

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
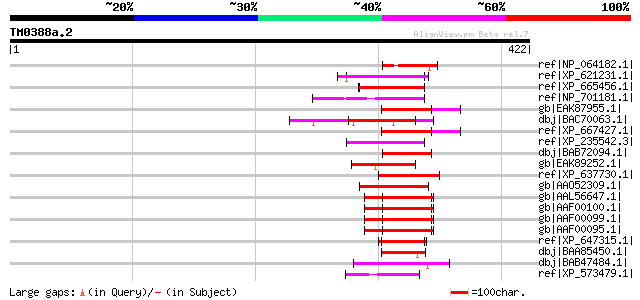
Query= TM0388a.2
(422 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_064182.1| pR83 [Murid herpesvirus 2] gi|9800297|gb|AAF991... 51 6e-05
ref|XP_621231.1| PREDICTED: similar to hypothetical protein [Mus... 51 8e-05
ref|XP_665456.1| hypothetical protein Chro.30409 [Cryptosporidiu... 50 1e-04
ref|NP_701181.1| hypothetical protein PF11_0321 [Plasmodium falc... 50 1e-04
gb|EAK87955.1| membrane associated thioredoxin [Cryptosporidium ... 50 2e-04
dbj|BAC70063.1| hypothetical protein [Streptomyces avermitilis M... 50 2e-04
ref|XP_667427.1| transmembrane protein 17 [Cryptosporidium homin... 50 2e-04
ref|XP_235542.3| PREDICTED: similar to leucine zipper, down-regu... 49 2e-04
dbj|BAB72094.1| histone acetyltransferase MORF [Macaca fascicula... 49 2e-04
gb|EAK89252.1| hypothetical protein with 4 or more possible tran... 49 3e-04
ref|XP_637730.1| structural maintenance of chromosome protein [D... 49 3e-04
gb|AAO52309.1| similar to Oryza sativa (japonica cultivar-group)... 49 4e-04
gb|AAL56647.1| histone acetyltransferase MOZ2 [Homo sapiens] 49 4e-04
gb|AAF00100.1| histone acetyltransferase MORF beta [Homo sapiens... 49 4e-04
gb|AAF00099.1| histone acetyltransferase MORF alpha [Homo sapiens] 49 4e-04
gb|AAF00095.1| histone acetyltransferase MORF [Homo sapiens] gi|... 49 4e-04
ref|XP_647315.1| hypothetical protein DDB0202150 [Dictyostelium ... 49 4e-04
dbj|BAA85450.1| S-locus protein 1 [Brassica rapa] 49 4e-04
dbj|BAB47484.1| KIAA1855 protein [Homo sapiens] 48 5e-04
ref|XP_573479.1| PREDICTED: similar to LRRGT00170 [Rattus norveg... 48 5e-04
>ref|NP_064182.1| pR83 [Murid herpesvirus 2] gi|9800297|gb|AAF99171.1| pR83 [rat
cytomegalovirus Maastricht]
Length = 647
Score = 51.2 bits (121), Expect = 6e-05
Identities = 28/48 (58%), Positives = 30/48 (62%), Gaps = 5/48 (10%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVT---PPRADPSP 348
EDEEE DP E GE++EEEEEEEEEEE EV PP DP P
Sbjct: 465 EDEEEADPAE--GEEEEEEEEEEEEEEEAMAEEEVAAASPPPDVDPDP 510
>ref|XP_621231.1| PREDICTED: similar to hypothetical protein [Mus musculus]
Length = 255
Score = 50.8 bits (120), Expect = 8e-05
Identities = 30/79 (37%), Positives = 46/79 (57%), Gaps = 5/79 (6%)
Query: 267 RARRPV-----HVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEE 321
RA+RPV ++I + + +E + +E +E+EEEE+ EE E++EE
Sbjct: 168 RAQRPVQSILLYLITAQAQLERKNHREEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 227
Query: 322 EEEEEEEEENVEVVNEVVT 340
EEEEEEEEE E N+++T
Sbjct: 228 EEEEEEEEEEEEEENKIIT 246
Score = 47.4 bits (111), Expect = 9e-04
Identities = 28/63 (44%), Positives = 35/63 (55%)
Query: 275 IEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEV 334
I + PV +L AQ E +E+EEEE+ EE E++EEEEEEEEEEE E
Sbjct: 167 IRAQRPVQSILLYLITAQAQLERKNHREEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 226
Query: 335 VNE 337
E
Sbjct: 227 EEE 229
Score = 42.4 bits (98), Expect = 0.027
Identities = 19/33 (57%), Positives = 25/33 (75%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
+E+EEEE+ EE E++EEEEEEEEEE + V
Sbjct: 215 EEEEEEEEEEEEEEEEEEEEEEEEEEENKIITV 247
>ref|XP_665456.1| hypothetical protein Chro.30409 [Cryptosporidium hominis]
gi|54656153|gb|EAL35225.1| hypothetical protein
Chro.30409 [Cryptosporidium hominis]
Length = 985
Score = 50.4 bits (119), Expect = 1e-04
Identities = 28/54 (51%), Positives = 35/54 (63%)
Query: 284 LLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
LL +RI Q+ KE +E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 412 LLLLQRINQMRKERRKLEYEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 465
Score = 48.9 bits (115), Expect = 3e-04
Identities = 26/53 (49%), Positives = 33/53 (62%)
Query: 285 LSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+ KER Y+E +E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 421 MRKERRKLEYEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 473
Score = 44.7 bits (104), Expect = 0.006
Identities = 26/63 (41%), Positives = 37/63 (58%), Gaps = 4/63 (6%)
Query: 275 IEKRLPVAPLLSKERIAQLYKE----WTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+ L V L E I ++K+ RM +E+EEEE+ EE +++EEEEEEE+EEE
Sbjct: 169 VNASLNVQELKIVENIHGVFKDEKIKGKGRMKEEEEEEEEEEEEEEKEEEEEEEEEKEEE 228
Query: 331 NVE 333
E
Sbjct: 229 GKE 231
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 447 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 481
