 1429 CVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAV 1488
+L+ Y+ ++ S D +K++ KP L R+ +W + LTEY + Y KAIKG +
Sbjct: 1580 ARRLRQYMLNHTTLLISKMDPVKYIFEKPALTGRVARWQMILTEYDIQYTSQKAIKGSIL 1639
Query: 1489 ADFLADHTV----------PKEIVTYVGIQP---------------WKLYFDGSSHKNGT 1523
+D+LA+ + P E + Y+ ++ W L FDG+ + NG
Sbjct: 1640 SDYLAEQPIEDYQPMMFEFPDEDIMYLKMKDCKEPLVEEGPDPDDKWTLMFDGAVNMNGN 1699
Query: 1524 GIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVK 1583
G+G +++P+GA F R+ + +NNE EYEA I G+E + L K + I GDS LVV
Sbjct: 1700 GVGAVLINPKGAHMPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKTLDIFGDSALVVN 1759
Query: 1584 QLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMVDKLKL 1643
Q+ ++ HL Y +L F ++ + HVPR ENQ A+ LA ++S V+
Sbjct: 1760 QVNGDWNTNQPHLIPYRDYTRRILTFFKKVKLYHVPRDENQMADALATLSSMIKVNWWNH 1819
Query: 1644 TELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVEYLQN---PVGST--DRKVKYRALSYT 1698
+ + P+ + + W I +L+ P G++ D+K R
Sbjct: 1820 VPHVAVNRLERPAYVFAAESVVIDEKPWYYDIKNFLKTQEYPEGASKNDKKTLRRLAGSF 1879
Query: 1699 ILGNE--LFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPT 1756
L + L+K+N + LL+C+ +A M + H+G G H G M L R G YW T
Sbjct: 1880 YLNQDDVLYKRNFDMVLLRCMDRPEADMLMQEVHEGSFGTHAGGHAMAKKLLRAGYYWMT 1939
Query: 1757 IMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQHKYI 1816
+ DC +YAR C CQ ++ HVP S L+ + PWPF W ID+IG+I P +S H++I
Sbjct: 1940 MESDCFKYARKCHKCQIYADRVHVPPSPLNVMNSPWPFAMWGIDMIGKIEPTASNGHRFI 1999
Query: 1817 IVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAAFAE 1876
+VA+DYFTKWVEA N+T++ V FI+ I+ R+G+PE I TD G+ + + +
Sbjct: 2000 LVAIDYFTKWVEAASYANITKQVVTRFIKKEIICRYGVPERIITDNGSNLNNKMMKELCK 2059
Query: 1877 SWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEA 1936
+ I+ S+PY + NG VEAANK + +++K V K WHE L L YR S +
Sbjct: 2060 DFKIEHHNSSPYRPKMNGAVEAANKNIKKIVRKMVVTY-KDWHEMLPFALHGYRTSVRTS 2118
Query: 1937 TGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYWNMMLDEMVNLDEERLLALDVL 1996
TG TP+ L YG EAVLP EV + S R+ ++ + +E+ ++E RL +
Sbjct: 2119 TGATPYSLVYGMEAVLPVEVEIPSLRVLLDVKLDEAEWIRTRFNELSLIEERRLAVVCHG 2178
Query: 1997 TRQKDRIAKAYNKKVRDRSFITGDYVWKVILPMDKKDRVYGKWTPNWEGPFTVEKTLPNN 2056
+ R+ +A+++KVR RS+ GD V K ILP +R GKWTPN+EGP+ V+K
Sbjct: 2179 QLYQRRMKRAFDQKVRPRSYQIGDLVLKRILPPGTDNR--GKWTPNYEGPYVVKKVFSGG 2236
Query: 2057 AYVIKELGNQRQCVTINGKYLKAY 2080
A ++ + + +N +K Y
Sbjct: 2237 ALMLTTMDGEDFPSPVNSDVVKKY 2260
>dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera]
Length = 1027
Score = 690 bits (1780), Expect = 0.0
Identities = 386/1041 (37%), Positives = 591/1041 (56%), Gaps = 24/1041 (2%)
Query: 1048 MPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLA 1107
M G++ + +L + +PV+Q RRFHPD I+ EI++LL+ FIR Y DWLA
Sbjct: 1 MKGIHPSITSHRLNVVSTARPVRQRIRRFHPDRQRVIRNEIDKLLEAGFIREVSYPDWLA 60
Query: 1108 NVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQI 1167
NVV V KK GK RVC+D+ +LN A PKD + +P + +VDS +G LS LD +SGY+QI
Sbjct: 61 NVVVVPKKEGKWRVCVDYTNLNNACPKDSFPLPRIDQIVDSTSGQGMLSFLDAFSGYHQI 120
Query: 1168 FIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQV 1227
++ +D K AF P L Y+ VMPFGLKNAG ATYQR+M IF I ++V
Sbjct: 121 PMSPDDEEKIAFITPHDLYCYK--VMPFGLKNAG-----ATYQRLMTKIFKPLIGHSVEV 173
Query: 1228 YIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEIN 1287
YIDDIVVKS +R+ H+LHL++ F +R+YG+K+NP KCAFGV A FLGF+V ++GIE++
Sbjct: 174 YIDDIVVKSKTREQHILHLQEVFYLLRRYGMKLNPSKCAFGVSARKFLGFMVSQRGIEVS 233
Query: 1288 KNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAE 1347
++ KA+++T PP +KK+LQ L GK+ L RFI ++ + F L ++K W
Sbjct: 234 PDQVKAVMETPPPRNKKELQRLTGKLVALGRFIARFIDELRPF--FLAIRKAGTHGWTDN 291
Query: 1348 HQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRV 1407
Q A + +K YL PP++ PI + +Y++ ++ I ++L + ++ I+Y+SR
Sbjct: 292 CQNALERIKHYLMQPPILSSPIPKEKLYMYLAVSEWAISAVLFRCPSPKEQKPIYYVSRA 351
Query: 1408 LNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWA 1467
L ETRY+ +E + L L + KL+ Y + V+V + ++++L KP L R+ +WA
Sbjct: 352 LADVETRYSKMELISLALRSAAQKLRPYFQAHPVIVLTDQP-LRNILHKPDLTGRMLQWA 410
Query: 1468 LALTEYSLTYAPLKAIKGQAVADFLADHT-VPKEIVTYVGIQPWKLYFDGSSHKNGTGIG 1526
+ L+E+ + + P ++KGQ +ADF+ +++ P + + W L DG+S +G+G+G
Sbjct: 411 IELSEFGIEFQPRLSMKGQVMADFVLEYSRKPGQHEGSRKKEWWTLRVDGASRSSGSGVG 470
Query: 1527 MFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVKQLT 1586
+ + SP G + R+ + SNNE EYEA++SGL++ LAL + I DS+LVVK +
Sbjct: 471 LLLQSPTGEHLEQAIRLGFSASNNEAEYEAILSGLDLALALSVSKLRIFSDSQLVVKHVQ 530
Query: 1587 KEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMVDKLKLTEL 1646
+EY+ +A+Y K N L +F E I + R +N+ A+ LA IA+ + + L
Sbjct: 531 EEYEAKDARMARYLAKVRNTLQQFTEWTIEKIKRADNRRADALAGIAASLSIKEAILLP- 589
Query: 1647 IKIKEKLSPSDLDIMCIDNLTSND---WRKPIVEYLQNPVGSTD----RKVKYRALSYTI 1699
I ++ S S++ I D W I EY++ D KV+ +A +T+
Sbjct: 590 IHVQTNPSVSEISICSTTEAPQADDQEWMNDITEYIRTGTLPGDPKQAHKVRVQAARFTL 649
Query: 1700 LGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMK 1759
+G L+K++ G L+CL ++A ++ H+G+ G H G + QG YWPT+ K
Sbjct: 650 IGGHLYKRSFTGPYLRCLGHSEAQYVLAELHEGIYGNHSGGRSLAHRAHSQGYYWPTMKK 709
Query: 1760 DCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQHKYIIVA 1819
+ Y + C CQ+++ I H+P++ L SI PWPF W +D++ + P + Q K+++VA
Sbjct: 710 EAAAYVKRCDKCQRYAPIPHMPSTTLKSISGPWPFAQWGMDIVRPL-PTAPAQKKFLLVA 768
Query: 1820 VDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAAFAESWG 1879
DYF+KWVEA + + V +F+ +I+ RFG+P++I D G F F
Sbjct: 769 TDYFSKWVEAEAYASTKDKDVTKFVWKNIICRFGIPQTIIADNGPQFDSIAFRNFCSELN 828
Query: 1880 IKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGT 1939
