

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.14
(76 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
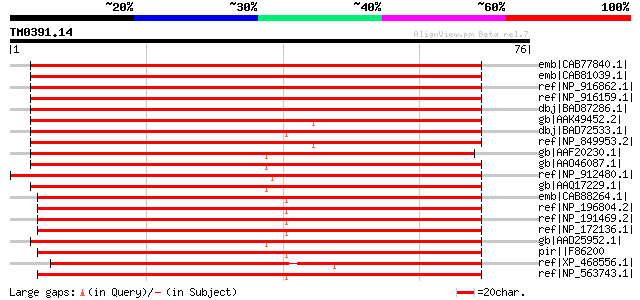
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB77840.1| putative glucan synthase component [Arabidopsis ... 95 4e-19
emb|CAB81039.1| AT4g04970 [Arabidopsis thaliana] gi|5732072|gb|A... 94 9e-19
ref|NP_916862.1| putative 1,3-beta-glucan synthase [Oryza sativa... 87 1e-16
ref|NP_916159.1| putative glucan synthase [Oryza sativa (japonic... 82 4e-15
dbj|BAD87286.1| putative callose synthase 1 catalytic subunit [O... 82 4e-15
gb|AAK49452.2| putative beta-1,3-glucan synthase [Nicotiana alata] 76 3e-13
dbj|BAD72533.1| putative callose synthase 1 catalytic subunit [O... 74 7e-13
ref|NP_849953.2| glycosyl transferase family 48 protein [Arabido... 73 2e-12
gb|AAF20230.1| putative glucan synthase [Arabidopsis thaliana] g... 73 2e-12
gb|AAO46087.1| putative callose synthase [Hordeum vulgare subsp.... 72 4e-12
ref|NP_912480.1| Putative glucan synthase [Oryza sativa (japonic... 70 1e-11
gb|AAQ17229.1| beta 1,3 glucan synthase [Lolium multiflorum] 70 1e-11
emb|CAB88264.1| callose synthase catalytic subunit-like protein ... 67 1e-10
ref|NP_196804.2| glycosyl transferase family 48 protein [Arabido... 67 1e-10
ref|NP_191469.2| glycosyl transferase family 48 protein [Arabido... 67 2e-10
ref|NP_172136.1| glycosyl transferase family 48 protein [Arabido... 65 3e-10
gb|AAD25952.1| putative callose synthase catalytic subunit [Goss... 65 3e-10
pir||F86200 protein F12K11.17 [imported] - Arabidopsis thaliana ... 65 3e-10
ref|XP_468556.1| putative callose synthase 1 catalytic subunit [... 62 5e-09
ref|NP_563743.1| callose synthase 1 (CALS1) / 1,3-beta-glucan sy... 60 1e-08
>emb|CAB77840.1| putative glucan synthase component [Arabidopsis thaliana]
gi|4206209|gb|AAD11597.1| putative glucan synthase
component [Arabidopsis thaliana]
gi|4263042|gb|AAD15311.1| putative glucan synthase
component [Arabidopsis thaliana] gi|25407209|pir||A85045
probable glucan synthase component [imported] -
Arabidopsis thaliana gi|15236339|ref|NP_192264.1|
glycosyl transferase family 48 protein [Arabidopsis
thaliana]
Length = 1780
Score = 95.1 bits (235), Expect = 4e-19
Identities = 41/66 (62%), Positives = 53/66 (80%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWY 63
F+FSYFLQ++P+I S+++ LKDV+Y+WH F + N V LLW+P VLIYLMDIQIWY
Sbjct: 502 FTFSYFLQVKPMIKPSKLLWNLKDVDYEWHQFYGDSNRFSVALLWLPVVLIYLMDIQIWY 561
Query: 64 AIYSSL 69
AIYSS+
Sbjct: 562 AIYSSI 567
>emb|CAB81039.1| AT4g04970 [Arabidopsis thaliana] gi|5732072|gb|AAD48971.1| contains
similarity to glucan synthases [Arabidopsis thaliana]
gi|25407306|pir||E85062 hypothetical protein AT4g04970
[imported] - Arabidopsis thaliana
gi|18412763|ref|NP_567278.1| callose synthase, putative
/ 1,3-beta-glucan synthase, putative [Arabidopsis
thaliana]
Length = 1768
Score = 94.0 bits (232), Expect = 9e-19
Identities = 39/66 (59%), Positives = 52/66 (78%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWY 63
F FSYFLQI+P+IA +R +L LKD Y WH F + + + VG+LW+P +L+YLMD+QIWY
Sbjct: 493 FIFSYFLQIRPLIAPTRALLNLKDATYNWHEFFGSTHRIAVGMLWLPVILVYLMDLQIWY 552