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 470 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 504
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 473 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKEE 507
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 452 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 486
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 440 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 474
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 455 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 489
Score = 44.3 bits (103), Expect = 0.007
Identities = 20/34 (58%), Positives = 27/34 (78%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVN 336
+E+EEEE+ EE E+++EEEEE+EEEE EV N
Sbjct: 488 EEEEEEEEEEEEEEEEEKEEEEEKEEEEEEEVYN 521
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 461 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 495
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 465 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 499
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 460 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 494
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 464 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 498
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 457 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 491
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 446 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 480
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 467 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 501
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 459 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 493
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 458 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 492
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 453 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 487
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 444 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 478
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 442 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 476
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 474 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKEEE 508
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 450 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 484
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 463 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 497
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 476 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEKEEEEE 510
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 469 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 503
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 443 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 477
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 472 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKE 506
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 441 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 475
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 449 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 483
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 462 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 496
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 468 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 502
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 456 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 490
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 466 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 500
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 445 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 479
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 451 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 485
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 448 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 482
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 454 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 488
Score = 43.5 bits (101), Expect = 0.012
Identities = 21/37 (56%), Positives = 27/37 (72%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
+E+EEEE+ EE E++EEEEE+EEEEE E E V
Sbjct: 483 EEEEEEEEEEEEEEEEEEEEEEKEEEEEKEEEEEEEV 519
Score = 42.7 bits (99), Expect = 0.021
Identities = 20/35 (57%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEE+EE E E
Sbjct: 480 EEEEEEEEEEEEEEEEEEEEEEEEEKEEEEEKEEE 514
Score = 42.7 bits (99), Expect = 0.021
Identities = 20/35 (57%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEE+EEEE E E
Sbjct: 482 EEEEEEEEEEEEEEEEEEEEEEEKEEEEEKEEEEE 516
Score = 42.7 bits (99), Expect = 0.021
Identities = 20/35 (57%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEE+E E E
Sbjct: 479 EEEEEEEEEEEEEEEEEEEEEEEEEEKEEEEEKEE 513
Score = 42.7 bits (99), Expect = 0.021
Identities = 20/35 (57%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEE+ E E
Sbjct: 478 EEEEEEEEEEEEEEEEEEEEEEEEEEEKEEEEEKE 512
Score = 39.7 bits (91), Expect = 0.18
Identities = 17/32 (53%), Positives = 25/32 (78%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEV 334
+E+EEEE+ EE E++E+EEEEEEE N ++
Sbjct: 493 EEEEEEEEEEEEKEEEEEKEEEEEEEVYNYDI 524
Score = 38.1 bits (87), Expect = 0.52
Identities = 17/31 (54%), Positives = 24/31 (76%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ EE E++EEEEE+EEE + E
Sbjct: 203 EEEEEEEEEEEEKEEEEEEEEEKEEEGKEKE 233
Score = 35.0 bits (79), Expect = 4.4
Identities = 14/33 (42%), Positives = 26/33 (78%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
+E+E+EE+ +E G+D+E++ E+EE EE+ + V
Sbjct: 221 EEEEKEEEGKEKEGDDEEDDGEKEETEEDRDFV 253
>ref|NP_701181.1| hypothetical protein PF11_0321 [Plasmodium falciparum 3D7]
gi|23496246|gb|AAN35905.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 773
Score = 50.1 bits (118), Expect = 1e-04
Identities = 32/91 (35%), Positives = 48/91 (52%), Gaps = 6/91 (6%)
Query: 247 DKHRPIFPHLICSLIHAVRDRARRPVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDE 306
D+ + I+ H++ + + R V VI KR K++ W + + +E+E
Sbjct: 159 DQRKDIYLHILTYVNKELYRYGSRTV-VITKRKEKTTTRGKKKFL-----WKSLIDEEEE 212
Query: 307 EEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
EEE+ EE E++EEEEEEEEEEE E E
Sbjct: 213 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 243
Score = 41.6 bits (96), Expect = 0.047
Identities = 18/31 (58%), Positives = 25/31 (80%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ EE E++EEEEEEEEEE+ +
Sbjct: 221 EEEEEEEEEEEEEEEEEEEEEEEEEEEKKYQ 251
Score = 37.0 bits (84), Expect = 1.2
Identities = 20/47 (42%), Positives = 31/47 (65%), Gaps = 3/47 (6%)
Query: 284 LLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
L+ +E + +E +E+EEEE+ EE E++EEEEEEEEE++
Sbjct: 206 LIDEEEEEEEEEEEEEEEEEEEEEEEEEEE---EEEEEEEEEEEEKK 249
Score = 36.2 bits (82), Expect = 2.0
Identities = 15/29 (51%), Positives = 24/29 (82%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+E+EEEE+ EE E++EEEEEEE++ ++
Sbjct: 224 EEEEEEEEEEEEEEEEEEEEEEEEKKYQH 252
Score = 35.0 bits (79), Expect = 4.4
Identities = 15/31 (48%), Positives = 23/31 (73%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ EE E++EEEEE++ + N E
Sbjct: 226 EEEEEEEEEEEEEEEEEEEEEEKKYQHGNYE 256
>gb|EAK87955.1| membrane associated thioredoxin [Cryptosporidium parvum]
gi|66357966|ref|XP_626161.1| membrane associated
thioredoxin [Cryptosporidium parvum]
Length = 363
Score = 49.7 bits (117), Expect = 2e-04
Identities = 23/41 (56%), Positives = 30/41 (73%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
+E+EEEE+ EE E++EEEEEEEEEEE E ++ TP R
Sbjct: 291 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAPKQLKTPQR 331
Score = 48.1 bits (113), Expect = 5e-04
Identities = 27/64 (42%), Positives = 36/64 (56%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRMFLRQ 362
+E+EEEE+ EE E++EEEEEEEEEEE E E P+ +P A +
Sbjct: 284 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAPKQLKTPQRGRRAAENIAKED 343
Query: 363 DRLH 366
DRL+
Sbjct: 344 DRLN 347
Score = 44.7 bits (104), Expect = 0.006
Identities = 22/46 (47%), Positives = 29/46 (62%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+E+EEEE+ EE E++EEEEEEEEEEE E E + +P
Sbjct: 278 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAP 323
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 272 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 306
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 276 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 310
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 268 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 302
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 275 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 309
