

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0544.8
(1753 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
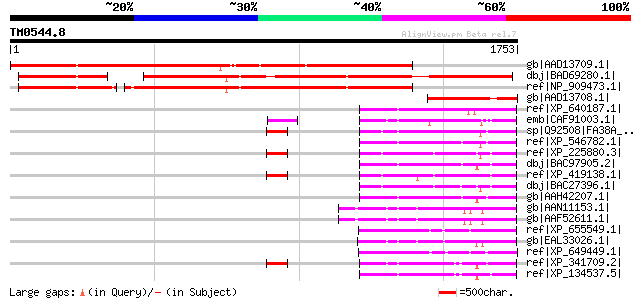
Score E
Sequences producing significant alignments: (bits) Value
gb|AAD13709.1| hypothetical protein [Arabidopsis thaliana] gi|25... 1554 0.0
dbj|BAD69280.1| hypothetical protein [Oryza sativa (japonica cul... 1326 0.0
ref|NP_909473.1| OSJNBb0008D07.21 [Oryza sativa (japonica cultiv... 1023 0.0
gb|AAD13708.1| unknown protein [Arabidopsis thaliana] gi|1522720... 364 2e-98
ref|XP_640187.1| hypothetical protein DDB0218401 [Dictyostelium ... 227 2e-57
emb|CAF91003.1| unnamed protein product [Tetraodon nigroviridis] 219 9e-55
sp|Q92508|FA38A_HUMAN Protein FAM38A gi|7662014|ref|NP_055560.1|... 216 4e-54
ref|XP_546782.1| PREDICTED: similar to Protein FAM38A [Canis fam... 216 7e-54
ref|XP_225880.3| PREDICTED: similar to family with sequence simi... 210 3e-52
dbj|BAC97905.2| mKIAA0233 protein [Mus musculus] 207 3e-51
ref|XP_419138.1| PREDICTED: similar to Hypothetical protein KIAA... 204 2e-50
dbj|BAC27396.1| unnamed protein product [Mus musculus] 204 3e-50
gb|AAH42207.1| 2310061F22Rik protein [Mus musculus] 203 5e-50
gb|AAN11153.1| CG8486-PB, isoform B [Drosophila melanogaster] gi... 200 4e-49
gb|AAF52611.1| CG8486-PA, isoform A [Drosophila melanogaster] gi... 200 4e-49
ref|XP_655549.1| conserved hypothetical protein [Entamoeba histo... 197 3e-48
gb|EAL33026.1| GA21110-PA [Drosophila pseudoobscura] 197 3e-48
ref|XP_649449.1| conserved hypothetical protein [Entamoeba histo... 192 1e-46
ref|XP_341709.2| PREDICTED: similar to Protein FAM38A [Rattus no... 189 6e-46
ref|XP_134537.5| PREDICTED: RIKEN cDNA 2310061F22 [Mus musculus] 189 7e-46
>gb|AAD13709.1| hypothetical protein [Arabidopsis thaliana] gi|25409089|pir||G84922
hypothetical protein At2g48050 [imported] - Arabidopsis
thaliana gi|15227204|ref|NP_182326.1| expressed protein
[Arabidopsis thaliana]
Length = 1500
Score = 1554 bits (4024), Expect = 0.0
Identities = 805/1417 (56%), Positives = 1024/1417 (71%), Gaps = 36/1417 (2%)
Query: 1 MKIYAVFVFVSIYSIGVVSSSDMWFPMIIDLQTAFGYNPAASMFQNIWESLAILVVMQLY 60
+K YAV VF+ IY + S +W IDL GYN A + N+WESLA+L+VMQLY
Sbjct: 82 LKAYAVLVFMFIYCLSSFVSLQLWLSGFIDLYFYLGYNSKAPLLDNVWESLAVLIVMQLY 141
Query: 61 SYERRQSKSSGSSDFDLPEVGPFPFTRRLLVRHTEKILFLALFYASLSPISAFGFLYLLG 120
SYERRQS L G F F R L H +KILF ALFYASLSPIS FGF+YLLG
Sbjct: 142 SYERRQSGHYIPGQSSLLHPGVFGFFERFLAWHGQKILFAALFYASLSPISVFGFVYLLG 201
Query: 121 LINCSRLPKSSQIPAKVFLVYSGLLIMVEYLFQMLGDQAEMFPGQKYSQLSLIMGLQLYK 180
L+ C+ PKSS IP+K FL+Y+G L+ EYLFQ+ G QA+MFPGQKY++LS +GL++Y+
Sbjct: 202 LVICTTFPKSSSIPSKSFLIYTGFLVSAEYLFQLWGMQAQMFPGQKYAELSFYLGLRVYE 261
Query: 181 PGFKGLESGFRGKVVVIVACILQYNVFRWLEKMQYVDPNGGKWNEPCPLFNMVDIPNETT 240
PGF G+ESG RGKV+V+ AC LQYNVFRWLE+ + GK+ EPCPLF V + T
Sbjct: 262 PGFWGIESGLRGKVLVVAACTLQYNVFRWLERTSGLTVIKGKYEEPCPLF--VSAEDTTA 319
Query: 241 ACTPQNKQAETST---SATIKR-LARSRSCPTVNSALSQGTD------SGIEGDSTKKLR 290
+ + N + +ST S ++K+ A S S P + +QG G E S++K
Sbjct: 320 SVSSSNGENPSSTDHASISMKQGEATSNSWPFFSPRGNQGAGFLHPKTGGSESGSSRKFS 379
Query: 291 QLHFWESSKDSLKWNRKRLLFLRKERLEMQKTALKVSLKFWIENMFTLFGLEINMIALLL 350
HFW S K+S +WNR+R+L L+KER E QK LK+ LKFWIENMF L+GLEINMIALLL
Sbjct: 380 FGHFWGSIKESHRWNRRRILALKKERFETQKNLLKIYLKFWIENMFNLYGLEINMIALLL 439
Query: 351 ASFAVLNAISLLYIASLAACVLLHRLLIKKLWPIFVFLFASVVTVEYLAIWMR-LTFMNL 409
ASFA+LNAIS++YIA LAACVLL R +I+KLWP+ VFLFAS++ +EY+A W L
Sbjct: 440 ASFALLNAISMVYIALLAACVLLRRRVIQKLWPVVVFLFASILAIEYVATWNSFLPSDQA 499
Query: 410 EIEEHVPCRDCWRVSDIYFSYCRKCWLGIIVDDPRMLIGYYGVFMFSCFKFRADKSSSLT 469
E V C DCW ++ +YF +CR+CWLG+ VDDPR LI Y+ VFM +CFK RAD SS +
Sbjct: 500 PSETSVHCHDCWSIAALYFKFCRECWLGVRVDDPRTLISYFVVFMLACFKLRADHISSFS 559
Query: 470 GLEMYQKIHSQWKSASVLSNLSFETKGYWTFLDHLRLYGYCHLLDFVLSLILITGTLEYD 529
Y ++ SQ K++ V +LSFETK WT LD+LRLY Y HLLD VL LILITGTLEYD
Sbjct: 560 ESSTYHQMKSQRKNSFVWRDLSFETKSMWTVLDYLRLYCYVHLLDVVLILILITGTLEYD 619
Query: 530 FLHLGYLGFALVFFRMRLKILKQGNNIFRYLRMYNFAVIVLSLAYQSPFLGDFSEIKSGF 589
LHLGYL FALVF RMRL+ILK+ N IFR+LR+YNF +I+ SLAYQSPF+G+F++ K
Sbjct: 620 ILHLGYLAFALVFARMRLEILKKKNKIFRFLRVYNFVLIIFSLAYQSPFVGNFNDGKCET 679
Query: 590 IEYINELVGFQKYDYGFRITSRSALVEIIIYMLVSLQSYMFSFPEFDYVSTYLEKEQIGA 649
++YI E++GF KYDYGFRIT+RSALVEIII+MLVSLQSYMFS EFDYVS YLE EQIGA
Sbjct: 680 VDYIYEVIGFYKYDYGFRITARSALVEIIIFMLVSLQSYMFSSQEFDYVSRYLEAEQIGA 739
Query: 650 ILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNAS 709
I+R+QEKKAA KT QL+ IR+AEE K R+LQVEKMKSEMLNL+ QLH M++D+N+ AS
Sbjct: 740 IVREQEKKAARKTEQLQQIREAEEKKRQRNLQVEKMKSEMLNLRVQLHRMNSDSNFGVAS 799
Query: 710 LKNDGLRERRNSFLENE------------FRKQALDTNTESIGTLDVNQSLLSEKSESPL 757
+ +GLR R++ +L + RK+ + +S + ++ +S E+
Sbjct: 800 PRTEGLRRRKSPYLIPDSGAASPEIDGVVHRKEEQPIDEDSQYPFEAHEFPVSTTPEALD 859
Query: 758 VPEYMKHPMGSPHGIVEVKERTKNNDFLDLEIRNRYKLPGKRNALVSAVHFIGSGISQAQ 817
PEY SP I EV++ + D + +E + K GK N L+SAV IG G+SQ Q
Sbjct: 860 SPEYSFG--ASPCEITEVQQ---DLDVMSMERERKQKSEGKENPLISAVQLIGDGVSQVQ 914
Query: 818 SLGNMAVNNLVNYLKIEREGQEPTDDSSEDEEYY-EIENQNSGAEPLEPTFSTNSINEHT 876
+GN AVNNLVN+L I E + + SS D+E Y E+E+Q P E + S+
Sbjct: 915 FIGNQAVNNLVNFLNISPENSDTNEQSSVDDEVYDEMESQKRKHTPFE---RSTSLQSDR 971
Query: 877 GSDTSCLQIGILVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNT 936
SD + QIG + RHIWSRM+SNN++VCYCCFI+ +LWNFSLLS+VYLAALFLYALC +T
Sbjct: 972 SSDGTSFQIGRIFRHIWSRMQSNNDIVCYCCFIIAFLWNFSLLSMVYLAALFLYALCVHT 1031
Query: 937 GPSYIFWVLILIYTEVCILLQYLYQIIIQHTEFEFPMSLLQELGFPAKKITSSFVTSNLP 996
GP++IFWV++L+YTE+ ILLQYLYQIIIQH LL ELGFP ++I SSFV S+LP
Sbjct: 1032 GPTHIFWVIMLMYTEIYILLQYLYQIIIQHCGLSIDAPLLHELGFPTQRIKSSFVVSSLP 1091
Query: 997 FFLIYIFTLLQISITMKGGGWTISVDPSFFRRRNQKSYLEDVKCSTYHERVQRLFLPLSN 1056
FLIYIFTL+Q SIT+K G W S D F RRN + +D+ +R+ +F L +
Sbjct: 1092 LFLIYIFTLIQSSITVKDGDWVPSAD--FTSRRNARGSQKDLTRIRLSQRILDVFKKLRD 1149
Query: 1057 ALKLLTRSLCRYWKSLTWGAETPPYFVQLSMEVNLWPKEGIQPKRIESKINKLLRILRHR 1116
+ KL+ RS+ RYW SLT GAE+PPYFVQ++M+V++WP++GIQP+R+E ++N+LLR++ +
Sbjct: 1150 SAKLVIRSIYRYWISLTRGAESPPYFVQVTMDVHMWPEDGIQPERVECRMNQLLRLVHNE 1209
Query: 1117 RCKEEKIFNLHSASRVRVQSIEKSEENENLCLVIFEVLYASPPNEFAAEEWYSSLTPAAD 1176
RC++ +SRV VQSIE+S E N LV+ EV YASP N ++ EWY SLTPA+D
Sbjct: 1210 RCEKGNPDLCPYSSRVHVQSIERSTETPNEALVVLEVEYASPTNGCSSAEWYKSLTPASD 1269
Query: 1177 VSYEIQKAQHAGILKEIGFPYRILSVIGGGKREVDLYAYIFGADLAVFFLIAIFYESIMK 1236
V+ EI+KAQH+G+ + GFPY ILSVIGGGKR+ DLYAYIFGADL VFFL+AIFY+S++K
Sbjct: 1270 VAKEIRKAQHSGLGEGTGFPYPILSVIGGGKRDTDLYAYIFGADLIVFFLVAIFYQSVIK 1329
Query: 1237 ANSEFLEVYQLEDQFPEDFVLVLMVVFFLIVLDRIIYLCSFATAKVIFYLFSLVLFTYSV 1296
SEF++VYQLEDQFP DFV++LMV+FFLIV+DR+IYLCSFAT KV++YLFSL+LFTY+V
Sbjct: 1330 NKSEFIDVYQLEDQFPFDFVIILMVIFFLIVVDRVIYLCSFATGKVVYYLFSLILFTYAV 1389
Query: 1297 TKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQAIQFHFGMPHKSILYRQFLTSSVSRIN 1356
T+YAW + P ++++ LALR I+ KA+SL LQAIQ +G+PHKS LYRQFLTS VSRIN
Sbjct: 1390 TEYAWSIYPTQQHAAGLALRIIFLAKAMSLALQAIQIRYGLPHKSTLYRQFLTSEVSRIN 1449
Query: 1357 VMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYDWLKL 1393
G+RLYRA+PFLYELRCVLDWSCT TSLTMYDWLK+
Sbjct: 1450 YYGYRLYRALPFLYELRCVLDWSCTATSLTMYDWLKV 1486
>dbj|BAD69280.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 1762
Score = 1326 bits (3431), Expect = 0.0
Identities = 708/1317 (53%), Positives = 893/1317 (67%), Gaps = 131/1317 (9%)
Query: 461 RADKSSSLTGLEMYQKIHSQWKSASVLSNLSFETKGYWTFLDHLRLYGYCHLLDFVLSLI 520
R+D+ S + + Y ++ SQ K+A V +LS ETK +WTFLD++RLY YCHLLD VL+LI
Sbjct: 481 RSDRFSGFSDSDTYHQMMSQRKNALVWRDLSLETKSFWTFLDYIRLYAYCHLLDIVLALI 540
Query: 521 LITGTLEYDFLHLGYLGFALVFFRMRLKILKQGNNIFRYLRMYNFAVIVLSLAYQSPFLG 580
ITGTLEYD LHLGYLGFALVFFRMRL+ILK+ N IF+YLRMYNFA+IVLSLAYQSP+ G
Sbjct: 541 AITGTLEYDVLHLGYLGFALVFFRMRLEILKRKNKIFKYLRMYNFALIVLSLAYQSPYFG 600
Query: 581 DFSEIKSGFIEYINELVGFQKYDYGFRITSRSALVEIIIYMLVSLQSYMFSFPEFDYVST 640
FS K I+YI E++GF KYDYGF+ITSRSA VEI+I++LVS+QSY+FS EFDYVS
Sbjct: 601 QFSSGKCDQIDYIYEIIGFYKYDYGFKITSRSAFVEIVIFLLVSIQSYIFSSGEFDYVSR 660
Query: 641 YLEKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMS 700
YLE EQIGA++ +QEKKA KT QL+H+R++EE K R++QVE+MKSEM NLQ+QL+ M+
Sbjct: 661 YLEAEQIGAMVHEQEKKALKKTEQLQHLRRSEEQKRERNMQVERMKSEMYNLQSQLNRMN 720
Query: 701 TDANYSNASLKNDGLRERRNSFLENEFRKQALDTNTES------IGTLDVNQSL------ 748
+ +NAS ++GLR RRN+ L + D+ S G+ D +QS