I+ STP Y Q+NGQ EA NK LI+ +KK + + +W E L VLWAYR +P TG
Sbjct: 829 IRNSYSTPRYPQSNGQAEATNKTLITALKKRLEQAKGKWVEELPGVLWAYRTTPGRPTGN 888
Query: 1940 TPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYWNMMLDEMVNLDEERLLALDVLTRQ 1999
TPF LAYG +AV+P E+ L + ++ + + LD DE R A +
Sbjct: 889 TPFALAYGMDAVIPIEIGLPTIWTNAAKQSDANMQLGRNLDW---TDEVRESASIRMADY 945
Query: 2000 KDRIAKAYNKKVRDRSFITGDYVWKVILPMDKKDRVYGKWTPNWEGPFTVEKTLPNNAYV 2059
+ R + YN+KVR RS G V + + + GK+ NWEGP+ V K N AY
Sbjct: 946 QQRASAHYNRKVRPRSLKNGTLVLRKFFE-NTTEVGAGKFQANWEGPYIVSKASDNGAYH 1004
Query: 2060 IKELGNQRQCVTINGKYLKAY 2080
+++L N LK Y
Sbjct: 1005 LQKLDGTPLLRPWNVSNLKQY 1025
>gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37531820|ref|NP_920212.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|22128689|gb|AAM92802.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 2017
Score = 686 bits (1771), Expect = 0.0
Identities = 404/1125 (35%), Positives = 627/1125 (54%), Gaps = 52/1125 (4%)
Query: 986 EKPVVDVSKKMEAQDPLEEVDLGDGL----EKRPTYISALIDPELKDRMVKL-------L 1034
+K D+++ E E++ L E T IS + E K + + L
Sbjct: 913 DKESCDMAQTREMASAREDIRLAAATASEGEVPATKISKSGESEAKTKKIPLDPSDPTKT 972
Query: 1035 KEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKC 1094
KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+ +LL
Sbjct: 973 ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAA 1032
Query: 1095 KFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEY 1154
FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS AG E
Sbjct: 1033 GFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCEL 1092
Query: 1155 LSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMN 1214
LS LD YSGY+QI + D KT+F P G Y +V MPF GLKNAGATYQR++
Sbjct: 1093 LSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPF-----GLKNAGATYQRMIQ 1145
Query: 1215 TIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1274
F I ++ Y+DD+VVK+ +D +L L ++F +R + +K+NP KC FGV +G
Sbjct: 1146 RCFSTQIGRNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFRMKLNPEKCTFGVPSGKL 1205
Query: 1275 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLL 1334
LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L
Sbjct: 1206 LGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL- 1264
Query: 1335 RLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQE-D 1393
LKK D F+W E QKAF++ KK L+ PPV+ P P+ LY+SAT + + ++L E +
Sbjct: 1265 -LKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVERE 1323
Query: 1394 EDGN----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDI 1449
E+G+ +R I+++S VL ++TRY ++KL + + KL +Y + V V + +
Sbjct: 1324 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP- 1382
Query: 1450 IKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTYVGIQP 1509
+ +L + RI KWAL L L++ P +IK QA+ADF+A+ T +E T ++
Sbjct: 1383 LGDILHNREANGRIAKWALELMSLDLSFKPRISIKSQALADFVAEWTECQEDTTVKKMEH 1442
Query: 1510 WKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGA 1569
W ++FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I ++LG
Sbjct: 1443 WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGI 1502
Query: 1570 KNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANEL 1629
K ++++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R +N+ A+ L
Sbjct: 1503 KRLIVRGDSQLVVNQVMKEWSCLDDNMKAYRQEVRKLEDKFDGLELSHVLRHDNEAADRL 1562
Query: 1630 A------QIASGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVEYLQNPV 1683
A ++A + + T + K+ + + + + DWR+P++ +L +
Sbjct: 1563 ANFGSKREVAPSDVFVEHLYTPTVPHKDTTQAAGIHDVA---MVETDWREPLIRFLTSQE 1619
Query: 1684 GSTDR----KVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQA 1739
D+ ++ R+ Y + EL+KK+ +G L +C+S + + H G+CG H A
Sbjct: 1620 LPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAA 1679
Query: 1740 GAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAI 1799
+ +RQG +WPT + D + R C+ CQ + H+PA EL +I WPF W +
Sbjct: 1680 ARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGL 1739
Query: 1800 DLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESIT 1859
D++G A + ++ VA+D F+KW+EA P+ +T + +F N IV+RFG+P I
Sbjct: 1740 DMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN-IVHRFGVPNRII 1797
Query: 1860 TDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPK--- 1916
TD GT F G F E +GIK+ ++ + +NGQVE AN +++ IK V + K
Sbjct: 1798 TDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYA 1857
Query: 1917 -RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYW 1975
+W + L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R + E E Y
Sbjct: 1858 GKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYE 1914
Query: 1976 NMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILPMDKKDRV 2035
+D++ L+E R AL R + + +N+ VR R+F+ GD V + I +DR
Sbjct: 1915 EDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI--QTTRDR- 1971
Query: 2036 YGKWTPNWEGPFTVEKTLPNNAYVIKELGNQRQCVTINGKYLKAY 2080
K +P WEGPF + + +Y +K + N +YL+ +
Sbjct: 1972 -HKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEYLRRF 2015
>ref|XP_475022.1| OSJNBb0093G06.3 [Oryza sativa (japonica cultivar-group)]
gi|38347562|emb|CAE04995.2| OSJNBb0093G06.3 [Oryza sativa
(japonica cultivar-group)]
Length = 1986
Score = 686 bits (1770), Expect = 0.0
Identities = 391/1053 (37%), Positives = 602/1053 (57%), Gaps = 45/1053 (4%)
Query: 1030 MVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1089
++ L+ KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+
Sbjct: 937 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 996
Query: 1090 RLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1149
