

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.11
(269 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
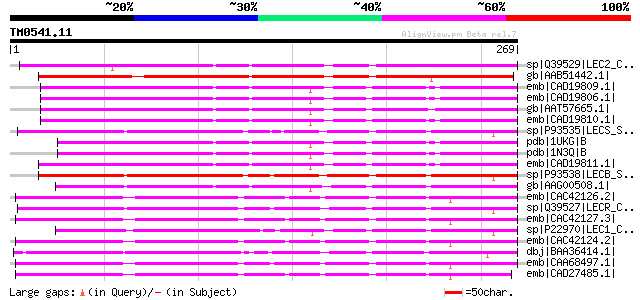
Score E
Sequences producing significant alignments: (bits) Value
sp|Q39529|LEC2_CLALU Agglutinin II precursor (ClAII) (LecClAII) ... 201 2e-50
gb|AAB51442.1| lectin precursor 200 3e-50
emb|CAD19809.1| lectin [Pterocarpus angolensis] gi|27368671|emb|... 183 5e-45
emb|CAD19806.1| lectin [Pterocarpus angolensis] 182 9e-45
gb|AAT57665.1| lectin [Pterocarpus rotundifolius] 182 1e-44
emb|CAD19810.1| lectin [Pterocarpus angolensis] gi|27368665|emb|... 181 2e-44
sp|P93535|LECS_SOPJA Seed lectin precursor (LECSJASG) gi|1755064... 179 7e-44
pdb|1UKG|B Chain B, Pterocarps Angolensis Lectin Pal In Complex ... 179 7e-44
pdb|1N3Q|B Chain B, Pterocarpus Angolensis Lectin Complexed With... 179 7e-44
emb|CAD19811.1| lectin [Pterocarpus angolensis] gi|27368661|emb|... 178 1e-43
sp|P93538|LECB_SOPJA Bark lectin precursor (LECSJABG) gi|1755070... 177 3e-43
gb|AAG00508.1| lectin [Sophora flavescens] 177 3e-43
emb|CAC42126.2| lectin [Lens ervoides] 170 4e-41
sp|Q39527|LECR_CLALU Lectin-related protein precursor (CLLRP) (L... 169 6e-41
emb|CAC42127.3| lectin [Lens nigricans] 167 3e-40
sp|P22970|LEC1_CYTSE Anti-H(O) lectin I (CSA-I) gi|228857|prf||1... 167 3e-40
emb|CAC42124.2| lectin [Lens culinaris] gi|26800840|emb|CAC42123... 166 6e-40
dbj|BAA36414.1| lectin [Robinia pseudoacacia] 166 6e-40
emb|CAA68497.1| lectin-precursor (AA -30 to 245) [Pisum sativum]... 166 8e-40
emb|CAD27485.1| lectin [Lathyrus sativus] 166 8e-40
>sp|Q39529|LEC2_CLALU Agglutinin II precursor (ClAII) (LecClAII)
gi|1141759|gb|AAC49137.1| lectin precursor
Length = 290
Score = 201 bits (511), Expect = 2e-50
Identities = 116/268 (43%), Positives = 158/268 (58%), Gaps = 13/268 (4%)
Query: 6 SISLLTTLLPFILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTGG--VLQLTKKDL 62
S+ LL + F++L+ V S DS SF F F D + ++ GDA + G LQLTK D
Sbjct: 16 SLPLLAFITLFLMLLNRVNSSDSLSFTFDNFRPDQRDLILQGDAKISSGGDSLQLTKTDT 75
Query: 63 YGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDF 122
G P + S G+ ++ +HL D+ T ++A F T F+FV++ + P GDG F+IA +
Sbjct: 76 SGKPVRGSVGRALYYTPLHLWDSSTNRLASFQTTFTFVLSSPTNNP-GDGIAFFIAPPET 134
Query: 123 DFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDINSIR 181
P GG LGLF P NA N S NQ+VAVEFD+F N WDPS+ H+GID+N+I+
Sbjct: 135 TIPPG-SSGGLLGLFSPDNALNNSLNQIVAVEFDTFVNNNWDPSHR----HIGIDVNTIK 189
Query: 182 SVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSD 241
S AT W + G + A ISY S +K LSV YPN+ + V+Y+VDL L +
Sbjct: 190 SSATVRWQRE---NGSLATAQISYNSDTKKLSVVSSYPNTQANEDYTVSYDVDLKTELPE 246
Query: 242 WVLVGFTAATGSVSETHDILSWAFNSFL 269
WV VGF+ +TG + H+ILSW FNS L
Sbjct: 247 WVRVGFSGSTGGYVQNHNILSWTFNSNL 274
>gb|AAB51442.1| lectin precursor
Length = 266
Score = 200 bits (509), Expect = 3e-50
Identities = 117/256 (45%), Positives = 156/256 (60%), Gaps = 18/256 (7%)
Query: 16 FILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQT 74
F+ L+ V S DS SF F F D + ++ GDA+ G LQLTK D G+ G+
Sbjct: 7 FLTLLNMVNSSDSLSFTFNNFGPDQRDLILQGDAHIPSGTLQLTKTDSSGV------GRA 60
Query: 75 TFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFL 134
++ VHL D+R G++A F T FSFV+ +G++ GDG F+IA + P GGFL
Sbjct: 61 LYYLPVHLWDSRRGRLASFETSFSFVITSQGTDDPGDGIAFFIAPPETTIPPR-SSGGFL 119