Sbjct: 721 SFTPINNAS-HSEGLRHRRNTKLYTDIDTPLQDSGIGSPRKEDKTGSTDSSQSFEFSVED 779
Query: 749 --------------------LSEKSESPLVPEYMKHPMGSPHGIVEVKERTK--NNDFLD 786
+ SE V + ++ +GS I EV+E N++ L
Sbjct: 780 AQKSLTDLMFRTPCDTPRSPIRGTSEEFKVTDNARNSLGSTSEITEVEENEGKVNHNLLK 839
Query: 787 LEIRNRYKLPGKRNALVSAVHFIGSGISQAQSLGNMAVNNLVNYLKIEREGQEPTDDSSE 846
L+ K N L SAV IG G+SQ QS GN AV N+V++L I+ E +D +E
Sbjct: 840 LQYGRGAV---KENPLKSAVQLIGDGVSQVQSFGNQAVTNIVSFLNIDPEEPHSSDHPAE 896
Query: 847 DEEYYEIENQNSGAEPLEPTFSTNSINEHTGSDTSC-LQIGILVRHIWSRMRSNNEVVCY 905
D+ Y +E+Q + T+S+ G+ +S + IG++
Sbjct: 897 DDIYDMVESQRETHDG--QLLRTHSVTSGNGTKSSANMPIGVI----------------- 937
Query: 906 CCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLILIYTEVCILLQYLYQIIIQ 965
F LLS+VYL ALFLYALC N GPSY+FWV++LIYTE+ IL QY+YQI+IQ
Sbjct: 938 ----------FRLLSMVYLGALFLYALCVNYGPSYLFWVIVLIYTELNILSQYIYQIVIQ 987
Query: 966 HTEFEFPMSLLQELGFPAKKITSSFVTSNLPFFLIYIFTLLQISITMKGGGWTISVDPSF 1025
H + LLQ LGFP KI +SFV S LP FL+YI TLLQ SIT K G W + SF
Sbjct: 988 HCGLNIHIPLLQRLGFPDDKIKASFVVSILPLFLVYISTLLQSSITAKDGEWVPVTEFSF 1047
Query: 1026 FRRRNQKSYLEDVKCSTYHERVQRLFLPLSNALKLLTRSLCRYWKSLTWGAETPPYFVQL 1085
RN + + + + +R++ + LP+ N ++++ R + RYW SLT GAE+PPYFVQ+
Sbjct: 1048 LSARNNVEEKQRMPYN-WRDRLKNIHLPVMNLIRMIGRGISRYWLSLTQGAESPPYFVQV 1106
Query: 1086 SMEVNLWPKEGIQPKRIESKINKLLRILRHRRCKEEKIFNLHSASRVRVQSIEKSEENEN 1145
+MEVN WP++GIQP+RIES IN++L I RC+ + HS SRVR+QSIE+S+EN +
Sbjct: 1107 TMEVNHWPEDGIQPERIESAINRVLAIAHEERCQANSPSSCHSCSRVRIQSIERSKENSS 1166
Query: 1146 LCLVIFEVLYASPPNEFAAEEWYSSLTPAADVSYEIQKAQHAGILKEIGFPYRILSVIGG 1205
+ L + EV+YA+P + +A WY SLTPAADV EI ++Q AG+ +++ FPY ++SVIGG
Sbjct: 1167 MALAVLEVVYAAPLDCQSAG-WYKSLTPAADVEKEIHESQKAGLFEDVNFPYPVVSVIGG 1225
Query: 1206 GKREVDLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQLEDQFPEDFVLVLMVVFFL 1265
GKRE+DLYAY FGADLAVFFL+A+FY+S++K SEFLEVYQLEDQFP++FV +LM++FFL
Sbjct: 1226 GKREIDLYAYYFGADLAVFFLVAMFYQSVLKNKSEFLEVYQLEDQFPKEFVFILMILFFL 1285
Query: 1266 IVLDRIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAIS 1325
IV+DRIIYL SFAT KVIFYLF+LVLFTYSVT+YAW M+ + R G LRAIY TK+IS
Sbjct: 1286 IVVDRIIYLWSFATGKVIFYLFNLVLFTYSVTEYAWGMELVHRNVGGFVLRAIYLTKSIS 1345
Query: 1326 LVLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSL 1385
L LQA+Q +G+P+KS LYRQFLTS V+++N GFRLYRA+PFLYELRCVLDWSCTTTSL
Sbjct: 1346 LALQALQIRYGIPNKSNLYRQFLTSKVTQVNYFGFRLYRALPFLYELRCVLDWSCTTTSL 1405
Query: 1386 TMYDWLKLEDIHASLFLVKCDVDLNRASHQQGQKQTKMTKFCNGICLFFVLMCVIWAPML 1445
TMYDWLK+
Sbjct: 1406 TMYDWLKI---------------------------------------------------- 1413
Query: 1446 MYSSGNPTNIANPIKDASARVHIKTLSGRLTLFETTLCEIISWETLEARTSLDPKGYLST 1505
YSSGNPTNIANPI D S ++ IK L GRLT F+TT+CE I W+ + A +DP YL
Sbjct: 1414 -YSSGNPTNIANPIIDVSVKIDIKALGGRLTFFKTTVCEKIPWKHMRAYDDVDPLDYLGG 1472
Query: 1506 YNEKDIQLICCQSDASTLWLVPPIVQARFMKSLS------WNMDLAFSWEFTRDRPRGKE 1559
YN +DIQLICCQ DAST+WL+P VQ RF++SL NM+L +W+F R RP+GKE
Sbjct: 1473 YNVEDIQLICCQPDASTMWLIPAPVQTRFIQSLEETEMIFGNMELILNWDFLRARPKGKE 1532
Query: 1560 VVKYELTIQEQDLPKSSEVTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSLEQSVELVSG 1619
+VKYE + P +V V NGT NSF + YPRYFRVTGSG+VR LE S++ VSG
Sbjct: 1533 LVKYESPVDRS--PSVDDVKRVLNGTINSFRITDAYPRYFRVTGSGEVRRLEASIDSVSG 1590
Query: 1620 DLVLNRGNPEWWSFYDLDISDAHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIWGLYIT 1679
+L+LN G P WWSFYD + SD GC GPMAI+VSEETPQGIIG+TLSKFSIW LYIT
Sbjct: 1591 ELLLNNGTPPWWSFYDTNPSDLAGCQGLNGPMAIVVSEETPQGIIGETLSKFSIWSLYIT 1650
Query: 1680 FVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLYWTL 1736
FVLAV RFIRLQCSDLRMRIP+ENLPSCDRL+ +CE IYAARAEGELEVEEVLYWTL
Sbjct: 1651 FVLAVARFIRLQCSDLRMRIPYENLPSCDRLLDICEGIYAARAEGELEVEEVLYWTL 1707
Score = 266 bits (679), Expect = 6e-69
Identities = 145/321 (45%), Positives = 201/321 (62%), Gaps = 15/321 (4%)
Query: 29 IDLQTAFGYNPAASMFQNIWESLAILVVMQLYSYERRQSKSSGSSDFDLPEVGPFPFTRR 88
+ L G++P AS+ N+W+SLA+LVVMQLYSYERRQ+ D E G F RR
Sbjct: 99 VKLYPDLGFDPEASLLMNVWQSLAVLVVMQLYSYERRQNSDKNFGVSDASESGLLGFLRR 158
Query: 89 LLVRHTEKILFLALFYASLSPISAFGFLYLLGLINCSRLPKSSQIPAKVFLVYSGLLIMV 148
LL+ H+EKIL + +FYA LS IS G +YLLGLI S LPK S+IP+KV+LVY+GLL
Sbjct: 159 LLIWHSEKILSVTVFYACLSSISLSGLIYLLGLIMFSILPKVSRIPSKVYLVYTGLLATS 218
Query: 149 EYLFQMLGDQAEMFPGQKYSQLSLIMGLQLYKPGFKGLESGFRGKVVVIVACILQYNVFR 208
EYLFQML + A+M PGQ++ LS+ +GL+ Y GF G+E G RGKV+VIVAC +QYNVF
Sbjct: 219 EYLFQMLCEPAQMCPGQQFHGLSVFLGLKHYDAGFWGVEYGLRGKVLVIVACTIQYNVFH 278
Query: 209 WLEKMQYVDPNGGKWNEPCPLFNMVDIPNETTACTPQNKQAETSTS------ATIKRLAR 262
WL+ M + GKW EPC LF I +T++ N + S++ + ++ L
Sbjct: 279 WLDLMPTSLLHEGKWEEPCQLF----ISGDTSSNARDNNKDSHSSNRFSSLFSKVQGLIG 334
Query: 263 SRSCPTVNSALSQGTDSGIE-----GDSTKKLRQLHFWESSKDSLKWNRKRLLFLRKERL 317
S S +++S + T ++ D K+ W SK+S KW++++++ LR+ER
Sbjct: 335 SSSSSSLSSGSTCQTSEPVQNETSGSDEGKRYSFSKIWGMSKESHKWDKRKIISLRRERF 394
Query: 318 EMQKTALKVSLKFWIENMFTL 338
E QKT K +KFW+EN+F L
Sbjct: 395 ETQKTTFKCYMKFWMENLFKL 415
>ref|NP_909473.1| OSJNBb0008D07.21 [Oryza sativa (japonica cultivar-group)]
Length = 1452
Score = 1023 bits (2645), Expect = 0.0
Identities = 543/1032 (52%), Positives = 713/1032 (68%), Gaps = 50/1032 (4%)
Query: 397 YLAIWMRLTFMNLEIEEHVPCRDCWRVSDIYFSYCRKCWLGIIVDDPRMLIGYYGVFMFS 456
Y+ WM F +E ++ ++C Y L IV + + G+ +
Sbjct: 408 YMKFWMENLFKLRGLEINMIVKECP-------GYTVSMTLKFIVANAGKIQGFSLHIAQN 460
Query: 457 CFKFRADKSSSLTGLEMYQKIHSQWKSASVLSNLSFETKGYWTFLDHLRLYGYCHLLDFV 516
R+D+ S + + Y ++ SQ K+A V +LS ETK +WTFLD++RLY YCHLLD V
Sbjct: 461 VGWLRSDRFSGFSDSDTYHQMMSQRKNALVWRDLSLETKSFWTFLDYIRLYAYCHLLDIV 520
Query: 517 LSLILITGTLEYDFLHLGYLGFALVFFRMRLKILKQGNNIFRYLRMYNFAVIVLSLAYQS 576
L+LI ITGTLEYD LHLGYLGFALVFFRMRL+ILK+ N IF+YLRMYNFA+IVLSLAYQS
Sbjct: 521 LALIAITGTLEYDVLHLGYLGFALVFFRMRLEILKRKNKIFKYLRMYNFALIVLSLAYQS 580
Query: 577 PFLGDFSEIKSGFIEYINELVGFQKYDYGFRITSRSALVEIIIYMLVSLQSYMFSFPEFD 636
P+ G FS K I+YI E++GF KYDYGF+ITSRSA VEI+I++LVS+QSY+FS EFD
Sbjct: 581 PYFGQFSSGKCDQIDYIYEIIGFYKYDYGFKITSRSAFVEIVIFLLVSIQSYIFSSGEFD 640
Query: 637 YVSTYLEKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQL 696
YVS YLE EQIGA++ +QEKKA KT QL+H+R++EE K R++QVE+MKSEM NLQ+QL
Sbjct: 641 YVSRYLEAEQIGAMVHEQEKKALKKTEQLQHLRRSEEQKRERNMQVERMKSEMYNLQSQL 700
Query: 697 HSMSTDANYSNASLKNDGLRERRNSFLENEFRKQALDTNTES------IGTLDVNQSL-- 748
+ M++ +NAS ++GLR RRN+ L + D+ S G+ D +QS
Sbjct: 701 NRMNSFTPINNAS-HSEGLRHRRNTKLYTDIDTPLQDSGIGSPRKEDKTGSTDSSQSFEF 759
Query: 749 ------------------------LSEKSESPLVPEYMKHPMGSPHGIVEVKERTK--NN 782
+ SE V + ++ +GS I EV+E N+
Sbjct: 760 SVEDAQKSLTDLMFRTPCDTPRSPIRGTSEEFKVTDNARNSLGSTSEITEVEENEGKVNH 819
Query: 783 DFLDLEIRNRYKLPGKRNALVSAVHFIGSGISQAQSLGNMAVNNLVNYLKIEREGQEPTD 842
+ L L+ K N L SAV IG G+SQ QS GN AV N+V++L I+ E +D
Sbjct: 820 NLLKLQYGRGAV---KENPLKSAVQLIGDGVSQVQSFGNQAVTNIVSFLNIDPEEPHSSD 876
Query: 843 DSSEDEEYYEIENQNSGAEPLEPTFSTNSINEHTGSDTSC-LQIGILVRHIWSRMRSNNE 901
+ED+ Y +E+Q + T+S+ G+ +S + IG++ R+IW +MRSN +
Sbjct: 877 HPAEDDIYDMVESQRETHDG--QLLRTHSVTSGNGTKSSANMPIGVIFRYIWYQMRSNYD 934
Query: 902 VVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLILIYTEVCILLQYLYQ 961
VCYCCF+L++LWNFSLLS+VYL ALFLYALC N GPSY+FWV++LIYTE+ IL QY+YQ
Sbjct: 935 YVCYCCFVLVFLWNFSLLSMVYLGALFLYALCVNYGPSYLFWVIVLIYTELNILSQYIYQ 994
Query: 962 IIIQHTEFEFPMSLLQELGFPAKKITSSFVTSNLPFFLIYIFTLLQISITMKGGGWTISV 1021
I+IQH + LLQ LGFP KI +SFV S LP FL+YI TLLQ SIT K G W
Sbjct: 995 IVIQHCGLNIHIPLLQRLGFPDDKIKASFVVSILPLFLVYISTLLQSSITAKDGEWVPVT 1054
Query: 1022 DPSFFRRRNQKSYLEDVKCSTYHERVQRLFLPLSNALKLLTRSLCRYWKSLTWGAETPPY 1081
+ SF RN + + + + +R++ + LP+ N ++++ R + RYW SLT GAE+PPY
Sbjct: 1055 EFSFLSARNNVEEKQRMPYN-WRDRLKNIHLPVMNLIRMIGRGISRYWLSLTQGAESPPY 1113
Query: 1082 FVQLSMEVNLWPKEGIQPKRIESKINKLLRILRHRRCKEEKIFNLHSASRVRVQSIEKSE 1141
FVQ++MEVN WP++GIQP+RIES IN++L I RC+ + HS SRVR+QSIE+S+
Sbjct: 1114 FVQVTMEVNHWPEDGIQPERIESAINRVLAIAHEERCQANSPSSCHSCSRVRIQSIERSK 1173
Query: 1142 ENENLCLVIFEVLYASPPNEFAAEEWYSSLTPAADVSYEIQKAQHAGILKEIGFPYRILS 1201
EN ++ L + EV+YA+P + +A WY SLTPAADV EI ++Q AG+ +++ FPY ++S
Sbjct: 1174 ENSSMALAVLEVVYAAPLDCQSAG-WYKSLTPAADVEKEIHESQKAGLFEDVNFPYPVVS 1232
Query: 1202 VIGGGKREVDLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQLEDQFPEDFVLVLMV 1261
VIGGGKRE+DLYAY FGADLAVFFL+A+FY+S++K SEFLEVYQLEDQFP++FV +LM+
Sbjct: 1233 VIGGGKREIDLYAYYFGADLAVFFLVAMFYQSVLKNKSEFLEVYQLEDQFPKEFVFILMI 1292
