

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0404.10
(989 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
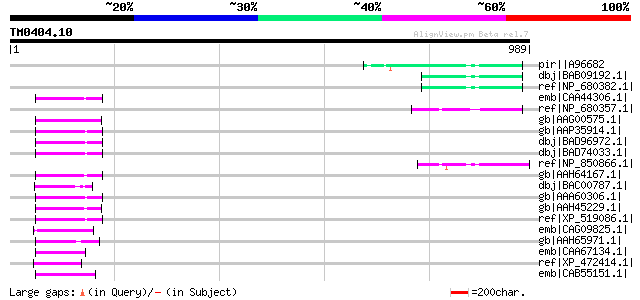
Score E
Sequences producing significant alignments: (bits) Value
pir||A96682 protein F1E22.12 [imported] - Arabidopsis thaliana g... 76 6e-12
dbj|BAB09192.1| non-LTR retroelement reverse transcriptase-like ... 66 5e-09
ref|NP_680382.1| hypothetical protein [Arabidopsis thaliana] 66 5e-09
emb|CAA44306.1| PR 264 [Gallus gallus] gi|266991|sp|P30352|SFRS2... 66 7e-09
ref|NP_680357.1| RNase H domain-containing protein [Arabidopsis ... 65 1e-08
gb|AAG00575.1| splicing factor arginine/serine rich 2 [Oryzias l... 65 1e-08
gb|AAP35914.1| splicing factor, arginine/serine-rich 2 [Homo sap... 64 2e-08
dbj|BAD96972.1| splicing factor, arginine/serine-rich 2 variant ... 64 2e-08
dbj|BAD74033.1| arginine/serine-rich 2 splicing factor [Pan trog... 64 2e-08
ref|NP_850866.1| reverse transcriptase-related [Arabidopsis thal... 64 2e-08
gb|AAH64167.1| Hypothetical protein MGC75633 [Xenopus tropicalis... 63 6e-08
dbj|BAC00787.1| glycine-rich RNA-binding protein [Physcomitrella... 63 6e-08
gb|AAA60306.1| splicing factor 61 2e-07
gb|AAH45229.1| Sfrs2-prov protein [Xenopus laevis] 61 2e-07
ref|XP_519086.1| PREDICTED: similar to Splicing factor, arginine... 59 6e-07
emb|CAG09825.1| unnamed protein product [Tetraodon nigroviridis] 59 8e-07
gb|AAH65971.1| Zgc:55876 protein [Danio rerio] 58 2e-06
emb|CAA67134.1| PR264/SC35 [Mus musculus] 58 2e-06
ref|XP_472414.1| OJ000126_13.13 [Oryza sativa (japonica cultivar... 58 2e-06
emb|CAB55151.1| Hypothetical protein Y116A8C.34 [Caenorhabditis ... 58 2e-06
>pir||A96682 protein F1E22.12 [imported] - Arabidopsis thaliana
gi|6686397|gb|AAF23831.1| F1E22.12 [Arabidopsis
thaliana]
Length = 1055
Score = 75.9 bits (185), Expect = 6e-12
Identities = 75/316 (23%), Positives = 120/316 (37%), Gaps = 36/316 (11%)
Query: 675 GAGWGMGKSTKRMVVKQKIRNSRAELVILQETKMDVHRNKVLLGWATSMD---------- 724
G GW K + N+R EL + + R++ L W S D
Sbjct: 329 GRGWDFAK-----IDPYTTNNTRLELRAVVLDLVTGARDR--LSWKFSQDGQFSVRSAYE 381
Query: 725 -MGLEIVPAVGSAGGLVTLWKTSLFQVSQQLMEC--NIAVKASLIRRGIPIENGGLCDIC 781
+ ++ VP A LWK + + + + N AV R + +C +C
Sbjct: 382 MLTVDEVPRPNMASFFNCLWKVRVPERVKTFLWLVGNQAVMTEEERHRRHLSASNVCQVC 441
Query: 782 GLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWEII 841
ES H++ CP +W ++ + F S+ L + R+ + I W I
Sbjct: 442 KGGVESMLHVLRDCPAQLGIWVRVVPQRRQQGFFSKSLFEWLYDNLGDRSGCEDIPWSTI 501
Query: 842 PYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKI 901
+ W W+ R +F + D + ++K W E + H +
Sbjct: 502 FAVIIWWGWKWRCGNIFGENTKCRDRVK---------FVKEWAVEV------YRAHSGNV 546
Query: 902 RIKVAAPKVHSA-DWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVG 960
+ + P+V W P G +K N DG+SRGN G++ GGVL+D G FS +G
Sbjct: 547 LVGITQPRVERMIGWVSPCVGWVKVNTDGASRGNPGLASAGGVLRDCTGAWCGGFSLNIG 606
Query: 961 STWAYVAERKALLEGL 976
A AE + GL
Sbjct: 607 RCSAPQAELWGVYYGL 622
>dbj|BAB09192.1| non-LTR retroelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 308
Score = 66.2 bits (160), Expect = 5e-09
Identities = 51/192 (26%), Positives = 75/192 (38%), Gaps = 16/192 (8%)
Query: 786 ESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWEIIPYAM 845