Query: 64 AIYSSL 69
+IYSSL
Sbjct: 553 SIYSSL 558
>ref|NP_916862.1| putative 1,3-beta-glucan synthase [Oryza sativa (japonica
cultivar-group)] gi|21644609|dbj|BAC01168.1|
1,3-beta-glucan synthase component-like [Oryza sativa
(japonica cultivar-group)] gi|18461174|dbj|BAB84371.1|
1,3-beta-glucan synthase component-like [Oryza sativa
(japonica cultivar-group)]
Length = 1769
Score = 87.0 bits (214), Expect = 1e-16
Identities = 37/66 (56%), Positives = 50/66 (75%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWY 63
F+FSYFLQI+P++ ++ + +LK ++Y WH F N V +LW+P VLIYLMDIQIWY
Sbjct: 483 FAFSYFLQIRPLVKPTQEIYKLKKIDYAWHEFFGKSNRFAVFVLWLPVVLIYLMDIQIWY 542
Query: 64 AIYSSL 69
AI+SSL
Sbjct: 543 AIFSSL 548
>ref|NP_916159.1| putative glucan synthase [Oryza sativa (japonica cultivar-group)]
gi|20160746|dbj|BAB89687.1| putative callose synthase 1
catalytic subunit [Oryza sativa (japonica
cultivar-group)]
Length = 1790
Score = 82.0 bits (201), Expect = 4e-15
Identities = 33/66 (50%), Positives = 51/66 (77%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWY 63
FSFSYFLQI+P++ ++V+ +L D++ W F+ + + V +LW+P ++IYLMDIQIWY
Sbjct: 511 FSFSYFLQIKPMVGPTKVIFKLHDIKRNWFEFMPHTERLAVIILWLPVIIIYLMDIQIWY 570
Query: 64 AIYSSL 69
A++SSL
Sbjct: 571 AVFSSL 576
>dbj|BAD87286.1| putative callose synthase 1 catalytic subunit [Oryza sativa
(japonica cultivar-group)]
Length = 1618
Score = 82.0 bits (201), Expect = 4e-15
Identities = 33/66 (50%), Positives = 51/66 (77%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWY 63
FSFSYFLQI+P++ ++V+ +L D++ W F+ + + V +LW+P ++IYLMDIQIWY
Sbjct: 339 FSFSYFLQIKPMVGPTKVIFKLHDIKRNWFEFMPHTERLAVIILWLPVIIIYLMDIQIWY 398
Query: 64 AIYSSL 69
A++SSL
Sbjct: 399 AVFSSL 404
>gb|AAK49452.2| putative beta-1,3-glucan synthase [Nicotiana alata]
Length = 1931
Score = 75.9 bits (185), Expect = 3e-13
Identities = 31/68 (45%), Positives = 49/68 (71%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVY--VGLLWIPAVLIYLMDIQI 61
F+FSYF+QI+P+I +++++++ V+Y WH F + + Y V LW P +L+Y MD QI
Sbjct: 689 FAFSYFIQIKPLIKPTKMIMDINRVQYAWHEFFPDARSNYGAVLSLWAPVILVYFMDAQI 748
Query: 62 WYAIYSSL 69
WYAI+S+L
Sbjct: 749 WYAIFSTL 756
>dbj|BAD72533.1| putative callose synthase 1 catalytic subunit [Oryza sativa
(japonica cultivar-group)]
Length = 1910
Score = 74.3 bits (181), Expect = 7e-13
Identities = 31/68 (45%), Positives = 48/68 (70%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQI 61
F+FSYF+QI+P+I ++ ++ + ++ Y+WH F N N V LW P +L+YLMD QI
Sbjct: 707 FAFSYFVQIKPLIKPTKDIMNVHNIHYEWHEFFPNASYNVGAVMSLWAPVLLVYLMDTQI 766
Query: 62 WYAIYSSL 69
WYAI+S++
Sbjct: 767 WYAIFSTI 774
>ref|NP_849953.2| glycosyl transferase family 48 protein [Arabidopsis thaliana]
Length = 1923
Score = 73.2 bits (178), Expect = 2e-12
Identities = 29/68 (42%), Positives = 47/68 (68%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVY--VGLLWIPAVLIYLMDIQI 61
F+FSYFLQ++ ++ + ++ ++ V+Y+WH F N Y V LW+P +L+Y MD QI
Sbjct: 676 FAFSYFLQVKLLVKPTNAIMSIRHVKYKWHEFFPNAEHNYGAVVSLWLPVILVYFMDTQI 735
Query: 62 WYAIYSSL 69
WYAI+S++
Sbjct: 736 WYAIFSTI 743
>gb|AAF20230.1| putative glucan synthase [Arabidopsis thaliana]