Query: 1262 VFFLIVLDRIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFT 1321
+FFLIV+DRIIYL SFAT KVIFYLF+LVLFTYSVT+YAW M+ + R G LRAIY T
Sbjct: 1293 LFFLIVVDRIIYLWSFATGKVIFYLFNLVLFTYSVTEYAWGMELVHRNVGGFVLRAIYLT 1352
Query: 1322 KAISLVLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCT 1381
K+ISL LQA+Q +G+P+KS LYRQFLTS V+++N GFRLYRA+PFLYELRCVLDWSCT
Sbjct: 1353 KSISLALQALQIRYGIPNKSNLYRQFLTSKVTQVNYFGFRLYRALPFLYELRCVLDWSCT 1412
Query: 1382 TTSLTMYDWLKL 1393
TTSLTMYDWLK+
Sbjct: 1413 TTSLTMYDWLKV 1424
Score = 278 bits (711), Expect = 1e-72
Identities = 156/351 (44%), Positives = 218/351 (61%), Gaps = 17/351 (4%)
Query: 29 IDLQTAFGYNPAASMFQNIWESLAILVVMQLYSYERRQSKSSGSSDFDLPEVGPFPFTRR 88
+ L G++P AS+ N+W+SLA+LVVMQLYSYERRQ+ D E G F RR
Sbjct: 103 VKLYPDLGFDPEASLLMNVWQSLAVLVVMQLYSYERRQNSDKNFGVSDASESGLLGFLRR 162
Query: 89 LLVRHTEKILFLALFYASLSPISAFGFLYLLGLINCSRLPKSSQIPAKVFLVYSGLLIMV 148
LL+ H+EKIL + +FYA LS IS G +YLLGLI S LPK S+IP+KV+LVY+GLL
Sbjct: 163 LLIWHSEKILSVTVFYACLSSISLSGLIYLLGLIMFSILPKVSRIPSKVYLVYTGLLATS 222
Query: 149 EYLFQMLGDQAEMFPGQKYSQLSLIMGLQLYKPGFKGLESGFRGKVVVIVACILQYNVFR 208
EYLFQML + A+M PGQ++ LS+ +GL+ Y GF G+E G RGKV+VIVAC +QYNVF
Sbjct: 223 EYLFQMLCEPAQMCPGQQFHGLSVFLGLKHYDAGFWGVEYGLRGKVLVIVACTIQYNVFH 282
Query: 209 WLEKMQYVDPNGGKWNEPCPLFNMVDIPNETTACTPQNKQAETSTS------ATIKRLAR 262
WL+ M + GKW EPC LF I +T++ N + S++ + ++ L
Sbjct: 283 WLDLMPTSLLHEGKWEEPCQLF----ISGDTSSNARDNNKDSHSSNRFSSLFSKVQGLIG 338
Query: 263 SRSCPTVNSALSQGTDSGIE-----GDSTKKLRQLHFWESSKDSLKWNRKRLLFLRKERL 317
S S +++S + T ++ D K+ W SK+S KW++++++ LR+ER
Sbjct: 339 SSSSSSLSSGSTCQTSEPVQNETSGSDEGKRYSFSKIWGMSKESHKWDKRKIISLRRERF 398
Query: 318 EMQKTALKVSLKFWIENMFTLFGLEINMIALLLASFAVLNAISLLYIASLA 368
E QKT K +KFW+EN+F L GLEINMI + V +++L +I + A
Sbjct: 399 ETQKTTFKCYMKFWMENLFKLRGLEINMIVKECPGYTV--SMTLKFIVANA 447
>gb|AAD13708.1| unknown protein [Arabidopsis thaliana] gi|15227203|ref|NP_182325.1|
expressed protein [Arabidopsis thaliana]
gi|25370846|pir||F84922 hypothetical protein At2g48040
[imported] - Arabidopsis thaliana
Length = 294
Score = 364 bits (934), Expect = 2e-98
Identities = 180/312 (57%), Positives = 226/312 (71%), Gaps = 25/312 (8%)
Query: 1446 MYSSGNPTNIANPIKDASARVHIKTLSGRLTLFETTLCEIISWETLEARTSLDPKGYLST 1505
MYSSGNPTNIANPIKDAS ++ +KT+ G+LTL++TTLCE IS + ++ L + +L T
Sbjct: 1 MYSSGNPTNIANPIKDASVQIDLKTVGGKLTLYQTTLCERISGDNIDLGLDLGSQSFLPT 60
Query: 1506 YNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSW--NMDLAFSWEFTRDRPRGKEVVKY 1563
YN+ DIQLICCQ+DAS LWLVP V RF++SL W +MD+ F+W RDRP+GKE VKY
Sbjct: 61 YNKNDIQLICCQADASVLWLVPDTVVTRFIQSLDWDTDMDITFTWVLNRDRPKGKETVKY 120
Query: 1564 ELTIQEQDLPKSSEVTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSLEQSVELVSGDLVL 1623
E ++ DLPK S++ V NG+ + F N+YP++FRVTGSGDVRS E + VS D+++
Sbjct: 121 ERSVDPLDLPKRSDIQMVLNGSMDGFRVHNLYPKFFRVTGSGDVRSFEDQTDEVSADILI 180
Query: 1624 NRGNPE-WWSFYDLDISD-AHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIWGLYITFV 1681
N N + WWSF++L S+ C GP+AII+SEETP
Sbjct: 181 NHANFKWWWSFHNLKASENISACEGMDGPVAIIMSEETPP-------------------- 220
Query: 1682 LAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLYWTLVKIYR 1741
VGRFIRLQCSDLRMRIP+ENLPSCDRL+A+CED+YAARAEGEL VEEVLYWTLVKIYR
Sbjct: 221 -PVGRFIRLQCSDLRMRIPYENLPSCDRLIAICEDLYAARAEGELGVEEVLYWTLVKIYR 279
Query: 1742 TPHMLLEYTQAE 1753
+PHMLLEYT+ +
Sbjct: 280 SPHMLLEYTKLD 291
>ref|XP_640187.1| hypothetical protein DDB0218401 [Dictyostelium discoideum]
gi|60468250|gb|EAL66260.1| hypothetical protein
DDB0218401 [Dictyostelium discoideum]
Length = 3080
Score = 227 bits (579), Expect = 2e-57
Identities = 159/590 (26%), Positives = 278/590 (46%), Gaps = 54/590 (9%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQLEDQFPEDFVLVLMVVFFLIVLDR 1270
D Y + D A F + IF ++ S + + ++ P ++++L+ F +I+LDR
Sbjct: 2456 DYYMPLLFTDFACLFFLVIFPQNFTGIPSSDIAEFLEQNVIPRQYIVILLAQFGVIILDR 2515
Query: 1271 IIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDM--DPLRRYSGRLALRAIYFTKAISLVL 1328
IIYL AK + + VL+ + Y D+ P + L Y K I L
Sbjct: 2516 IIYLYKSVKAKFVLQIVLTVLYHVFLFFYFPDLIVKPFS-FGYTWPLVVFYLMKCIYLYY 2574
Query: 1329 QAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLTMY 1388
A+Q +G P S +FL S + +G+ LY+AIPF+YELR +LDW T T++ Y
Sbjct: 2575 SALQICYGYPILS--QNRFLMDGYSDFHNIGYALYKAIPFVYELRTLLDWIATDTTMLFY 2632
Query: 1389 DWLKLEDIHASLFLVKCDVD-LNRASHQQGQKQTKMTKFCNGICLFFVLMCVIWAPMLMY 1447
DWLK ED+++++F VKC ++ + R Q+G KQ K KF G+ F L+ ++W P+++
Sbjct: 2633 DWLKFEDLYSTIFSVKCRLEWIKRQGRQKGHKQPKFEKFVTGVTFFIGLVILLWFPLIIL 2692
Query: 1448 SSGNPTNIANPIKDASARVHIKTLSGRLTLFETTLCEIISWETLEARTSL----DPKGYL 1503
SSG P + P+ + V + + L + + E ++ + S + +L
Sbjct: 2693 SSGLPGSNIEPVNNIQIEVSVVGWNPFLKINQDLSLESGDGDSTMDQNSFNNLKEDYSFL 2752
Query: 1504 STYNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSWNMDLAFSWEFTRDRPRG-KEVVK 1562
+T + + IQ I + + +W + P +A+ + L N L +T R G V+
Sbjct: 2753 TTDDRQGIQNISINTFSEEIWNLSPPAKAQLINYLLTNTSLQIEVSYTLTRSGGVNSVIV 2812
Query: 1563 YELTIQ---EQDLPKSSEVTEVFNGT---------------------------SNSFSAF 1592
++Q +Q+L + +T V + SNSF
Sbjct: 2813 GSNSVQLKPDQELNFYNILTRVQSNNSNNSNNPNENSSSGSDDNNNNSNNNIGSNSFIVE 2872
Query: 1593 NIYPRYFRVTG-----------SGDVRSLEQSVELVSGDLVLNRGNPEW-WSFYDLDISD 1640
++ ++ ++ G +G+V +L+ + S + + P++ W D +
Sbjct: 2873 GLFYKFIKLPGVSGNPIYPTDNNGNVLALDALFTMNSTNFNNSNQPPQYFWQVNAYDSTK 2932
Query: 1641 AHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIP 1700
+ +S + P GI TL I GLY++ VL+VGRF+RL + + ++I
Sbjct: 2933 TYNISTLESIQFYTISSKLPNGIT-STLVSAGIIGLYVSVVLSVGRFLRLSITQISIKIQ 2991
Query: 1701 FENLPSCDRLMALCEDIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYT 1750
ENL SCD ++ + +D++ AR G+L +EE LY L++++R P +L T
Sbjct: 2992 LENLDSCDEILKMIDDVFIAREYGDLVLEEELYHELIQVFRQPQLLYSLT 3041
Score = 41.6 bits (96), Expect = 0.25
Identities = 61/290 (21%), Positives = 119/290 (41%), Gaps = 45/290 (15%)
Query: 320 QKTALKVSLKFWIENMFTLFGLEINMIALLLASFAVLNAISLLYIASLAACVLLHRLLIK 379
+K LK L ++I N + L+GLE + L A F LN + ++Y+ +A + + + +
Sbjct: 1254 KKENLKKRLFWYISNFYELYGLECVFLVLAFALFWRLNILGMIYLIIIAVGLNIDKRNLH 1313
Query: 380 KLWPIFV-FLFASVVTVEYLAIWMRLTFMNLEIEEHVPCRDCWRVSDIYFSY-CRKCWLG 437
KL I+V L A + ++YL I + T E P W + ++ L
Sbjct: 1314 KL--IYVSALLAPTILIQYLLILVVPT-----KENSYP----WLDHPFFLNHKTIDNLLL 1362
Query: 438 IIVDDPRMLIGYYGVFMFSCFKFRADKSSSL-----------TGLEMYQKIHSQWKSASV 486
+ + D +L+ + V FS F+ L ++ Q+ H KS +
Sbjct: 1363 LSIPDRYVLVIDFLVLFFSMLLFKQRNGYYLYKDFELHQQQQLNQQLNQQQHDSLKSIAS 1422
Query: 487 LSNLSFE---------------------TKGYWTFLDHLRLYGYCHLLDFVLSLILITGT 525
++ S + TK ++ + LR + +L +I + GT
Sbjct: 1423 SNSSSQKPLHPKNFFKFNSSIDGGDDDFTKEPRSWSNELRYMIIRYSSQVILIVIFLAGT 1482
Query: 526 LEYDFLHLGYLGFALVFFRMRLKILKQGNNIFRYLRMYNFAVIVLSLAYQ 575
E D L Y+ F++ ++ + +++ L +YN+ V++ + +Q
Sbjct: 1483 AECDILSCFYVFFSVYVLFSGNAHSRKWSYLWKSLHIYNWLVLMAQIIFQ 1532
Score = 40.8 bits (94), Expect = 0.42
Identities = 20/47 (42%), Positives = 33/47 (69%), Gaps = 1/47 (2%)
Query: 917 SLLSVVYLAALFLYA-LCQNTGPSYIFWVLILIYTEVCILLQYLYQI 962
S++S+VYL A+FLY L ++ PS FW ++ Y+ + I L+Y++QI
Sbjct: 2086 SIISLVYLLAVFLYGRLFESPRPSKNFWRFMIGYSSLIICLKYVFQI 2132
>emb|CAF91003.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1183
Score = 219 bits (557), Expect = 9e-55
Identities = 155/573 (27%), Positives = 277/573 (48%), Gaps = 46/573 (8%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F D+ F +I + + K ++ L EDQ PE F+ +L++ F +++D
Sbjct: 623 DVYALMFLTDVIDFIVIIFGFWAFGKHSAAADIASSLSEDQVPEAFLAMLLIQFSTMIID 682
Query: 1270 RIIYLCSFATAKVIFYL---FSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISL 1326
R +YL K+IF + F + L+ + + + R ++ + YF K I
Sbjct: 683 RALYLRKAVLGKLIFQVILVFGIHLWMFFILPAVTE----RMFNQNFVAQLWYFVKCIYF 738
Query: 1327 VLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLT 1386
L A+Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L+
Sbjct: 739 GLSALQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLS 796
Query: 1387 MYDWLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPML 1445
+ +W+ +EDI+A++F++KC + + Q +GQK+ K+ K+ G + F L+C+IW P+L
Sbjct: 797 LSNWMCVEDIYANIFIIKCSRETEKKYPQPKGQKKKKIVKYGMGGLIIFFLICIIWFPLL 856
Query: 1446 MYSS--------GNPTNIANPIKDASARVHIKTLSGRLTLFETTLCEIISWETLEARTSL 1497
S +P ++ IK + T+S + + T + S
Sbjct: 857 FISLVRSVVGVVNHPIDVTVTIK-LGGYEPLFTMSVQQQSIQPFTKAEYEQLTQKFHDSA 915