ES H+ CP +W + R F S+ L + R+ + I W I +
Sbjct: 25 ESVLHVFRDCPAQLGIWVRFVPRRRQQGFFSKSLFEWLYDNLCDRSSCEDIPWSTIFAVI 84
Query: 846 FWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIRIKV 905
W W+ R +F + D + ++K W E + H +
Sbjct: 85 IWWGWKWRCSNIFGENTKCRDRVK---------FVKEWVVEV------YRAHLGNALVGS 129
Query: 906 AAPKVHSA-DWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVGSTWA 964
P+V W P G +K N DG+SRGN G++ GGVL+D G FS +G A
Sbjct: 130 TQPRVERLIGWVLPCVGWVKVNTDGASRGNPGLASAGGVLRDCEGAWCGGFSLNIGRCSA 189
Query: 965 YVAERKALLEGL 976
AE + GL
Sbjct: 190 QHAELWGVYYGL 201
>ref|NP_680382.1| hypothetical protein [Arabidopsis thaliana]
Length = 258
Score = 66.2 bits (160), Expect = 5e-09
Identities = 51/192 (26%), Positives = 75/192 (38%), Gaps = 16/192 (8%)
Query: 786 ESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWEIIPYAM 845
ES H+ CP +W + R F S+ L + R+ + I W I +
Sbjct: 25 ESVLHVFRDCPAQLGIWVRFVPRRRQQGFFSKSLFEWLYDNLCDRSSCEDIPWSTIFAVI 84
Query: 846 FWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIRIKV 905
W W+ R +F + D + ++K W E + H +
Sbjct: 85 IWWGWKWRCSNIFGENTKCRDRVK---------FVKEWVVEV------YRAHLGNALVGS 129
Query: 906 AAPKVHSA-DWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVGSTWA 964
P+V W P G +K N DG+SRGN G++ GGVL+D G FS +G A
Sbjct: 130 TQPRVERLIGWVLPCVGWVKVNTDGASRGNPGLASAGGVLRDCEGAWCGGFSLNIGRCSA 189
Query: 965 YVAERKALLEGL 976
AE + GL
Sbjct: 190 QHAELWGVYYGL 201
>emb|CAA44306.1| PR 264 [Gallus gallus] gi|266991|sp|P30352|SFRS2_CHICK Splicing
factor, arginine/serine-rich 2 (Splicing factor SC35)
(SC-35) (Splicing component, 35 kDa) (PR264 protein)
gi|47604918|ref|NP_001001305.1| arginine/serine-rich2
splicing factor [Gallus gallus] gi|228503|prf||1805195A
RNA-binding protein PR264
Length = 221
Score = 65.9 bits (159), Expect = 7e-09
Identities = 42/131 (32%), Positives = 67/131 (51%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G S +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRY-GSSGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>ref|NP_680357.1| RNase H domain-containing protein [Arabidopsis thaliana]
Length = 633
Score = 65.1 bits (157), Expect = 1e-08
Identities = 55/216 (25%), Positives = 90/216 (41%), Gaps = 26/216 (12%)
Query: 767 RRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEW 826
RR + G+C +C E+ H++ CP I +W ++ R ++ F S+I L+W
Sbjct: 318 RRRRHLSASGVCQVCKGGDETILHVLRDCPSIAGIWGRLVPRGKITAFFASNI----LDW 373
Query: 827 --SSLRAISD--PIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYI-WDLHLIRVMWWIK 881
+L +++ W + + W W+ R VF + D + + + R +W
Sbjct: 374 VYQNLSDVTEIRGCPWATLFAIVVWWAWKWRCGNVFGENGRCRDRVRFVVDQAREIW--- 430
Query: 882 SWWKECPFTVLDFSQHFEKIRIKVAAPKVH-SADWCPPPPGCLKFNVDGSSRGNSGVSGI 940
H R + +V S W PP G K N DG+SRGN G++
Sbjct: 431 -------------IAHLNLRRGAMRGSEVEMSIKWTPPSTGWFKLNTDGASRGNPGLATA 477
Query: 941 GGVLKDANSTMLGRFSCAVGSTWAYVAERKALLEGL 976
GGV++D F +G A +AE + GL
Sbjct: 478 GGVVRDGEGQWCVGFVLNIGICSAPLAELWGVYYGL 513
>gb|AAG00575.1| splicing factor arginine/serine rich 2 [Oryzias latipes]
Length = 211
Score = 64.7 bits (156), Expect = 1e-08
Identities = 39/129 (30%), Positives = 65/129 (50%), Gaps = 3/129 (2%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 1 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYSKESRGFAFVRFFDKRDAEDAMDA 60