gi|15231404|ref|NP_187372.1| glycosyl transferase family
48 protein [Arabidopsis thaliana]
Length = 1931
Score = 73.2 bits (178), Expect = 2e-12
Identities = 31/67 (46%), Positives = 48/67 (71%), Gaps = 2/67 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFI--KNGNTVYVGLLWIPAVLIYLMDIQI 61
FSF+YFLQI+P++ +R++++ ++ Y WH F+ KN N + V LW P V IYL+DI I
Sbjct: 696 FSFAYFLQIKPLVGPTRMIVKQNNIPYSWHDFVSRKNYNALTVASLWAPVVAIYLLDIHI 755
Query: 62 WYAIYSS 68
+Y I+S+
Sbjct: 756 FYTIFSA 762
>gb|AAO46087.1| putative callose synthase [Hordeum vulgare subsp. vulgare]
Length = 1915
Score = 72.0 bits (175), Expect = 4e-12
Identities = 30/68 (44%), Positives = 47/68 (69%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFI--KNGNTVYVGLLWIPAVLIYLMDIQI 61
FSF+YFLQI+P++ +R+++ K ++YQWH F+ N N + + LW P IYL+DI +
Sbjct: 675 FSFTYFLQIRPLVKPTRLIISFKGLQYQWHDFVSKNNHNAITILSLWAPVASIYLLDIHV 734
Query: 62 WYAIYSSL 69
+Y I S+L
Sbjct: 735 FYTIMSAL 742
>ref|NP_912480.1| Putative glucan synthase [Oryza sativa (japonica cultivar-group)]
gi|20330757|gb|AAM19120.1| Putative glucan synthase
[Oryza sativa (japonica cultivar-group)]
Length = 1642
Score = 70.1 bits (170), Expect = 1e-11
Identities = 30/71 (42%), Positives = 48/71 (67%), Gaps = 2/71 (2%)
Query: 1 IIFFSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIK--NGNTVYVGLLWIPAVLIYLMD 58
+I F+FSY+++I+P++ ++ +++L +QWH F NGN V LW P +L+Y MD
Sbjct: 345 LIKFAFSYYVEIKPLVEPTKDIMKLPIHTFQWHEFFPKANGNIGVVIALWAPIILVYFMD 404
Query: 59 IQIWYAIYSSL 69
QIWY I+S+L
Sbjct: 405 TQIWYTIFSTL 415
>gb|AAQ17229.1| beta 1,3 glucan synthase [Lolium multiflorum]
Length = 1906
Score = 70.1 bits (170), Expect = 1e-11
Identities = 29/68 (42%), Positives = 47/68 (68%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFI--KNGNTVYVGLLWIPAVLIYLMDIQI 61
FSF+YFLQI+P++ +++++ +D++YQWH F N N + LW P V IYL+DI +
Sbjct: 674 FSFTYFLQIKPLVEPTQLIISFRDLQYQWHDFFSKNNHNAFTILSLWAPVVSIYLLDIHV 733
Query: 62 WYAIYSSL 69
+Y I S++
Sbjct: 734 FYTIMSAI 741
>emb|CAB88264.1| callose synthase catalytic subunit-like protein [Arabidopsis
thaliana] gi|11357214|pir||T49914 callose synthase
catalytic subunit-like protein - Arabidopsis thaliana
Length = 1963
Score = 67.0 bits (162), Expect = 1e-10
Identities = 27/67 (40%), Positives = 45/67 (66%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+FSY+++I+P++A ++ +++ + +QWH F N V LW P +L+Y MD QIW
Sbjct: 684 AFSYYIEIRPLVAPTQAIMKARVTNFQWHEFFPRAKNNIGVVIALWAPIILVYFMDSQIW 743
Query: 63 YAIYSSL 69
YAI+S+L
Sbjct: 744 YAIFSTL 750
>ref|NP_196804.2| glycosyl transferase family 48 protein [Arabidopsis thaliana]
Length = 1889
Score = 67.0 bits (162), Expect = 1e-10
Identities = 27/67 (40%), Positives = 45/67 (66%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+FSY+++I+P++A ++ +++ + +QWH F N V LW P +L+Y MD QIW
Sbjct: 684 AFSYYIEIRPLVAPTQAIMKARVTNFQWHEFFPRAKNNIGVVIALWAPIILVYFMDSQIW 743
Query: 63 YAIYSSL 69
YAI+S+L
Sbjct: 744 YAIFSTL 750
>ref|NP_191469.2| glycosyl transferase family 48 protein [Arabidopsis thaliana]
Length = 1934
Score = 66.6 bits (161), Expect = 2e-10
Identities = 29/67 (43%), Positives = 45/67 (66%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+F+Y+++I P+I +++++ L YQWH F + N V +W P VL+YLMD QIW