+LL FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS
Sbjct: 997 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1056
Query: 1150 AGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATY 1209
AG E LS LD YSGY+QI + D KT+F P G Y +V MPF GLKNAGATY
Sbjct: 1057 AGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPF-----GLKNAGATY 1109
Query: 1210 QRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGV 1269
QR++ F I ++ Y+DD+VVK+ +D + L ++F +R + +K+NP KC FGV
Sbjct: 1110 QRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGV 1169
Query: 1270 IAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKS 1329
+G LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+
Sbjct: 1170 PSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMP 1229
Query: 1330 FSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSML 1389
F L LKK D F+W E QKAF++ K+ L+ PPV+ P P+ LY+SAT + + ++L
Sbjct: 1230 FFKL--LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQVVSTVL 1287
Query: 1390 AQE-DEDGN----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVF 1444
E +E+G+ +R I+++S VL ++TRY ++KL + + KL +Y + V V
Sbjct: 1288 VVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1347
Query: 1445 SHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTY 1504
+ + + +L + RI KWAL L +++ P +IK QA+ADF+A+ T +E
Sbjct: 1348 TSFP-LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQEDTPV 1406
Query: 1505 VGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEIL 1564
I+ W ++FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I
Sbjct: 1407 EKIEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIA 1466
Query: 1565 LALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQ 1624
++LG K ++++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R N+
Sbjct: 1467 ISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNE 1526
Query: 1625 EANELAQIA-------SGYMVDKLKLTELIKIKEKLSPSDL-DIMCIDNLTSNDWRKPIV 1676
A+ LA S V+ L T + K+ +D D++ ++ DWR+P +
Sbjct: 1527 AADRLANFGSKREAAPSDVFVEHL-YTPTVPHKDTTQDADTHDVVMVE----ADWREPFI 1581
Query: 1677 EYLQNPVGSTDR----KVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDG 1732
+L + D+ ++ R+ Y + +EL+KK+ +G L +C+S + + H G
Sbjct: 1582 RFLSSQELPQDKDEAERISRRSKLYVMHESELYKKSPSGILQRCVSLEEGRQLLKDIHSG 1641
Query: 1733 LCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPW 1792
+CG H A + +RQG +WPT + D + R C+ CQ + H+PA EL +I W
Sbjct: 1642 ICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSW 1701
Query: 1793 PFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRF 1852
PF W +D++G A + ++ VA+D F+KW+EA P+ +T + +F N IV+RF
Sbjct: 1702 PFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN-IVHRF 1759
Query: 1853 GLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVG 1912
G+P I TD GT F G F E +GIK+ ++ + +NGQVE AN +++ IK V
Sbjct: 1760 GVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVF 1819
Query: 1913 RKPK----RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEE 1968
+ K +W + L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R + E
Sbjct: 1820 DRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE 1879
Query: 1969 IPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILP 2028
E Y +D++ L+E R AL R + + +N+ VR R+F+ GD V + I
Sbjct: 1880 ---ERYEEDRVDDLHRLEEAREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI-- 1934
Query: 2029 MDKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIK 2061
+DR K +P WEGPF + + +Y +K
Sbjct: 1935 QTTRDR--HKLSPLWEGPFIISEVTRPGSYRLK 1965
>gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||H86337 protein
F5M15.26 [imported] - Arabidopsis thaliana
Length = 1838
Score = 682 bits (1761), Expect = 0.0
Identities = 394/1093 (36%), Positives = 594/1093 (54%), Gaps = 30/1093 (2%)
Query: 1003 EEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPI 1062
E V++ + R + A I P ++ ++ LLK FAW ++M G++ + +L +
Sbjct: 759 EMVNIDESDPTRCVGVGAEISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNV 818
Query: 1063 KEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVC 1122
KPVKQ R+ P+ + EE+E+LLK I +Y +WLAN V V KKNGK RVC
Sbjct: 819 DPTFKPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVC 878
Query: 1123 IDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCP 1182
+D+ DLN A PKD Y +P + +V++ +G+ LS +D +SGYNQI + +D KT+F
Sbjct: 879 VDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTD 938
Query: 1183 GALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGH 1242
GTY + VM FGLKNAG ATYQR +N + D I ++VYIDD++VKS + H
Sbjct: 939 R--GTYCYKVMSFGLKNAG-----ATYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDH 991
Query: 1243 LLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTS 1302
+ HL K F+ + YG+K+NP KC FGV +G+FLG+VV K+GIE N + +AIL+ P +
Sbjct: 992 VEHLSKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIRAILELPSPRN 1051
Query: 1303 KKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNP 1362
+++Q L G+I L RFI+ ++K F +LL+ + + F W+ + ++AF++LK YLS P
Sbjct: 1052 AREVQRLTGRIAALNRFISRSTDKCLPFYNLLKRRAQ--FDWDKDSEEAFEKLKDYLSTP 1109
Query: 1363 PVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLC 1422
P+++ P G + LYI+ +D + S+L +ED G +R IFY S+ L AETRY +IEK
Sbjct: 1110 PILVKPEVGETLYLYIAVSDHAVSSVLVREDR-GEQRPIFYTSKSLVEAETRYPVIEKAA 1168
Query: 1423 LCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKA 1482
L + S KL+ Y + + V + + + L P R+ KWA+ L+EY + + P A
Sbjct: 1169 LAVVTSARKLRPYFQSHTIAVLTDQPL-RVALHSPSQSGRMTKWAVELSEYDIDFRPRPA 1227
Query: 1483 IKGQAVADFLADHTVPKEIVTYVGI--QPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKFK 1540