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 271 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 305
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 267 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 301
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 265 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 299
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 263 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 297
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 264 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 298
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 262 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 296
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 269 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 303
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 273 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 307
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 266 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 300
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 274 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 308
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 270 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 304
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 277 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 311
Score = 43.1 bits (100), Expect = 0.016
Identities = 21/34 (61%), Positives = 24/34 (69%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
ED EEE+ EE E++EEEEEEEEEEE E E
Sbjct: 259 EDHEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 292
Score = 42.0 bits (97), Expect = 0.036
Identities = 20/35 (57%), Positives = 25/35 (71%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
++ EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 259 EDHEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 293
Score = 39.7 bits (91), Expect = 0.18
Identities = 20/33 (60%), Positives = 23/33 (69%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
D+E ED EE E++EEEEEEEEEEE E E
Sbjct: 255 DDECEDHEEEEEEEEEEEEEEEEEEEEEEEEEE 287
Score = 38.1 bits (87), Expect = 0.52
Identities = 19/34 (55%), Positives = 23/34 (66%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+DE E+ EE E++EEEEEEEEEEE E E
Sbjct: 255 DDECEDHEEEEEEEEEEEEEEEEEEEEEEEEEEE 288
Score = 33.9 bits (76), Expect = 9.7
Identities = 17/35 (48%), Positives = 22/35 (62%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+ D+ +D E E++EEEEEEEEEEE E E
Sbjct: 249 KRDKYSDDECEDHEEEEEEEEEEEEEEEEEEEEEE 283
>dbj|BAC70063.1| hypothetical protein [Streptomyces avermitilis MA-4680]
gi|29828894|ref|NP_823528.1| hypothetical protein
SAV2352 [Streptomyces avermitilis MA-4680]
Length = 376
Score = 49.7 bits (117), Expect = 2e-04
Identities = 25/55 (45%), Positives = 35/55 (63%)
Query: 276 EKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
E+ P A +E + ++ +E+EEEE+ EEP GE++EEEEEEEEEEE
Sbjct: 148 EEEEPEAAYEEEEEEEEEEEQPEGEYEEEEEEEEEEEEPEGEEEEEEEEEEEEEE 202
Score = 47.8 bits (112), Expect = 7e-04
Identities = 44/128 (34%), Positives = 59/128 (45%), Gaps = 12/128 (9%)
Query: 228 MDVATIIAGRIYSLYNRS-------KDKHRPIFPHLICSLIHAVRDRARRPVHVIEKR-- 278
+ +AT +AGR + L R + P F L L V D R+ V R
Sbjct: 29 LSIATYLAGRRFGLEPRQLATEGMRRLGEMPQFAQLQEQLRGEVLDAGRKAVTAAADRGM 88
Query: 279 LPVAPLLSKERIAQLYKEWTARMPQEDEEEED--PEEPAGEDKEEEEEEEEEEENVEVVN 336
+A LS +R A+L K+ +E+E EE+ P+E E +EEEEEEEEE E E
Sbjct: 89 SSLADALS-DRTARLGKKGEEEEGEEEEGEEEYAPDEEGAEGEEEEEEEEEEPEEEEPEE 147
Query: 337 EVVTPPRA 344
E P A
Sbjct: 148 EEEEPEAA 155
Score = 41.6 bits (96), Expect = 0.047
Identities = 18/29 (62%), Positives = 23/29 (79%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
P+E+E EE+ EEP +EEEEEEEEEE+
Sbjct: 140 PEEEEPEEEEEEPEAAYEEEEEEEEEEEQ 168
Score = 40.8 bits (94), Expect = 0.080
Identities = 21/36 (58%), Positives = 26/36 (71%), Gaps = 5/36 (13%)
Query: 303 QEDEEEEDPEEPAGEDKEEE-----EEEEEEEENVE 333
+E+EEEE+PEE E++EEE EEEEEEEE E
Sbjct: 132 EEEEEEEEPEEEEPEEEEEEPEAAYEEEEEEEEEEE 167
Score = 38.5 bits (88), Expect = 0.40
Identities = 23/50 (46%), Positives = 29/50 (58%), Gaps = 7/50 (14%)
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEE-------EEEEENVEVVNE 337
+E A +E+EEEE+ E+P GE +EEEEEE EEEE E E
Sbjct: 150 EEPEAAYEEEEEEEEEEEQPEGEYEEEEEEEEEEEEPEGEEEEEEEEEEE 199
Score = 35.8 bits (81), Expect = 2.6
