

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.6
(1591 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
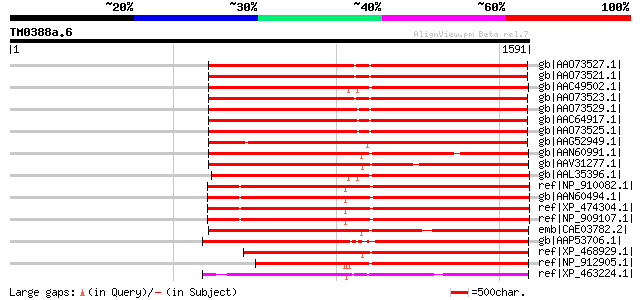
Sequences producing significant alignments: (bits) Value
gb|AAO73527.1| gag-pol polyprotein [Glycine max] 960 0.0
gb|AAO73521.1| gag-pol polyprotein [Glycine max] 957 0.0
gb|AAC49502.1| Pol [Zea mays] gi|7489803|pir||T04112 pol protein... 954 0.0
gb|AAO73523.1| gag-pol polyprotein [Glycine max] 953 0.0
gb|AAO73529.1| gag-pol polyprotein [Glycine max] 949 0.0
gb|AAC64917.1| gag-pol polyprotein [Glycine max] 938 0.0
gb|AAO73525.1| gag-pol polyprotein [Glycine max] 937 0.0
gb|AAG52949.1| gag/pol polyprotein [Arabidopsis thaliana] 919 0.0
gb|AAN60991.1| Putative Zea mays retrotransposon Opie-2 [Oryza s... 919 0.0
gb|AAV31277.1| putative polyprotein [Oryza sativa (japonica cult... 916 0.0
gb|AAL35396.1| Opie2a pol [Zea mays] 915 0.0
ref|NP_910082.1| putative gag-pol polyprotein [Oryza sativa (jap... 908 0.0
gb|AAN60494.1| Putative Zea mays retrotransposon Opie-2 [Oryza s... 904 0.0
ref|XP_474304.1| OSJNBb0004A17.2 [Oryza sativa (japonica cultiva... 904 0.0
ref|NP_909107.1| unnamed protein product [Oryza sativa (japonica... 904 0.0
emb|CAE03782.2| OSJNBa0063G07.6 [Oryza sativa (japonica cultivar... 900 0.0
gb|AAP53706.1| hypothetical protein [Oryza sativa (japonica cult... 884 0.0
ref|XP_468929.1| putative polyprotein [Oryza sativa (japonica cu... 866 0.0
ref|NP_912905.1| unnamed protein product [Oryza sativa (japonica... 834 0.0
ref|XP_463224.1| putative integrase [Oryza sativa (japonica cult... 792 0.0
>gb|AAO73527.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 960 bits (2482), Expect = 0.0
Identities = 476/979 (48%), Positives = 654/979 (66%), Gaps = 6/979 (0%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI+G G + D P + VLL
Sbjct: 560 EDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHDGLPSLNKVLL 619
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E +
Sbjct: 620 VKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQETSYSS 679
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ VRG+PNLK +C CQ GK K+
Sbjct: 680 T-CLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKM 738
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TSR LELLH+DL GP++ ES+GGKRY ++VDD+SR+TWV F+ K E+
Sbjct: 739 SHQKLRHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSETFE 798
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN +F S GI H+FS TPQQNG+VER
Sbjct: 799 VFKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSAAITPQQNGIVER 858
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE AR ML + + W EA+NTACYI NR+++R T YE+WK KP++ +
Sbjct: 859 KNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPSVKH 918
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 919 FHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 978
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRITAAHPME 1087
+ + + E L NV+D K+ E AE D +E+ + +RI HP E
Sbjct: 979 SPARKKDVEEDVRTLGDNVADAAKSGENAE-NSDSATDESNINQPDKRSSTRIQKMHPKE 1037
Query: 1088 LIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSK 1147
LI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF +
Sbjct: 1038 LIIGDPNRGVTTRSR----EVEIVSNSCFVSKIEPKNVKEALTDEFWINAMQEELEQFKR 1093
Query: 1148 NDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIA 1207
N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP+A
Sbjct: 1094 NEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAPVA 1153
Query: 1208 RLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKS 1267
RLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D PDHV++LKK+
Sbjct: 1154 RLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADPTHPDHVYRLKKA 1213
Query: 1268 LYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQS 1327
LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 1214 LYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSNE 1273
Query: 1328 LCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAK 1387
+ + F + MQ+EFEMS++GELTYFLG+QV Q + ++ QS+Y K ++KKF M ++ +
Sbjct: 1274 MLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIVKKFGMENASHKR 1333
Query: 1388 TPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLT 1447
TP L K++A V Q LYR MIGSLLYLTASRPDI ++V +CAR+Q++P+ +HLT
Sbjct: 1334 TPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISHLT 1393
Query: 1448 AVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWE 1507
VKRIL+Y+ T++ G+MY S L GYCDAD+AG +RKSTSG C +LG+NL+SW
Sbjct: 1394 QVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADDRKSTSGGCFYLGNNLISWF 1453
Query: 1508 SKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPI 1567
SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKNP+
Sbjct: 1454 SKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINISKNPV 1513
Query: 1568 LHSRAKHIEVKYHFIRDYV 1586
HSR KHI++++H+IRD V
Sbjct: 1514 QHSRTKHIDIRHHYIRDLV 1532
>gb|AAO73521.1| gag-pol polyprotein [Glycine max]
Length = 1574
Score = 957 bits (2475), Expect = 0.0
Identities = 475/979 (48%), Positives = 653/979 (66%), Gaps = 6/979 (0%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI+G G + D P + VLL
Sbjct: 558 EDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHDGLPSLNKVLL 617
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E +
Sbjct: 618 VKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQETSYSS 677
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ VRG+PNLK +C CQ GK K+
Sbjct: 678 T-CLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKM 736
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TSR LELLH+DL GP++ ES+GGKRY ++VDD+SR+TWVKF+ K E+
Sbjct: 737 SHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVKFIREKSETFE 796
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN + S GI H+FS TPQQNG+VER
Sbjct: 797 VFKELSLRLQREKDCVIKRIRSDHGREFENSRLTEFCTSEGITHEFSAAITPQQNGIVER 856
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE AR ML + + W EA+NTACYI NR+++R T YE+WK KP++ +
Sbjct: 857 KNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPSVKH 916
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 917 FHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 976
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRITAAHPME 1087
+ + + E NV+D K+ E AE D +E+ + +RI HP E
Sbjct: 977 SPARKKDVEEDVRTSGDNVADAAKSGENAE-NSDSATDESNINQPDKRSSTRIQKMHPKE 1035
Query: 1088 LIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSK 1147
LI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF +
Sbjct: 1036 LIIGDPNRGVTTRSR----EVEIVSNSCFVSKIEPKNVKEALTDEFWINAMQEELEQFKR 1091
Query: 1148 NDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIA 1207
N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP+A
Sbjct: 1092 NEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAPVA 1151
Query: 1208 RLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKS 1267
RLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D PDHV++LKK+
Sbjct: 1152 RLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADPTHPDHVYRLKKA 1211
Query: 1268 LYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQS 1327
LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 1212 LYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSNE 1271
Query: 1328 LCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAK 1387
+ + F + MQ+EFEMS++GELTYFLG+QV Q + ++ QS+Y K ++KKF M ++ +
Sbjct: 1272 MLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIVKKFGMENASHKR 1331
Query: 1388 TPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLT 1447
TP L K++A V Q LYR MIGSLLYLTASRPDI ++V +CAR+Q++P+ +HLT
Sbjct: 1332 TPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISHLT 1391
Query: 1448 AVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWE 1507
VKRIL+Y+ T++ G+MY S L GYCDAD+AG +RKSTSG C +LG+NL+SW
Sbjct: 1392 QVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADDRKSTSGGCFYLGNNLISWF 1451
Query: 1508 SKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPI 1567
SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKNP+
Sbjct: 1452 SKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINISKNPV 1511
Query: 1568 LHSRAKHIEVKYHFIRDYV 1586
HSR KHI++++H+IRD V
Sbjct: 1512 QHSRTKHIDIRHHYIRDLV 1530
>gb|AAC49502.1| Pol [Zea mays] gi|7489803|pir||T04112 pol protein homolog - maize
retrotransposon Opie-2
Length = 1068
Score = 954 bits (2465), Expect = 0.0
Identities = 500/1023 (48%), Positives = 657/1023 (63%), Gaps = 48/1023 (4%)
Query: 609 SWYLDSGCSRHMTGERRMFQE-LKLKPGGE-VGFGGNEKGKIVGTGTICVDSSPCIYNVL 666
SW +DSGC+ HMTGE++MF +K K + + FG +GK+ G G I + + I NV
Sbjct: 10 SWIIDSGCTNHMTGEKKMFTSYVKNKDSQDSIIFGDGNQGKVKGLGKIAISNEHSISNVF 69
Query: 667 LVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQ 726
LV+ L +NLLS+SQL + GY+ +F + DGS+ F +Y + ++ EA
Sbjct: 70 LVESLGYNLLSVSQLCNMGYNCLFTNIDVSVFRRCDGSLAFKGVLDGKLYLVDFAKEEAG 129
Query: 727 NVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTK 786
CL++ W+WHRRL H M+ + +L K V GL N++F D C ACQ GK
Sbjct: 130 LDACLIAKTSMGWLWHRRLAHVGMKNLHKLLKGEHVIGLTNVQFEKDRPCAACQAGKQVG 189
Query: 787 VPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESH 846
KNV++TSRPLE+LH+DLFGPV SIGG +YG++IVDD+SR+TWV FL K E+
Sbjct: 190 GSHHTKNVMTTSRPLEMLHMDLFGPVAYLSIGGSKYGLVIVDDFSRFTWVFFLQEKSETQ 249