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P ++R P ++ G +G S+ G+ R R+
Sbjct: 61 MDGALLDGRELRVQMARYGRPPDSMYSRRGPPPRRYGGGGGYGRRSRSRSGSPRRRRRSR 120
Query: 167 IPLKRNSQS 175
+ S+S
Sbjct: 121 SRSRSRSRS 129
>gb|AAP35914.1| splicing factor, arginine/serine-rich 2 [Homo sapiens]
gi|6755478|ref|NP_035488.1| splicing factor,
arginine/serine-rich 2 [Mus musculus]
gi|455419|emb|CAA53383.1| PR264/SC35 [Homo sapiens]
gi|35597|emb|CAA44307.1| PR 264 [Homo sapiens]
gi|61359244|gb|AAX41688.1| splicing factor
arginine/serine-rich 2 [synthetic construct]
gi|61821161|ref|XP_585074.1| PREDICTED: similar to
Splicing factor, arginine/serine-rich 2 (Splicing factor
SC35) (SC-35) (Splicing component, 35 kDa) [Bos taurus]
gi|67969334|dbj|BAE01019.1| unnamed protein product
[Macaca fascicularis] gi|47123339|gb|AAH70086.1|
Splicing factor, arginine/serine-rich 2 [Homo sapiens]
gi|47271443|ref|NP_003007.2| splicing factor,
arginine/serine-rich 2 [Homo sapiens]
gi|12654915|gb|AAH01303.1| SFRS2 protein [Homo sapiens]
gi|12653143|gb|AAH00339.1| SFRS2 protein [Homo sapiens]
gi|60416437|sp|Q01130|SFRS2_HUMAN Splicing factor,
arginine/serine-rich 2 (Splicing factor SC35) (SC-35)
(Splicing component, 35 kDa) (PR264 protein)
gi|18280933|sp|Q62093|SFRS2_MOUSE Splicing factor,
arginine/serine-rich 2 (Splicing factor SC35) (SC-35)
(Splicing component, 35 kDa) (PR264 protein)
gi|52783335|sp|Q6PDU1|SFRS2_RAT Splicing factor,
arginine/serine-rich 2 (Splicing factor SC35) (SC-35)
(Splicing component, 35 kDa) gi|3335676|gb|AAC71000.1|
splicing factor SC35 [Mus musculus]
gi|57528425|ref|NP_001009720.1| hypothetical protein
LOC494445 [Rattus norvegicus] gi|539663|pir||A42701
splicing factor SFRS2 - human
gi|26352962|dbj|BAC40111.1| unnamed protein product [Mus
musculus] gi|26351947|dbj|BAC39610.1| unnamed protein
product [Mus musculus] gi|13529557|gb|AAH05493.1| Sfrs2
protein [Mus musculus] gi|34849641|gb|AAH58508.1|
Similar to splicing factor, arginine/serine-rich 2
[Rattus norvegicus] gi|228504|prf||1805195B RNA-binding
protein PR264
Length = 221
Score = 64.3 bits (155), Expect = 2e-08
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>dbj|BAD96972.1| splicing factor, arginine/serine-rich 2 variant [Homo sapiens]
Length = 221
Score = 64.3 bits (155), Expect = 2e-08
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>dbj|BAD74033.1| arginine/serine-rich 2 splicing factor [Pan troglodytes]
gi|60414777|sp|Q5R1W5|SFRS2_PANTR Splicing factor,
arginine/serine-rich 2 (Splicing factor SC35) (SC-35)
(Splicing component, 35 kDa)
Length = 221
Score = 64.3 bits (155), Expect = 2e-08
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>ref|NP_850866.1| reverse transcriptase-related [Arabidopsis thaliana]
Length = 411
Score = 64.3 bits (155), Expect = 2e-08
Identities = 58/218 (26%), Positives = 89/218 (40%), Gaps = 24/218 (11%)
Query: 777 LCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLR----AI 832
LC++C ES H++ CP + +W I+ + + F S L EW A+
Sbjct: 120 LCEVCKGGVESAMHILRDCPTMEGIWLRIVPPRKRHEFFKKS----LFEWVYGNLGDDAV 175
Query: 833 SDPIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVL 892