Query: 135 GLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDINSIRSVATAPWPIDLV 193
GLF P+ A N+S N VVAVEFD+F N WDPSY H+GID+NSI+S A A W
Sbjct: 120 GLFSPETALNSSLNPVVAVEFDTFINEDWDPSYW----HIGIDVNSIKSSAAARWERK-- 173
Query: 194 PQGEIGKALISYRSASKVLSVFVVYPNSP--VKIETFVAYEVDLGAVLSDWVLVGFTAAT 251
G A ISY S+SK LSV YPN+ V+++ V+Y++DL VL +WV +GF+A+T
Sbjct: 174 -SGRKFTAHISYNSSSKKLSVVSSYPNTNCLVRVDYTVSYDIDLTTVLPEWVRIGFSAST 232
Query: 252 GSVSETHDILSWAFNS 267
G E H ILSW+F+S
Sbjct: 233 GYKIEEHSILSWSFSS 248
>emb|CAD19809.1| lectin [Pterocarpus angolensis] gi|27368671|emb|CAD19808.1| lectin
[Pterocarpus angolensis] gi|27368669|emb|CAD19807.1|
lectin [Pterocarpus angolensis]
gi|27368663|emb|CAD19804.1| lectin [Pterocarpus
angolensis]
Length = 260
Score = 183 bits (464), Expect = 5e-45
Identities = 109/257 (42%), Positives = 148/257 (57%), Gaps = 16/257 (6%)
Query: 17 ILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTF 76
+LL DS SF FP F SD K ++ GDA T +QLTK D G P ++ G+ F
Sbjct: 1 MLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQTKNNAVQLTKTDSNGNPVASTVGRILF 60
Query: 77 FGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGL 136
VHL + + +VA+F ++FSF + S DG F+IA D P GG LGL
Sbjct: 61 SAQVHLWEKSSSRVANFQSQFSFSLKSPLSNG-ADGIAFFIAPPDTTIPSG-SGGGLLGL 118
Query: 137 FDPKNAFNTSANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDL 192
F P A NTSANQV+AVEFD+F +N WDP+YP H+GID+NSIRSV T W
Sbjct: 119 FAPGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYP----HIGIDVNSIRSVKTVKWD--- 171
Query: 193 VPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATG 252
G+ L+++ +++ L V Y + + E V+YEVD+ +VL +WV VGF+AA+G
Sbjct: 172 RRDGQSLNVLVTFNPSTRNLDVVATYSDG-TRYE--VSYEVDVRSVLPEWVRVGFSAASG 228
Query: 253 SVSETHDILSWAFNSFL 269
+TH + SW+F S L
Sbjct: 229 EQYQTHTLESWSFTSTL 245
>emb|CAD19806.1| lectin [Pterocarpus angolensis]
Length = 260
Score = 182 bits (462), Expect = 9e-45
Identities = 109/257 (42%), Positives = 148/257 (57%), Gaps = 16/257 (6%)
Query: 17 ILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTF 76
+LL DS SF FP F SD K ++ GDA T +QLTK D G P ++ G+ F
Sbjct: 1 MLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQTKNNAVQLTKTDSNGNPVASTVGRILF 60
Query: 77 FGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGL 136
VHL + + +VA+F ++FSF + S DG F+IA D P GG LGL
Sbjct: 61 SAQVHLWEKSSSRVANFQSQFSFSLKSPLSNG-ADGIAFFIAPPDTTIPSG-SGGGLLGL 118
Query: 137 FDPKNAFNTSANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDL 192
F P A NTSANQV+AVEFD+F +N WDP+YP H+GID+NSIRSV T W
Sbjct: 119 FAPGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYP----HIGIDVNSIRSVKTVKWD--- 171
Query: 193 VPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATG 252
G+ L+++ +++ L V Y + + E V+YEVD+ +VL +WV VGF+AA+G
Sbjct: 172 RRDGQSLNVLVTFNPSTRNLDVVATYSDG-TRYE--VSYEVDVRSVLPEWVGVGFSAASG 228
Query: 253 SVSETHDILSWAFNSFL 269
+TH + SW+F S L
Sbjct: 229 EQYQTHTLESWSFTSTL 245
>gb|AAT57665.1| lectin [Pterocarpus rotundifolius]
Length = 249
Score = 182 bits (461), Expect = 1e-44
Identities = 107/257 (41%), Positives = 146/257 (56%), Gaps = 16/257 (6%)
Query: 17 ILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTF 76
+LL SDSF F F F+ D + ++ GDA VLQLTK D G P +++ G+ +
Sbjct: 1 MLLNKAYSSDSFPFGFFNFDQDERNLIYQGDARAQNNVLQLTKTDSNGNPVRSTVGRILY 60
Query: 77 FGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGL 136
V L + T +VA+F ++FS ++ S P DG F+IA D P GG LGL