Query: 847 AVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVE 906
F+ + QNE ++ ++RSD+G EF+N + E + GI H+FS P TPQQNGVVE
Sbjct: 250 GTLKRFLRRAQNEFELKVKKIRSDNGSEFKNLQVEEFLEEEGIKHEFSAPYTPQQNGVVE 309
Query: 907 RKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNIS 966
RKNRTL +MARTML E + FWTEAVNTAC+ NR+ + IL T YEL KPN+S
Sbjct: 310 RKNRTLIDMARTMLGEFKTPECFWTEAVNTACHAINRVYLHRILKNTSYELLTGNKPNVS 369
Query: 967 YFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDD 1026
YF FG CY+L K R KF K+ + LLGY +K +R +N + +E S V FD+
Sbjct: 370 YFRVFGSKCYILVKKGRNSKFAPKAVEGFLLGYDSNTKAYRVFNKSSGLVEVSGDVVFDE 429
Query: 1027 KLDSDQSKLV-------EKFADLSINVSDKGKAPEEAEPEEDEP---------------- 1063
S + ++V E +I G+ + + E ++P
Sbjct: 430 TNGSPREQVVDCDDVDEEDIPTAAIRTMAIGEVRPQEQDEREQPSPSTMVHPPTQDDEQV 489
Query: 1064 -----------------EEEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPS 1106
EEEA P+ T ++ I HP++ I+G+ + V TRS
Sbjct: 490 HQQEVCDQGGAQDDHVLEEEAQPA-PPTQVRAMIQRDHPVDQILGDISKGVTTRSRL--- 545
Query: 1107 EETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTK 1166
VS IEP ++EAL D DW+LAM+EELN F +N+VW+LV +P+ +V+GTK
Sbjct: 546 -VNFCEHNSFVSSIEPFRVEEALLDPDWVLAMQEELNNFKRNEVWTLVPRPKQ-NVVGTK 603
Query: 1167 WVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVL 1226
WVFRNK +E+G V RNKARLVA+GY+Q G+D+ ETFAP+ARLE+IR+L++++ +H+ L
Sbjct: 604 WVFRNKQDERGVVTRNKARLVAKGYAQVAGLDFEETFAPVARLESIRILLAYAAHHSFRL 663
Query: 1227 HQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFL 1286
+QMDVKSAFLNG I EEVYV QPPGFEDE+ PDHV KL K+LYGLKQAPRAWYE L FL
Sbjct: 664 YQMDVKSAFLNGPIKEEVYVEQPPGFEDERYPDHVCKLSKALYGLKQAPRAWYECLRDFL 723
Query: 1287 LENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMG 1346
+ N F GK D TLF KT D+ V QIYVDDIIFGS NQ C+EFS++M +FEMSMMG
Sbjct: 724 IANAFKVGKADPTLFTKTCDGDLFVCQIYVDDIIFGSTNQKSCEEFSRVMTQKFEMSMMG 783
Query: 1347 ELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVC 1406
EL YFLG QV Q +GT+I Q+KYT++LLK+F M ++ AKTPM + V
Sbjct: 784 ELNYFLGFQVKQLKDGTFISQTKYTQDLLKRFGMKDAKPAKTPMGTDGHTDLNKGGKSVD 843
Query: 1407 QKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMY 1466
QK YR MIGSLLYL ASRPDI+ SV +CARFQSDP+E HL AVKRILRYL T GL Y
Sbjct: 844 QKAYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKECHLVAVKRILRYLVATPCFGLWY 903
Query: 1467 KKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISA 1526
K S + L GY D+DYAG + +RKSTSG CQFLG +LVSW SK+Q+++ALSTAEAEY++A
Sbjct: 904 PKGSTFDLVGYSDSDYAGCKVDRKSTSGTCQFLGRSLVSWNSKKQTSVALSTAEAEYVAA 963
Query: 1527 AICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV 1586
C Q LWM+ L D+ S +P+ CDN +AI +++NP+ HSR KHI++++HF+RD+
Sbjct: 964 GQCCAQLLWMRQTLRDFGYNLSKVPLLCDNESAIRMAENPVEHSRTKHIDIRHHFLRDHQ 1023
Query: 1587 *KG 1589
KG
Sbjct: 1024 QKG 1026
>gb|AAO73523.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 953 bits (2464), Expect = 0.0
Identities = 473/979 (48%), Positives = 652/979 (66%), Gaps = 6/979 (0%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI+G G + D P + VLL
Sbjct: 560 EDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHDGLPSLNKVLL 619
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E +
Sbjct: 620 VKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQETSYSS 679
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ + VRG+PNLK +C CQ GK K+
Sbjct: 680 T-CLSSKEDEVRIWHQRFGHLHLRGMKKILDKSAVRGIPNLKIEEGRICGECQIGKQVKM 738
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TSR LELLH+DL GP++ ES+GGKRY ++VDD+SR+TWV F+ K +
Sbjct: 739 SHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSGTFE 798
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN +F S GI H+FS TPQQNG+VER
Sbjct: 799 VFKKLSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSAAITPQQNGIVER 858
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE AR ML + + W EA+NTACYI NR+++R T YE+WK KP++ +
Sbjct: 859 KNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPSVKH 918
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 919 FHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 978
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRITAAHPME 1087
+ + + E NV+D K+ E AE D +E+ + +RI HP E
Sbjct: 979 SPARKKDVEEDVRTSGDNVADAAKSGENAE-NSDSATDESNINQPDKRSSTRIQKMHPKE 1037
Query: 1088 LIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSK 1147
LI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF +
Sbjct: 1038 LIIGDPNRGVTTRSR----EVEIVSNSCFVSKIEPKNVKEALTDEFWINAMQEELEQFKR 1093
Query: 1148 NDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIA 1207
N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP+A
Sbjct: 1094 NEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAPVA 1153
Query: 1208 RLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKS 1267
RLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D PDHV++LKK+
Sbjct: 1154 RLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADPTHPDHVYRLKKA 1213
Query: 1268 LYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQS 1327
LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 1214 LYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSNE 1273
Query: 1328 LCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAK 1387
+ + F + MQ+EFEMS++GELTYFLG+QV Q + ++ QS+Y K ++KKF M ++ +
Sbjct: 1274 MLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIVKKFGMENASHKR 1333
Query: 1388 TPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLT 1447
TP L K++A V QK YR MIGSLLYLTASRPDI ++V +CAR+Q++P+ +HL
Sbjct: 1334 TPAPTHLKLSKDEAGTSVDQKPYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISHLN 1393
Query: 1448 AVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWE 1507
VKRIL+Y+ T++ G+MY S L GYCDAD+AG +RKSTSG C +LG+NL+SW
Sbjct: 1394 QVKRILKYVNGTSDYGIMYCHCSSSMLVGYCDADWAGSADDRKSTSGGCFYLGNNLISWF 1453
Query: 1508 SKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPI 1567
SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKNP+
Sbjct: 1454 SKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINISKNPV 1513
Query: 1568 LHSRAKHIEVKYHFIRDYV 1586
HSR KHI++++H+IRD V
Sbjct: 1514 QHSRTKHIDIRHHYIRDLV 1532
>gb|AAO73529.1| gag-pol polyprotein [Glycine max]
Length = 1577
Score = 949 bits (2454), Expect = 0.0
Identities = 474/981 (48%), Positives = 653/981 (66%), Gaps = 10/981 (1%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI G G + + P + VLL
Sbjct: 561 EDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHEGLPSLNKVLL 620
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E + +
Sbjct: 621 VKGLTVNLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQE-SSHS 679
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ VRG+PNLK +C CQ GK K+
Sbjct: 680 STCLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKM 739
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TSR LELLH+DL GP++ ES+GGKRY ++VDD+SR+TWV F+ K ++
Sbjct: 740 SHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSDTFE 799
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN KF S GI H+FS TPQQNG+VER
Sbjct: 800 VFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQQNGIVER 859
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE AR ML + + W EA+NTACYI NR+++R T YE+WK KP + +
Sbjct: 860 KNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPTVKH 919
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 920 FHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 979
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKS--RITAAHP 1085
+ + + E NV+D K+ E AE + +E P+ +Q K+ RI HP
Sbjct: 980 TPARKKDVEEDVRTSGDNVADTAKSAENAENSDSATDE---PNINQPDKRPSIRIQKMHP 1036
Query: 1086 MELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQF 1145
ELI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF
Sbjct: 1037 KELIIGDPNRGVTTRSR----EIEIVSNSCFVSKIEPKNVKEALTDEFWINAMQEELEQF 1092
Query: 1146 SKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAP 1205
+N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP
Sbjct: 1093 KRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAP 1152
Query: 1206 IARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLK 1265
+ARLE+IRLL+ + L+QMDVKSAFLNGY++EE YV QP GF D PDHV++LK
Sbjct: 1153 VARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFVDPTHPDHVYRLK 1212
Query: 1266 KSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSAN 1325
K+LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 1213 KALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMS 1272
Query: 1326 QSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTV 1385
+ + F + MQ+EFEMS++GELTYFLG+QV Q + ++ QSKY K ++KKF M ++
Sbjct: 1273 NEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSKYAKNIVKKFGMENASH 1332
Query: 1386 AKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETH 1445
+TP L K++A V Q LYR MIGSLLYLTASRPDI ++V +CAR+Q++P+ +H
Sbjct: 1333 KRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISH 1392
Query: 1446 LTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVS 1505
L VKRIL+Y+ T++ G+MY S L GYCDAD+AG +RKSTSG C +LG+NL+S
Sbjct: 1393 LNQVKRILKYVNGTSDYGIMYCHCSGSMLVGYCDADWAGSADDRKSTSGGCFYLGNNLIS 1452
Query: 1506 WESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKN 1565
W SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKN
Sbjct: 1453 WFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINISKN 1512