D + W ++ W W+ R VF + D + ++K+ KE
Sbjct: 176 VDGVPWSVMFAMGVWWGWKWRCGNVFGENRKCRDRVQ---------FVKNVAKEV----- 221
Query: 893 DFSQHFEKIRIKVAAPKVHSA-DWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTM 951
F H + +V W P G +K N DG+SRGN G++ GGVL+D
Sbjct: 222 -FQAHGSRAGNDRGRNRVERLIGWGSPGVGWVKLNTDGASRGNPGLATAGGVLRDEEGNW 280
Query: 952 LGRFSCAVGSTWAYVAERKALLEGLLLCGKIYSCKLRL 989
G F+ +G A +AE + GL I +L L
Sbjct: 281 RGGFAANIGRCSAPLAELWEVYYGLYFAWAIRIPQLEL 318
>gb|AAH64167.1| Hypothetical protein MGC75633 [Xenopus tropicalis]
gi|45361503|ref|NP_989328.1| hypothetical protein
LOC394953 [Xenopus tropicalis]
Length = 220
Score = 62.8 bits (151), Expect = 6e-08
Identities = 40/131 (30%), Positives = 65/131 (49%), Gaps = 6/131 (4%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H R P ++ ++G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHGRRGPPPRRY---GDYGRRSRSPRRRRRSRSRSK 131
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 132 SRSRSRSRSRY 142
>dbj|BAC00787.1| glycine-rich RNA-binding protein [Physcomitrella patens]
Length = 155
Score = 62.8 bits (151), Expect = 6e-08
Identities = 43/113 (38%), Positives = 58/113 (51%), Gaps = 10/113 (8%)
Query: 48 GKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAM 107
G LFV GL TT +K AF++FG + V + + R FGFV F+ D DAE A+
Sbjct: 41 GSKLFVGGLAWGTTDDNIKEAFSAFGEVTEVKIICDRDTGRSRGFGFVTFATDQDAEAAL 100
Query: 108 RRMNGSILNGATINVSLARLPQDQHTRPRPTDNQWKGVSEWG--EHSKPNKGN 158
+ ++G L G TI V+ A T+ P D Q G +G +HS N GN
Sbjct: 101 QALDGRDLAGRTIRVNYA-------TKQSPQDRQ-SGGGMYGRRDHSGGNSGN 145
>gb|AAA60306.1| splicing factor
Length = 221
Score = 61.2 bits (147), Expect = 2e-07
Identities = 40/131 (30%), Positives = 65/131 (49%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F + R+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYRRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>gb|AAH45229.1| Sfrs2-prov protein [Xenopus laevis]
Length = 215
Score = 61.2 bits (147), Expect = 2e-07
Identities = 39/129 (30%), Positives = 64/129 (49%), Gaps = 6/129 (4%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H R P ++ ++G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHGRRGPAPRRY---GDYGRRSRSPRRRRRSRSRSK 131
Query: 167 IPLKRNSQS 175
+ S+S
Sbjct: 132 SRSRSRSRS 140
>ref|XP_519086.1| PREDICTED: similar to Splicing factor, arginine/serine-rich, 46kD
[Pan troglodytes]
Length = 583
Score = 59.3 bits (142), Expect = 6e-07
Identities = 38/131 (29%), Positives = 65/131 (49%), Gaps = 5/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
+L VD LT +T+ ++ F +GR+ V++ + F FVRF SDA+ A
Sbjct: 305 TLKVDNLTYRTSPDSLRRVFEKYGRVGDVYIPREPHTKAPRGFAFVRFHDRSDAQDAEAA 364
Query: 110 MNGSILNGATINVSLARLPQ---DQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR + + ++ P+ W G +G S+ +G RS +
Sbjct: 365 MDGAVLDGRELRVQMARYGRRDLPRSSQEEPSGRSWGG--RYGRRSRSPRGRHRSQSRGP 422
Query: 167 IPLKRNSQSDY 177
K S+S Y
Sbjct: 423 SYSKSRSRSHY 433
>emb|CAG09825.1| unnamed protein product [Tetraodon nigroviridis]
Length = 391
Score = 58.9 bits (141), Expect = 8e-07
Identities = 39/118 (33%), Positives = 56/118 (47%), Gaps = 4/118 (3%)
Query: 46 RIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEV 105
R GK LF+ GL +TT ++ F+ +GRI V + + + F FV F SDA+