Query: 1498 DPKGYLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSWN---MDLAFSWEFTRDR 1554
+++ YN +DI + + ++W + P + +K L + + L +W F RD
Sbjct: 916 VAMQFITLYNYEDIVTARIEGSSGSVWRISPPSRQEVIKELLTSPAHLTLRLAWNFQRDL 975
Query: 1555 PRGKEVV----KYELTIQEQDLPKSSEVTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSL 1610
+G V K+ + ++ + ++ + + S + +P Y R + + +
Sbjct: 976 GKGGTVEHSFDKHSVPLEPGNPVRAELASLLLGNRSEPAHVPHFFPNYLRAPNGAEAKPV 1035
Query: 1611 EQ-----SVELVSGDLVLNRGNP-------EWWSFYDLDISDAHGCGKFPGPMAIIVSEE 1658
Q + ++ L L P EWW D+ I A G P M I +
Sbjct: 1036 SQLYQGGEEDYLNISLSLKSDRPQRSGVTQEWW---DVAIEGASSSGVLP--MVIFNDKV 1090
Query: 1659 TPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIY 1718
+P + L+ + I GLY++ VL +G+F+R S++ I FE LP DR++ LC DI+
Sbjct: 1091 SPPSL--GFLAGYGIMGLYVSVVLVIGKFVRGFFSEISHSIMFEELPCVDRILKLCTDIF 1148
Query: 1719 AARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
R GELE+EE LY L+ +YR+P ++++T+
Sbjct: 1149 LVRETGELELEEELYCKLIFLYRSPETMIKWTR 1181
Score = 50.1 bits (118), Expect = 7e-04
Identities = 26/103 (25%), Positives = 56/103 (54%)
Query: 892 IWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLILIYTE 951
+++ + +++EV CY +L + S++S+V +FL+A+ P+ FW+ ++YTE
Sbjct: 325 MYNLLAAHSEVACYFIMVLNNMVTASVISLVLPILVFLWAMLSVPRPTKRFWMTAIVYTE 384
Query: 952 VCILLQYLYQIIIQHTEFEFPMSLLQELGFPAKKITSSFVTSN 994
V ++++Y++Q + M+L ++ F +I T N
Sbjct: 385 VMVVVKYVFQFGFFPWNSVYEMALNEDKPFFPPRILGLEKTDN 427
>sp|Q92508|FA38A_HUMAN Protein FAM38A gi|7662014|ref|NP_055560.1| family with sequence
similarity 38, member A [Homo sapiens]
gi|1510143|dbj|BAA13240.1| similar to C.elegans protein
encoded in cosmid T20D3 (Z68220). [Homo sapiens]
Length = 2035
Score = 216 bits (551), Expect = 4e-54
Identities = 155/571 (27%), Positives = 282/571 (49%), Gaps = 41/571 (7%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F AD+ F +I + + K ++ L +DQ PE F+++L++ F +V+D
Sbjct: 1473 DVYALMFLADVVDFIIIIFGFWAFGKHSAATDITSSLSDDQVPEAFLVMLLIQFSTMVVD 1532
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQ 1329
R +YL K+ F + +LVL + + R ++ + + YF K I L
Sbjct: 1533 RALYLRKTVLGKLAFQV-ALVLAIHLWMFFILPAVTERMFNQNVVAQLWYFVKCIYFALS 1591
Query: 1330 AIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYD 1389
A Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L++
Sbjct: 1592 AYQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLSLSS 1649
Query: 1390 WLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPMLMYS 1448
W+ +EDI+A++F++KC + + Q +GQK+ K+ K+ G + L+ +IW P+L S
Sbjct: 1650 WMCVEDIYANIFIIKCSRETEKKYPQPKGQKKKKIVKYGMGGLIILFLIAIIWFPLLFMS 1709
Query: 1449 S-GNPTNIANPIKDASARVHIKTLSGRLTLF--ETTLCEIISWETLEARTSLDPKG---- 1501
+ + N D + + + T+ + ++ + E DP+
Sbjct: 1710 LVRSVVGVVNQPIDVTVTLKLGGYEPLFTMSAQQPSIIPFTAQAYEELSRQFDPQPLAMQ 1769
Query: 1502 YLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSL---SWNMDLAFSWEFTRDRPRGK 1558
++S Y+ +DI + + LW + P +A+ + L + ++ L F+W F RD +G
Sbjct: 1770 FISQYSPEDIVTAQIEGSSGALWRISPPSRAQMKRELYNGTADITLRFTWNFQRDLAKGG 1829
Query: 1559 EVV----KYELTIQEQDLPKSSEVTEVFNGTSNSFSAF-NIYPRYFRVTGSGDVRSLEQ- 1612
V K+ L + + ++ + GTS+ N++P+Y R + ++Q
Sbjct: 1830 TVEYANEKHMLALAPNSTARR-QLASLLEGTSDQSVVIPNLFPKYIRAPNGPEANPVKQL 1888
Query: 1613 ----SVELVSGDLVLNR-------GNPEWWSFYDLDISDAH-GCGKFPGPMAIIVSEETP 1660
+ + + L R G EWW +++ + C P +I S++
Sbjct: 1889 QPNEEADYLGVRIQLRREQGAGATGFLEWWV---IELQECRTDCNLLP---MVIFSDKVS 1942
Query: 1661 QGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAA 1720
+G L+ + I GLY++ VL +G+F+R S++ I FE LP DR++ LC+DI+
Sbjct: 1943 PPSLG-FLAGYGIMGLYVSIVLVIGKFVRGFFSEISHSIMFEELPCVDRILKLCQDIFLV 2001
Query: 1721 RAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
R ELE+EE LY L+ +YR+P ++++T+
Sbjct: 2002 RETRELELEEELYAKLIFLYRSPETMIKWTR 2032
Score = 50.8 bits (120), Expect = 4e-04
Identities = 22/74 (29%), Positives = 47/74 (62%)
Query: 888 LVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLIL 947
L+R ++ + +++E++CY IL ++ S S+V +FL+A+ PS FW+ +
Sbjct: 1189 LLRAVYQCVAAHSELLCYFIIILNHMVTASAGSLVLPVLVFLWAMLSIPRPSKRFWMTAI 1248
Query: 948 IYTEVCILLQYLYQ 961
++TE+ ++++YL+Q
Sbjct: 1249 VFTEIAVVVKYLFQ 1262
>ref|XP_546782.1| PREDICTED: similar to Protein FAM38A [Canis familiaris]
Length = 2681
Score = 216 bits (549), Expect = 7e-54
Identities = 159/574 (27%), Positives = 281/574 (48%), Gaps = 46/574 (8%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F AD+ F +I + + K ++ L +DQ PE F+++L++ F +V+D
Sbjct: 2118 DVYALMFLADVVDFIIIIFGFWAFGKHSAATDITSSLSDDQVPEAFLVMLLIQFSTMVID 2177
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQ 1329
R +YL K+ F + LVL + + R ++ + YF K I L
Sbjct: 2178 RALYLRKTVLGKLAFQVV-LVLAIHLWMFFILPAVTERMFNQNAVAQLWYFVKCIYFSLS 2236
Query: 1330 AIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYD 1389
A Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L++
Sbjct: 2237 AYQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLSLSS 2294
Query: 1390 WLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPMLMYS 1448
W+ +EDI+A++F++KC + + Q +GQK+ K+ K+ G + L+ +IW P+L S
Sbjct: 2295 WMCVEDIYANIFIIKCSRETEKKYPQPKGQKKKKIVKYGMGGLIILFLVAIIWFPLLFMS 2354
Query: 1449 S-GNPTNIANPIKDASARVHIKTLSGRLTLF--ETTLCEIISWETLEARTSLDPKG---- 1501
+ + N D + + + T+ + ++ E DP
Sbjct: 2355 LVRSVVGVVNQPIDVTVTLKLGGYEPLFTMSAQQPSIVPFTQQAYEELSRQFDPNPLAMQ 2414
Query: 1502 YLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSL---SWNMDLAFSWEFTRDRPRGK 1558
++S Y+ +DI + + LW + P +A+ + L + ++ L F+W F RD +G
Sbjct: 2415 FISQYSPEDIVTAQIEGSSGALWRISPPSRAQMKRELYNGTADITLRFTWNFQRDLAKGG 2474
Query: 1559 EVVKYELTIQEQDL---PKSSE---VTEVFNGTSNSFSAF-NIYPRYFRVTGSGDVRSLE 1611
V E T ++ L P S+E + + GTS+ +++P+Y R + ++
Sbjct: 2475 TV---EYTNEKHTLDWAPNSTERRQLASLLEGTSDQSVVIPHLFPKYIRAPNGPEANPVK 2531
Query: 1612 Q-----SVELVSGDLVLNR--------GNPEWWSFYDLDISDAHG-CGKFPGPMAIIVSE 1657
Q + + + L R G EWW +++ D C P +I S+
Sbjct: 2532 QLQPNEEADYLGVRIQLRRERVGSGAAGFLEWWV---IELQDCQAECNLLP---MVIFSD 2585
Query: 1658 ETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDI 1717
+ +G L+ + I GLY++ VL +G+F+R S++ I FE LP DR++ LC+DI
Sbjct: 2586 KVSPPSLG-FLAGYGIMGLYVSIVLVIGKFVRGFFSEISHSIMFEELPCVDRILKLCQDI 2644
Query: 1718 YAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
+ R ELE+EE LY L+ +YR+P ++++T+
Sbjct: 2645 FLVRETRELELEEELYAKLIFLYRSPETMIKWTR 2678
Score = 48.5 bits (114), Expect = 0.002
Identities = 23/74 (31%), Positives = 45/74 (60%)
Query: 888 LVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLIL 947
L++ + + +++E++CY IL ++ S S+V +FL+A+ PS FW+ +
Sbjct: 1831 LLQAAYQCVAAHSELLCYFIIILNHMVTASATSLVLPVLVFLWAMLSIPRPSKRFWMTAI 1890
Query: 948 IYTEVCILLQYLYQ 961
I+TEV ++ +YL+Q
Sbjct: 1891 IFTEVSVVTKYLFQ 1904
>ref|XP_225880.3| PREDICTED: similar to family with sequence similarity 38, member B
[Rattus norvegicus]
Length = 2826
Score = 210 bits (535), Expect = 3e-52
Identities = 159/576 (27%), Positives = 280/576 (48%), Gaps = 50/576 (8%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+Y +F AD F +I + + K ++ L EDQ P F++++++ F +V+D
Sbjct: 2262 DVYVLMFLADTVDFIIIVFGFWAFGKHSAAADITSSLSEDQVPGPFLVMVLIQFGTMVVD 2321
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPL---RRYSGRLALRAIYFTKAISL 1326
R +YL KVIF V+ + + + + + P R++S L + YF K +
Sbjct: 2322 RALYLRKTVLGKVIFQ----VILVFGIHFWMFFILPSVTERKFSQNLVAQLWYFVKCVYF 2377
Query: 1327 VLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLT 1386
L A Q G P + + FLT S + +N+ F+ +R +PFL ELR V+DW T T+L+
Sbjct: 2378 GLSAYQIRCGYPTRVL--GNFLTKSYNYVNLFLFQGFRLVPFLTELRAVMDWVWTDTTLS 2435
Query: 1387 MYDWLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPML 1445
+ W+ +EDI+A +F++KC + + Q +GQK+ K K+ G + +L+C++W P+L
Sbjct: 2436 LSSWICVEDIYAHIFILKCWRESEKRYPQPRGQKKKKAVKYGMGGMIIVLLICIVWFPLL 2495
Query: 1446 MYSS-GNPTNIANPIKDASARVHIKTLSGRLTLF-----ETTLCEIISWETLEARTSLDP 1499
S + + N D S + TL G +F ++ L + + E S P
Sbjct: 2496 FMSLIKSVAGVINQPLDVSVTI---TLGGYQPIFTMSAQQSQLKVMDGTKYNEFIKSFGP 2552
Query: 1500 KG----YLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSWN---MDLAFSWEFTR 1552
+L Y +DI + + ++++LW + P + + ++ L+ + FSW R
Sbjct: 2553 NSGAMQFLENYERQDITVAELEGNSNSLWTISPPSKQKMIQELTDPNSCFSVVFSWSIQR 2612
Query: 1553 DRPRGK--EVVKYELTIQEQDLPKSSEVTEVFNGTSNSFSA----FNIYPRYFRVTGSGD 1606
+ G E+ +L+ D +SS + + S + IYP Y + +
Sbjct: 2613 NMTLGAKAEIATDKLSFPLTDATRSSIAKMIAGNDTESSNTPVTIEKIYPYYVKAPSDSN 2672