+K Q +ADFL + + G + W LY DGSS G+GIG+ ++SP +
Sbjct: 1228 MKSQVLADFLIELPLQSAERAVSGNRGEEWSLYVDGSSSARGSGIGIRLVSPTAEVLEQS 1287
Query: 1541 FRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYY 1600
FR+ +NN EYE LI+GL + + + DS+L+ QL+ EY+ +E + Y
Sbjct: 1288 FRLRFVATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYL 1347
Query: 1601 VKANNLLAKFNEIGIGHVPRIENQEANELAQIASGYMVDKLKLTELIKIKEKLSPSDLDI 1660
+ F + +PR +N A+ LA +A D ++ + I + S +
Sbjct: 1348 KIVQLMTKDFENFKLSKIPRGDNAPADALAALALTSDSDLRRIIPVESIDKPSIDSTDAV 1407
Query: 1661 MCIDNLTSN---------DWRKPIVEYLQNPVGSTD----RKVKYRALSYTILGNELFKK 1707
++ + S+ DWR I +YL + +D R+++ +A YT++ L K
Sbjct: 1408 EIVNTIRSSNAPDPADPTDWRVEIRDYLSDGTLPSDKWTARRLRIKAAKYTLMKEHLLKV 1467
Query: 1708 NINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARG 1767
+ G +L CL + + H+G G H G + L + G YWPT++ DC +
Sbjct: 1468 SAFGAMLNCLHGTEINEIMKETHEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAK 1527
Query: 1768 CQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWV 1827
C+ CQ+H+ H P L + + P+PF WA+D++G + +S+Q ++I+V DYFTKWV
Sbjct: 1528 CEQCQRHAPTIHQPTELLRAGVAPYPFMRWAMDIVGPM--PASRQKRFILVMTDYFTKWV 1585
Query: 1828 EAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTP 1887
EA + V F+ I+ R GLP I TD G+ F+ F SW I+L STP
Sbjct: 1586 EAESYATIRANDVQNFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTP 1645
Query: 1888 YYAQANGQVEAANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGTTPFRLAYG 1947
Y Q NGQ EA NK ++S +KK + K W + L VLW+YR +P AT TPF AYG
Sbjct: 1646 RYPQGNGQAEATNKTILSGLKKRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYG 1705
Query: 1948 QEAVLPAEVYLQSCRIQRQEEIPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAY 2007
EA+ PAEV S R + P E+ MMLD + +L+E R AL + + AK Y
Sbjct: 1706 MEAMAPAEVGYSSLRRSMMVKNP-ELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHY 1764
Query: 2008 NKKVRDRSFITGDYVWKVILPMDKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIKELGNQR 2067
N+KV +R F GD V + + + GK NWEG + V K + Y + +
Sbjct: 1765 NQKVHNRHFDVGDLVLRKVFENTAEINA-GKLGANWEGSYQVSKIVRPGDYELLTMSGTA 1823
Query: 2068 QCVTINGKYLKAY 2080
T N +LK Y
Sbjct: 1824 VPRTWNSMHLKRY 1836
>gb|AAP52501.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37531824|ref|NP_920214.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|22128685|gb|AAM92798.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 1986
Score = 682 bits (1761), Expect = 0.0
Identities = 389/1052 (36%), Positives = 598/1052 (55%), Gaps = 43/1052 (4%)
Query: 1030 MVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1089
++ L+ KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+
Sbjct: 937 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 996
Query: 1090 RLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1149
+LL FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS
Sbjct: 997 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1056
Query: 1150 AGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATY 1209
AG E LS LD YSGY+QI + D KT+F P G Y +V MPF GLKNAGATY
Sbjct: 1057 AGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPF-----GLKNAGATY 1109
Query: 1210 QRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGV 1269
QR++ F I ++ Y+DD+VVK+ +D + L ++F +R + +K+NP KC FGV
Sbjct: 1110 QRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGV 1169
Query: 1270 IAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKS 1329
+G LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+
Sbjct: 1170 PSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMP 1229
Query: 1330 FSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSML 1389
F L LKK D F+W E QKAF++ K+ L+ PPV+ P P+ LY+SAT + + ++L
Sbjct: 1230 FFKL--LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQVVSTVL 1287
Query: 1390 AQE-DEDGN----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVF 1444
E +E+G+ +R I+++S VL ++ RY ++KL + + KL +Y + V V
Sbjct: 1288 VVEREEEGHVQKVQRPIYFVSEVLADSKARYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1347
Query: 1445 SHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTY 1504
+ + + +L + RI KWAL L +++ P +IK QA+ADF+A+ T +E
Sbjct: 1348 TSFP-LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQEDTPV 1406
Query: 1505 VGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEIL 1564
I+ W ++FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I
Sbjct: 1407 EKIEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIA 1466
Query: 1565 LALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQ 1624
++LG K ++++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R N+
Sbjct: 1467 ISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNE 1526
Query: 1625 EANELAQIA-------SGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVE 1677
A+ LA S V+ L T + K+ +D + + DWR+P +
Sbjct: 1527 AADRLANFGSKRETAPSDVFVEHL-YTPTVPHKDTTQDADTRNIA---MVEADWREPFIR 1582
Query: 1678 YLQNPVGSTDR----KVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGL 1733
+L + D+ ++ R+ Y + +EL+KK+ +G L +C+S + + H G+
Sbjct: 1583 FLTSQELPQDKDEAERISRRSKLYVLHESELYKKSPSGILQRCVSLEEGRQLLKDIHSGI 1642
Query: 1734 CGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWP 1793
CG H A + +RQG +WPT + D + R C+ CQ + H+PA EL +I WP
Sbjct: 1643 CGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWP 1702
Query: 1794 FRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFG 1853
F W +D++G A + ++ VA+D F+KW+EA P+ +T + +F N IV+RFG
Sbjct: 1703 FAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN-IVHRFG 1760
Query: 1854 LPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGR 1913