Sbjct: 697 AFNYYVEILPLITPTKMIMNLHIGHYQWHEFFPHATNNIGVVIAIWAPIVLVYLMDTQIW 756
Query: 63 YAIYSSL 69
YAI+S+L
Sbjct: 757 YAIFSTL 763
>ref|NP_172136.1| glycosyl transferase family 48 protein [Arabidopsis thaliana]
Length = 1933
Score = 65.5 bits (158), Expect = 3e-10
Identities = 27/67 (40%), Positives = 45/67 (66%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+FSY+++I P++ ++++ ++ V Y+WH F N N + +W P VL+Y MD QIW
Sbjct: 693 AFSYYVEILPLVNPTKLIWDMHVVNYEWHEFFPNATHNIGVIIAIWGPIVLVYFMDTQIW 752
Query: 63 YAIYSSL 69
YAI+S+L
Sbjct: 753 YAIFSTL 759
>gb|AAD25952.1| putative callose synthase catalytic subunit [Gossypium hirsutum]
Length = 1899
Score = 65.5 bits (158), Expect = 3e-10
Identities = 27/68 (39%), Positives = 45/68 (65%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFI--KNGNTVYVGLLWIPAVLIYLMDIQI 61
F+F+Y QI+P++ +R V+ + ++EY WH F+ N N V V LW P + +YL+DI I
Sbjct: 673 FAFAYSFQIKPLVKPTRTVIAMDNIEYSWHDFVSRNNHNAVTVVCLWAPVIAMYLLDIYI 732
Query: 62 WYAIYSSL 69
+Y + S++
Sbjct: 733 FYTVLSAV 740
>pir||F86200 protein F12K11.17 [imported] - Arabidopsis thaliana
gi|6692688|gb|AAF24822.1| F12K11.17 [Arabidopsis
thaliana]
Length = 1930
Score = 65.5 bits (158), Expect = 3e-10
Identities = 27/67 (40%), Positives = 45/67 (66%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+FSY+++I P++ ++++ ++ V Y+WH F N N + +W P VL+Y MD QIW
Sbjct: 690 AFSYYVEILPLVNPTKLIWDMHVVNYEWHEFFPNATHNIGVIIAIWGPIVLVYFMDTQIW 749
Query: 63 YAIYSSL 69
YAI+S+L
Sbjct: 750 YAIFSTL 756
>ref|XP_468556.1| putative callose synthase 1 catalytic subunit [Oryza sativa
(japonica cultivar-group)] gi|48716406|dbj|BAD23015.1|
putative callose synthase 1 catalytic subunit [Oryza
sativa (japonica cultivar-group)]
Length = 1969
Score = 61.6 bits (148), Expect = 5e-09
Identities = 26/66 (39%), Positives = 45/66 (67%), Gaps = 4/66 (6%)
Query: 7 SYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGL---LWIPAVLIYLMDIQIWY 63
SY+++I+P++ ++ +++ +QWH F +GN +G+ LW P +L+Y MD QIWY
Sbjct: 707 SYYVEIKPLVRPTKDIMKEPIRTFQWHEFFPHGNN-NIGIVIALWAPIILVYFMDTQIWY 765
Query: 64 AIYSSL 69
AI+S+L
Sbjct: 766 AIFSTL 771
>ref|NP_563743.1| callose synthase 1 (CALS1) / 1,3-beta-glucan synthase 1
[Arabidopsis thaliana]
Length = 1922
Score = 60.5 bits (145), Expect = 1e-08
Identities = 26/67 (38%), Positives = 41/67 (60%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+FSY+ +I+P++ ++ ++ + Y WH F + N V LW P +L+Y MD QIW
Sbjct: 647 AFSYYAEIKPLVGPTKDIMRIHISVYSWHEFFPHAKNNLGVVIALWSPVILVYFMDTQIW 706
Query: 63 YAIYSSL 69
YAI S+L
Sbjct: 707 YAIVSTL 713
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.333 0.146 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 128,231,833
Number of Sequences: 2540612
Number of extensions: 4233504
Number of successful extensions: 14804
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 14742
Number of HSP's gapped (non-prelim): 40
length of query: 76
length of database: 863,360,394
effective HSP length: 52
effective length of query: 24
effective length of database: 731,248,570
effective search space: 17549965680
effective search space used: 17549965680
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0391.14