Query: 1566 PILHSRAKHIEVKYHFIRDYV 1586
P+ HSR KHI++++H+IR+ V
Sbjct: 1513 PVQHSRTKHIDIRHHYIRELV 1533
>gb|AAC64917.1| gag-pol polyprotein [Glycine max]
Length = 1550
Score = 938 bits (2425), Expect = 0.0
Identities = 472/981 (48%), Positives = 647/981 (65%), Gaps = 10/981 (1%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI G G + D P + VLL
Sbjct: 534 EDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHDGLPSLNKVLL 593
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E +
Sbjct: 594 VKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQETSYSS 653
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ VRG+PNLK +C CQ GK K+
Sbjct: 654 T-CLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKM 712
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TSR LELLH+DL GP++ ES+G KRY ++VDD+SR+TWV F+ K ++
Sbjct: 713 SNQKLQHQTTSRVLELLHMDLMGPMQVESLGRKRYAYVVVDDFSRFTWVNFIREKSDTFE 772
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN KF S GI H+FS TPQQNG+VER
Sbjct: 773 VFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQQNGIVER 832
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE AR ML + + W EA+NTACYI NR+++R T YE+WK KP + +
Sbjct: 833 KNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPTVKH 892
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH G CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 893 FHICGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 952
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKS--RITAAHP 1085
+ + + E NV+D K+ E AE + +E P+ +Q K+ RI HP
Sbjct: 953 TPARKKDVEEDVRTSGDNVADTAKSAENAENSDSATDE---PNINQPDKRPSIRIQKMHP 1009
Query: 1086 MELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQF 1145
ELI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF
Sbjct: 1010 KELIIGDPNRGVTTRSR----EIEIISNSCFVSKIEPKNVKEALTDEFWINAMQEELEQF 1065
Query: 1146 SKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAP 1205
+N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP
Sbjct: 1066 KRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAP 1125
Query: 1206 IARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLK 1265
ARLE+IRLL+ + L+QMDVKSAFLNGY++EE YV QP GF D PDHV++LK
Sbjct: 1126 GARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFVDPTHPDHVYRLK 1185
Query: 1266 KSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSAN 1325
K+LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 1186 KALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMS 1245
Query: 1326 QSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTV 1385
+ + F + MQ+EFEMS++GELTYFLG+QV Q + ++ QSKY K ++KKF M ++
Sbjct: 1246 NEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSKYAKNIVKKFGMENASH 1305
Query: 1386 AKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETH 1445
+TP L K++A V Q LYR MIGSLLYLTASRPDI ++V CAR+Q++P+ +H
Sbjct: 1306 KRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGGCARYQANPKISH 1365
Query: 1446 LTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVS 1505
L VKRIL+Y+ T++ G+MY S+ L GYCDAD+AG +RKST G C +LG+N +S
Sbjct: 1366 LNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCDADWAGSVDDRKSTFGGCFYLGTNFIS 1425
Query: 1506 WESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKN 1565
W SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKN
Sbjct: 1426 WFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNLSAINISKN 1485
Query: 1566 PILHSRAKHIEVKYHFIRDYV 1586
P+ HSR KHI++++H+IRD V
Sbjct: 1486 PVQHSRTKHIDIRHHYIRDLV 1506
>gb|AAO73525.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 937 bits (2421), Expect = 0.0
Identities = 470/981 (47%), Positives = 645/981 (64%), Gaps = 10/981 (1%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI G G + D P + VLL
Sbjct: 560 EDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHDGLPSLNKVLL 619
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E +
Sbjct: 620 VKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQETSYSS 679
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ VRG+PNLK +C CQ GK K+
Sbjct: 680 T-CLSSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKM 738
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TS LELLH+DL GP++ ES+GGKRY ++VDD+SR+TWV F+ K ++
Sbjct: 739 SHQKLQHQTTSMVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSDTFE 798
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN KF S GI H+FS TPQQNG+VER
Sbjct: 799 VFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQQNGIVER 858
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE R ML + + W EA+NTACYI NR+++R T YE+WK KP + +
Sbjct: 859 KNRTLQEATRVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPTVKH 918
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 919 FHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 978
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKS--RITAAHP 1085
+ + + E NV+D K+ E AE + +E P+ +Q K RI P
Sbjct: 979 TPARKKDVEEDVRTSEDNVADTAKSAENAEKSDSTTDE---PNINQPDKSPFIRIQKMQP 1035
Query: 1086 MELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQF 1145
ELI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF
Sbjct: 1036 KELIIGDPNRGVTTRSR----EIEIVSNSCFVSKIEPKNVKEALTDEFWINAMQEELEQF 1091
Query: 1146 SKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAP 1205
+N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP
Sbjct: 1092 KRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAP 1151
Query: 1206 IARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLK 1265
+ARLE+IRLL+ + L+QMDVKSAFLNGY++EE YV QP GF D DHV++LK
Sbjct: 1152 VARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFVDPTHLDHVYRLK 1211
Query: 1266 KSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSAN 1325
K+LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 1212 KALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMS 1271
Query: 1326 QSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTV 1385
+ + F MQ+EFEMS++GEL YFLG+QV Q + ++ QSKY K ++KKF M ++
Sbjct: 1272 NEMLRHFVPQMQSEFEMSLVGELHYFLGLQVKQMEDSIFLSQSKYAKNIVKKFGMENASH 1331
Query: 1386 AKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETH 1445
+TP L K++A V Q LYR MIGSLLYLTASRPDI F+V +CAR+Q++P+ +H
Sbjct: 1332 KRTPAPTHLKLSKDEAGTSVDQNLYRSMIGSLLYLTASRPDITFAVGVCARYQANPKISH 1391
Query: 1446 LTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVS 1505
L VKRIL+Y+ T++ G+MY S+ L GYCDAD+AG +RK TSG C +LG+NL+S
Sbjct: 1392 LNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCDADWAGSADDRKCTSGGCFYLGTNLIS 1451
Query: 1506 WESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKN 1565
W SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKN
Sbjct: 1452 WFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINISKN 1511
Query: 1566 PILHSRAKHIEVKYHFIRDYV 1586
P+ H+R KHI++++H+IRD V
Sbjct: 1512 PVQHNRTKHIDIRHHYIRDLV 1532
>gb|AAG52949.1| gag/pol polyprotein [Arabidopsis thaliana]
Length = 1643
Score = 919 bits (2375), Expect = 0.0
Identities = 464/997 (46%), Positives = 641/997 (63%), Gaps = 28/997 (2%)
Query: 610 WYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLLVD 669
WY DSG SRHMTG + V FGG KG+I G G + P + NV V+
Sbjct: 614 WYFDSGASRHMTGSQANLNNYSSVKESNVMFGGGAKGRIKGKGDLTETEKPHLTNVYFVE 673
Query: 670 GLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQNVK 729
GLT NL+S+SQL D+G V FN+ C A ++ + + L + NN Y ++
Sbjct: 674 GLTANLISVSQLCDEGLTVSFNKVKCWATNERNQNTLTGVRTGNNCYMWEEPKI------ 727
Query: 730 CLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPF 789
CL + E+ +WH+RLGH + R +S+L +VRG+P LK +C AC +GK +V
Sbjct: 728 CLRAEKEDPVLWHQRLGHMNARSMSKLVNKEMVRGVPELKHIEKIVCGACNQGKQIRVQH 787
Query: 790 KAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHAVF 849
K + T++ L+L+H+DL GP+++ESI GKRY ++VDD+SR+ WV+F+ K E+ F
Sbjct: 788 KRVEGIQTTQVLDLIHMDLMGPMQTESIAGKRYVFVLVDDFSRYAWVRFIREKSETANSF 847
Query: 850 STFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKN 909
Q++NEK I ++RSD GGEF N+ F S +S GI H +S PRTPQ NGVVERKN
Sbjct: 848 KILALQLKNEKKMGIKQIRSDRGGEFMNEAFNSFCESQGIFHQYSAPRTPQSNGVVERKN 907
Query: 910 RTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISYFH 969
RTLQEMAR M+ G+ + FW EA++TACY+ NR+ VR +KTPYE+WK KPN+SYF
Sbjct: 908 RTLQEMARAMIHGNGVPEKFWAEAISTACYVINRVYVRLGSDKTPYEIWKGKKPNLSYFR 967
Query: 970 PFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDKLD 1029
FGCVCY++N KD+L KFD++S + LGY+ S +R +N IEES++V FDD
Sbjct: 968 VFGCVCYIMNDKDQLGKFDSRSEEGFFLGYATNSLAYRVWNKQRGKIEESMNVVFDDGSM 1027
Query: 1030 SDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRITAAHPMELI 1089
+ +V + ++S+ ++ ++G + + +++ H + I
Sbjct: 1028 PELQIIVRNRNEPQTSISNNHGEERNDNQFDNGDINKSGEESDEEVPPAQVHRDHASKDI 1087
Query: 1090 MGNKD-------------------EPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQ 1130
+G+ + R ++F EE + S VS++EPK++ EAL+
Sbjct: 1088 IGDPSGERVTRGVKQDYRQLAGIKQKHRVMASFACFEEIMFSC--FVSIVEPKNVKEALE 1145
Query: 1131 DKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQG 1190
D WILAMEEEL +FS++ VW LV +P V+VIGTKW+F+NK +E G++ RNKARLVAQG
Sbjct: 1146 DHFWILAMEEELEEFSRHQVWDLVPRPPQVNVIGTKWIFKNKFDEVGNITRNKARLVAQG 1205
Query: 1191 YSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPP 1250
Y+Q EG+D+ ETFAP+ARLE IR L+ + LHQMDVK AFLNG I EEVYV QP
Sbjct: 1206 YTQVEGLDFDETFAPVARLECIRFLLGTACGMGFKLHQMDVKCAFLNGIIEEEVYVEQPK 1265