+P I TD GT F G F E +GIK+ ++ + +NGQVE AN +++ IK V
Sbjct: 1761 VPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFD 1820
Query: 1914 KPK----RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEI 1969
+ K +W + L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R + E
Sbjct: 1821 RLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE- 1879
Query: 1970 PSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILPM 2029
E Y +D++ L+E R AL R + + +N+ VR R+F+ GD V + I
Sbjct: 1880 --ERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI--Q 1935
Query: 2030 DKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIK 2061
+DR K +P WEGPF + + +Y +K
Sbjct: 1936 TTRDR--HKLSPLWEGPFIISEVTRPGSYRLK 1965
>ref|XP_475587.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46391119|gb|AAS90646.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1991
Score = 681 bits (1758), Expect = 0.0
Identities = 390/1053 (37%), Positives = 599/1053 (56%), Gaps = 45/1053 (4%)
Query: 1030 MVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1089
++ L+ KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+
Sbjct: 942 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 1001
Query: 1090 RLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1149
+LL FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS
Sbjct: 1002 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1061
Query: 1150 AGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATY 1209
AG E LS LD YSGY+QI + D KT+F P G Y +V MPF GLKNAGATY
Sbjct: 1062 AGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPF-----GLKNAGATY 1114
Query: 1210 QRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGV 1269
QR++ F I ++ Y+DD+VVK+ +D +L L ++F +R + +K+NP KC FGV
Sbjct: 1115 QRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFRMKLNPEKCTFGV 1174
Query: 1270 IAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKS 1329
+G LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+
Sbjct: 1175 PSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMP 1234
Query: 1330 FSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSML 1389
F L LKK D F+W E QKAF++ K+ L+ PPV+ P P+ LY+SAT + + ++L
Sbjct: 1235 FFKL--LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQVVSTVL 1292
Query: 1390 AQE-DEDGN----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVF 1444
E +E+G+ +R I+++S VL ++TRY ++KL + + KL +Y + V V
Sbjct: 1293 VVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1352
Query: 1445 SHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTY 1504
+ + + +L + RI KWAL L +++ P +IK QA+ADF+A+ T +E
Sbjct: 1353 TSFP-LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQEDTPV 1411
Query: 1505 VGIQPWKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEIL 1564
I+ W ++FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I
Sbjct: 1412 EKIEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIA 1471
Query: 1565 LALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQ 1624
++LG K ++++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R N+
Sbjct: 1472 ISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNE 1531
Query: 1625 EANELAQIA-------SGYMVDKLKLTELIKIKEKLSPSDL-DIMCIDNLTSNDWRKPIV 1676
A+ LA S V+ L + + K+ +D DI ++ DWR+P++
Sbjct: 1532 AADRLANFGSKREAAPSDVFVEHL-YSPTVPHKDATQVADTHDIAMVE----ADWREPLI 1586
Query: 1677 EYLQN----PVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDG 1732
+L + ++ R+ Y + EL+KK+ +G L +C+S + + H G
Sbjct: 1587 RFLTSQELPQYKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSG 1646
Query: 1733 LCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPW 1792
+CG H A + +RQG +WPT + D + R C+ CQ + H+PA EL +I W
Sbjct: 1647 ICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSW 1706
Query: 1793 PFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRF 1852
PF W +D++G A + ++ +A+D F+KW+EA P+ +T + +F N IV+RF
Sbjct: 1707 PFAVWGLDMVGPFKKAVG-GYTHLFMAIDKFSKWIEAKPVVTITADNARDFFIN-IVHRF 1764
Query: 1853 GLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVG 1912
G+P I TD GT F G F E +GIK+ ++ + +NGQVE AN +++ IK V
Sbjct: 1765 GVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQRIKARVF 1824
Query: 1913 RKPK----RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEE 1968
+ K +W L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R + E
Sbjct: 1825 DRLKPYAGKWVSQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE 1884
Query: 1969 IPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILP 2028
E Y +D++ L+E R AL R + + +N+ VR R+F+ GD V + I
Sbjct: 1885 ---ERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI-- 1939
Query: 2029 MDKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIK 2061
+DR K +P WEGPF + + +Y +K
Sbjct: 1940 QTTRDR--HKLSPLWEGPFIISEVTRPGSYRLK 1970
>ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultivar-group)]
gi|38347303|emb|CAE02298.2| OSJNBa0042F21.5 [Oryza sativa
(japonica cultivar-group)]
Length = 1950
Score = 681 bits (1756), Expect = 0.0
Identities = 389/1044 (37%), Positives = 594/1044 (56%), Gaps = 43/1044 (4%)
Query: 1038 KDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFI 1097
KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+ +LL FI
Sbjct: 909 KDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFI 968
Query: 1098 RTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSL 1157
+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS AG E LS
Sbjct: 969 KEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSF 1028
Query: 1158 LDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIF 1217