Identities = 21/36 (58%), Positives = 25/36 (69%), Gaps = 6/36 (16%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEE-----EEEEENVE 333
+E+EEEE+ EEP E+ EEEEEE EEEEE E
Sbjct: 130 EEEEEEEE-EEPEEEEPEEEEEEPEAAYEEEEEEEE 164
Score = 35.8 bits (81), Expect = 2.6
Identities = 20/38 (52%), Positives = 24/38 (62%), Gaps = 7/38 (18%)
Query: 303 QEDEEEEDPEEPAGEDK-------EEEEEEEEEEENVE 333
+E+EEEE EE E++ EEEEEEEEEEE E
Sbjct: 133 EEEEEEEPEEEEPEEEEEEPEAAYEEEEEEEEEEEQPE 170
>ref|XP_667427.1| transmembrane protein 17 [Cryptosporidium hominis]
gi|54658565|gb|EAL37199.1| transmembrane protein 17
[Cryptosporidium hominis]
Length = 366
Score = 49.7 bits (117), Expect = 2e-04
Identities = 23/41 (56%), Positives = 30/41 (73%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
+E+EEEE+ EE E++EEEEEEEEEEE E ++ TP R
Sbjct: 294 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAPKQLKTPQR 334
Score = 48.1 bits (113), Expect = 5e-04
Identities = 27/64 (42%), Positives = 36/64 (56%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRMFLRQ 362
+E+EEEE+ EE E++EEEEEEEEEEE E E P+ +P A +
Sbjct: 287 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAPKQLKTPQRGRRAAENIAKED 346
Query: 363 DRLH 366
DRL+
Sbjct: 347 DRLN 350
Score = 44.7 bits (104), Expect = 0.006
Identities = 22/46 (47%), Positives = 29/46 (62%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+E+EEEE+ EE E++EEEEEEEEEEE E E + +P
Sbjct: 281 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAP 326
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 279 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 313
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 266 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 300
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 268 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 302
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 269 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 303
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 276 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 310
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 278 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 312
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 272 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 306
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 275 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 309
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 263 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 297
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 265 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 299
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 280 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 314
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 267 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 301
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 271 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 305
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 273 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 307
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 270 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 304
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 262 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 296
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 277 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 311
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 274 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 308
Score = 44.3 bits (103), Expect = 0.007
Identities = 21/35 (60%), Positives = 26/35 (74%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 264 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 298
Score = 41.2 bits (95), Expect = 0.061
Identities = 20/34 (58%), Positives = 24/34 (69%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+ EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 260 DHEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 293
Score = 41.2 bits (95), Expect = 0.061
Identities = 20/33 (60%), Positives = 23/33 (69%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
D EEE+ EE E++EEEEEEEEEEE E E
Sbjct: 260 DHEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 292
Score = 37.0 bits (84), Expect = 1.2
Identities = 19/33 (57%), Positives = 22/33 (66%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