Query: 1251 GFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDIL 1310
GFE+ + P++V+KLKK+LYGLKQAPRAWYERL++FL+ + RG VD TLF K I+
Sbjct: 1266 GFENLEFPEYVYKLKKALYGLKQAPRAWYERLTTFLIVQGYTRGSVDKTLFVKNDVHGII 1325
Query: 1311 VVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKY 1370
++QIYVDDI+FG + L K F K M EF MSM+GEL YFLG+Q++QT EG I QS Y
Sbjct: 1326 IIQIYVDDIVFGGTSDKLVKTFVKTMTTEFRMSMVGELKYFLGLQINQTDEGITISQSTY 1385
Query: 1371 TKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFS 1430
+ L+K+F M S A TPM T L K++ KV +KLYRGMIGSLLYLTA+RPD+ S
Sbjct: 1386 AQNLVKRFGMCSSKPAPTPMSTTTKLFKDEKGVKVDEKLYRGMIGSLLYLTATRPDLCLS 1445
Query: 1431 VHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERK 1490
V LCAR+QS+P+ +HL AVKRI++Y+ T N GL Y + + L GYCDAD+ G+ +R+
Sbjct: 1446 VGLCARYQSNPKASHLLAVKRIIKYVSGTINYGLNYTRDTSLVLVGYCDADWGGNLDDRR 1505
Query: 1491 STSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDY-QILESN 1549
ST+G FLGSNL+SW SK+Q+ ++LS+ ++EYI+ C TQ LWM+ DY
Sbjct: 1506 STTGGVFFLGSNLISWHSKKQNCVSLSSTQSEYIALGSCCTQLLWMRQMGLDYGMTFPDP 1565
Query: 1550 IPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV 1586
+ + CDN +AI++SKNP+ HS KHI +++HF+R+ V
Sbjct: 1566 LLVKCDNESAIAISKNPVQHSVTKHIAIRHHFVRELV 1602
>gb|AAN60991.1| Putative Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)] gi|34902324|ref|NP_912508.1| Putative
Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)]
Length = 2011
Score = 919 bits (2374), Expect = 0.0
Identities = 486/1008 (48%), Positives = 647/1008 (63%), Gaps = 49/1008 (4%)
Query: 610 WYLDSGCSRHMTGERRMFQELKLKPGG----EVGFGGNEKGKIVGTGTICVDSSPCIYNV 665
W LDSGC++HMTG+R MF ++ GG +V FG N KGK++G G I + + I NV
Sbjct: 558 WVLDSGCTQHMTGDRAMFTTFEV--GGNEQEKVTFGDNSKGKVIGLGKIAISNDLSIDNV 615
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEA 725
LV L NLLS++Q+ D F + S +D S +F R N+Y + EA
Sbjct: 616 SLVKSLNFNLLSVAQICDLVLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYLGDFNSSEA 675
Query: 726 QNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFT 785
CL++ W+WHRRL H M ++S+LSK +LV GL ++KF D LC ACQ GK
Sbjct: 676 NLKTCLVAKTSLGWLWHRRLAHVGMNQLSKLSKRDLVVGLKDVKFEKDKLCSACQAGKQV 735
Query: 786 KVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDES 845
K+++STSRPLELLH++ FGP +SIGG + ++IVDDYSR+TW+ FL K
Sbjct: 736 ACSHPTKSIMSTSRPLELLHMEFFGPTTYKSIGGNSHCLVIVDDYSRYTWMFFLHDKSIV 795
Query: 846 HAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVV 905
+F F + QNE C +V++RSD+G +F+N E D GI H+ S +PQQNGVV
Sbjct: 796 AELFKKFAKRGQNEFNCTLVKIRSDNGSKFKNTNIEDYCDDLGIKHELSATYSPQQNGVV 855
Query: 906 ERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNI 965
E KNRTL EMARTML E G++ FW EA+NTAC+ NR+ + +L KT YEL KPN+
Sbjct: 856 EMKNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRLLKKTSYELIVGRKPNV 915
Query: 966 SYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFD 1025
+YF FGC CY+ RL KF+++ + LLGY+ SK +R YN + +EE+ V+FD
Sbjct: 916 AYFRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYRVYNKNKGIVEETADVQFD 975
Query: 1026 DKLDSDQS-KLVEKFADLSI-----NVSDKGKAPEEAEPE-----EDEPEEEAGPSDSQT 1074
+ S + + ++ D + N+S P E E + +DEP A PS +Q
Sbjct: 976 ETNGSQEGHENLDDVGDEGLMRAMKNMSIGDVKPIEVEDKPSTSTQDEPSTSATPSQAQV 1035
Query: 1075 LKK--------------SRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLI 1120
+ + ++ HP++ ++G+ + V+TRS ++ VS +
Sbjct: 1036 EVEEEKAQDLPMPPRIHTALSKDHPIDQVLGDISKGVQTRSRV----ASICEHYSFVSCL 1091
Query: 1121 EPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVV 1180
EPK +DEAL D DWI AM +ELN F++N VW+LV++ +VIGTKWVFRNK +E G VV
Sbjct: 1092 EPKHVDEALCDPDWINAMHKELNNFARNKVWTLVERLRDHNVIGTKWVFRNKQDENGLVV 1151
Query: 1181 RNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYI 1240
RNKAR VAQG++Q EG+D+ ETFAP+ RLEAI +L++F+ NI L QMDVKSAFLNG I
Sbjct: 1152 RNKARFVAQGFTQVEGLDFGETFAPVTRLEAICILLAFASCFNIKLFQMDVKSAFLNGEI 1211
Query: 1241 SEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTL 1300
+E V+V QPPGFED K P+HV+KL K+LYGLKQAPRAWYERL FLL +F GKVDTTL
Sbjct: 1212 AELVFVEQPPGFEDPKYPNHVYKLSKALYGLKQAPRAWYERLRDFLLSKDFKIGKVDTTL 1271
Query: 1301 FCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTP 1360
F K DD V QIYVDDIIFG N+ CKEF MM EFEMSM+GEL++F G+Q+ Q
Sbjct: 1272 FTKIIGDDFFVCQIYVDDIIFGCTNEVFCKEFGDMMSREFEMSMIGELSFFHGLQIKQLK 1331
Query: 1361 EGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYL 1420
+GT F + ++ KTPM L+ ++ V KLYR MIGSLLYL
Sbjct: 1332 DGT--------------FGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYL 1377
Query: 1421 TASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDA 1480
TASRPDI+FSV +CARFQ+ P+E HL AVKRILRYLK ++ +GL Y K +++KL GY D+
Sbjct: 1378 TASRPDIMFSVCMCARFQAAPKECHLVAVKRILRYLKHSSTIGLWYPKGAKFKLVGYSDS 1437
Query: 1481 DYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQL 1540
DYAG + +RKSTSG+CQ LG +LVSW SK+Q+ +AL AEAEY+SA C Q LWMK L
Sbjct: 1438 DYAGCKVDRKSTSGSCQMLGRSLVSWSSKKQNFVALFIAEAEYVSAGSCCAQLLWMKQIL 1497
Query: 1541 EDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
DY I + P+ C+N +AI ++ NP+ HSR KHI++++HF+RD+V K
Sbjct: 1498 LDYGISFTKTPLLCENDSAIKIANNPVQHSRTKHIDIRHHFLRDHVAK 1545
>gb|AAV31277.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1577
Score = 916 bits (2368), Expect = 0.0
Identities = 481/1006 (47%), Positives = 643/1006 (63%), Gaps = 47/1006 (4%)
Query: 610 WYLDSGCSRHMTGERRMFQ--ELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
W LDSGC++HMTG+R MF E+ +V FG N KGK++G G I + + I NV L
Sbjct: 549 WVLDSGCTQHMTGDRAMFTTFEVGRNEQEKVTFGDNSKGKVIGLGKIAISNDLSIDNVSL 608
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V L NLLS++Q+ D F + S +D S +F R N+Y + + EA
Sbjct: 609 VKSLNFNLLSVAQICDLSLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYLVDFNSSEANL 668
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL++ W+WHRRL H M ++S+ SK +LV GL ++KF D LC ACQ GK
Sbjct: 669 KTCLVAKTSLGWLWHRRLAHVGMNQLSKFSKRDLVMGLKDVKFEKDKLCSACQAGKQVAC 728
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
K+++STS+PLELLH+DLF P +SIGG + ++IVDDYSR+TWV FL K
Sbjct: 729 SHPTKSIMSTSKPLELLHMDLFDPTTYKSIGGNSHCLVIVDDYSRYTWVFFLHDKSIVAD 788
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
+F F + QNE +C +V++RS+ G EF+N E D GI H+ +PQQNGVVER
Sbjct: 789 LFKKFAKRAQNEFSCTLVKIRSNIGSEFKNTNIEDYCDDLGIKHELFATYSPQQNGVVER 848
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTL EMARTML E G++ FW EA+NTAC+ NR+ + +L KT YE+ KPNI+Y
Sbjct: 849 KNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRVLKKTSYEVIVGRKPNIAY 908
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
F FGC CY+ RL KF+++ + LLGY+ +SK +R YN + +EE+ V+FD+
Sbjct: 909 FRVFGCKCYIHRKGVRLTKFESRCDEGFLLGYASKSKAYRVYNKNKGIVEETADVQFDET 968
Query: 1028 LDSDQS-KLVEKFADLSI-----NVSDKGKAPEEAEPE-----EDEPEEEAGPSDSQTLK 1076
S + + ++ D + N+S P E E + +DEP A PS +Q
Sbjct: 969 NGSQEGHENLDDVGDEGLMRVMKNMSIGDVKPIEVEDKPSTSTQDEPSTSAMPSQAQVEV 1028
Query: 1077 KSR--------------ITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEP 1122
+ ++ HP++ ++G+ + V+TRS ++ VS +E
Sbjct: 1029 EEEKAQEPPMPPRIHTALSKDHPIDQVLGDISKGVQTRSRVT----SICEHYSFVSCLER 1084
Query: 1123 KSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRN 1182
K +DEAL D DW+ AM EEL F++N VW+LV++P +VIGTKWVFRNK +E G VVRN
Sbjct: 1085 KHVDEALCDPDWMNAMHEELKNFARNKVWTLVERPRDHNVIGTKWVFRNKQDENGLVVRN 1144
Query: 1183 KARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISE 1242
KARLVAQG++Q EG+D+ ETFAP+ARLEAI +L++F+ +I L QMDVKSAFLN
Sbjct: 1145 KARLVAQGFTQVEGLDFGETFAPVARLEAICILLAFASCFDIKLFQMDVKSAFLN----- 1199
Query: 1243 EVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFC 1302
D K P+HV+KL K+LYGL+QAPRAWYERL FLL +F GKVD TLF
Sbjct: 1200 -----------DTKYPNHVYKLSKALYGLRQAPRAWYERLRDFLLSKDFKIGKVDITLFT 1248
Query: 1303 KTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEG 1362
K DD V QIYVDDIIFGS N+ CKEF MM EFEMSM+GEL++FLG+Q+ Q G
Sbjct: 1249 KIIGDDFFVYQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIGELSFFLGLQIKQLKNG 1308
Query: 1363 TYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTA 1422
T++ Q+KY K+LLK+F + ++ KTPM L+ ++ V KLYR MIGSLLYLT
Sbjct: 1309 TFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYLTV 1368
Query: 1423 SRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADY 1482
SRPDI+FSV +CARFQ+ P+E HL AVKRILRYLK ++ +GL Y K +++KL GY D DY
Sbjct: 1369 SRPDIMFSVCMCARFQAAPKECHLVAVKRILRYLKHSSTIGLWYPKGAKFKLVGYSDPDY 1428
Query: 1483 AGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLED 1542
AG + +RKSTS +CQ LG +LVSW SK+Q+++ALSTAE EY+SA C Q LWMK L D
Sbjct: 1429 AGCKVDRKSTSSSCQMLGRSLVSWSSKKQNSVALSTAETEYVSAGSCCAQLLWMKQTLLD 1488
Query: 1543 YQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
Y I + P+ CDN AI ++ NP+ HSR KHI++++HF+RD+V K
Sbjct: 1489 YGISFTKTPLLCDNDGAIKIANNPVQHSRTKHIDIRHHFLRDHVAK 1534
>gb|AAL35396.1| Opie2a pol [Zea mays]
Length = 1048
Score = 915 bits (2366), Expect = 0.0
Identities = 482/1013 (47%), Positives = 640/1013 (62%), Gaps = 46/1013 (4%)
Query: 620 MTGERRMFQE-LKLKPGGE-VGFGGNEKGKIVGTGTICVDSSPCIYNVLLVDGLTHNLLS 677
MTGE++MF +K K + + FG +GK+ G G I + S I NV LV+ L +N LS
Sbjct: 1 MTGEKKMFTSYVKNKDSQDSIIFGDGNQGKVKGLGKIAISSEHSISNVFLVESLGYNFLS 60
Query: 678 ISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQNVKCLLSVNEE 737
+SQL + GY+ +F + DGS+ F +Y + ++ EA CL++