Query: 1607 VRSLEQSVELVSGDLVLN-----------RGNPEWWSFYDLDISDAHGCGKFPGPMAIIV 1655
+ ++Q L+S + +N + N EWW +L S G + +
Sbjct: 2673 SKPIKQ---LLSENNFMNITIILFRDNVTKSNSEWWVL-NLTGSRIFNQGSQALELVVFN 2728
Query: 1656 SEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCE 1715
+ +P + L+ + I GLY + VL +G+F+R S + I FE LP+ DR++ LC
Sbjct: 2729 DKVSPPSL--GFLAGYGIMGLYASVVLVIGKFVREFFSGISHSIMFEELPNVDRILKLCT 2786
Query: 1716 DIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
DI+ R GELE+EE LY L+ +YR+P ++++T+
Sbjct: 2787 DIFLVRETGELELEEDLYAKLIFLYRSPETMIKWTR 2822
Score = 52.4 bits (124), Expect = 1e-04
Identities = 22/75 (29%), Positives = 49/75 (65%)
Query: 887 ILVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLI 946
+L +++ + + +E+VCY IL ++ + S+++++ +FL+A+ PS FW++
Sbjct: 1983 LLFYAMYNTLVARSEMVCYFVIILNHMISASIITLLLPILIFLWAMLSVPRPSRRFWMMA 2042
Query: 947 LIYTEVCILLQYLYQ 961
++YTEV I+++Y +Q
Sbjct: 2043 IVYTEVAIVVKYFFQ 2057
>dbj|BAC97905.2| mKIAA0233 protein [Mus musculus]
Length = 1482
Score = 207 bits (526), Expect = 3e-51
Identities = 156/582 (26%), Positives = 280/582 (47%), Gaps = 54/582 (9%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F AD+ +I + + K ++ L +DQ P+ F+ +L+V F +V+D
Sbjct: 911 DVYALMFLADIVDIIIIIFGFWAFGKHSAATDIASSLSDDQVPQAFLFMLLVQFGTMVID 970
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQ 1329
R +YL K+ F + LV+ + + R +S + YF K I L
Sbjct: 971 RALYLRKTVLGKLAFQVV-LVVAIHIWMFFILPAVTERMFSQNAVAQLWYFVKCIYFALS 1029
Query: 1330 AIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYD 1389
A Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L++ +
Sbjct: 1030 AYQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLSLSN 1087
Query: 1390 WLKLEDIHASLFLVKCDVDLNRASHQQGQKQTKMTKFCNGICLFFVLMCVIWAPMLMYS- 1448
W+ +EDI+A++F++KC + + Q+G+K+ K+ K+ G + L+ +IW P+L S
Sbjct: 1088 WMCVEDIYANIFIIKCSRETEKVPGQRGRKKKKIVKYGMGGLIILFLIAIIWFPLLFMSL 1147
Query: 1449 SGNPTNIANPIKDASARVHIKTLSGRLTLF--ETTLCEIISWETLEARTSLDP----KGY 1502
+ + N D + + + T+ + ++ E DP +
Sbjct: 1148 IRSVVGVVNQPIDVTVTLKLGGYEPLFTMSAQQPSIVPFTPQAYEELSQQFDPYPLAMQF 1207
Query: 1503 LSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSL---SWNMDLAFSWEFTRDRPRGKE 1559
+S Y+ +DI + + LW + P +A+ + L + ++ L F+W F RD +G
Sbjct: 1208 ISQYSPEDIVTAQIEGSSGALWRISPPSRAQMKQELYNGTADITLRFTWNFQRDLAKGGT 1267
Query: 1560 VVKYELTIQEQDL---PKSS---EVTEVFNG-TSNSFSAFNIYPRYFRVTGSGDVRSLEQ 1612
V E T ++ L P S+ ++ ++ G S +++P+Y R + ++Q
Sbjct: 1268 V---EYTNEKHTLELAPNSTARRQLAQLLEGRPDQSVVIPHLFPKYIRAPNGPEANPVKQ 1324
Query: 1613 ------------SVEL--------VSGDLVLNRGNP--EWWSFYDLDISDAHG-CGKFPG 1649
++L SG+ + + EWW +++ D C
Sbjct: 1325 LQPDEEEDYLGVRIQLRREQVGTGASGEQAGTKASDFLEWWV---IELQDCKADCNLL-- 1379
Query: 1650 PMAIIVSEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDR 1709
PM I + +P + L+ + I GLY++ VL VG+F+R S++ I FE LP DR
Sbjct: 1380 PMVIFSDKVSPPSL--GFLAGYGIVGLYVSIVLVVGKFVRGFFSEISHSIMFEELPCVDR 1437
Query: 1710 LMALCEDIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
++ LC+DI+ R ELE+EE LY L+ +YR+P ++++T+
Sbjct: 1438 ILKLCQDIFLVRETRELELEEELYAKLIFLYRSPETMIKWTR 1479
Score = 48.1 bits (113), Expect = 0.003
Identities = 23/74 (31%), Positives = 45/74 (60%)
Query: 888 LVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLIL 947
L+R + + +++E++CY IL ++ S S+V +FL+A+ PS FW+ +
Sbjct: 611 LLRAGYQCVAAHSELLCYFIIILNHMVTASAASLVLPVLVFLWAMLTIPRPSKRFWMTAI 670
Query: 948 IYTEVCILLQYLYQ 961
++TEV ++ +YL+Q
Sbjct: 671 VFTEVMVVTKYLFQ 684
>ref|XP_419138.1| PREDICTED: similar to Hypothetical protein KIAA0233 [Gallus gallus]
Length = 3198
Score = 204 bits (519), Expect = 2e-50
Identities = 158/586 (26%), Positives = 276/586 (46%), Gaps = 58/586 (9%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+Y +F AD F +I + + K ++ L EDQ PE F++++++ F +V+D
Sbjct: 2622 DVYVLMFLADTVDFIIIVFGFWAFGKHSAAADITSSLSEDQVPEAFLVMVLIQFGTMVVD 2681
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDP---LRRYSGRLALRAIYFTKAISL 1326
R +YL KVIF V+ + + + + + P R++S + YF K +
Sbjct: 2682 RALYLKKTVMGKVIFQ----VILVFGIHFWMFFILPGVTERKFSQNTVAQLWYFVKCVYF 2737
Query: 1327 VLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLT 1386
L A Q G P + + FLT S + +N+ F+ +R +PFL ELR V+DW T T+L+
Sbjct: 2738 GLSAYQIRCGYPTRVL--GNFLTKSYNYVNLFLFQGFRLVPFLTELRAVMDWVWTDTTLS 2795
Query: 1387 MYDWLKLEDIHASLFLVKCDVD------------------LNRASHQQGQKQTKMTKFCN 1428
+ W+ +EDI+A +F++KC D L R +GQK+ K+ K+
Sbjct: 2796 LSSWICVEDIYAHIFILKCWRDRRPESLRLIHSSNRCCLLLQRYPQPRGQKKKKVVKYGM 2855
Query: 1429 GICLFFVLMCVIWAPMLMYSS-GNPTNIANPIKDASARVHIKTLSGRLTLF----ETTLC 1483
G + +L+C++W P+L S + I N D S + TL G +F + +
Sbjct: 2856 GGMIIVLLICIVWFPLLFMSLIKSVAGITNKPLDVSITI---TLGGYQPIFTMSAQQSQL 2912
Query: 1484 EIISWETLEA-----RTSLDPKGYLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSL 1538
+ ++ A R + +L Y ++DI L + ++++LW + P + + ++ L
Sbjct: 2913 KDLNQTGFSAFLGSYRGNTAALQFLEGYGKEDITLADLEGNSNSLWTISPPSREKMIQGL 2972
Query: 1539 ---SWNMDLAFSWEFTRDRPRGK--EVVKYELTIQEQDLPKSSEVTEVFNGTSNSFSAFN 1593
S + SW R+ G E+ +LT + + T + +
Sbjct: 2973 LDFSAEFTVVLSWSIQRNLTLGAKAEIASDKLTFGLPEKTRRDIATMMSGKPLEKVTLET 3032
Query: 1594 IYPRYFRVTGSGDVRSLEQSV-----ELVSGDLVLN---RGNPEWWSFYDLDISDAHGCG 1645
+YP Y + + ++Q + E ++ LV N G EWW L +
Sbjct: 3033 VYPYYIKAPSDSLAKPIKQLLTDCRWENITVSLVKNVSEEGVREWWVLNQL--GKRYKTN 3090
Query: 1646 KFPGPMAIIVSEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLP 1705
+ + I + +P + L+ + I GLY + VL +G+F+R S + I FE LP
Sbjct: 3091 EESLELFIFSDKVSPPSL--GFLAGYGIMGLYASVVLVIGKFVREFFSGISHSIMFEELP 3148
Query: 1706 SCDRLMALCEDIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
+ DR++ LC DI+ R GELE+EE LY L+ +YR+P ++++T+
Sbjct: 3149 NVDRILKLCTDIFLVRETGELELEEDLYAKLIFLYRSPETMIKWTR 3194
Score = 54.7 bits (130), Expect = 3e-05
Identities = 23/75 (30%), Positives = 49/75 (64%)
Query: 887 ILVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLI 946
+L+ +++ + + +E+VCY IL ++ + S++++V +FL+A+ PS FW+
Sbjct: 2343 LLIYALYNTLVARSEMVCYFVIILNHMISASMITLVLPILIFLWAMLSVPRPSKRFWMTA 2402
Query: 947 LIYTEVCILLQYLYQ 961
++YTEV I+++Y +Q
Sbjct: 2403 IVYTEVAIVIKYFFQ 2417
Score = 47.4 bits (111), Expect = 0.004
Identities = 92/403 (22%), Positives = 154/403 (37%), Gaps = 82/403 (20%)
Query: 328 LKFWIENMFTLFGLE------INMIALLLASFAVLNAISLLYIASLAACVLLHRLLIKKL 381
+K++I F FGLE +N+I + +A+++A L+ A R I ++
Sbjct: 1567 VKYFINYFFYKFGLETCFLLSVNVIGQRMDFYAMIHAFWLI-----AVLYRRRRKAIAEI 1621
Query: 382 WPIFVFLFASVVTVEYLAIWMRLTFMNLEIEEHVPCRDCWRVSDIYFSYCRKCWLGIIVD 441
WP + A ++T +Y F+ + I PC+D D C D
Sbjct: 1622 WPKYCCFLACIITFQY--------FLCIGIPP-APCKD-----DFMLLLCASLQRQTFED 1667
Query: 442 DPRMLIGYY-GVFMFSCFKFRADKSSSLTGLEMYQKIHSQWKSASVLSNLSFETKGYWTF 500
+ + + G + C A S + + IH + ++
Sbjct: 1668 ENKAAVRIMAGDNVEICMNLEAASFSQHNPVPDF--IHCR------------------SY 1707
Query: 501 LDHLRLYGYCHLLDFVLSLILITGTLEYDFLHLGYL--GFALVFFRMRLKILKQGNNIFR 558
LD ++ + +L FVL++I ITGT +GYL F + F L +LK +I R
Sbjct: 1708 LDMYKVIIFSYLFWFVLTIIFITGTTRISIFCMGYLVACFYFLLFGGDL-LLKPIRSILR 1766
Query: 559 Y---LRMYNFAVI----VLSLAYQSPFLGDFSEIKSGFIEYINELVG-----FQKYDY-- 604
Y L YN VI +LS++ F K G YI L+ Q +
Sbjct: 1767 YWDWLIAYNVFVITMKNILSVSSSCVSSEQFRGCKIGACGYIESLIQNSCWLIQAFSLAC 1826
Query: 605 ---GFRITSRSA-----------LVEIIIYMLVSLQSYMFSFPEFDYVSTYLEKEQIGA- 649
G+RI + +A + + I + + LQ +F F +V ++ QI A
Sbjct: 1827 TVKGYRIPTNNADCKLPSGEAGIIWDSICFAFLLLQRRVFMSYYFLHVVADIKASQILAS 1886
Query: 650 ----ILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSE 688
I +Q+K K L + + E +R + E + E
Sbjct: 1887 RMDRIKARQQKYKKGKERMLSMTQDSTEGPEIRKVSEEDDEGE 1929
>dbj|BAC27396.1| unnamed protein product [Mus musculus]
Length = 607
Score = 204 bits (518), Expect = 3e-50
Identities = 156/579 (26%), Positives = 279/579 (47%), Gaps = 56/579 (9%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+Y +F AD F +I + + K ++ L EDQ P F++++++ F +V+D
Sbjct: 43 DVYVLMFLADTVDFIIIVFGFWAFGKHSAAADITSSLSEDQVPGPFLVMVLIQFGTMVVD 102
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDP---LRRYSGRLALRAIYFTKAISL 1326
R +YL KVIF V+ + + + + + P R++S L + YF K +
Sbjct: 103 RALYLRKTVLGKVIFQ----VILVFGIHFWMFFILPGVTERKFSQNLVAQLWYFVKCVYF 158
Query: 1327 VLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLT 1386
L A Q G P + + FLT S + +N+ F+ +R +PFL ELR V+DW T T+L+
Sbjct: 159 GLSAYQIRCGYPTRVL--GNFLTKSYNYVNLFLFQGFRLVPFLTELRAVMDWVWTDTTLS 216