LD YSGY+QI + D KT+F P G Y +V MPF GLKNAGATYQR++ F
Sbjct: 1029 LDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPF-----GLKNAGATYQRMIQRCF 1081
Query: 1218 HDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGF 1277
I ++ Y+DD+VVK+ +D + L ++F +R + +K+NP KC FGV +G LGF
Sbjct: 1082 STQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGF 1141
Query: 1278 VVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLK 1337
+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L LK
Sbjct: 1142 MVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LK 1199
Query: 1338 KEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQE-DEDG 1396
K D F+W E QKAF++ KK L+ PP++ P P+ LY+SAT + + ++L E +E+G
Sbjct: 1200 KTDNFQWGPEAQKAFEDFKKLLTEPPILASPHPQEPLLLYVSATSQVVSTVLVVEREEEG 1259
Query: 1397 N----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKH 1452
+ +R I+++S VL ++TRY ++KL + + KL +Y + V V + + +
Sbjct: 1260 HVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP-LGD 1318
Query: 1453 MLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTYVGIQPWKL 1512
+L + RI KWAL L +++ P +IK QA+ADF+A+ T +E ++ W +
Sbjct: 1319 ILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAENMEHWTM 1378
Query: 1513 YFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGAKNV 1572
+FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I ++LG K +
Sbjct: 1379 HFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRL 1438
Query: 1573 VIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANELAQI 1632
+++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R N+ A+ LA
Sbjct: 1439 IVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANF 1498
Query: 1633 A-------SGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVEYLQNPVGS 1685
S V+ L T + K+ +D + L DWR+P + +L +
Sbjct: 1499 GSKREMAPSDVFVEHL-YTPTVPHKDTTQDADTHDVA---LVEADWREPFIRFLTSQELP 1554
Query: 1686 TDR----KVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGA 1741
D+ ++ R+ Y + EL+KK+ +G L +C+S + + H G+CG H A
Sbjct: 1555 QDKDEAERISRRSKLYAMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAAR 1614
Query: 1742 KMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDL 1801
+ +RQG +WPT + D + R C+ CQ + H+PA EL +I WPF W +D+
Sbjct: 1615 TIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDM 1674
Query: 1802 IGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTD 1861
+G A + ++ VA+D F+KW+EA P+ +T + +F N IV+RFG+P I TD
Sbjct: 1675 VGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN-IVHRFGVPNRIITD 1732
Query: 1862 QGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPK----R 1917
GT F G F E +GIK+ ++ + +NGQVE AN +++ IK V + K +
Sbjct: 1733 NGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGK 1792
Query: 1918 WHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYWNM 1977
W + L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R + E E Y
Sbjct: 1793 WVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEED 1849
Query: 1978 MLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILPMDKKDRVYG 2037
+D++ L+E R AL R + + +N+ VR R+F+ GD V + I +DR
Sbjct: 1850 RVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI--QTTRDR--H 1905
Query: 2038 KWTPNWEGPFTVEKTLPNNAYVIK 2061
K +P WEGPF + + +Y +K
Sbjct: 1906 KLSPLWEGPFIISEVTRPGSYRLK 1929
>ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultivar-group)]
gi|38344177|emb|CAE03508.2| OSJNBa0053K19.16 [Oryza
sativa (japonica cultivar-group)]
Length = 2010
Score = 681 bits (1756), Expect = 0.0
Identities = 399/1118 (35%), Positives = 617/1118 (54%), Gaps = 76/1118 (6%)
Query: 986 EKPVVDVSKKMEAQDPLEEVDLGDGL----EKRPTYISALIDPELKDRMVKL-------L 1034
+K D+++ E E++ L E+ T IS + E K + + L
Sbjct: 906 DKESCDMAQTREMASAREDIRLAAATASEGEEPATKISKSGESEAKTKKIPLDPSDPAKT 965
Query: 1035 KEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKC 1094
KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+ +LL
Sbjct: 966 ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAA 1025
Query: 1095 KFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEY 1154
FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS AG E
Sbjct: 1026 GFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGREL 1085
Query: 1155 LSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMN 1214
LS LD YSGY+QI + D KT+F P G Y +V MPF GLKNAGATYQR++
Sbjct: 1086 LSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPF-----GLKNAGATYQRMIQ 1138
Query: 1215 TIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1274
F I ++ Y+DD+VVK+ +D + L ++F +R + +K+NP KC FGV +G
Sbjct: 1139 RCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKL 1198
Query: 1275 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLL 1334
+GF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L
Sbjct: 1199 VGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL- 1257
Query: 1335 RLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQE-D 1393
LKK D F+W E QKAF++ KK L+ PPV+ P P+ LY+SAT + + ++L E +
Sbjct: 1258 -LKKTDNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVERE 1316
Query: 1394 EDGN----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDI 1449
E+G+ +R I+++S VL ++TRY ++KL + + KL +Y + V V + +
Sbjct: 1317 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP- 1375
Query: 1450 IKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTYVGIQP 1509
+ +L + RI KWAL L +++ P +IK QA+ADF+A+ T +E ++
Sbjct: 1376 LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAENMEH 1435
Query: 1510 WKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGA 1569
W ++FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I ++LG