D+E D EE E++EEEEEEEEEEE E E
Sbjct: 255 DDECVDHEEEEEEEEEEEEEEEEEEEEEEEEEE 287
Score = 35.4 bits (80), Expect = 3.3
Identities = 18/34 (52%), Positives = 22/34 (63%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+DE + EE E++EEEEEEEEEEE E E
Sbjct: 255 DDECVDHEEEEEEEEEEEEEEEEEEEEEEEEEEE 288
>ref|XP_235542.3| PREDICTED: similar to leucine zipper, down-regulated in cancer
1-like protein [Rattus norvegicus]
Length = 916
Score = 49.3 bits (116), Expect = 2e-04
Identities = 28/63 (44%), Positives = 38/63 (59%)
Query: 275 IEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEV 334
I+ +P L ++E + KE +E+EEEE+ EE E++EEEEEEEEEEE E
Sbjct: 712 IQAWMPSMFLRTEEEEEEEDKEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEGEEE 771
Query: 335 VNE 337
V E
Sbjct: 772 VKE 774
Score = 46.6 bits (109), Expect = 0.001
Identities = 25/52 (48%), Positives = 31/52 (59%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAA 354
+E+EEEE+ EE E++EEEEEEEEEEE E E V + EAA
Sbjct: 736 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEGEEEVKEEEVEEEEGEEEAA 787
Score = 39.7 bits (91), Expect = 0.18
Identities = 21/47 (44%), Positives = 29/47 (61%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
KE + +E +E+EEEE+ EE E++EEE EEE +EE VE
Sbjct: 732 KEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEGEEEVKEEEVE 778
Score = 39.3 bits (90), Expect = 0.23
Identities = 18/35 (51%), Positives = 24/35 (68%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE GE++ +EEE EEEE E E
Sbjct: 755 EEEEEEEEEEEEEGEEEVKEEEVEEEEGEEEAAGE 789
>dbj|BAB72094.1| histone acetyltransferase MORF [Macaca fascicularis]
Length = 1784
Score = 49.3 bits (116), Expect = 2e-04
Identities = 24/40 (60%), Positives = 29/40 (72%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
E+EE+E+ EE ED+EEEEEEEEEEE E N +PPR
Sbjct: 787 EEEEDEEEEEEEEEDEEEEEEEEEEEEEEEEENIQSSPPR 826
Score = 46.6 bits (109), Expect = 0.001
Identities = 21/41 (51%), Positives = 31/41 (75%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRA 344
E+EEEE+ E+ E++EEEEEEEEEEEN++ +T P++
Sbjct: 792 EEEEEEEEEDEEEEEEEEEEEEEEEEENIQSSPPRLTKPQS 832
Score = 45.1 bits (105), Expect = 0.004
Identities = 23/46 (50%), Positives = 28/46 (60%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+E+EEEED EE E++EEEEE+EEEEE E E SP
Sbjct: 779 EEEEEEEDEEEEDEEEEEEEEEDEEEEEEEEEEEEEEEEENIQSSP 824
Score = 37.7 bits (86), Expect = 0.67
Identities = 18/44 (40%), Positives = 29/44 (65%), Gaps = 1/44 (2%)
Query: 295 KEWTARMPQEDEEEEDPEEPA-GEDKEEEEEEEEEEENVEVVNE 337
KE +AR+ +EEEE+ EEP+ ED + ++E++ E+ EV E
Sbjct: 1110 KEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKE 1153
Score = 35.8 bits (81), Expect = 2.6
Identities = 19/32 (59%), Positives = 23/32 (71%), Gaps = 5/32 (15%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P+E+EEEE+ EE +EE EEEEEE NVE
Sbjct: 1064 PEEEEEEEEEEE-----EEEGEEEEEEGGNVE 1090
>gb|EAK89252.1| hypothetical protein with 4 or more possible transmembrane domains
[Cryptosporidium parvum] gi|66359466|ref|XP_626911.1|
hypothetical protein cgd3_3620 [Cryptosporidium parvum]
Length = 1581
Score = 48.9 bits (115), Expect = 3e-04
Identities = 27/58 (46%), Positives = 37/58 (63%), Gaps = 6/58 (10%)
Query: 279 LPVAPLLSKERIAQLYKE------WTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
L V L E I +++K+ RM +E+EEEE+ EE GE++EE+EEEEEEEE
Sbjct: 832 LNVHELKMVENIQEVFKDEKVKGKGKGRMKEEEEEEEEEEEEEGEEEEEKEEEEEEEE 889
Score = 38.9 bits (89), Expect = 0.30
Identities = 25/53 (47%), Positives = 32/53 (60%), Gaps = 7/53 (13%)
Query: 284 LLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVN 336
LL +RI Q+ KE +E EEEE E++EEEEEE+E+EE EV N
Sbjct: 1074 LLLLQRINQMRKERRKLEYEEVEEEE-------EEEEEEEEEKEKEEGEEVYN 1119
Score = 36.2 bits (82), Expect = 2.0
Identities = 14/33 (42%), Positives = 26/33 (78%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
+E+EEEE+ +E G+D+E++ E+ E EE+++ V
Sbjct: 883 EEEEEEEEGKEKEGDDEEDDGEKGETEEDIDFV 915
>ref|XP_637730.1| structural maintenance of chromosome protein [Dictyostelium
discoideum] gi|60466163|gb|EAL64226.1| structural
maintenance of chromosome protein [Dictyostelium
discoideum]
Length = 1415
Score = 48.9 bits (115), Expect = 3e-04
Identities = 23/49 (46%), Positives = 32/49 (64%)
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPA 349
M DEEEE+ EE E++EEE+EEEEEEE E + + PP++ P+
Sbjct: 28 MKDIDEEEEEEEEEEEEEEEEEKEEEEEEEEEETPKKPMPPPQSKAKPS 76
Score = 41.6 bits (96), Expect = 0.047