Query: 1387 MYDWLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPML 1445
+ W+ +EDI+A +F++KC + + Q +GQK+ K K+ G + +L+C++W P+L
Sbjct: 217 LSSWICVEDIYAHIFILKCWRESEKRYPQPRGQKKKKAVKYGMGGMIIVLLICIVWFPLL 276
Query: 1446 MYSS-GNPTNIANPIKDASARVHIKTLSGRLTLF-----ETTLCEIISWETLEARTSLDP 1499
S + + N D S + TL G +F ++ L + + + E S P
Sbjct: 277 FMSLIKSVAGVINQPLDVSVTI---TLGGYQPIFTMSAQQSQLKVMDNSKYNEFLKSFGP 333
Query: 1500 KG----YLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSWN---MDLAFSWEFTR 1552
+L Y +D+ + + ++++LW + P + + ++ L+ + FSW R
Sbjct: 334 NSGAMQFLENYEREDVTVAELEGNSNSLWTISPPSKQKMIQELTDPNSCFSVVFSWSIQR 393
Query: 1553 DRPRGKEVVKYELTIQEQDLPKS----SEVTEVFNGTSNSFSAF-----NIYPRYFRVTG 1603
+ G K E+ + P + + + ++ G S IYP Y +
Sbjct: 394 NMTLG---AKAEIATDKLSFPLAVATRNSIAKMIAGNDTESSNTPVTIEKIYPYYVKAPS 450
Query: 1604 SGDVRSLEQSVELVSGDLVLN-----------RGNPEWWSFYDLDISDAHGCGKFPGPMA 1652
+ + ++Q L+S + +N + N EWW +L S G +
Sbjct: 451 DSNSKPIKQ---LLSENNFMNITIILFRDNVTKSNSEWWVL-NLTGSRIFNQGSQALELV 506
Query: 1653 IIVSEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMA 1712
+ + +P + L+ + I GLY + VL +G+F+R S + I FE LP+ DR++
Sbjct: 507 VFNDKVSPPSL--GFLAGYGIMGLYASVVLVIGKFVREFFSGISHSIMFEELPNVDRILK 564
Query: 1713 LCEDIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
LC DI+ R GELE+EE LY L+ +YR+ ++++T+
Sbjct: 565 LCTDIFLVRETGELELEEDLYAKLIFLYRSQETMIKWTR 603
>gb|AAH42207.1| 2310061F22Rik protein [Mus musculus]
Length = 1535
Score = 203 bits (516), Expect = 5e-50
Identities = 156/583 (26%), Positives = 281/583 (47%), Gaps = 55/583 (9%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F AD+ +I + + K ++ L +DQ P+ F+ +L+V F +V+D
Sbjct: 963 DVYALMFLADIVDIIIIIFGFWAFGKHSAATDIASSLSDDQVPQAFLFMLLVQFGTMVID 1022
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQ 1329
R +YL K+ F + LV+ + + R +S + YF K I L
Sbjct: 1023 RALYLRKTVLGKLAFQVV-LVVAIHIWMFFILPAVTERMFSQNAVAQLWYFVKCIYFALS 1081
Query: 1330 AIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYD 1389
A Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L++ +
Sbjct: 1082 AYQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLSLSN 1139
Query: 1390 WLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPMLMYS 1448
W+ +EDI+A++F++KC + + Q +GQK+ K+ K+ G + L+ +IW P+L S
Sbjct: 1140 WMCVEDIYANIFIIKCSRETEKKYPQPKGQKKKKIVKYGMGGLIILFLIAIIWFPLLFMS 1199
Query: 1449 S-GNPTNIANPIKDASARVHIKTLSGRLTLF--ETTLCEIISWETLEARTSLDP----KG 1501
+ + N D + + + T+ + ++ E DP
Sbjct: 1200 LIRSVVGVVNQPIDVTVTLKLGGYEPLFTMSAQQPSIVPFTPQAYEELSQQFDPYPLAMQ 1259
Query: 1502 YLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSL---SWNMDLAFSWEFTRDRPRGK 1558
++S Y+ +DI + + LW + P +A+ + L + ++ L F+W F RD +G
Sbjct: 1260 FISQYSPEDIVTAQIEGSSGALWRISPPSRAQMKQELYNGTADITLRFTWNFQRDLAKGG 1319
Query: 1559 EVVKYELTIQEQDL---PKSS---EVTEVFNGTSNSFSAF-NIYPRYFRVTGSGDVRSLE 1611
V E T ++ L P S+ ++ ++ G + +++P+Y R + ++
Sbjct: 1320 TV---EYTNEKHTLELAPNSTARRQLAQLLEGRPDQSVVIPHLFPKYIRAPNGPEANPVK 1376
Query: 1612 Q------------SVEL--------VSGDLVLNRGNP--EWWSFYDLDISDAHG-CGKFP 1648
Q ++L SG+ + + EWW +++ D C P
Sbjct: 1377 QLQPDEEEDYLGVRIQLRREQVGTGASGEQAGTKASDFLEWWV---IELQDCKADCNLLP 1433
Query: 1649 GPMAIIVSEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCD 1708
+I S++ +G L+ + I GLY++ VL VG+F+R S++ I FE LP D
Sbjct: 1434 ---MVIFSDKVSPPSLG-FLAGYGIVGLYVSIVLVVGKFVRGFFSEISHSIMFEELPCVD 1489
Query: 1709 RLMALCEDIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
R++ LC+DI+ R ELE+EE LY L+ +YR+P ++++T+
Sbjct: 1490 RILKLCQDIFLVRETRELELEEELYAKLIFLYRSPETMIKWTR 1532
Score = 48.1 bits (113), Expect = 0.003
Identities = 23/74 (31%), Positives = 45/74 (60%)
Query: 888 LVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLIL 947
L+R + + +++E++CY IL ++ S S+V +FL+A+ PS FW+ +
Sbjct: 663 LLRAGYQCVAAHSELLCYFIIILNHMVTASAASLVLPVLVFLWAMLTIPRPSKRFWMTAI 722
Query: 948 IYTEVCILLQYLYQ 961
++TEV ++ +YL+Q
Sbjct: 723 VFTEVMVVTKYLFQ 736
>gb|AAN11153.1| CG8486-PB, isoform B [Drosophila melanogaster]
gi|24582718|ref|NP_723354.1| CG8486-PB, isoform B
[Drosophila melanogaster]
Length = 2720
Score = 200 bits (508), Expect = 4e-49
Identities = 170/676 (25%), Positives = 310/676 (45%), Gaps = 77/676 (11%)
Query: 1136 SIEKSEENENLCLVIFEVLYASPPNEFAAEEWYSSLTPAADVSY----EIQKAQHAGILK 1191
SI++ ++++NL L + + + +SL + + S ++++ ++ L
Sbjct: 2029 SIDERDDSDNLSQPDSRQLNDDAAQKLSLQVSQASLPGSPEFSKTGINQLERTKYTSSLY 2088
Query: 1192 EIGFPYRILSVIGGGKREVDLYAYIFGADLAVFFLIAIFYESI---MKANSEFLEVYQLE 1248
+ F S++ + D+YA +F D FF++ + + + E ++ Y E
Sbjct: 2089 KFFF-----SLVHKSRLATDVYALMFLCDFVNFFVLLFGFTAFGTQQTESDEGVQTYLAE 2143
Query: 1249 DQFPEDFVLVLMVVFFLIVLDRIIYLCSFATAKVIFYLFSLV---LFTYSVTKYAWDMDP 1305
++ P F+++L+V F LIV+DR +YL K+IF+ FS++ ++ + V +
Sbjct: 2144 NKVPIPFLIMLLVQFLLIVIDRALYLRKALVNKIIFHFFSVIGIHIWMFFVVPAVTE--- 2200
Query: 1306 LRRYSGRLALRAIYFTKAISLVLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRA 1365
R ++ Y K ++L + Q G P + + F T S +N++ F++Y
Sbjct: 2201 -RTFNSLAPPIIFYVIKCFYMLLSSYQIKSGYPKR--ILGNFFTKGFSMVNMIAFKVYMQ 2257
Query: 1366 IPFLYELRCVLDWSCTTTSLTMYDWLKLEDIHASLFLVKCDVDLNRASH-----QQGQKQ 1420
IPFLYELR +LDW C +++T++DWLK+EDI ++++L++C R S + QK+
Sbjct: 2258 IPFLYELRTILDWVCIDSTMTIFDWLKMEDIFSNIYLIRC----TRQSETDFPAMRAQKK 2313
Query: 1421 TKMTKFCNGICLFFVLMCVIWAPMLMYSSGNPTNIANPIKDASARVH------IKTLSGR 1474
++K G + +++ IW P+ +++ GN +N S + I T +
Sbjct: 2314 ASLSKLIMGGTIVLLIVICIWGPLCLFALGNAVGTSNVPFHVSLSIRIGPYDPIYTTNNY 2373
Query: 1475 LTLFETTLCEIISWETLEARTSLDPKGYLSTYNEKDIQLICCQSDASTLWLVPPIVQARF 1534
++FE E+ S T +++ Y+ D+ + ++ +LW + P + R
Sbjct: 2374 DSIFEIN-PEMYSQMTNAYIKEKQALTFIAGYDATDVAAVRLAGNSPSLWNIAPPDRQRL 2432
Query: 1535 MKSLSWNMDL--AFSWEFTRDRPRG--KEVVKYELTIQEQD-----------LPKSSEVT 1579
+ L N L FS+ TR P KE V E I + L ++ +V
Sbjct: 2433 LNDLRNNHTLKARFSYSLTRKAPAKGLKENVGDEHAISLDESFEGRAALIHMLSETHDVE 2492
Query: 1580 EVF-NGTSNSFSAF--------NIYPRYFRVTGSGD--VRSLEQSVELVSGDLVL----- 1623
++ NGT+N + + P++ +V SGD V S+ LV+
Sbjct: 2493 PIYSNGTTNGTTPEVEEVVVIPGMIPKFIKVLNSGDAAVVSVLSPKHYDYRPLVIKMHRD 2552
Query: 1624 NRGNPEWW--------SFYDLDISDAHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIWG 1675
N N WW +FY+ +S G + +++ L+ I G
Sbjct: 2553 NETNGLWWEIRDYCNDTFYNETLSKFAYSNCTSGIVMYTFNDKKFPSTF-SFLTAGGIIG 2611
Query: 1676 LYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLYWT 1735
LY TFVL RF++ +I FE+LP DR++ LC DIY R E +EE L+
Sbjct: 2612 LYTTFVLLASRFMKSFIGGQNRKIMFEDLPYVDRVLQLCLDIYLVREALEFALEEDLFAK 2671
Query: 1736 LVKIYRTPHMLLEYTQ 1751
L+ +YR+P L+++T+
Sbjct: 2672 LLFLYRSPETLIKWTR 2687
Score = 45.8 bits (107), Expect = 0.013
Identities = 30/117 (25%), Positives = 54/117 (45%), Gaps = 9/117 (7%)
Query: 893 WSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLILIYTEV 952
W + +N +++CY + + N SL+S+ +FL+ P+ FWV ++ YT+
Sbjct: 1890 WYALLANTDLICYIVVFINQVVNASLISLPLPIMVFLWGTLSLPRPTKTFWVTLIAYTQA 1949
Query: 953 CILLQYLYQI-IIQHTEFEFPMSLLQELGFPAKKITSSFVTSNLPFFLIYIFTLLQI 1008
+L++ ++Q +I + P L PAK F N + IY LL +
Sbjct: 1950 IVLIKCIFQFKLIWSNYHQLPNQPLT----PAK----IFGVENKAHYAIYDLILLLV 1998
Score = 42.0 bits (97), Expect = 0.19
Identities = 25/93 (26%), Positives = 49/93 (51%), Gaps = 1/93 (1%)
Query: 308 RLLFLRKERLEMQKTALKVSLKFWIENMFTLFGLEINMIALLLASFAVLNAISLLYIASL 367
+LLF R + +K + + +K+ + F FG+EI++IAL+ + ++++Y L
Sbjct: 1019 KLLFPNIIRADAEKDLVGL-VKYLLNFGFYKFGIEISLIALVSTITYRQDIVAVVYALWL 1077
Query: 368 AACVLLHRLLIKKLWPIFVFLFASVVTVEYLAI 400
+LL R K+W +F FA + +Y+ +
Sbjct: 1078 VVLLLLRRSQCAKIWGVFQAFFAISILTQYIVL 1110
>gb|AAF52611.1| CG8486-PA, isoform A [Drosophila melanogaster]
gi|24582716|ref|NP_609188.1| CG8486-PA, isoform A
[Drosophila melanogaster]
Length = 2771
Score = 200 bits (508), Expect = 4e-49
Identities = 170/676 (25%), Positives = 310/676 (45%), Gaps = 77/676 (11%)
Query: 1136 SIEKSEENENLCLVIFEVLYASPPNEFAAEEWYSSLTPAADVSY----EIQKAQHAGILK 1191
SI++ ++++NL L + + + +SL + + S ++++ ++ L
Sbjct: 2080 SIDERDDSDNLSQPDSRQLNDDAAQKLSLQVSQASLPGSPEFSKTGINQLERTKYTSSLY 2139
Query: 1192 EIGFPYRILSVIGGGKREVDLYAYIFGADLAVFFLIAIFYESI---MKANSEFLEVYQLE 1248
+ F S++ + D+YA +F D FF++ + + + E ++ Y E
Sbjct: 2140 KFFF-----SLVHKSRLATDVYALMFLCDFVNFFVLLFGFTAFGTQQTESDEGVQTYLAE 2194
Query: 1249 DQFPEDFVLVLMVVFFLIVLDRIIYLCSFATAKVIFYLFSLV---LFTYSVTKYAWDMDP 1305
++ P F+++L+V F LIV+DR +YL K+IF+ FS++ ++ + V +
Sbjct: 2195 NKVPIPFLIMLLVQFLLIVIDRALYLRKALVNKIIFHFFSVIGIHIWMFFVVPAVTE--- 2251