Sbjct: 61 VSQLCNMGYNCLFTNVDVSVFRRSDGSLAFKGVLDGKLYLVDFAKEEAGLDACLIAKTSM 120
Query: 738 QWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVST 797
W+WHRRL H M+ + +L K V GL N++F D C ACQ GK + KNV++T
Sbjct: 121 GWLWHRRLAHVGMKNLHKLLKGEHVIGLTNVQFKKDRPCVACQAGKQVRGSHHTKNVMTT 180
Query: 798 SRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHAVFSTFIAQVQ 857
SRPLE+LH+DLFGPV SIGG +YG++IVDD+SR+TWV FL K E+ ++ + Q
Sbjct: 181 SRPLEMLHMDLFGPVAYLSIGGSKYGLVIVDDFSRFTWVFFLQDKSETQGTLKRYLRRAQ 240
Query: 858 NEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEMAR 917
NE ++ ++RSD+G EF+N + E GI H+FS P TPQQNGVVERKNRTL +MAR
Sbjct: 241 NEFELKVKKIRSDNGSEFKNLQVEEFLVEEGIKHEFSAPYTPQQNGVVERKNRTLMDMAR 300
Query: 918 TMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISYFHPFGCVCYV 977
TML E + FW+EAVNTAC+ NR+ + +L T YEL KPN+SYF FG CY+
Sbjct: 301 TMLGEFKTPERFWSEAVNTACHSINRVYLHRLLKNTSYELLTGNKPNVSYFRVFGSKCYI 360
Query: 978 LNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDKLDSDQSKLV- 1036
L K R KF K+ + LLGY +K +R +N + +E S V FD+ S + ++V
Sbjct: 361 LVKKGRTSKFAPKAVEGFLLGYDSNTKAYRVFNKSSGLVEVSSDVVFDETNGSLREQVVN 420
Query: 1037 ------EKFADLSINVSDKGKAPEEAEPEEDEP--------------------------- 1063
E ++ G + E+D+P
Sbjct: 421 LDDVDEEDVPTAAMRTMAIGDVRPQEHLEQDQPSSTTMVHPPTQDDEQAPQVGAHDQGGA 480
Query: 1064 -----EEEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVS 1118
EEE P T ++ I HP+ I+G+ + V TRS VS
Sbjct: 481 QDVQVEEEEAPQAPPTQVRATIQRNHPVNQILGDISKGVTTRSRL----VNFCEHYSFVS 536
Query: 1119 LIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGD 1178
IEP ++E L D DW+LAM+EELN F +N+VW+LV +P+ +V+GTKWVFRNK +E G
Sbjct: 537 SIEPFRVEEVLLDPDWVLAMQEELNNFKRNEVWTLVPRPKQ-NVVGTKWVFRNKQDEHGV 595
Query: 1179 VVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNG 1238
V RNKARLVA+GY+Q G+D+ ETFAP+ARL++IR+ ++++ +H+ L+QMDVKSAFLNG
Sbjct: 596 VTRNKARLVAKGYAQVAGLDFEETFAPVARLKSIRIWLAYAAHHSFRLYQMDVKSAFLNG 655
Query: 1239 YISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDT 1298
I EEVYV QPPGFEDE+ PDHV KL K+LYGLKQAPRAWYE L FL+ N F GK D
Sbjct: 656 PIKEEVYVEQPPGFEDERFPDHVCKLSKALYGLKQAPRAWYECLRDFLIANAFKVGKADP 715
Query: 1299 TLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQ 1358
TLF KT D+ V QIYVDDIIFGS NQ+ C+EFS++M +FEMSMMGEL+YFLG QV Q
Sbjct: 716 TLFTKTCDGDLFVCQIYVDDIIFGSTNQNSCEEFSRVMTQKFEMSMMGELSYFLGFQVRQ 775
Query: 1359 TPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLL 1418
+GT+I Q+KYT++L+K+F M ++ AKTPM + V QK YR IGSLL
Sbjct: 776 LKDGTFISQTKYTQDLIKRFGMKDAKPAKTPMGTDGHTDLNKGGKSVDQKAYRSTIGSLL 835
Query: 1419 YLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYC 1478
YL ASRPDI+ SV +CARFQSDPRE HL AVKRILRYL T G+ Y K S + L GY
Sbjct: 836 YLCASRPDIMLSVCMCARFQSDPRECHLVAVKRILRYLVATPCFGIWYPKGSTFDLIGYS 895
Query: 1479 DADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKH 1538
D+DYA + +RKSTS CQFLG +LVSW SK+Q+++ALSTAEAEY++ C Q LWM+
Sbjct: 896 DSDYARCKVDRKSTSRMCQFLGRSLVSWNSKKQTSVALSTAEAEYVAVGQCCAQLLWMRQ 955
Query: 1539 QLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
L D+ S +P+ CDN +AI +++NP+ HSR KHI++++HF+RD+ KGV+
Sbjct: 956 TLRDFGYNLSEVPLLCDNESAIRIAENPVEHSRTKHIDIRHHFLRDHQQKGVI 1008
>ref|NP_910082.1| putative gag-pol polyprotein [Oryza sativa (japonica cultivar-group)]
gi|28269414|gb|AAO37957.1| putative gag-pol polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 1969
Score = 908 bits (2346), Expect = 0.0
Identities = 478/1015 (47%), Positives = 640/1015 (62%), Gaps = 38/1015 (3%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGE-VGFGGNEKGKIVGTGTICVDSSPCIYN 664
+ SW +DSGCSRHMTGE + F L G E + FG G+++ GTI V+ + +
Sbjct: 750 RSNSWLVDSGCSRHMTGEAKWFTSLTRASGDETITFGDASSGRVMAKGTIKVNDKFMLKD 809
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELE 724
V LV L +NLLS+SQL D+ +V F + R + + V F+ R ++
Sbjct: 810 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 868
Query: 725 AQNVKCLL-SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+CL+ S N + + WHRRLGH +S++S ++L+RGLP LK D +C C+ GK
Sbjct: 869 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVQKDLVCAPCRHGK 928
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKD 843
T K +V T P +LLH+D GP + +S+GGK Y +++VDD+SR++WV FL K+
Sbjct: 929 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 988
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F + + E + +RSD+G EF+N FES DS G+ H FS P PQQNG
Sbjct: 989 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1048
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERKNRTL EMARTML E + FWTEA++ AC+I NR+ +R IL+KTPYEL +P
Sbjct: 1049 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1108
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVR 1023
+S+ FGC C+VL + + L KF+++S + LGY+ S+ +R Y I E+ V
Sbjct: 1109 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1167
Query: 1024 FDDKL---------------------DSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDE 1062
FD+ D D + D + V + G +P P D
Sbjct: 1168 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETG-SPSTTSPSGDA 1226
Query: 1063 PE------EEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGL 1116
P EE S I HP + ++G E V ++ L
Sbjct: 1227 PTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYE------LVNSAF 1280
Query: 1117 VSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEK 1176
V+ EPK++ AL D++W+ AM EEL F +N VWSLV+ P +VIGTKWVF+NKL E
Sbjct: 1281 VASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGED 1340
Query: 1177 GDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1236
G +VRNKARLVAQG++Q EG+D+ ETFAP+ARLEAIR+L++F+ + L QMDVKSAFL
Sbjct: 1341 GSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFL 1400
Query: 1237 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1296
NG I EEVYV QPPGFE+ K P+HVFKL+K+LYGLKQAPRAWYERL +FLL+N F G V
Sbjct: 1401 NGVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAV 1460
Query: 1297 DTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQV 1356
D TLF D L+VQIYVDDIIFG ++ +L +FS +M EFEMSMMGELT+FLG+Q+
Sbjct: 1461 DKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQI 1520
Query: 1357 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGS 1416
QT EG ++HQ+KY+KELLKKF+M + TPM T L ++ +V Q+ YR MIGS
Sbjct: 1521 KQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGS 1580
Query: 1417 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSG 1476
LLYLTASRPDI FSV LCARFQ+ PR +H AVKRI RY+K T G+ Y +S +
Sbjct: 1581 LLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRIFRYIKSTLEYGIWYSCSSALSVRA 1640
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWM 1536
+ DAD+AG + +RKSTSG C FLG++LVSW S++QS++A STAEAEY++AA +Q LWM
Sbjct: 1641 FSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWM 1700
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
L+DY + S +P+ CDNT+AI+++KNP+ HSR KHIE++YHF+RD V KG +
Sbjct: 1701 ISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTI 1755
>gb|AAN60494.1| Putative Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)] gi|34902378|ref|NP_912535.1| Putative
Zea mays retrotransposon Opie-2 [Oryza sativa (japonica
cultivar-group)]
Length = 2145
Score = 904 bits (2337), Expect = 0.0
Identities = 476/1015 (46%), Positives = 639/1015 (62%), Gaps = 38/1015 (3%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGE-VGFGGNEKGKIVGTGTICVDSSPCIYN 664
+ SW +DSGCSRHMTGE + F L E + FG G+++ GTI V+ + +
Sbjct: 750 RSNSWLVDSGCSRHMTGEAKWFTSLTRASSDETITFGDASSGRVMAKGTIKVNDKFMLKD 809
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELE 724
V LV L +NLLS+SQL D+ +V F + R + + V F+ R ++
Sbjct: 810 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 868
Query: 725 AQNVKCLL-SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+CL+ S N + + WHRRLGH +S++S ++L+RGLP LK D +C C+ GK
Sbjct: 869 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPCRHGK 928
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKD 843
T K +V T P +LLH+D GP + +S+GGK Y +++VDD+SR++WV FL K+
Sbjct: 929 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 988
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F + + E + +RSD+G EF+N FES DS G+ H FS P PQQNG
Sbjct: 989 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1048
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERKNRTL EMARTML E + FWTEA++ AC+I NR+ +R IL+KTPYEL +P
Sbjct: 1049 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1108
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVR 1023
+S+ FGC C+VL + + L KF+++S + LGY+ S+ +R Y I E+ V
Sbjct: 1109 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1167
Query: 1024 FDDKL---------------------DSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDE 1062
FD+ D D + D + V + G +P P D
Sbjct: 1168 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETG-SPSTTSPSGDA 1226
Query: 1063 PE------EEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGL 1116
P EE S I HP + ++G E V ++ L
Sbjct: 1227 PTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYE------LVNSAF 1280
Query: 1117 VSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEK 1176
V+ EPK++ AL D++W+ AM EEL F +N VWSLV+ P +VIGTKWVF+NKL E
Sbjct: 1281 VASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGED 1340
Query: 1177 GDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1236
G +VRNKARLVAQG++Q EG+D+ ETFAP+ARLEAIR+L++F+ + L QMDVKSAFL