Sbjct: 1436 WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGI 1495
Query: 1570 KNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANEL 1629
K ++++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R N+ A+ L
Sbjct: 1496 KRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRL 1555
Query: 1630 AQIASGYMVDKLKLTELIKIKEKLSPSDLDIMCIDN------------------LTSNDW 1671
A S K +++PSD+ + + + DW
Sbjct: 1556 ANFGS---------------KREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVETDW 1600
Query: 1672 RKPIVEYLQNPVGSTDR----KVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVS 1727
R+P++ +L + D+ ++ R+ Y + EL+KK+ +G L +C+S + +
Sbjct: 1601 REPLIRFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLQ 1660
Query: 1728 AAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHS 1787
H G+CG H A + +RQG +WPT + D + R C+ CQ + H+PA EL +
Sbjct: 1661 DIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQT 1720
Query: 1788 IIKPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNH 1847
I WPF W +D++G A + ++ VA+D F+KW+EA P+ +T + +F N
Sbjct: 1721 IPLSWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN- 1778
Query: 1848 IVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLI 1907
IV+RFG+P I TD GT F G F E +GIK+ ++ + +NGQVE AN +++ I
Sbjct: 1779 IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGI 1838
Query: 1908 KKHVGRKPK----RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRI 1963
K V + K +W + L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R
Sbjct: 1839 KARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRF 1898
Query: 1964 QRQEEIPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVW 2023
+ E E Y +D++ L+E R AL R + + +N+ VR R+F+ GD V
Sbjct: 1899 RNFRE---ERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVL 1955
Query: 2024 KVILPMDKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIK 2061
+ I +DR K +P WEGPF + + +Y +K
Sbjct: 1956 RKI--QTTRDR--HKLSPLWEGPFIISEVTRPGSYRLK 1989
>ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46063437|gb|AAS79740.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1756
Score = 676 bits (1745), Expect = 0.0
Identities = 387/1055 (36%), Positives = 594/1055 (55%), Gaps = 65/1055 (6%)
Query: 1038 KDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFI 1097
KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+ +LL FI
Sbjct: 715 KDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFVQDRKDAIKEELTKLLAAGFI 774
Query: 1098 RTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSL 1157
+ + DWLAN V V KK G+ R+C+D+ DLN PKD + +P + +VDS AG E LS
Sbjct: 775 KEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCELLSF 834
Query: 1158 LDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIF 1217
LD YSGY+QI + D KT+F P +G Y +V MPF GLKNAGATYQR++ F
Sbjct: 835 LDCYSGYHQIRLKESDCLKTSFITP--IGAYCYVTMPF-----GLKNAGATYQRMIQRCF 887
Query: 1218 HDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGF 1277
I ++ Y+DD+V+K+ +D + L ++F +R + +K+NP KC FGV +G LGF
Sbjct: 888 STQIGRNVEAYVDDVVIKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGF 947
Query: 1278 VVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLK 1337
+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L LK
Sbjct: 948 MVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LK 1005
Query: 1338 KEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQE-DEDG 1396
K D F+W E QKAF++ KK+L+ PPV+ P P+ LY+SAT + + ++L E +E+G
Sbjct: 1006 KTDDFQWGPEAQKAFEDFKKFLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEG 1065
Query: 1397 N----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKH 1452
+ +R I+++S VL ++TRY ++KL + + KL +Y + V V + + +
Sbjct: 1066 HVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP-LGD 1124
Query: 1453 MLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTYVGIQPWKL 1512
+L + RI KWAL L +++ P +IK QA+ADF+A+ T +E ++ W +
Sbjct: 1125 ILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAEKMEHWTM 1184
Query: 1513 YFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGAKNV 1572
+FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I ++LG K +
Sbjct: 1185 HFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRL 1244
Query: 1573 VIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANELAQI 1632
+++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R N+ A+ LA
Sbjct: 1245 IVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANF 1304
Query: 1633 ASGYMVDKLKLTELIKIKEKLSPSDLDIMCIDN------------------LTSNDWRKP 1674
S K +++PSD+ + + L DWR+P
Sbjct: 1305 GS---------------KREVAPSDVFVEHLYTPTVPHKDTTQIAGTHDVALVEADWREP 1349
Query: 1675 IVEYLQNPVGSTDR----KVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAH 1730
+ +L + D+ ++ R+ Y + EL+KK+ +G L +C+S + + H
Sbjct: 1350 FIRFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIH 1409
Query: 1731 DGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIK 1790
G+CG H A + +RQG +WPT + D + R C+ CQ + H+PA EL +I
Sbjct: 1410 SGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPL 1469
Query: 1791 PWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVY 1850
WPF W +D++G A + ++ VA+D F+KW+EA P+ +T + +F N IV+
Sbjct: 1470 SWPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN-IVH 1527
Query: 1851 RFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKH 1910
RFG+P I TD G F G F E +GIK+ ++ + +NGQVE AN +++ IK
Sbjct: 1528 RFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKAR 1587
Query: 1911 VGRKPK----RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQ 1966
V + K +W + L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R +