Identities = 22/61 (36%), Positives = 35/61 (57%)
Query: 288 ERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPS 347
++I++ +E ++E ++ EE E++EEEEEEEEE+E E E TP + P
Sbjct: 9 DQISEEEEEKKKNNDSDEEMKDIDEEEEEEEEEEEEEEEEEKEEEEEEEEEETPKKPMPP 68
Query: 348 P 348
P
Sbjct: 69 P 69
>gb|AAO52309.1| similar to Oryza sativa (japonica cultivar-group). Putative histone
acetyltransferase [Dictyostelium discoideum]
gi|66820374|ref|XP_643810.1| hypothetical protein
DDB0167422 [Dictyostelium discoideum]
gi|60471835|gb|EAL69789.1| hypothetical protein
DDB0167422 [Dictyostelium discoideum]
Length = 639
Score = 48.5 bits (114), Expect = 4e-04
Identities = 24/56 (42%), Positives = 35/56 (61%)
Query: 285 LSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVT 340
L +E +E T +E++EEE+ EE E++EEEEEEEEE+E +EV + T
Sbjct: 82 LEEEMEEDKEEEETEEKDEEEQEEEEEEEEEEEEEEEEEEEEEEDEEIEVEKPIET 137
Score = 43.1 bits (100), Expect = 0.016
Identities = 22/47 (46%), Positives = 29/47 (60%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
K +L +E +E+ EE+D EE E++EEEEEEEEEEE E
Sbjct: 76 KVETEELEEEMEEDKEEEETEEKDEEEQEEEEEEEEEEEEEEEEEEE 122
Score = 42.7 bits (99), Expect = 0.021
Identities = 19/30 (63%), Positives = 25/30 (83%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
E+EEEE+ E+ E++EEEE+EEEEEE VE
Sbjct: 49 EEEEEEEEEKVVEEEEEEEEDEEEEEEKVE 78
Score = 42.0 bits (97), Expect = 0.036
Identities = 21/39 (53%), Positives = 26/39 (65%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTP 341
+E +EEE EE E++EEEEEEEEEEE + EV P
Sbjct: 96 EEKDEEEQEEEEEEEEEEEEEEEEEEEEEEDEEIEVEKP 134
Score = 42.0 bits (97), Expect = 0.036
Identities = 21/54 (38%), Positives = 33/54 (60%)
Query: 284 LLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
++ +E + +E +E+EEEE+ EE +++ EE EEEEEEE +VV E
Sbjct: 9 IMEEEEEDEEVEEEEEEQEEEEEEEEEKEEMVEDEEIEEPEEEEEEEEEKVVEE 62
Score = 40.8 bits (94), Expect = 0.080
Identities = 21/43 (48%), Positives = 26/43 (59%)
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E M ++ EEEE E+ E +EEEEEEEEEEE E E
Sbjct: 80 EELEEEMEEDKEEEETEEKDEEEQEEEEEEEEEEEEEEEEEEE 122
Score = 40.4 bits (93), Expect = 0.10
Identities = 19/41 (46%), Positives = 27/41 (65%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRAD 345
+EEEED E E+++EEEEEEEEE+ V +E + P +
Sbjct: 11 EEEEEDEEVEEEEEEQEEEEEEEEEKEEMVEDEEIEEPEEE 51
Score = 40.4 bits (93), Expect = 0.10
Identities = 19/40 (47%), Positives = 27/40 (67%)
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
T + +E EE+++ EE +D+EE+EEEEEEEE E E
Sbjct: 79 TEELEEEMEEDKEEEETEEKDEEEQEEEEEEEEEEEEEEE 118
Score = 39.7 bits (91), Expect = 0.18
Identities = 19/35 (54%), Positives = 25/35 (71%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E EEEE+ EE ++EEEEEE+EEEE +V E
Sbjct: 46 EEPEEEEEEEEEKVVEEEEEEEEDEEEEEEKVETE 80
Score = 38.9 bits (89), Expect = 0.30
Identities = 18/38 (47%), Positives = 26/38 (68%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTP 341
E+EEE++ E E++EEEEEEEEE+E + E+ P
Sbjct: 11 EEEEEDEEVEEEEEEQEEEEEEEEEKEEMVEDEEIEEP 48
Score = 38.5 bits (88), Expect = 0.40
Identities = 17/44 (38%), Positives = 30/44 (67%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E ++ ++ P+E+EEEE+ + E++EEE+EEEEEE+
Sbjct: 33 EEEKEEMVEDEEIEEPEEEEEEEEEKVVEEEEEEEEDEEEEEEK 76
Score = 38.5 bits (88), Expect = 0.40
Identities = 21/39 (53%), Positives = 27/39 (68%), Gaps = 2/39 (5%)
Query: 304 EDEEEEDPEEPAGEDKEE--EEEEEEEEENVEVVNEVVT 340
EDEE E+PEE E++E+ EEEEEEEE+ E +V T
Sbjct: 41 EDEEIEEPEEEEEEEEEKVVEEEEEEEEDEEEEEEKVET 79
Score = 37.7 bits (86), Expect = 0.67
Identities = 21/39 (53%), Positives = 27/39 (68%), Gaps = 4/39 (10%)
Query: 303 QEDEEEEDPEEPAGEDKE----EEEEEEEEEENVEVVNE 337
+E+EEEE+ +E ED+E EEEEEEEEE+ VE E
Sbjct: 27 EEEEEEEEEKEEMVEDEEIEEPEEEEEEEEEKVVEEEEE 65
Score = 37.0 bits (84), Expect = 1.2
Identities = 21/42 (50%), Positives = 25/42 (59%), Gaps = 7/42 (16%)
Query: 303 QEDEEEEDP-------EEPAGEDKEEEEEEEEEEENVEVVNE 337
+EDEEEE+ EE EDKEEEE EE++EE E E
Sbjct: 67 EEDEEEEEEKVETEELEEEMEEDKEEEETEEKDEEEQEEEEE 108
Score = 37.0 bits (84), Expect = 1.2
Identities = 21/51 (41%), Positives = 29/51 (56%)
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E ++ +E + +E+EEEE E E+ EE EEEEEEEE V E
Sbjct: 13 EEEDEEVEEEEEEQEEEEEEEEEKEEMVEDEEIEEPEEEEEEEEEKVVEEE 63
Score = 35.8 bits (81), Expect = 2.6
Identities = 16/35 (45%), Positives = 25/35 (70%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E++EEE+ E+ E+ EEE EE++EEE E +E