Sbjct: 1341 GSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFL 1400
Query: 1237 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1296
NG I EEVYV QPPGFE+ K P+HVFKL+K+LYGLKQAPRAWYERL +FLL+N F G V
Sbjct: 1401 NGVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAV 1460
Query: 1297 DTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQV 1356
D TLF D L+VQIYVDDIIFG ++ +L +FS +M EFEMSMMGELT+FLG+Q+
Sbjct: 1461 DKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQI 1520
Query: 1357 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGS 1416
QT EG ++HQ+KY+KELLKKF+M + TPM T L ++ +V Q+ YR MIGS
Sbjct: 1521 KQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGS 1580
Query: 1417 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSG 1476
LLYLTASRPDI FSV LCARFQ+ PR +H AVKR+ RY+K T G+ Y +S +
Sbjct: 1581 LLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRA 1640
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWM 1536
+ DAD+AG + +RKSTSG C FLG++LVSW S++QS++A STAEAEY++AA +Q LWM
Sbjct: 1641 FSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWM 1700
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
L+DY + S +P+ CDNT+AI+++KNP+ HSR KHIE++YHF+RD V KG +
Sbjct: 1701 ISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTI 1755
>ref|XP_474304.1| OSJNBb0004A17.2 [Oryza sativa (japonica cultivar-group)]
gi|32488723|emb|CAE03600.1| OSJNBb0004A17.2 [Oryza sativa
(japonica cultivar-group)]
Length = 1877
Score = 904 bits (2337), Expect = 0.0
Identities = 476/1015 (46%), Positives = 639/1015 (62%), Gaps = 38/1015 (3%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGE-VGFGGNEKGKIVGTGTICVDSSPCIYN 664
+ SW +DSGCSRHMTGE + F L E + FG G+++ GTI V+ + +
Sbjct: 832 RSNSWLVDSGCSRHMTGEAKWFTSLTRASSDETITFGDASSGRVMAKGTIKVNDKFMLKD 891
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELE 724
V LV L +NLLS+SQL D+ +V F + R + + V F+ R ++
Sbjct: 892 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 950
Query: 725 AQNVKCLL-SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+CL+ S N + + WHRRLGH +S++S ++L+RGLP LK D +C C+ GK
Sbjct: 951 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPCRHGK 1010
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKD 843
T K +V T P +LLH+D GP + +S+GGK Y +++VDD+SR++WV FL K+
Sbjct: 1011 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 1070
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F + + E + +RSD+G EF+N FES DS G+ H FS P PQQNG
Sbjct: 1071 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1130
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERKNRTL EMARTML E + FWTEA++ AC+I NR+ +R IL+KTPYEL +P
Sbjct: 1131 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1190
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVR 1023
+S+ FGC C+VL + + L KF+++S + LGY+ S+ +R Y I E+ V
Sbjct: 1191 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1249
Query: 1024 FDDKL---------------------DSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDE 1062
FD+ D D + D + V + G +P P D
Sbjct: 1250 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETG-SPSTTSPSGDA 1308
Query: 1063 PE------EEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGL 1116
P EE S I HP + ++G E V ++ L
Sbjct: 1309 PTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYE------LVNSAF 1362
Query: 1117 VSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEK 1176
V+ EPK++ AL D++W+ AM EEL F +N VWSLV+ P +VIGTKWVF+NKL E
Sbjct: 1363 VASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGED 1422
Query: 1177 GDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1236
G +VRNKARLVAQG++Q EG+D+ ETFAP+ARLEAIR+L++F+ + L QMDVKSAFL
Sbjct: 1423 GSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFL 1482
Query: 1237 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1296
NG I EEVYV QPPGFE+ K P+HVFKL+K+LYGLKQAPRAWYERL +FLL+N F G V
Sbjct: 1483 NGVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAV 1542
Query: 1297 DTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQV 1356
D TLF D L+VQIYVDDIIFG ++ +L +FS +M EFEMSMMGELT+FLG+Q+
Sbjct: 1543 DKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQI 1602
Query: 1357 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGS 1416
QT EG ++HQ+KY+KELLKKF+M + TPM T L ++ +V Q+ YR MIGS
Sbjct: 1603 KQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGS 1662
Query: 1417 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSG 1476
LLYLTASRPDI FSV LCARFQ+ PR +H AVKR+ RY+K T G+ Y +S +
Sbjct: 1663 LLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRA 1722
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWM 1536
+ DAD+AG + +RKSTSG C FLG++LVSW S++QS++A STAEAEY++AA +Q LWM
Sbjct: 1723 FSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWM 1782
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
L+DY + S +P+ CDNT+AI+++KNP+ HSR KHIE++YHF+RD V KG +
Sbjct: 1783 ISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTI 1837
>ref|NP_909107.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 1410
Score = 904 bits (2335), Expect = 0.0
Identities = 477/1015 (46%), Positives = 639/1015 (61%), Gaps = 38/1015 (3%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGE-VGFGGNEKGKIVGTGTICVDSSPCIYN 664
+ SW +DSGCSRHMTGE + F L G E + FG G+++ GTI V+ + +
Sbjct: 365 RSNSWLVDSGCSRHMTGEAKWFTSLTRASGDETITFGDASSGRVMAKGTIKVNDKFMLKD 424
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELE 724
V LV L +NLLS+SQL D+ +V F + R + + V F+ R ++
Sbjct: 425 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 483
Query: 725 AQNVKCLL-SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+CL+ S N + + WHRRLGH +S++S ++L+RGLP LK D +C C+ GK
Sbjct: 484 PGPSRCLVASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKAPKDLVCAPCRHGK 543
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKD 843
T K +V T P +LLH++ GP + +S+GGK Y +++VDD+SR++WV FL K+
Sbjct: 544 MTSSSHKPVTMVMTDGPGQLLHMNTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 603
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F + + E + +RSD+G EF+N FES DS G+ H FS P PQQNG
Sbjct: 604 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 663
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERKNRTL EMARTML E + FWTEA++ AC+I NR+ +R IL+KTPYEL +P
Sbjct: 664 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 723
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVR 1023
+S+ FGC C+VL + + L KF+++S + LGY+ S+ +R Y I E+ V
Sbjct: 724 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 782
Query: 1024 FDDKL---------------------DSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDE 1062
FD+ D D + D + V + G +P P D
Sbjct: 783 FDEASPGARPEISGVLDESIFVDEDSDDDDDDSIPPPLDSTPPVQETG-SPSTTSPSGDA 841
Query: 1063 PE------EEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGL 1116
P EE S I HP + ++G E V ++ L
Sbjct: 842 PTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYD------LVNSAF 895
Query: 1117 VSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEK 1176
V+ EPK++ AL D++W+ AM EEL F +N VWSLV+ P +VIGTKWVF+NKL E
Sbjct: 896 VASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGED 955
Query: 1177 GDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1236
G +VRNKARLVAQG++Q EG+D+ ETFAP+ARLEAIR+L++F+ + L QMDVKSAFL
Sbjct: 956 GSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFL 1015
Query: 1237 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1296
NG I EEVYV QPPGFE+ K P+HVFKL K+LYGLKQAPRAWYERL +FLL+N F G V
Sbjct: 1016 NGVIEEEVYVKQPPGFENPKFPNHVFKLDKALYGLKQAPRAWYERLKTFLLQNGFEMGAV 1075
Query: 1297 DTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQV 1356
D TLF D L+VQIYVDDIIFG ++ +L +FS +M EFEMSMMGELT+FLG+Q+
Sbjct: 1076 DKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQI 1135
Query: 1357 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGS 1416
QT EG ++HQ+KY+KELLKKF+M + TPM T L ++ +V Q+ YR MIGS
Sbjct: 1136 KQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGS 1195
Query: 1417 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSG 1476
LLYLTASRPDI FSV LCARFQ+ PR +H AVKRI RY+K T G+ Y +S +
Sbjct: 1196 LLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRIFRYIKSTLEYGIWYSCSSALSVRA 1255
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWM 1536
+ DAD+AG + +RKSTSG C FLG++LVSW S++QS++A STAEAEY++AA +Q LWM
Sbjct: 1256 FSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWM 1315
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
L+DY + S +P+ CDNT+AI+++KNP+ HSR KHIE++YHF+RD V KG +
Sbjct: 1316 ISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTI 1370
>emb|CAE03782.2| OSJNBa0063G07.6 [Oryza sativa (japonica cultivar-group)]
gi|50923031|ref|XP_471876.1| OSJNBa0063G07.6 [Oryza
sativa (japonica cultivar-group)]
Length = 1539
Score = 900 bits (2327), Expect = 0.0
Identities = 478/1008 (47%), Positives = 641/1008 (63%), Gaps = 64/1008 (6%)
Query: 610 WYLDSGCSRHMTGERRMFQELKLKPGG----EVGFGGNEKGKIVGTGTICVDSSPCIYNV 665
W LDSGC++HMTG+R MF ++ GG +V F N K K++G G I + + I NV
Sbjct: 524 WVLDSGCTQHMTGDRAMFTTFEV--GGNEQEKVTFVDNSKRKVIGLGKIAISNDLSIDNV 581
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEA 725
V L NLLS++Q+ D G F + S +D S +F R N+Y + + EA
Sbjct: 582 SFVKSLNFNLLSVAQICDLGLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYFVDFNSSEA 641