Sbjct: 61 TAQVRLWEKSTNRVANFQSQFSLHLSSSLSNP-ADGIAFFIAPPDTTIPSG-SGGGLLGL 118
Query: 137 FDPKNAFNTSANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDL 192
F P A NTSANQV+AVEFD+F +N WDP+Y H+GID+NSIRS T W
Sbjct: 119 FAPGTAQNTSANQVLAVEFDTFYAQDSNTWDPNYQ----HIGIDVNSIRSARTVRWE--- 171
Query: 193 VPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATG 252
GE L++Y +++ L V YP+ V+YEVD+ +VL +WV VGF+AA+G
Sbjct: 172 RRDGETLNVLVTYNPSTRTLDVVATYPDGQ---RYEVSYEVDVRSVLPEWVRVGFSAASG 228
Query: 253 SVSETHDILSWAFNSFL 269
+TH + SW+F S L
Sbjct: 229 EQYQTHSLESWSFTSTL 245
>emb|CAD19810.1| lectin [Pterocarpus angolensis] gi|27368665|emb|CAD19805.1| lectin
[Pterocarpus angolensis]
Length = 260
Score = 181 bits (458), Expect = 2e-44
Identities = 108/257 (42%), Positives = 147/257 (57%), Gaps = 16/257 (6%)
Query: 17 ILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTF 76
+LL DS SF FP F SD K ++ GDA +QLTK D G P ++ G+ F
Sbjct: 1 MLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQIKNNAVQLTKTDSNGNPVASTVGRILF 60
Query: 77 FGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGL 136
VHL + + +VA+F ++FSF + S DG F+IA D P GG LGL
Sbjct: 61 SAQVHLWEKSSSRVANFQSQFSFSLKSPLSNG-ADGIAFFIAPPDTTIPSG-SGGGLLGL 118
Query: 137 FDPKNAFNTSANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDL 192
F P A NTSANQV+AVEFD+F +N WDP+YP H+GID+NSIRSV T W
Sbjct: 119 FAPGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYP----HIGIDVNSIRSVKTVKWD--- 171
Query: 193 VPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATG 252
G+ L+++ +++ L V Y + + E V+YEVD+ +VL +WV VGF+AA+G
Sbjct: 172 RRDGQSLNVLVTFNPSTRNLDVVATYSDG-TRYE--VSYEVDVRSVLPEWVRVGFSAASG 228
Query: 253 SVSETHDILSWAFNSFL 269
+TH + SW+F S L
Sbjct: 229 EQYQTHTLESWSFTSTL 245
>sp|P93535|LECS_SOPJA Seed lectin precursor (LECSJASG) gi|1755064|gb|AAB51441.1| lectin
precursor
Length = 292
Score = 179 bits (454), Expect = 7e-44
Identities = 110/270 (40%), Positives = 160/270 (58%), Gaps = 19/270 (7%)
Query: 5 FSISLLTTLLPFILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDL 62
FS+ L ++ F+LL+ V S + SF+FP+F S+ + +L GDA + G LQLT +
Sbjct: 17 FSVVLAISITFFLLLLNKVNSAEILSFSFPKFASNQEDLLLQGDALVSSKGELQLTTVE- 75
Query: 63 YGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDF 122
G+P NS G+ ++ VH+ D TG+VA F T FSFVV + DG F++A +
Sbjct: 76 NGVPIWNSTGRALYYAPVHIWDKSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPPNN 135
Query: 123 DFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRS 182
GG LGLF + +N S+ Q++AV+FD+ N WDP N+ H+GID+NSI S
Sbjct: 136 QI--QGPGGGHLGLFH-SSGYN-SSYQIIAVDFDTHINAWDP----NTRHIGIDVNSINS 187
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDW 242
T W GE+ LISY++A++ L+V + YP+S + ++ VDL ++L +W
Sbjct: 188 TKTVTWGWQ---NGEVANVLISYQAATETLTVSLTYPSS--QTSYILSAAVDLKSILPEW 242
Query: 243 VLVGFTAATGSVS---ETHDILSWAFNSFL 269
V VGFTAATG + ETHD+LSW+F S L
Sbjct: 243 VRVGFTAATGLTTQYVETHDVLSWSFTSTL 272
>pdb|1UKG|B Chain B, Pterocarps Angolensis Lectin Pal In Complex With Methyl-
Alpha-Mannose gi|46015823|pdb|1UKG|A Chain A, Pterocarps
Angolensis Lectin Pal In Complex With Methyl-
Alpha-Mannose gi|46015356|pdb|1Q8V|B Chain B,
Pterocarpus Angolensis Lectin (Pal) In Complex With The
Trimannoside [man(Alpha1-3)]man(Alpha1-6)man
gi|46015355|pdb|1Q8V|A Chain A, Pterocarpus Angolensis
Lectin (Pal) In Complex With The Trimannoside
[man(Alpha1-3)]man(Alpha1-6)man gi|46015354|pdb|1Q8S|B