Sbjct: 1588 VFDRLKPYAGKWVKQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNF 1647
Query: 1967 EEIPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVI 2026
E E Y +D++ L+E R AL R + + +N+ VR R+F+ GD V + I
Sbjct: 1648 RE---ERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI 1704
Query: 2027 LPMDKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIK 2061
+DR K +P WEGPF + + +Y +K
Sbjct: 1705 --QTTRDR--HKLSPLWEGPFIISEVTRPGSYRLK 1735
>gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37533012|ref|NP_920808.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|19881590|gb|AAM00991.1| Putative retroelement [Oryza
sativa]
Length = 2017
Score = 676 bits (1744), Expect = 0.0
Identities = 401/1118 (35%), Positives = 613/1118 (53%), Gaps = 76/1118 (6%)
Query: 986 EKPVVDVSKKMEAQDPLEEVDLGDGL----EKRPTYISALIDPELKDRMVKL-------L 1034
+K D+++ E E++ L E T IS + E K + + L
Sbjct: 913 DKESCDMAQTREMASAREDIRLAAATASEGEVPATKISKSGESEAKTKKIPLDPSDSTKT 972
Query: 1035 KEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKC 1094
KD FAW +MPG+ RE++E L +KED KP+KQ RRF D IKEE+ +LL
Sbjct: 973 ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAA 1032
Query: 1095 KFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEY 1154
FI+ + DWLAN V V KK G+ R+C+D+ DLN PKD + +P + +VDS AG E
Sbjct: 1033 GFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCEL 1092
Query: 1155 LSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMN 1214
LS LD YSGY+QI + D KT+F P G Y +V MPF GLKNAGATYQR++
Sbjct: 1093 LSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPF-----GLKNAGATYQRMIQ 1145
Query: 1215 TIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1274
F I ++ Y+DD+VVK+ +D + L ++F +R + +K+NP KC FGV +G
Sbjct: 1146 RCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKL 1205
Query: 1275 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLL 1334
LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L
Sbjct: 1206 LGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL- 1264
Query: 1335 RLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQE-D 1393
LKK D F W E QKAF++ KK L+ PPV+ P P+ LY+SAT + + ++L E +
Sbjct: 1265 -LKKTDDFHWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVERE 1323
Query: 1394 EDGN----ERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDI 1449
EDG+ +R I+++S VL ++TRY ++KL + + KL +Y + V V + +
Sbjct: 1324 EDGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHLVTVVTSFP- 1382
Query: 1450 IKHMLSKPILHSRIGKWALALTEYSLTYAPLKAIKGQAVADFLADHTVPKEIVTYVGIQP 1509
+ +L + RI KWAL L +++ P +IK QA+ADF+A+ T +E ++
Sbjct: 1383 LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAEKVEH 1442
Query: 1510 WKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGA 1569
W ++FDGS +GTG G+ ++SP G + I + S+N EYEAL+ GL I ++LG
Sbjct: 1443 WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGI 1502
Query: 1570 KNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANEL 1629
K ++++GDS+LVV Q+ KE+ C+ +++ Y + L KF+ + + HV R N+ A+ L
Sbjct: 1503 KRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNKAADRL 1562
Query: 1630 AQIASGYMVDKLKLTELIKIKEKLSPSDLDIMCI------------------DNLTSNDW 1671
A S K +++PSD+ + + + DW
Sbjct: 1563 ANFGS---------------KREVAPSDVFVEHLYAPTVPHKDTTQVAGTHDVAMVEADW 1607
Query: 1672 RKPIVEYLQNPVGSTDR----KVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVS 1727
R+P++ +L + D+ ++ R+ Y + EL+KK+ +G L +C+S + +
Sbjct: 1608 REPLIRFLTSQELPQDKDEAERISRRSRLYVMHEAELYKKSPSGILQRCVSLEEGRQLLK 1667
Query: 1728 AAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHS 1787
H G+CG H A + +RQG +WPT + D + R C+ CQ + H+PA EL +
Sbjct: 1668 DIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQT 1727
Query: 1788 IIKPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNH 1847
I WPF W +D++G A + ++ VA+D F+KW+EA P+ +T + +F N
Sbjct: 1728 IPLSWPFAVWGLDMVGPFKKAFG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFIN- 1785
Query: 1848 IVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLI 1907
IV+RFG+P I TD G F G F E +GIK+ ++ + +NGQVE AN +++ I
Sbjct: 1786 IVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGI 1845
Query: 1908 KKHVGRKPK----RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRI 1963
K V + K +W E L VLW+ R +P+ ATG +PF L YG EA+LP+EV +S R
Sbjct: 1846 KARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRF 1905
Query: 1964 QRQEEIPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVW 2023
+ E E Y +D++ L+E R AL R + + +N+ VR R+F+ GD V
Sbjct: 1906 RNFRE---ERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVL 1962
Query: 2024 KVILPMDKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIK 2061
+ I +DR K +P WEGPF + + +Y +K
Sbjct: 1963 RKI--QTTRDR--HKLSPLWEGPFIISEVTRPGSYRLK 1996
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.333 0.143 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,268,422,405
Number of Sequences: 2540612
Number of extensions: 132602651
Number of successful extensions: 514740
Number of sequences better than 10.0: 15461
Number of HSP's better than 10.0 without gapping: 2849
Number of HSP's successfully gapped in prelim test: 12614
Number of HSP's that attempted gapping in prelim test: 494893
Number of HSP's gapped (non-prelim): 21243
length of query: 2087
length of database: 863,360,394
effective HSP length: 144
effective length of query: 1943
effective length of database: 497,512,266
effective search space: 966666332838
effective search space used: 966666332838
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 83 (36.6 bits)
Lotus: description of TM0545.3