Query: 726 QNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFT 785
CL++ W+WHRRL H M ++S+LSK +LV GL ++KF D LC ACQ GK
Sbjct: 642 NLKTCLVAKTSLGWLWHRRLAHVGMNQLSKLSKRDLVVGLKDVKFEKDKLCSACQAGKQV 701
Query: 786 KVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDES 845
K+++STSRPLELLH+DLFGP +SIGG + ++IVDDYSR+TWV FL K
Sbjct: 702 ACSHPTKSIMSTSRPLELLHMDLFGPTTYKSIGGNSHCLVIVDDYSRYTWVFFLHDKSIV 761
Query: 846 HAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVV 905
+F + QNE +C +V++RSD+G EF+N E D GI H+ S +PQQNGVV
Sbjct: 762 AELFKKIAKRAQNEFSCTLVKIRSDNGSEFKNTNIEDYCDDLGIKHELSATYSPQQNGVV 821
Query: 906 ERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNI 965
ERKNRTL EMARTML E G++ FW EA+NTAC+ NR + +L T YEL KPN+
Sbjct: 822 ERKNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRFYLHRLLKNTSYELIVGRKPNV 881
Query: 966 SYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFD 1025
+YF FGC CY+ RL KF+++ + LLGY+ SK +R YN + T+EE+ V+FD
Sbjct: 882 AYFRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYRVYNKNKGTVEETADVQFD 941
Query: 1026 DKLDSDQS-KLVEKFADLSI-----NVSDKGKAPEEAEPE-----EDEPEEEAGPSDSQT 1074
+ S + + ++ D + N+S P E E + +DEP A PS +Q
Sbjct: 942 ETNGSQEGHENLDDVGDEGLIRAMKNMSFGDVKPIEVEDKPSTSTQDEPSTFATPSQAQV 1001
Query: 1075 LKK--------------SRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLI 1120
+ + ++ HP++ ++G+ + V+T S ++ VS +
Sbjct: 1002 EVEEEKAQDPPIPPRIHTTLSKDHPIDQVLGDISKGVQTLSRV----ASICEHYSFVSCL 1057
Query: 1121 EPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVV 1180
EPK +DEAL D DW+ AM EELN F++N VW+LV++P +VIGTKWVFRNK +E G VV
Sbjct: 1058 EPKHVDEALCDPDWMNAMHEELNNFARNKVWTLVERPRDHNVIGTKWVFRNKQDENGLVV 1117
Query: 1181 RNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYI 1240
RNKARLVAQG++Q EG+D+ ETFAP+ARLEAI +L++F+ +I L QMDVKSAFLNG I
Sbjct: 1118 RNKARLVAQGFTQVEGLDFGETFAPVARLEAICILLAFASWFDIKLFQMDVKSAFLNGEI 1177
Query: 1241 SEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTL 1300
+E V+V QPPGFED K P+H FK+ GKVDTTL
Sbjct: 1178 AELVFVEQPPGFEDPKYPNHDFKI-----------------------------GKVDTTL 1208
Query: 1301 FCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTP 1360
F K DD V QIYVDDIIFGS N+ CKEF MM EFEMSM+GEL++FLG+Q+ Q
Sbjct: 1209 FTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIGELSFFLGLQIKQLK 1268
Query: 1361 EGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYL 1420
+GT++ Q+KY K+LLK+F + ++ KTPM L+ ++ V KLYR MIGSLLYL
Sbjct: 1269 DGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYL 1328
Query: 1421 TASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDA 1480
TASRPDI+FSV +CA FQ+ P+E HL AVKRILRYLK ++ +GL Y K +++KL GY D+
Sbjct: 1329 TASRPDIMFSVCMCAWFQAAPKECHLVAVKRILRYLKYSSTIGLWYPKGAKFKLVGYSDS 1388
Query: 1481 DYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQL 1540
DYAG + +R STSG+CQ LG +LVSW SK+Q+++ALSTAEAEY+SA C Q LWMK L
Sbjct: 1389 DYAGCKVDRNSTSGSCQMLGRSLVSWSSKKQNSVALSTAEAEYVSAGSCCAQLLWMKQTL 1448
Query: 1541 EDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
DY I + P+ CDN +AI ++ NP+ HSR KHI++++HF+RD+V K
Sbjct: 1449 LDYGISFTKTPLLCDNDSAIKIANNPVQHSRTKHIDIRHHFLRDHVAK 1496
>gb|AAP53706.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37534234|ref|NP_921419.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1419
Score = 884 bits (2285), Expect = 0.0
Identities = 461/1001 (46%), Positives = 642/1001 (64%), Gaps = 36/1001 (3%)
Query: 591 RTRLSMLQISFIAPLKHQSWYLDSGCSRHMTGERRMFQELKLKPGGE-VGFGGNEKGKIV 649
++++ +L+++ +A K W +DSG SRHMTG++ F LK E + F +V
Sbjct: 411 QSKVLVLEVALVAR-KENVWIVDSGDSRHMTGDKNWFSSLKKASKTESIVFADASTSAVV 469
Query: 650 GTGTICVDSSPCIYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNS 709
TG + V+ + NV LV L +NLLS+SQ+ D+ ++V F + + SVL N
Sbjct: 470 ATGLVKVNEKFELKNVALVKDLKYNLLSVSQIVDENFEVHFKKTGSKVFDSCGDSVL-NI 528
Query: 710 KRKNNIYKIRLSELEAQNVKCLLS-VNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNL 768
R ++K + + CL++ +++ WHRRLGH +++LS L+LVRGL L
Sbjct: 529 SRYERVFKADFENSVSLVITCLVAKFDKDVMFWHRRLGHVGFDHLTRLSGLDLVRGLSKL 588
Query: 769 KFASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVD 828
K D +C C+ K V T P +LLH+D GP + +S+GGK Y ++IVD
Sbjct: 589 KKDLDLICTPCRHAKMVSTSHAPIVSVMTDAPGQLLHMDTVGPARVQSVGGKWYVLVIVD 648
Query: 829 DYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYG 888
D+SR++WV F+T KDE+ F +++ E + R+RSD+GGEF+N FE + G
Sbjct: 649 DFSRYSWVFFMTTKDEAFQHFRGLFLRLELEFPGSLKRIRSDNGGEFKNASFEQFCNERG 708
Query: 889 IAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRP 948
+ H+FS PR PQQN VVERKNR L EMARTML E + FW EA+NTACYI NR+ +R
Sbjct: 709 LEHEFSSPRVPQQNSVVERKNRVLVEMARTMLDEYKTTRKFWAEAINTACYISNRVFLRS 768
Query: 949 ILNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRF 1008
L KT YEL +P +S+ FGC C+VL + + L KF+A+S+ L LGY ++G+R
Sbjct: 769 KLGKTSYELRFGHQPKVSHLRVFGCKCFVLKSGN-LDKFEARSTDGLFLGYPAHTRGYRV 827
Query: 1009 YNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAG 1068
I E+ V FD+ + + + + + G+ E+ +D+ +E G
Sbjct: 828 LILGTNKIVETCEVSFDEASPGTRPDIAGTLSQVQ---GEDGRIFEDESDYDDD--DEVG 882
Query: 1069 PSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEA 1128
+ +T + S++T V SAF V+ EPK + A
Sbjct: 883 SAGERTTR-SKVTT------------HDVCANSAF-------------VASFEPKDVSHA 916
Query: 1129 LQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVA 1188
L D+ WI AM EEL F +N VW+LV+ P +VIGTKWVF+NK NE G +VRNKARLVA
Sbjct: 917 LTDESWINAMHEELENFERNKVWTLVEPPSGHNVIGTKWVFKNKQNEDGLIVRNKARLVA 976
Query: 1189 QGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQ 1248
QG++Q EG+D+ ETFAP+AR+EAIRLL++F+ + L+QMDVKSAFLNG+I EEVYV Q
Sbjct: 977 QGFTQVEGLDFDETFAPVARIEAIRLLLAFAASKGFKLYQMDVKSAFLNGFIQEEVYVKQ 1036
Query: 1249 PPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDD 1308
PPGFE+ P+HVFKL K+LYGLKQAPRAWY+RL +FLL F GKVD TLF + D+
Sbjct: 1037 PPGFENPDFPNHVFKLSKALYGLKQAPRAWYDRLKNFLLAKGFTMGKVDKTLFVLKHGDN 1096
Query: 1309 ILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQS 1368
L VQIYVDDIIFG + ++ +F++ M+ EFEMSMMGEL+YFLG+Q+ QTP+GT++HQ+
Sbjct: 1097 QLFVQIYVDDIIFGCSTHAVVVDFAETMRREFEMSMMGELSYFLGLQIKQTPQGTFVHQT 1156
Query: 1369 KYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDIL 1428
KYTK+LL++F M TP+ T +L+ ++ V QK YR MIGSLLYLTASRPDI
Sbjct: 1157 KYTKDLLRRFKMENCKPISTPIGSTAVLDPDEDGEAVDQKEYRSMIGSLLYLTASRPDIQ 1216
Query: 1429 FSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTE 1488
F+V LCARFQ+ PR +H AVKRI+RYL T G+ Y +S LSGY DAD+ G R +
Sbjct: 1217 FAVCLCARFQASPRASHRQAVKRIMRYLNHTLEFGIWYSTSSSICLSGYSDADFGGCRID 1276
Query: 1489 RKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILES 1548
RKSTSG C FLG++L++W S++QS++A STAE+EY++AA C +Q LW+ L+DY I
Sbjct: 1277 RKSTSGTCHFLGTSLIAWSSRKQSSVAQSTAESEYVAAASCCSQILWLLSTLKDYGITFE 1336
Query: 1549 NIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KG 1589
+P++CDNT+AI+++KNP+ HSR KHI++++HF+RD+V KG
Sbjct: 1337 KVPLFCDNTSAINIAKNPVQHSRTKHIDIRFHFLRDHVEKG 1377
>ref|XP_468929.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|28209450|gb|AAO37468.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1128
Score = 866 bits (2237), Expect = 0.0
Identities = 448/899 (49%), Positives = 602/899 (66%), Gaps = 31/899 (3%)
Query: 717 KIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALC 776
K+ + +NV L++ W+WHRRL H M ++S+LSK +LV GL ++KF D LC
Sbjct: 191 KVTFGDNSKRNVIGLVAKTSFGWLWHRRLAHVGMNQLSKLSKRDLVVGLKDVKFEKDKLC 250
Query: 777 EACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWV 836
ACQ K K+++STSRPLELLH+DLFGP +SIGG + ++IVDDYS +TWV
Sbjct: 251 SACQASKQVACSHPTKSIMSTSRPLELLHMDLFGPTTYKSIGGNSHCLVIVDDYSCYTWV 310
Query: 837 KFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCP 896
FL K +F F + QNE +C +V++RSD+G +F+N E D I H+ S
Sbjct: 311 FFLHDKCIVAELFKKFAKRAQNEFSCTLVKIRSDNGSKFKNTNIEDYCDDLSIKHELSAT 370
Query: 897 RTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYE 956
+PQQNGVVERKNRTL EMARTML E G++ FW EA+NTAC+ NR+ + +L KT YE
Sbjct: 371 YSPQQNGVVERKNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRLLKKTSYE 430
Query: 957 LWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTI 1016
L KPN++YF FGC CY+ RL KF+++ + LLGY+ SK +R YN + +
Sbjct: 431 LIVGRKPNVAYFRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYRVYNKNKGIV 490
Query: 1017 EESIHVRFDDKLDSDQS-KLVEKFADLSI-----NVSDKGKAPEEAEPE-----EDEPEE 1065
EE+ V+FD+ S + + ++ D + N+S P E E + +DEP
Sbjct: 491 EETADVQFDETNGSQEGHENLDDVGDEGLMRAMKNMSIGDVKPIEVEDKPSTSTQDEPST 550
Query: 1066 EAGPSDSQT-LKKSR-------------ITAAHPMELIMGNKDEPVRTRSAFRPSEETLL 1111
A PS +Q ++K + ++ HP++ ++G+ + V+TRS ++
Sbjct: 551 SASPSQAQVEVEKEKAQDPPMPPRIYTALSKDHPIDQVLGDISKGVQTRSPVA----SIC 606
Query: 1112 SLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRN 1171
VS +EPK +DEAL D DW+ A+ EELN F++N VW+LV++P +VIGTKWVFRN
Sbjct: 607 EHYSFVSCLEPKHVDEALYDPDWMNAIHEELNNFARNKVWTLVERPRDHNVIGTKWVFRN 666
Query: 1172 KLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDV 1231
K +E VVRNKARLVAQG++Q E +D+ ETF P+ARLEAIR+L++F+ +I L QMDV
Sbjct: 667 KQDENRLVVRNKARLVAQGFTQVEDLDFGETFGPVARLEAIRILLAFASCFDIKLFQMDV 726
Query: 1232 KSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEF 1291
KSAFLNG I+E V+V QPPGF+D K P+HV+KL K+LYGLKQAPRAWYERL FLL +F
Sbjct: 727 KSAFLNGEIAELVFVEQPPGFDDPKYPNHVYKLSKALYGLKQAPRAWYERLRDFLLSKDF 786