Chain B, Pterocarpus Angolensis Lectin (Pal) In Complex
With The Dimannoside Man(Alpha1-6)man
gi|46015353|pdb|1Q8S|A Chain A, Pterocarpus Angolensis
Lectin (Pal) In Complex With The Dimannoside
Man(Alpha1-6)man gi|46015352|pdb|1Q8Q|B Chain B,
Pterocarpus Angolensis Lectin (Pal) In Complex With The
Dimannoside Man(Alpha1-4)man gi|46015351|pdb|1Q8Q|A
Chain A, Pterocarpus Angolensis Lectin (Pal) In Complex
With The Dimannoside Man(Alpha1-4)man
gi|46015350|pdb|1Q8P|B Chain B, Pterocarpus Angolensis
Lectin Pal In Complex With The Dimannoside
Man(Alpha1-3)man gi|46015349|pdb|1Q8P|A Chain A,
Pterocarpus Angolensis Lectin Pal In Complex With The
Dimannoside Man(Alpha1-3)man gi|46015348|pdb|1Q8O|B
Chain B, Pterocartpus Angolensis Lectin Pal In Complex
With The Dimmanoside Man(Alpha1-2)man
gi|46015347|pdb|1Q8O|A Chain A, Pterocartpus Angolensis
Lectin Pal In Complex With The Dimmanoside
Man(Alpha1-2)man
Length = 252
Score = 179 bits (454), Expect = 7e-44
Identities = 106/248 (42%), Positives = 144/248 (57%), Gaps = 16/248 (6%)
Query: 26 DSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLSDN 85
DS SF FP F SD K ++ GDA +QLTK D G P ++ G+ F VHL +
Sbjct: 2 DSLSFGFPTFPSDQKNLIFQGDAQIKNNAVQLTKTDSNGNPVASTVGRILFSAQVHLWEK 61
Query: 86 RTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGLFDPKNAFNT 145
+ +VA+F ++FSF + S DG F+IA D P GG LGLF P A NT
Sbjct: 62 SSSRVANFQSQFSFSLKSPLSNG-ADGIAFFIAPPDTTIPSG-SGGGLLGLFAPGTAQNT 119
Query: 146 SANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKA 201
SANQV+AVEFD+F +N WDP+YP H+GID+NSIRSV T W G+
Sbjct: 120 SANQVIAVEFDTFYAQDSNTWDPNYP----HIGIDVNSIRSVKTVKWD---RRDGQSLNV 172
Query: 202 LISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDIL 261
L+++ +++ L V Y + + E V+YEVD+ +VL +WV VGF+AA+G +TH +
Sbjct: 173 LVTFNPSTRNLDVVATYSDG-TRYE--VSYEVDVRSVLPEWVRVGFSAASGEQYQTHTLE 229
Query: 262 SWAFNSFL 269
SW+F S L
Sbjct: 230 SWSFTSTL 237
>pdb|1N3Q|B Chain B, Pterocarpus Angolensis Lectin Complexed With Turanose
gi|27065992|pdb|1N3Q|A Chain A, Pterocarpus Angolensis
Lectin Complexed With Turanose gi|27065990|pdb|1N3P|B
Chain B, Pterocarpus Angolensis Lectin In Complex With
Sucrose gi|27065989|pdb|1N3P|A Chain A, Pterocarpus
Angolensis Lectin In Complex With Sucrose
gi|27065986|pdb|1N3O|B Chain B, Pterocarcpus Angolensis
Lectin In Complex With Alpha-Methyl Glucose
gi|27065985|pdb|1N3O|A Chain A, Pterocarcpus Angolensis
Lectin In Complex With Alpha-Methyl Glucose
gi|60593453|pdb|1S1A|B Chain B, Pterocarpus Angolensis
Seed Lectin (Pal) With One Binding Site Free And One
Binding Site Containing The Disaccharide Man(A1-3)manme
gi|60593452|pdb|1S1A|A Chain A, Pterocarpus Angolensis
Seed Lectin (Pal) With One Binding Site Free And One
Binding Site Containing The Disaccharide Man(A1-3)manme
Length = 252
Score = 179 bits (454), Expect = 7e-44
Identities = 106/248 (42%), Positives = 144/248 (57%), Gaps = 16/248 (6%)
Query: 26 DSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLSDN 85
DS SF FP F SD K ++ GDA +QLTK D G P ++ G+ F VHL +
Sbjct: 2 DSLSFGFPTFPSDQKNLIFQGDAQIKNNAVQLTKTDSNGNPVASTVGRILFSAQVHLWEK 61
Query: 86 RTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGLFDPKNAFNT 145
+ +VA+F ++FSF + S DG F+IA D P GG LGLF P A NT
Sbjct: 62 SSSRVANFQSQFSFSLKSPLSNG-ADGIAFFIAPPDTTIPSG-SGGGLLGLFAPGTAQNT 119
Query: 146 SANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKA 201