Query: 1306 LRRYSGRLALRAIYFTKAISLVLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRA 1365
R ++ Y K ++L + Q G P + + F T S +N++ F++Y
Sbjct: 2252 -RTFNSLAPPIIFYVIKCFYMLLSSYQIKSGYPKR--ILGNFFTKGFSMVNMIAFKVYMQ 2308
Query: 1366 IPFLYELRCVLDWSCTTTSLTMYDWLKLEDIHASLFLVKCDVDLNRASH-----QQGQKQ 1420
IPFLYELR +LDW C +++T++DWLK+EDI ++++L++C R S + QK+
Sbjct: 2309 IPFLYELRTILDWVCIDSTMTIFDWLKMEDIFSNIYLIRC----TRQSETDFPAMRAQKK 2364
Query: 1421 TKMTKFCNGICLFFVLMCVIWAPMLMYSSGNPTNIANPIKDASARVH------IKTLSGR 1474
++K G + +++ IW P+ +++ GN +N S + I T +
Sbjct: 2365 ASLSKLIMGGTIVLLIVICIWGPLCLFALGNAVGTSNVPFHVSLSIRIGPYDPIYTTNNY 2424
Query: 1475 LTLFETTLCEIISWETLEARTSLDPKGYLSTYNEKDIQLICCQSDASTLWLVPPIVQARF 1534
++FE E+ S T +++ Y+ D+ + ++ +LW + P + R
Sbjct: 2425 DSIFEIN-PEMYSQMTNAYIKEKQALTFIAGYDATDVAAVRLAGNSPSLWNIAPPDRQRL 2483
Query: 1535 MKSLSWNMDL--AFSWEFTRDRPRG--KEVVKYELTIQEQD-----------LPKSSEVT 1579
+ L N L FS+ TR P KE V E I + L ++ +V
Sbjct: 2484 LNDLRNNHTLKARFSYSLTRKAPAKGLKENVGDEHAISLDESFEGRAALIHMLSETHDVE 2543
Query: 1580 EVF-NGTSNSFSAF--------NIYPRYFRVTGSGD--VRSLEQSVELVSGDLVL----- 1623
++ NGT+N + + P++ +V SGD V S+ LV+
Sbjct: 2544 PIYSNGTTNGTTPEVEEVVVIPGMIPKFIKVLNSGDAAVVSVLSPKHYDYRPLVIKMHRD 2603
Query: 1624 NRGNPEWW--------SFYDLDISDAHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIWG 1675
N N WW +FY+ +S G + +++ L+ I G
Sbjct: 2604 NETNGLWWEIRDYCNDTFYNETLSKFAYSNCTSGIVMYTFNDKKFPSTF-SFLTAGGIIG 2662
Query: 1676 LYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLYWT 1735
LY TFVL RF++ +I FE+LP DR++ LC DIY R E +EE L+
Sbjct: 2663 LYTTFVLLASRFMKSFIGGQNRKIMFEDLPYVDRVLQLCLDIYLVREALEFALEEDLFAK 2722
Query: 1736 LVKIYRTPHMLLEYTQ 1751
L+ +YR+P L+++T+
Sbjct: 2723 LLFLYRSPETLIKWTR 2738
Score = 45.8 bits (107), Expect = 0.013
Identities = 30/117 (25%), Positives = 54/117 (45%), Gaps = 9/117 (7%)
Query: 893 WSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLILIYTEV 952
W + +N +++CY + + N SL+S+ +FL+ P+ FWV ++ YT+
Sbjct: 1941 WYALLANTDLICYIVVFINQVVNASLISLPLPIMVFLWGTLSLPRPTKTFWVTLIAYTQA 2000
Query: 953 CILLQYLYQI-IIQHTEFEFPMSLLQELGFPAKKITSSFVTSNLPFFLIYIFTLLQI 1008
+L++ ++Q +I + P L PAK F N + IY LL +
Sbjct: 2001 IVLIKCIFQFKLIWSNYHQLPNQPLT----PAK----IFGVENKAHYAIYDLILLLV 2049
Score = 42.0 bits (97), Expect = 0.19
Identities = 25/93 (26%), Positives = 49/93 (51%), Gaps = 1/93 (1%)
Query: 308 RLLFLRKERLEMQKTALKVSLKFWIENMFTLFGLEINMIALLLASFAVLNAISLLYIASL 367
+LLF R + +K + + +K+ + F FG+EI++IAL+ + ++++Y L
Sbjct: 1019 KLLFPNIIRADAEKDLVGL-VKYLLNFGFYKFGIEISLIALVSTITYRQDIVAVVYALWL 1077
Query: 368 AACVLLHRLLIKKLWPIFVFLFASVVTVEYLAI 400
+LL R K+W +F FA + +Y+ +
Sbjct: 1078 VVLLLLRRSQCAKIWGVFQAFFAISILTQYIVL 1110
>ref|XP_655549.1| conserved hypothetical protein [Entamoeba histolytica HM-1:IMSS]
gi|56472698|gb|EAL50163.1| conserved hypothetical protein
[Entamoeba histolytica HM-1:IMSS]
Length = 2422
Score = 197 bits (501), Expect = 3e-48
Identities = 155/558 (27%), Positives = 263/558 (46%), Gaps = 28/558 (5%)
Query: 1207 KREVDLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQLEDQFPEDFVLVLMVVFFLI 1266
K+ D Y +F ++ F + IF + F++ + +D P +VL L + F +I
Sbjct: 1880 KQGADFYIPLFISEALCFLFLIIFQGTFTNIQGSFIKFFT-DDYLPITYVLGLFLQFAMI 1938
Query: 1267 VLDRIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISL 1326
+ DRIIYLC AK+ FSL+L+ + + + L Y K +
Sbjct: 1939 LADRIIYLCKSIKAKLFMQYFSLLLYHILIVIVYPSFVETKTKIVKACLTIFYLFKVVYW 1998
Query: 1327 VLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLT 1386
++ +Q G + + ++ L ++ S + M F Y +PF+YE+R +LDW + TS+
Sbjct: 1999 IISGLQIRAG--YNILSSKRILMTNYSFYSQMIFHTYYTLPFVYEIRTILDWCLSKTSMF 2056
Query: 1387 MYDWLKLEDIHASLFLVKCD--VDLNRASHQQGQKQTKMTKFCNGICLFFVLMCVIWAPM 1444
WLK+EDIHA L++ +CD ++ NR +H GQ + + + +G + VL+ ++W P+
Sbjct: 2057 YKQWLKVEDIHAELYMNQCDRVIERNR-NHVYGQPRGIVERITSGFIMVIVLLAILWFPL 2115
Query: 1445 LMYSSGNPTNIANPIKDASARVHIKTLS-GRLTLFETTLCEIIS---WETLEARTSLDPK 1500
L SS P N P K + V + + G L + ++ + I+ W L+ S
Sbjct: 2116 LFMSSAAP-NFTQP-KPTNIEVVLNFVGQGELYKQQQSVFKEITKTEWNELQKDHSS--- 2170
Query: 1501 GYLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSL--SWNMDLAFSWEFTRDRPRGK 1558
LS + + I SDA T WL+ P + +L + L +S +R+
Sbjct: 2171 --LSLLSGEAIYSSQLSSDAMTYWLLTPQKKKELNTNLLADETLTLTYSISMSRESSAAS 2228
Query: 1559 EVVKYELTIQEQDLPKSSEVTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSLE--QSVEL 1616
K+ + + K T + NGTS+ + +I+P+ ++ + L+ +V +
Sbjct: 2229 NSFKFSGSYEMNTEEKKKLSTILNNGTSDLY--LSIFPQILKMPTLKEKELLDAFDNVNI 2286
Query: 1617 VSGDLVL--NRGNPEWWSFYD-LDISDAHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSI 1673
++ + N +WSF +++ + C + G I S + P I LS I
Sbjct: 2287 TLKPILHFDSVTNLYYWSFQQCVNVDELWNCSE--GIKFFISSPKVPNSGILSALSSLGI 2344
Query: 1674 WGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLY 1733
GLY VLA+ I+ I F+NLP+C L+ LC+DI AR +G+L +EE L
Sbjct: 2345 IGLYTVVVLALYSLIKSDYVGQAHTIMFKNLPNCLGLLQLCDDIIIARQDGDLRLEEDLV 2404
Query: 1734 WTLVKIYRTPHMLLEYTQ 1751
L+ IYRTP +L E T+
Sbjct: 2405 NELITIYRTPSLLFEKTK 2422
>gb|EAL33026.1| GA21110-PA [Drosophila pseudoobscura]
Length = 2728
Score = 197 bits (500), Expect = 3e-48
Identities = 154/594 (25%), Positives = 277/594 (45%), Gaps = 59/594 (9%)
Query: 1201 SVIGGGKREVDLYAYIFGADLAVFFLIAIFYESI---MKANSEFLEVYQLEDQFPEDFVL 1257
S++ + D+YA +F D FF++ + + + E ++ Y E++ P F++
Sbjct: 2131 SLVHKSRLATDVYALMFLCDFINFFVLLFGFTAFGTQQTESDEGVQTYLAENKVPIPFLI 2190
Query: 1258 VLMVVFFLIVLDRIIYLCSFATAKVIFYLFSLV---LFTYSVTKYAWDMDPLRRYSGRLA 1314
+L+V F LIV+DR +YL K+IF+ FS++ ++ + V + R ++
Sbjct: 2191 MLLVQFLLIVIDRALYLRKALVNKIIFHFFSVIGIHIWMFFVVPAVTE----RTFNSLAP 2246
Query: 1315 LRAIYFTKAISLVLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRC 1374
Y K ++L + Q G P + + F T S +N++ F++Y IPFLYELR
Sbjct: 2247 PIIFYVIKCFYMLLSSYQIKSGYPKR--ILGNFFTKGFSMVNMIAFKVYMQIPFLYELRT 2304
Query: 1375 VLDWSCTTTSLTMYDWLKLEDIHASLFLVKCDVDLNRASH-----QQGQKQTKMTKFCNG 1429
+LDW C +++T++DWLK+EDI ++++L++C R S + QK+ ++K G
Sbjct: 2305 ILDWVCIDSTMTIFDWLKMEDIFSNIYLIRC----TRQSETDFPAMRAQKKASISKLIMG 2360
Query: 1430 ICLFFVLMCVIWAPMLMYSSGNPTNIANPIKDASARVH------IKTLSGRLTLFE--TT 1481
+ +++ IW P+ +++ GN +N S + I T + ++FE T
Sbjct: 2361 GTIVLLIVICIWGPLCLFALGNAVGTSNVPFQVSLSIRIGPYDPIYTTNNYDSIFEIDPT 2420
Query: 1482 LCEIISWETLEARTSLDPKGYLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSWN 1541
+ ++ ++ + +L +++ Y+ D+ + ++ +LW + P + R + L N
Sbjct: 2421 MYSQMTNAYIKDKQALT---FITGYDATDVAAVKLAGNSPSLWNIAPPDRQRLLNDLRNN 2477
Query: 1542 MDL--AFSWEFTRDRPRGK---EVVKYELTIQEQDLPKSSEVTEVFNGTSNSFSAFNI-- 1594
L FS+ TR P + + DL + T NGTS SF +
Sbjct: 2478 HTLKARFSYTLTRKAPXXXFEGRAALINMLNETHDLEPTGNDTST-NGTS-SFEEVVVLP 2535
Query: 1595 --YPRYFRVTGSGDVRSL-------EQSVELVSGDLVLNRGNPEWW--------SFYDLD 1637
P++ +V SGD + E+ LV N WW +FY+
Sbjct: 2536 AMIPKFIKVLNSGDAAVVTVLSPKNEEYRPLVIKMHRDKETNGLWWEIRDFCNDTFYNTT 2595
Query: 1638 ISDAHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRM 1697
+ + G + +++ L+ I GLY TFVL RF++
Sbjct: 2596 LKEFAYSNCTSGIVMYTFNDKKFPSTF-SFLTAGGIIGLYTTFVLLASRFMKSFIGGQNR 2654
Query: 1698 RIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
+I FE+LP DR++ LC DIY R E +EE L+ L+ +YR+P L+++T+
Sbjct: 2655 KIMFEDLPYVDRVLQLCLDIYLVREALEFALEEDLFAKLLFLYRSPETLIKWTR 2708
Score = 45.1 bits (105), Expect = 0.022
Identities = 18/69 (26%), Positives = 38/69 (54%)
Query: 893 WSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLILIYTEV 952
W + +N +++CY + + N SL+S+ +FL+ P+ FWV ++ YT+
Sbjct: 1925 WYALLANTDLICYIVVFINQVVNASLISLPLPIMVFLWGTLSLPRPTKTFWVTLIAYTQA 1984
Query: 953 CILLQYLYQ 961
+L++ ++Q
Sbjct: 1985 IVLIKCIFQ 1993
Score = 42.0 bits (97), Expect = 0.19
Identities = 25/93 (26%), Positives = 48/93 (50%), Gaps = 1/93 (1%)
Query: 308 RLLFLRKERLEMQKTALKVSLKFWIENMFTLFGLEINMIALLLASFAVLNAISLLYIASL 367
+LLF R + +K + + +K+ + F FG+EI++IAL+ + +++ Y L
Sbjct: 996 KLLFPNIIRADAEKDLVGL-VKYLLNYAFYKFGIEISLIALVSTITYRQDIVAVAYALWL 1054
Query: 368 AACVLLHRLLIKKLWPIFVFLFASVVTVEYLAI 400
+LL R K+W +F FA + +Y+ +
Sbjct: 1055 VVLLLLRRSQCAKIWGVFQAFFAISILTQYIIL 1087
>ref|XP_649449.1| conserved hypothetical protein [Entamoeba histolytica HM-1:IMSS]
gi|56465894|gb|EAL44063.1| conserved hypothetical protein
[Entamoeba histolytica HM-1:IMSS]
Length = 2249
Score = 192 bits (487), Expect = 1e-46
Identities = 148/559 (26%), Positives = 255/559 (45%), Gaps = 35/559 (6%)