Query: 1292 VRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYF 1351
GKVDTTLF K DD V QIYVDDIIFGS N+ CKEF MM EFEMSM+ EL++F
Sbjct: 787 KIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIEELSFF 846
Query: 1352 LGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYR 1411
LG+Q+ Q +GT++ Q+KY K+LLK+F + ++ KTPM L+ ++ V KLYR
Sbjct: 847 LGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNWHLDLDEGGKPVDLKLYR 906
Query: 1412 GMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSE 1471
MIGSLLYLTASRPDI+FSV + ARFQ+ P+E HL AVKRILRYLK ++ + L Y K ++
Sbjct: 907 SMIGSLLYLTASRPDIMFSVCMYARFQAAPKECHLVAVKRILRYLKHSSTISLWYPKGAK 966
Query: 1472 YKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICST 1531
+KL GY D+DYAG + +RKSTSG+CQ LG +LVSW SK+Q+++ALSTAEAEYISA C
Sbjct: 967 FKLVGYSDSDYAGYKVDRKSTSGSCQMLGRSLVSWSSKKQNSVALSTAEAEYISAGSCCA 1026
Query: 1532 QKLWMKHQLEDYQI--LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
Q LWMK L DY I E+ P+ C+N + I ++ NP+ H R KHI++++HF+ D+V K
Sbjct: 1027 QLLWMKQILLDYGISFTETQTPLLCNNDSTIKIANNPVQHFRTKHIDIRHHFLTDHVAK 1085
Score = 38.9 bits (89), Expect = 1.4
Identities = 19/43 (44%), Positives = 27/43 (62%), Gaps = 2/43 (4%)
Query: 610 WYLDSGCSRHMTGERRMFQ--ELKLKPGGEVGFGGNEKGKIVG 650
W LDS C++ MTG+R MF E++ K +V FG N K ++G
Sbjct: 162 WVLDSVCTQRMTGDRAMFTTFEVEGKEQEKVTFGDNSKRNVIG 204
>ref|NP_912905.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 940
Score = 834 bits (2154), Expect = 0.0
Identities = 439/871 (50%), Positives = 574/871 (65%), Gaps = 41/871 (4%)
Query: 754 SQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVK 813
SQ SK + GL N++F D +C ACQ GK KNV++T+RPLELLH+DLFGP+
Sbjct: 34 SQTSKARHILGLTNIQFEKDRVCSACQAGKQIGAHHPVKNVMTTTRPLELLHMDLFGPIA 93
Query: 814 SESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGG 873
SIGG +YG++IVDD+S +TWV FL K E+ A+F F + QNE +I +RSD+
Sbjct: 94 YLSIGGNKYGLVIVDDFSCFTWVFFLHDKSETQAIFKKFARRAQNEFDLKIKNIRSDNVK 153
Query: 874 EFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEA 933
EF+N ES D GI H+FS P +PQQNGV ERKNRTL E+ARTML E + FW EA
Sbjct: 154 EFKNTCIESFLDEEGIKHEFSAPYSPQQNGVAERKNRTLIEIARTMLDEYKTSDRFWAEA 213
Query: 934 VNTACYIQNRISVRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSK 993
VNT C+ NR+ + +L KTPYEL KPN+SYF FG CY+LN K R KF K
Sbjct: 214 VNTVCHDINRLYLHRLLKKTPYELLTGNKPNVSYFRVFGSKCYILNKKARSSKFAPKVDG 273
Query: 994 CLLLGYSERSKGFRFYNTDAKTIEESIHVRFD-------DKLDS-------DQSKLVEKF 1039
LLGY +R +N + +E + V FD +++DS D + +++
Sbjct: 274 GFLLGYGSNECAYRVFNKTSGIVEIAPDVTFDETNGSQVEQVDSHVLGEEEDPREAIKRL 333
Query: 1040 A-----------------DLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRITA 1082
A + S + P + ++ E E+ PS S L RI
Sbjct: 334 ALGDVRPREPQQGASSSTQVEPPTSTQANDPSTSSLDQGEEGEQVPPS-SINLAHPRIHQ 392
Query: 1083 A----HPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAM 1138
+ HP + I+G+ ++ V TRS VS +EP ++EAL D DW++AM
Sbjct: 393 SIQRDHPTDNILGDINKGVSTRSHI----ANFCEHYSFVSSLEPLRVEEALNDPDWVMAM 448
Query: 1139 EEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGID 1198
+EELN F++N+VW+LV++ +VIGTKW+FRNK +E G V+RNKARLVAQG++Q EG+D
Sbjct: 449 QEELNNFTRNEVWTLVERSRQ-NVIGTKWIFRNKQDEAGVVIRNKARLVAQGFTQIEGLD 507
Query: 1199 YTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKP 1258
+ ETFAP+ARLE+IR+L++F+ N N L+QMDVKSAFLNG I+E VYV QPPGF+D K P
Sbjct: 508 FGETFAPVARLESIRILLTFATNLNFKLYQMDVKSAFLNGLINELVYVEQPPGFKDPKYP 567
Query: 1259 DHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDD 1318
+HV+KL K+LY LKQAPRAWYE L +FL++N F GK D+TLF K + +DI V QIYVDD
Sbjct: 568 NHVYKLHKALYELKQAPRAWYECLRNFLVKNGFEIGKADSTLFTKRHDNDIFVCQIYVDD 627
Query: 1319 IIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKF 1378
IIFGS N+S +EFS+MM FEMSMMGEL +FLG+Q+ Q EGT+I Q+KY K++LKKF
Sbjct: 628 IIFGSTNKSFSEEFSRMMTKRFEMSMMGELKFFLGLQIKQLKEGTFICQTKYLKDMLKKF 687
Query: 1379 NMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQ 1438
M + TPM L+ + V QK+YR +IGSLLYL ASRPDI+ SV +CARFQ
Sbjct: 688 GMENAKPIHTPMPSNGHLDLNEQGKDVDQKVYRSIIGSLLYLCASRPDIMLSVCMCARFQ 747
Query: 1439 SDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQF 1498
+ P+E HL AVKRILRYL T NLGL Y K + + L GY DADYAG + +RKSTSG CQF
Sbjct: 748 AAPKECHLVAVKRILRYLVHTPNLGLWYPKGARFDLIGYADADYAGCKVDRKSTSGTCQF 807
Query: 1499 LGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTA 1558
LG +LVSW SK+Q+++ALSTAEAEYIS C Q +WMK L DY + S IP+ CDN +
Sbjct: 808 LGRSLVSWSSKKQNSVALSTAEAEYISTGSCCAQLIWMKQTLRDYGLNVSKIPLLCDNES 867
Query: 1559 AISLSKNPILHSRAKHIEVKYHFIRDYV*KG 1589
AI ++ NP+ HSR KHI++++HF+RD+ +G
Sbjct: 868 AIKIANNPVQHSRTKHIDIRHHFLRDHSTRG 898
>ref|XP_463224.1| putative integrase [Oryza sativa (japonica cultivar-group)]
gi|40737034|gb|AAR89047.1| putative integrase [Oryza
sativa (japonica cultivar-group)]
Length = 1507
Score = 792 bits (2046), Expect = 0.0
Identities = 432/1026 (42%), Positives = 615/1026 (59%), Gaps = 92/1026 (8%)
Query: 591 RTRLSMLQISFIAPLKHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVG 650
++++ +L+++ +A K W +DSGC RHMTG ++ ++ +LK
Sbjct: 505 QSKVLVLEVALVAR-KENVWIVDSGCLRHMTGMVKVNEKFELK----------------- 546
Query: 651 TGTICVDSSPCIYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSK 710
NV LV+ L +NLLS+ Q+ + ++V F + + SVL N
Sbjct: 547 -------------NVALVEDLNYNLLSVLQIVYENFEVHFKKTRSKVFDSCGDSVL-NIS 592
Query: 711 RKNNIYKIRLSELEAQNVKCLLS-VNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLK 769
R ++K + + CL++ +++ WHRRLGH +++LS L+LVRGLP LK
Sbjct: 593 RYGRVFKADFENPVSPVITCLVAKFDKDVMFWHRRLGHVGFDHLTRLSGLDLVRGLPKLK 652
Query: 770 FASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDD 829
D C C K V T P +LLH+D GP + +S+GGK Y ++I+DD
Sbjct: 653 KDLDLDCAPCHHAKMVASSHAPIVSVMTDAPRQLLHMDTVGPARVQSVGGKWYVLVIIDD 712
Query: 830 YSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGI 889
+SR++WV F+ KDE+ F +++ E + R+RSD+G E++N FE + G+
Sbjct: 713 FSRYSWVFFMATKDEAFQHFRGLFLRLELEFPGSLKRIRSDNGSEYKNASFEQFCNERGL 772
Query: 890 AHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPI 949
H+FS PR PQQNGVVERKN L EM RTML E + FW EA+NTA YI NR+ +R
Sbjct: 773 EHEFSSPRVPQQNGVVERKNHVLVEMVRTMLDEYKTPRKFWAEAINTAYYISNRVFLRSK 832
Query: 950 LNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFY 1009
L K+ YEL +P +S+ FGC C+VL + + L KF+A+S+ L LGY ++G+R
Sbjct: 833 LGKSSYELRFGHQPKVSHLRVFGCKCFVLKSGN-LDKFEARSTDGLFLGYPAHTRGYRTL 891
Query: 1010 NT----DAKTIEESIHVRFDDKLDS----------------------DQSKLVEKFADLS 1043
+ D + E+ DD++ S ++S A+ S
Sbjct: 892 SQVQGEDGRIFEDESDDNDDDEVGSAGQTGRQAGQTAGTPPVRPAHEERSDRPGLSAEGS 951
Query: 1044 INVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAF 1103
++ G P E E S S+ + I HP E I+GN E TRS
Sbjct: 952 VDAVRDG--PLEITTSTSTDTEHG--STSEVVAPLHIQQRHPPEQIIGNIGERT-TRS-- 1004
Query: 1104 RPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVI 1163
+ + + + V+ EPK + AL D+ WI AM EEL F +N VW+LV+ P ++I
Sbjct: 1005 KVTTHDVCANYAFVASFEPKDVSHALTDESWINAMHEELENFERNKVWTLVEPPSGHNII 1064
Query: 1164 GTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHN 1223
GTKWVF+NK NE +VRNKARLVAQG++Q EG+D+ ETFAP+AR+EAIRL ++F+ +
Sbjct: 1065 GTKWVFKNKQNEDDLIVRNKARLVAQGFTQVEGLDFDETFAPVARIEAIRLFLAFASSKG 1124
Query: 1224 IVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1283
L+QMDVKSAFLNG+I EEVYV QPPGFE+ P++VFKL K+LYGLKQAPRAWY+RL
Sbjct: 1125 FKLYQMDVKSAFLNGFIQEEVYVKQPPGFENPDFPNYVFKLSKALYGLKQAPRAWYDRLK 1184
Query: 1284 SFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMS 1343
+FLL F GKVD TLF L +F++ M+ EFEMS
Sbjct: 1185 NFLLAKGFTMGKVDKTLFV-------------------------LKHDFAETMRREFEMS 1219
Query: 1344 MMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASG 1403
MMGEL+YFLG+Q+ QTP+GT++HQ+KYTK+LL++F M TP+ T +L+ ++
Sbjct: 1220 MMGELSYFLGLQIKQTPQGTFVHQTKYTKDLLRRFKMENCKPISTPIDSTAVLDPDEDGE 1279
Query: 1404 KVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLG 1463
V QK YR MIGSLLYLTASRP+I F+V LCARFQ+ PR +H AVKRI+RYL T G
Sbjct: 1280 AVDQKEYRSMIGSLLYLTASRPEIQFAVCLCARFQASPRASHRQAVKRIMRYLNHTLEFG 1339
Query: 1464 LMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEY 1523
+ Y +S LSGY DA++ G R +RKSTSG C FLG++L++W S++QS++A STAE+EY
Sbjct: 1340 IWYSTSSSLCLSGYSDANFGGCRIDRKSTSGTCHFLGTSLIAWSSRKQSSVAQSTAESEY 1399
Query: 1524 ISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIR 1583
++AA C +Q LW+ L++Y + +P++CDNT+AI+++KN + HSR KH+++++HF+R
Sbjct: 1400 VAAASCCSQILWLLSTLKNYGLTFEKVPLFCDNTSAINIAKNLVQHSRTKHVDIRFHFLR 1459
Query: 1584 DYV*KG 1589
D+V KG
Sbjct: 1460 DHVEKG 1465
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.332 0.142 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,376,623,122
Number of Sequences: 2540612
Number of extensions: 93817362
Number of successful extensions: 428897
Number of sequences better than 10.0: 4046
Number of HSP's better than 10.0 without gapping: 2283
Number of HSP's successfully gapped in prelim test: 1769
Number of HSP's that attempted gapping in prelim test: 418981
Number of HSP's gapped (non-prelim): 7120
length of query: 1591
length of database: 863,360,394
effective HSP length: 142
effective length of query: 1449
effective length of database: 502,593,490
effective search space: 728257967010
effective search space used: 728257967010
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0388a.6