SANQV+AVEFD+F +N WDP+YP H+GID+NSIRSV T W G+
Sbjct: 120 SANQVIAVEFDTFYAQDSNTWDPNYP----HIGIDVNSIRSVKTVKWD---RRDGQSLNV 172
Query: 202 LISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDIL 261
L+++ +++ L V Y + + E V+YEVD+ +VL +WV VGF+AA+G +TH +
Sbjct: 173 LVTFNPSTRNLDVVATYSDG-TRYE--VSYEVDVRSVLPEWVRVGFSAASGEQYQTHTLE 229
Query: 262 SWAFNSFL 269
SW+F S L
Sbjct: 230 SWSFTSTL 237
>emb|CAD19811.1| lectin [Pterocarpus angolensis] gi|27368661|emb|CAD19803.1| lectin
[Pterocarpus angolensis]
Length = 272
Score = 178 bits (452), Expect = 1e-43
Identities = 108/259 (41%), Positives = 149/259 (56%), Gaps = 17/259 (6%)
Query: 16 FILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQT 74
F++L+ S DS SF FP F SD K ++ GDA +QLTK D G P ++ G+
Sbjct: 11 FLMLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQIKNNAVQLTKTDSNGNPVASTVGRI 70
Query: 75 TFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFL 134
F VHL + + +VA+F ++FSF + S DG F+IA D P GG
Sbjct: 71 LFSAQVHLWEKSSSRVANFQSQFSFSLKSPLSNG-ADGIAFFIAPPDTTIPSG-SGGGLP 128
Query: 135 GLFDPKNAFNTSANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPI 190
GLF P A NTSANQV+AVEFD+F +N WDP+YP H+GID+NSIRSV T W
Sbjct: 129 GLFAPGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYP----HIGIDVNSIRSVKTVKWD- 183
Query: 191 DLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAA 250
G+ L+++ +++ L V Y + + E V+YEVD+ +VL +WV VGF+AA
Sbjct: 184 --RRDGQSLNVLVTFNPSTRNLDVVATYSDG-TRYE--VSYEVDVRSVLPEWVRVGFSAA 238
Query: 251 TGSVSETHDILSWAFNSFL 269
+G +TH + SW+F S L
Sbjct: 239 SGEQYQTHTLESWSFTSTL 257
>sp|P93538|LECB_SOPJA Bark lectin precursor (LECSJABG) gi|1755070|gb|AAB51458.1| lectin
precursor
Length = 270
Score = 177 bits (449), Expect = 3e-43
Identities = 105/259 (40%), Positives = 159/259 (60%), Gaps = 19/259 (7%)
Query: 16 FILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDLYGIPSKNSFGQ 73
F+LL+ V S + SF+FP+F S+ + +L GDA + G LQLT + G+P NS G+
Sbjct: 6 FLLLLNKVNSAEILSFSFPKFVSNQEDLLLQGDALVSSEGELQLTTVE-NGVPVWNSTGR 64
Query: 74 TTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGF 133
++ VH+ DN TG+VA F T FSFVV + DG F++A L+ GG
Sbjct: 65 ALYYAPVHIWDNSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPLNNQI--HGAGGGL 122
Query: 134 LGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLV 193
GLF+ ++ +S+ Q+VAVEFD+ TN WDP N+ H+GID+NS++S T W +
Sbjct: 123 YGLFN--SSSYSSSYQIVAVEFDTHTNAWDP----NTRHIGIDVNSVKSTKTVTWGWE-- 174
Query: 194 PQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGS 253
GE+ LI+Y++A+++L+V + YP++ + ++ VDL ++L +WV VGFTA TG
Sbjct: 175 -NGEVANVLITYQAATEMLTVSLTYPSN--QTSYILSAAVDLKSILPEWVRVGFTATTGL 231
Query: 254 VS---ETHDILSWAFNSFL 269
+ ET+D+LSW+F S L
Sbjct: 232 TTQYVETNDVLSWSFTSTL 250
>gb|AAG00508.1| lectin [Sophora flavescens]
Length = 284
Score = 177 bits (449), Expect = 3e-43
Identities = 107/250 (42%), Positives = 149/250 (58%), Gaps = 17/250 (6%)
Query: 25 SDSFSFNFPRFESDTKTILSDGDAN-TTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLS 83
+DS SF F F+ + + +L GDA+ T+ +LQLTK G+P +N+ G+ F +HL
Sbjct: 31 ADSLSFTFSDFDPNGEDLLFQGDAHVTSNNILQLTKTS-NGVPLQNTVGRALFSTPIHLW 89
Query: 84 DNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGLFDPKNAF 143
+ T +++ F + F+FV+ S P DGF F+IA D PE +GG LGLF P+NA
Sbjct: 90 EKSTNRLSSFESTFTFVLTSPQSNP-ADGFAFFIAPPDTTIPEG-SDGGLLGLFSPENAL 147
Query: 144 NTSANQVVAVEFDSF----TNVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIG 199