Query: 1207 KREVDLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQLEDQFPEDFVLVLMVVFFLI 1266
KR DLY +F +L + IF + + ++ F+E + D P +VL L++ F +I
Sbjct: 1714 KRGKDLYIPMFICELLCLLFLVIFQGTFINSDGSFIEFFT-SDYLPIFYVLGLLLQFIMI 1772
Query: 1267 VLDRIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISL 1326
++DRIIYLC AK+I FSL+L+ + + + + L Y K +
Sbjct: 1773 LIDRIIYLCKSIKAKLIMQYFSLILYHVLIFIIYPSILQTKTRLSSICLTIFYVFKVVYW 1832
Query: 1327 VLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLT 1386
+ +Q G + + ++ L ++ S I+ + F Y ++PF+YE+R +LDW+ TS+
Sbjct: 1833 TISGLQIKSG--YLILSSKRILMANYSYISSLIFSTYYSLPFVYEIRTILDWTFAKTSMF 1890
Query: 1387 MYDWLKLEDIHASLFLVKCDVDLNRA-SHQQGQKQTKMTKFCNGICLFFVLMCVIWAPML 1445
WLK+EDIHA L++ +CD ++ +A +H G+ + M K G + +++ ++W P+L
Sbjct: 1891 YKLWLKVEDIHAELYMNQCDREIEKARNHVYGESRGIMEKLTGGCIMIIIMLSILWFPLL 1950
Query: 1446 MYSSGNPTNIANPIKDASARVHIK-TLSGRLTLFETTLCEIIS-----WETLEARTSLDP 1499
+ SS P N I+ V I +L+G F+ I W+ L L
Sbjct: 1951 LMSSAAP----NFIQPKPTNVEITFSLAGNGKFFDQEDSNFIELTGAEWKVLGQTHDLKK 2006
Query: 1500 KGYLSTYNEKDIQLICCQSDAS-TLWLVPPIVQARFMKSLSWNMDLAFSWEFTRDRPRGK 1558
L + C S S + W + P Q +++ + + R
Sbjct: 2007 DASLWIFG-------CSLSLKSMSYWSLTPDKQEDLNETVCNGKQVILKTKIFMSREDSA 2059
Query: 1559 EVVKYELTIQEQDLPKSSE-VTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSLEQ---SV 1614
+EL + K + ++ NG N+F+ + +P + L ++
Sbjct: 2060 VANTFELNEKTSLSKKELQGFCDLLNGKENNFT-LSFFPEIVSMPTLSQTELLRHDSVNI 2118
Query: 1615 ELVSGDLVLNRGNPEWWSFYDLDISDAHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIW 1674
L+ + N+ +W F ++ S + S P + +LS I
Sbjct: 2119 SLIPKKGIDNKTQLSYWWFESINGSTDVDW--------YVSSPNVPDTSVISSLSSKGII 2170
Query: 1675 GLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLYW 1734
GLY V+ + I+ S L I F+NLP+C L+ LC+DI AR +G+L++EE L
Sbjct: 2171 GLYTVVVITLYSLIKSDYSGLSHTIMFKNLPNCHGLLQLCDDIIMARQDGDLKLEEDLVD 2230
Query: 1735 TLVKIYRTPHMLLEYTQAE 1753
L+ IYRTP +LLE T+ +
Sbjct: 2231 ELIMIYRTPSLLLEKTKID 2249
Score = 39.7 bits (91), Expect = 0.93
Identities = 20/77 (25%), Positives = 39/77 (49%), Gaps = 1/77 (1%)
Query: 887 ILVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVY-LAALFLYALCQNTGPSYIFWVL 945
+L+ IW EV FI+ +L+N ++LS Y + + +C+ PS + W +
Sbjct: 1350 MLLWGIWFYCSEQTEVFIQILFIINHLFNQNILSSFYPIIGFSIIMMCKRPNPSKVIWKI 1409
Query: 946 ILIYTEVCILLQYLYQI 962
IL Y + + ++ L ++
Sbjct: 1410 ILWYCFILVFIRMLLEL 1426
>ref|XP_341709.2| PREDICTED: similar to Protein FAM38A [Rattus norvegicus]
Length = 3507
Score = 189 bits (481), Expect = 6e-46
Identities = 151/578 (26%), Positives = 270/578 (46%), Gaps = 62/578 (10%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F AD+ +I + + K ++ L +DQ P+ F+ +L+V F +V+D
Sbjct: 2952 DVYALMFLADIVDIVVIIFGFWAFGKHSAATDIASSLSDDQVPQAFLFMLLVQFGTMVID 3011
Query: 1270 RIIYLCSFATAKVIFYLFSLV---LFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISL 1326
R +YL K+ F + +V L+ + + + R + + YF K I
Sbjct: 3012 RALYLRKTVLGKLAFQVVLVVAIHLWMFFILPAVTE----RMFRQNAVAQLWYFVKCIYF 3067
Query: 1327 VLQAIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLT 1386
L A Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L+
Sbjct: 3068 ALSAYQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLS 3125
Query: 1387 MYDWLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPML 1445
+ +W+ +EDI+A++F++KC + + Q +GQK+ K+ K+ G + L+ +IW P+L
Sbjct: 3126 LSNWMCVEDIYANIFIIKCSRETEKKYPQPKGQKKKKIVKYGMGGLIILFLIAIIWFPLL 3185
Query: 1446 MYSS-GNPTNIANPIKDASARVHIKTLSGRLTLF--ETTLCEIISWETLEARTSLDP--- 1499
S + + N D + + + T+ + ++ + E DP
Sbjct: 3186 FMSLIRSVVGVVNQPIDVTVTLKLGGYEPLFTMSAQQPSIVPFTPEDYEELSQQFDPYPL 3245
Query: 1500 -KGYLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSWNMDLAF--SWEFTRDRPR 1556
++S Y+ +DI + + LW + P +A+ L DLA S E+T +
Sbjct: 3246 AMQFISQYSPEDIVTAQIEGSSGALWRISPPSRAQMKHEL----DLAKGGSVEYTNE--- 3298
Query: 1557 GKEVVKYELTIQEQDLPKSSEVTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSLEQ---- 1612
K+ L + + + S +++P+Y R + ++Q
Sbjct: 3299 -----KHTLELAPNSTARRQLAQLLEGRPDQSVVIPHLFPKYIRAPNGPEANPVKQLQPD 3353
Query: 1613 --------SVEL--------VSGDLVLNRGNP--EWWSFYDLDISDAHG-CGKFPGPMAI 1653
++L SG+ + + EWW +++ D C P +
Sbjct: 3354 EEEDYLGVRIQLRREQVGTGTSGEQAGTKASDFLEWWV---IELQDCQAECNLLP---MV 3407
Query: 1654 IVSEETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMAL 1713
I S++ +G L+ + I GLY++ VL VG+F+R SD+ I FE LP DR++ L
Sbjct: 3408 IFSDKVSPPSLG-FLAGYGIVGLYVSIVLVVGKFVRGFFSDISHSIMFEELPCVDRILKL 3466
Query: 1714 CEDIYAARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
C+DI+ R ELE+EE LY L+ +YR+P ++++T+
Sbjct: 3467 CQDIFLVRETRELELEEELYAKLIFLYRSPETMIKWTR 3504
Score = 50.1 bits (118), Expect = 7e-04
Identities = 23/74 (31%), Positives = 46/74 (62%)
Query: 888 LVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLIL 947
L+R ++ + +++E++CY IL ++ S S+V +FL+A+ PS FW+ +
Sbjct: 2651 LLRAMYQCVAAHSELLCYFIIILNHMVTASAASLVLPVLVFLWAMLTIPRPSKRFWMTAI 2710
Query: 948 IYTEVCILLQYLYQ 961
++TEV ++ +YL+Q
Sbjct: 2711 VFTEVMVVTKYLFQ 2724
>ref|XP_134537.5| PREDICTED: RIKEN cDNA 2310061F22 [Mus musculus]
Length = 2573
Score = 189 bits (480), Expect = 7e-46
Identities = 147/573 (25%), Positives = 267/573 (45%), Gaps = 52/573 (9%)
Query: 1211 DLYAYIFGADLAVFFLIAIFYESIMKANSEFLEVYQL-EDQFPEDFVLVLMVVFFLIVLD 1269
D+YA +F AD+ +I + + K ++ L +DQ P+ F+ +L+V F +V+D
Sbjct: 2018 DVYALMFLADIVDIIIIIFGFWAFGKHSAATDIASSLSDDQVPQAFLFMLLVQFGTMVID 2077
Query: 1270 RIIYLCSFATAKVIFYLFSLVLFTYSVTKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQ 1329
R +YL K+ F + LV+ + + R +S + YF K I L
Sbjct: 2078 RALYLRKTVLGKLAFQVV-LVVAIHIWMFFILPAVTERMFSQNAVAQLWYFVKCIYFALS 2136
Query: 1330 AIQFHFGMPHKSILYRQFLTSSVSRINVMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYD 1389
A Q G P + + FLT + +N+ F+ +R +PFL ELR V+DW T T+L++ +
Sbjct: 2137 AYQIRCGYPTR--ILGNFLTKKYNHLNLFLFQGFRLVPFLVELRAVMDWVWTDTTLSLSN 2194
Query: 1390 WLKLEDIHASLFLVKCDVDLNRASHQ-QGQKQTKMTKFCNGICLFFVLMCVIWAPMLMYS 1448
W+ +EDI+A++F++KC + + Q +GQK+ K+ K+ G + L+ +IW P+L S
Sbjct: 2195 WMCVEDIYANIFIIKCSRETEKKYPQPKGQKKKKIVKYGMGGLIILFLIAIIWFPLLFMS 2254
Query: 1449 -SGNPTNIANPIKDASARVHIKTLSGRLTLF--ETTLCEIISWETLEARTSLDP----KG 1501
+ + N D + + + T+ + ++ E DP
Sbjct: 2255 LIRSVVGVVNQPIDVTVTLKLGGYEPLFTMSAQQPSIVPFTPQAYEELSQQFDPYPLAMQ 2314
Query: 1502 YLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSWNMDLAFSWEFTRDRPRGKEVV 1561
++S Y+ +DI + + LW + P +A+ + L ++ + E+T +
Sbjct: 2315 FISQYSPEDIVTAQIEGSSGALWRISPPSRAQMKQEL--DLAKGGTVEYTNE-------- 2364
Query: 1562 KYELTIQEQDLPKSSEVTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSLEQ--------- 1612
K+ L + + + S +++P+Y R + ++Q
Sbjct: 2365 KHTLELAPNSTARRQLAQLLEGRPDQSVVIPHLFPKYIRAPNGPEANPVKQLQPDEEEDY 2424
Query: 1613 ---SVEL--------VSGDLVLNRGNP--EWWSFYDLDISDAHG-CGKFPGPMAIIVSEE 1658
++L SG+ + + EWW +++ D C PM I +
Sbjct: 2425 LGVRIQLRREQVGTGASGEQAGTKASDFLEWWV---IELQDCKADCNLL--PMVIFSDKV 2479
Query: 1659 TPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIY 1718
+P + L+ + I GLY++ VL VG+F+R S++ I FE LP DR++ LC+DI+
Sbjct: 2480 SPPSL--GFLAGYGIVGLYVSIVLVVGKFVRGFFSEISHSIMFEELPCVDRILKLCQDIF 2537
Query: 1719 AARAEGELEVEEVLYWTLVKIYRTPHMLLEYTQ 1751
R ELE+EE LY L+ +YR+P ++++T+
Sbjct: 2538 LVRETRELELEEELYAKLIFLYRSPETMIKWTR 2570
Score = 48.1 bits (113), Expect = 0.003
Identities = 23/74 (31%), Positives = 45/74 (60%)
Query: 888 LVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLIL 947
L+R + + +++E++CY IL ++ S S+V +FL+A+ PS FW+ +
Sbjct: 1718 LLRAGYQCVAAHSELLCYFIIILNHMVTASAASLVLPVLVFLWAMLTIPRPSKRFWMTAI 1777
Query: 948 IYTEVCILLQYLYQ 961
++TEV ++ +YL+Q
Sbjct: 1778 VFTEVMVVTKYLFQ 1791
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,864,980,908
Number of Sequences: 2540612
Number of extensions: 119971986
Number of successful extensions: 393101
Number of sequences better than 10.0: 94
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 45
Number of HSP's that attempted gapping in prelim test: 392641
Number of HSP's gapped (non-prelim): 357
length of query: 1753
length of database: 863,360,394
effective HSP length: 142
effective length of query: 1611
effective length of database: 502,593,490
effective search space: 809678112390
effective search space used: 809678112390
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 83 (36.6 bits)
Lotus: description of TM0544.8