Sbjct: 6 RPGK-LFIGGLNTETTEKALEQFFSKYGRIAEVILMKDRETNKSRGFAFVTFESPSDAKD 64
Query: 106 AMRRMNGSILNGATINVSLARLPQDQ---HTRPRPTDNQWKGVSEWGEHSKPNKGNRR 160
A R MNG L+G I V A PQ + P P ++ +G+ S+ G R
Sbjct: 65 AAREMNGKSLDGKNIKVEQATKPQFESGGRRGPPPMHSRSRGLPRGMRGSRGGPGGMR 122
>gb|AAH65971.1| Zgc:55876 protein [Danio rerio]
Length = 220
Score = 57.8 bits (138), Expect = 2e-06
Identities = 41/123 (33%), Positives = 61/123 (49%), Gaps = 8/123 (6%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARLPQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRR-SARKEWIP 168
M+G+IL+G + V +AR RP D+ + G S G+RR AR+ P
Sbjct: 75 MDGAILDGRELRVQMARY-------GRPPDSYYGGGGGGRRGSGGGGGSRRHGARRSRSP 127
Query: 169 LKR 171
+R
Sbjct: 128 RRR 130
>emb|CAA67134.1| PR264/SC35 [Mus musculus]
Length = 121
Score = 57.8 bits (138), Expect = 2e-06
Identities = 33/98 (33%), Positives = 52/98 (52%), Gaps = 3/98 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKG 144
M+G++L+G + V +AR P H+R P ++ G
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG 112
>ref|XP_472414.1| OJ000126_13.13 [Oryza sativa (japonica cultivar-group)]
gi|38346331|emb|CAE02067.2| OJ000126_13.13 [Oryza sativa
(japonica cultivar-group)] gi|38344030|emb|CAE01512.2|
OJ991214_12.1 [Oryza sativa (japonica cultivar-group)]
Length = 137
Score = 57.8 bits (138), Expect = 2e-06
Identities = 31/93 (33%), Positives = 47/93 (50%)
Query: 45 RRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAE 104
R I LF+ GL+ T + AF+ +G++L + K R FGFV+F+ + A
Sbjct: 33 RGITYRLFIGGLSQFATEDSLAEAFSQYGQVLEATIVTDKMTNRPKGFGFVKFASEEAAN 92
Query: 105 VAMRRMNGSILNGATINVSLARLPQDQHTRPRP 137
A MNG +LNG I V +A+ ++ T P
Sbjct: 93 KAKEEMNGKVLNGRVIYVDIAKAKMNRTTDSSP 125
>emb|CAB55151.1| Hypothetical protein Y116A8C.34 [Caenorhabditis elegans]
gi|17539498|ref|NP_503034.1| CYcloPhilin (36.4 kD)
(cyp-13) [Caenorhabditis elegans] gi|7509375|pir||T31517
hypothetical protein Y116A8C.34 - Caenorhabditis elegans
Length = 331
Score = 57.8 bits (138), Expect = 2e-06
Identities = 36/117 (30%), Positives = 57/117 (47%), Gaps = 3/117 (2%)
Query: 49 KSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMR 108
++L+V G T+ T + AF FG ++ + + + FGFV F DA +A+
Sbjct: 11 RTLYVGGFTEDVTEKVLMAAFIPFGDVVAISIPMDYESGKHRGFGFVEFDMAEDAAMAID 70
Query: 109 RMNGSILNGATINVSLARLPQ--DQHTRPRPTDNQW-KGVSEWGEHSKPNKGNRRSA 162
MN S L G TI V+ AR P+ ++ +P D++W K GE + G+ A
Sbjct: 71 NMNESELFGKTIRVNFARPPKATERSQKPVWADDEWLKKYGRGGEAAAEEDGDAEKA 127
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.134 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,773,579,508
Number of Sequences: 2540612
Number of extensions: 77626941
Number of successful extensions: 210798
Number of sequences better than 10.0: 2186
Number of HSP's better than 10.0 without gapping: 964
Number of HSP's successfully gapped in prelim test: 1222
Number of HSP's that attempted gapping in prelim test: 207923
Number of HSP's gapped (non-prelim): 3721
length of query: 989
length of database: 863,360,394
effective HSP length: 138
effective length of query: 851
effective length of database: 512,755,938
effective search space: 436355303238
effective search space used: 436355303238
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0404.10