N ANQVVAVEFD+F +N WDP+Y H+GID+N I+S AT W +G IG
Sbjct: 148 NPKANQVVAVEFDTFYDKSSNSWDPNY----VHIGIDVNQIKSSATVRWD---RKEGVIG 200
Query: 200 KALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHD 259
A I+Y +A++ LSV YP + V+Y VDL L ++V VGF+A+TG + H
Sbjct: 201 TARINYNAATRNLSVVSSYPGG--SQDYVVSYVVDLRTKLPEFVRVGFSASTGQQYQVHS 258
Query: 260 ILSWAFNSFL 269
I SW F+S L
Sbjct: 259 IRSWFFSSSL 268
>emb|CAC42126.2| lectin [Lens ervoides]
Length = 275
Score = 170 bits (430), Expect = 4e-41
Identities = 104/271 (38%), Positives = 155/271 (56%), Gaps = 22/271 (8%)
Query: 4 SFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLY 63
SF + L+ LL I +++ SF+ +F D + ++ GD TT L LTK
Sbjct: 10 SFYLIFLSILLTTIFFFKVNSTETTSFSITKFSPDQQNLIFQGDGYTTKEKLTLTKA--- 66
Query: 64 GIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
KN+ G+ + +H+ D TG VA+F T F+FV+N S DGFTF+IA +D
Sbjct: 67 ---VKNTVGRALYSTPIHIWDRDTGNVANFVTSFTFVINAPNSYNVADGFTFFIAPVD-- 121
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRS 182
+ GG+LG+F+ K+ TS Q VAVEFD+F N WDPS + H+GID+NSI+S
Sbjct: 122 -TKPQTGGGYLGVFNSKDYDKTS--QTVAVEFDTFYNAAWDPS--NKDRHIGIDVNSIKS 176
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYE----VDLGAV 238
V+T W + GE +I++ +A+ VL+V + YPNS ++ E +Y V + V
Sbjct: 177 VSTKSWNLQ---NGERANVVIAFNAATNVLTVTLTYPNS-LEEENVTSYTLNEVVPMKDV 232
Query: 239 LSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
L +WV +GF+A TG+ H++LSW+F+S L
Sbjct: 233 LPEWVRIGFSATTGAEFAAHEVLSWSFHSEL 263
>sp|Q39527|LECR_CLALU Lectin-related protein precursor (CLLRP) (LRPCL)
gi|1141755|gb|AAC49150.1| storage protein precursor
Length = 290
Score = 169 bits (429), Expect = 6e-41
Identities = 106/270 (39%), Positives = 158/270 (58%), Gaps = 20/270 (7%)
Query: 5 FSISLLTTLLPFILLITTVKSD-SFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKKDL 62
FS+ L ++ ++LL+ V S+ + SF F +F S+ +L GDA + G LQLT+ +
Sbjct: 16 FSVVLAISITFYLLLLNKVNSEEALSFTFTKFVSNQDELLLQGDALVSSKGELQLTRVE- 74
Query: 63 YGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDF 122
G P +S G+ + VH+ D+ TG VA F T F+FVV DG F++A D
Sbjct: 75 NGQPIPHSVGRALYSDPVHIWDSSTGSVASFVTSFTFVVEAPNENKTADGIAFFLAPPD- 133
Query: 123 DFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRS 182
+ GGFLGLF+ ++ S+NQ++AVEFD+F+N WDP+ + H+GID+NSI S
Sbjct: 134 --TQVQSLGGFLGLFN--SSVYNSSNQILAVEFDTFSNSWDPT----ARHIGIDVNSIES 185
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDW 242
TA W GE+ LI+Y + ++ L + YP+S + ++ VDL ++L +W
Sbjct: 186 TRTATWG---WRNGEVAIVLITYVAPAETLIASLTYPSS--QTSYILSAAVDLKSILPEW 240
Query: 243 VLVGFTAATGSVS---ETHDILSWAFNSFL 269
V VGF+AATG + ETHD+LSW+F S L
Sbjct: 241 VRVGFSAATGRSAGYVETHDVLSWSFTSTL 270
>emb|CAC42127.3| lectin [Lens nigricans]
Length = 275
Score = 167 bits (423), Expect = 3e-40
Identities = 103/271 (38%), Positives = 155/271 (57%), Gaps = 22/271 (8%)
Query: 4 SFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLY 63
SF + L+ LL I +++ SF+ +F D + I+ GD TT L LTK
Sbjct: 10 SFYLIFLSILLTTIFFFKVNSTETTSFSITKFSPDQQNIIFQGDGYTTKEKLTLTKA--- 66
Query: 64 GIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
K++ G+ + +H+ D TG VA+F T F+FV+N S DGFTF+IA +D
Sbjct: 67 ---VKSTVGRALYSTPIHIWDRYTGNVANFVTSFTFVINAPNSYNVADGFTFFIAPVD-- 121
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRS 182
+ GG+LG+F+ K+ TS Q VAVEFD+F N WDPS + H+GID+NSI+S
Sbjct: 122 -SKPQTGGGYLGVFNSKDYDKTS--QTVAVEFDTFYNAAWDPS--NKDRHIGIDVNSIKS 176
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYE----VDLGAV 238
++T W + GE +I++ +A+ VL+V + YPNS ++ E +Y V L V
Sbjct: 177 LSTKSWNLQ---NGEQANVVIAFNAATNVLTVTLTYPNS-LEEENVTSYTLNEVVPLKDV 232
Query: 239 LSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
+ +WV +GF+A TG+ H++LSW+F+S L
Sbjct: 233 VPEWVRIGFSATTGAEFAAHEVLSWSFHSEL 263
>sp|P22970|LEC1_CYTSE Anti-H(O) lectin I (CSA-I) gi|228857|prf||1813204A anti-H(O) lectin
Length = 244
Score = 167 bits (423), Expect = 3e-40
Identities = 101/253 (39%), Positives = 148/253 (57%), Gaps = 23/253 (9%)
Query: 25 SDSFSFNFPRFESDTKTILSDGDAN-TTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHLS 83
+D SFNF +F + IL G+A+ +T GVLQ+TK P+ S G+ + VH+
Sbjct: 2 NDHLSFNFDKFVPNQNNILFQGEASVSTTGVLQVTKVSK---PATRSIGRALYAAPVHIW 58
Query: 84 DNRTGKVADFTTEFSFVVNPKGSEPHG-DGFTFYIAGLDFDFPEDPKEGGFLGLFDPKNA 142
D+ TG+VA F T FSFVV + + +G DG TF++A + P G F GLF+ +
Sbjct: 59 DSTTGRVASFETSFSFVVKDEPEKSNGVDGLTFFLAPANSQIPSGSSAGLF-GLFNSSD- 116
Query: 143 FNTSANQVVAVEFDSFT----NVWDPSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEI 198
N S+NQ++AVEFD++ N WDP + H+G+D+NSI+S+ T W GE+
Sbjct: 117 -NKSSNQIIAVEFDTYFGKTYNPWDPDFK----HIGVDVNSIKSIKTVKWD---WRNGEV 168
Query: 199 GKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVS--E 256
+I+YR+ +K L+V + YP+ + V VDL A+L +WV VGF+A G+ + E
Sbjct: 169 ANVVITYRAPTKSLTVSLSYPSD--QTSNIVTASVDLKAILPEWVSVGFSAGVGNAAEFE 226
Query: 257 THDILSWAFNSFL 269
THD+LSW F S L
Sbjct: 227 THDVLSWYFTSNL 239
>emb|CAC42124.2| lectin [Lens culinaris] gi|26800840|emb|CAC42123.2| lectin [Lens
culinaris]
Length = 275
Score = 166 bits (420), Expect = 6e-40
Identities = 103/271 (38%), Positives = 153/271 (56%), Gaps = 22/271 (8%)
Query: 4 SFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLY 63
SF + L+ LL I +++ SF+ +F D K ++ GD TT G L LTK
Sbjct: 10 SFYLIFLSILLTTIFFFKVNSTETTSFSITKFSPDQKNLIFQGDGYTTKGKLTLTKA--- 66
Query: 64 GIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
K++ G+ + +H+ D TG VA+F T F+FV++ S DGFTF+IA +D
Sbjct: 67 ---VKSTVGRALYSTPIHIWDRDTGNVANFVTSFTFVIDAPSSYNVADGFTFFIAPVD-- 121
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRS 182
+ GG+LG+F+ K TS Q VAVEFD+F N WDPS + H+GID+NSI+S
Sbjct: 122 -TKPQTGGGYLGVFNSKEYDKTS--QTVAVEFDTFYNAAWDPS--NKERHIGIDVNSIKS 176
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYE----VDLGAV 238
V T W + GE +I++ +A+ VL+V + YPNS ++ E +Y V L V
Sbjct: 177 VNTKSWNLQ---NGERANVVIAFNAATNVLTVTLTYPNS-LEEENVTSYTLNEVVPLKDV 232
Query: 239 LSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
+ +WV +GF+A TG+ H++ SW+F+S L
Sbjct: 233 VPEWVRIGFSATTGAEFAAHEVHSWSFHSEL 263
>dbj|BAA36414.1| lectin [Robinia pseudoacacia]
Length = 285
Score = 166 bits (420), Expect = 6e-40
Identities = 109/273 (39%), Positives = 157/273 (56%), Gaps = 22/273 (8%)
Query: 3 NSFSISLLTTLLPFILLITTVKSD-SFSFNFPRFESDTKTILSDGDANTTG-GVLQLTKK 60
NSF + LL+ F+LL+ V S S SF+FP+F + ++ GDA T GVLQLT
Sbjct: 10 NSFPL-LLSISFFFLLLLNKVNSTGSLSFSFPKFAPNQPYLILQGDALVTSTGVLQLTNV 68
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGL 120
+ G+PS+ S G+ + + D+ TG VA F T FSF++ DG F++A +
Sbjct: 69 -VNGVPSRKSLGRALYAAPFQIWDSTTGNVASFVTSFSFIIQAPNPATTADGLAFFLAPV 127
Query: 121 DFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINS 179
D P D GG LG+F KN + +NQ+VAVEFD+F+N WDP+ H+GI++NS
Sbjct: 128 DTQ-PLD--LGGMLGIF--KNGYFNKSNQIVAVEFDTFSNRHWDPT----GRHLGINVNS 178
Query: 180 IRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVL 239
I+SV T PW GE+ ISY +++K L+ +VYP+ ++ V VD+ VL
Sbjct: 179 IKSVRTVPWN---WTNGEVANVFISYEASTKSLTASLVYPS--LETSFIVHAIVDVKDVL 233
Query: 240 SDWVLVGFTAATG---SVSETHDILSWAFNSFL 269
+WV GF+A TG +T+D+LSW+F S L
Sbjct: 234 PEWVRFGFSATTGIDKGYVQTNDVLSWSFESNL 266
>emb|CAA68497.1| lectin-precursor (AA -30 to 245) [Pisum sativum]
gi|20804|emb|CAA47011.1| Psl lectin [Pisum sativum]
gi|126148|sp|P02867|LEC_PEA Lectin precursor
gi|169113|gb|AAA33676.1| lectin
Length = 275
Score = 166 bits (419), Expect = 8e-40
Identities = 103/271 (38%), Positives = 151/271 (55%), Gaps = 22/271 (8%)
Query: 4 SFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLY 63
SF L+ LL IL +++ SF +F D + ++ GD TT L LTK
Sbjct: 10 SFYAIFLSILLTTILFFKVNSTETTSFLITKFSPDQQNLIFQGDGYTTKEKLTLTKA--- 66
Query: 64 GIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
KN+ G+ + +H+ D TG VA+F T F+FV+N S DGFTF+IA +D
Sbjct: 67 ---VKNTVGRALYSSPIHIWDRETGNVANFVTSFTFVINAPNSYNVADGFTFFIAPVD-- 121
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRS 182
+ GG+LG+F+ +A Q VAVEFD+F N WDPS + H+GID+NSI+S
Sbjct: 122 -TKPQTGGGYLGVFN--SAEYDKTTQTVAVEFDTFYNAAWDPS--NRDRHIGIDVNSIKS 176
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYE----VDLGAV 238
V T W + GE +I++ +A+ VL+V + YPNS ++ E +Y V L V
Sbjct: 177 VNTKSWKLQ---NGEEANVVIAFNAATNVLTVSLTYPNS-LEEENVTSYTLSDVVSLKDV 232
Query: 239 LSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
+ +WV +GF+A TG+ H++LSW+F+S L
Sbjct: 233 VPEWVRIGFSATTGAEYAAHEVLSWSFHSEL 263
>emb|CAD27485.1| lectin [Lathyrus sativus]
Length = 251
Score = 166 bits (419), Expect = 8e-40
Identities = 103/268 (38%), Positives = 149/268 (55%), Gaps = 22/268 (8%)
Query: 4 SFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLY 63
SF L+ LL IL +++ SF +F D + ++ GD TT L LTK
Sbjct: 1 SFYAIFLSILLTTILFFKVNSTETTSFLITKFSPDQQNLIFQGDGYTTKEKLTLTKA--- 57
Query: 64 GIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
KN+ G+ + +H+ D+ TG VA F T F+F++N S DGFTF+IA +D
Sbjct: 58 ---VKNTVGRALYSSPIHIWDSTTGNVASFVTSFTFIINAPNSYNVADGFTFFIAPVD-- 112
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTN-VWDPSYPSNSPHVGIDINSIRS 182
+ GG+LG+F+ K+ TS Q VAVEFD+F N WDPS + H+GID+NSI+S
Sbjct: 113 -TKPQTGGGYLGVFNSKDYDKTS--QTVAVEFDTFYNAAWDPS--NGDRHIGIDVNSIKS 167
Query: 183 VATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYE----VDLGAV 238
V T W + G +I++ AS VL+V + YPNS V+ E +Y V L V
Sbjct: 168 VNTKSWNLQ---NGAEANVVIAFNGASNVLTVSLTYPNS-VEEENVTSYTLNEVVPLKDV 223
Query: 239 LSDWVLVGFTAATGSVSETHDILSWAFN 266
+ +WV +GF+A TG+ H++LSW+F+
Sbjct: 224 VPEWVRIGFSATTGAEFAAHEVLSWSFH 251
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 473,431,649
Number of Sequences: 2540612
Number of extensions: 21090398
Number of successful extensions: 43359
Number of sequences better than 10.0: 492
Number of HSP's better than 10.0 without gapping: 327
Number of HSP's successfully gapped in prelim test: 165
Number of HSP's that attempted gapping in prelim test: 41551
Number of HSP's gapped (non-prelim): 562
length of query: 269
length of database: 863,360,394
effective HSP length: 126
effective length of query: 143
effective length of database: 543,243,282
effective search space: 77683789326
effective search space used: 77683789326
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0541.11