Sbjct: 66 EEEDEEEEEEKVETEELEEEMEEDKEEEETEEKDE 100
Score = 34.7 bits (78), Expect = 5.7
Identities = 18/39 (46%), Positives = 25/39 (63%)
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E ++ +E+EEEE+ EE E E EE EEE EE+ E
Sbjct: 53 EEEEEKVVEEEEEEEEDEEEEEEKVETEELEEEMEEDKE 91
Score = 34.7 bits (78), Expect = 5.7
Identities = 18/37 (48%), Positives = 23/37 (61%)
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
M ++DE+ EE E+ EEEEEE+EEEE E E
Sbjct: 1 MDKQDEDIIMEEEEEDEEVEEEEEEQEEEEEEEEEKE 37
>gb|AAL56647.1| histone acetyltransferase MOZ2 [Homo sapiens]
Length = 2072
Score = 48.5 bits (114), Expect = 4e-04
Identities = 25/55 (45%), Positives = 36/55 (65%), Gaps = 2/55 (3%)
Query: 289 RIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
++ +L KE + +E++EEE+ EE E+ EEEEEEEEEEE E + +PPR
Sbjct: 1059 QLLELSKESSEEEEEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEENIQS--SPPR 1111
Score = 48.1 bits (113), Expect = 5e-04
Identities = 22/41 (53%), Positives = 31/41 (74%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRA 344
E+EEEE+ EE E++EEEEEEEEEEEN++ +T P++
Sbjct: 1077 EEEEEEEEEEEEDEEEEEEEEEEEEEENIQSSPPRLTKPQS 1117
Score = 38.5 bits (88), Expect = 0.40
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 5/53 (9%)
Query: 296 EWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
E + +E+EEEED EE +EEEEEEEE+EE E E SP
Sbjct: 1062 ELSKESSEEEEEEEDEEE-----EEEEEEEEEDEEEEEEEEEEEEEENIQSSP 1109
Score = 36.6 bits (83), Expect = 1.5
Identities = 19/32 (59%), Positives = 24/32 (74%), Gaps = 2/32 (6%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P+E+EEEE+ EE E++EEE EEEE NVE
Sbjct: 1349 PEEEEEEEEEEEE--EEEEEEGEEEEGGGNVE 1378
Score = 34.7 bits (78), Expect = 5.7
Identities = 18/44 (40%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Query: 295 KEWTARMPQ-EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
KE +AR+ E+EEEED E ED + ++E++ E+ EV E
Sbjct: 1398 KEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKE 1441
Score = 34.3 bits (77), Expect = 7.5
Identities = 15/26 (57%), Positives = 21/26 (80%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEE 330
++ E+D +P E++EEEEEEEEEEE
Sbjct: 1340 EKPEDDLIKPEEEEEEEEEEEEEEEE 1365
>gb|AAF00100.1| histone acetyltransferase MORF beta [Homo sapiens]
gi|6912512|ref|NP_036462.1| MYST histone
acetyltransferase (monocytic leukemia) 4 [Homo sapiens]
Length = 2073
Score = 48.5 bits (114), Expect = 4e-04
Identities = 25/55 (45%), Positives = 36/55 (65%), Gaps = 2/55 (3%)
Query: 289 RIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
++ +L KE + +E++EEE+ EE E+ EEEEEEEEEEE E + +PPR
Sbjct: 1060 QLLELSKESSEEEEEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEENIQS--SPPR 1112
Score = 48.1 bits (113), Expect = 5e-04
Identities = 22/41 (53%), Positives = 31/41 (74%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRA 344
E+EEEE+ EE E++EEEEEEEEEEEN++ +T P++
Sbjct: 1078 EEEEEEEEEEEEDEEEEEEEEEEEEEENIQSSPPRLTKPQS 1118
Score = 38.5 bits (88), Expect = 0.40
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 5/53 (9%)
Query: 296 EWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
E + +E+EEEED EE +EEEEEEEE+EE E E SP
Sbjct: 1063 ELSKESSEEEEEEEDEEE-----EEEEEEEEEDEEEEEEEEEEEEEENIQSSP 1110
Score = 36.6 bits (83), Expect = 1.5
Identities = 19/32 (59%), Positives = 24/32 (74%), Gaps = 2/32 (6%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P+E+EEEE+ EE E++EEE EEEE NVE
Sbjct: 1350 PEEEEEEEEEEEE--EEEEEEGEEEEGGGNVE 1379
Score = 34.7 bits (78), Expect = 5.7
Identities = 18/44 (40%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Query: 295 KEWTARMPQ-EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
KE +AR+ E+EEEED E ED + ++E++ E+ EV E
Sbjct: 1399 KEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKE 1442
Score = 34.3 bits (77), Expect = 7.5
Identities = 15/26 (57%), Positives = 21/26 (80%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEE 330
++ E+D +P E++EEEEEEEEEEE
Sbjct: 1341 EKPEDDLIKPEEEEEEEEEEEEEEEE 1366
>gb|AAF00099.1| histone acetyltransferase MORF alpha [Homo sapiens]
Length = 1890
Score = 48.5 bits (114), Expect = 4e-04
Identities = 25/55 (45%), Positives = 36/55 (65%), Gaps = 2/55 (3%)
Query: 289 RIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
++ +L KE + +E++EEE+ EE E+ EEEEEEEEEEE E + +PPR
Sbjct: 877 QLLELSKESSEEEEEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEENIQS--SPPR 929
Score = 48.1 bits (113), Expect = 5e-04
Identities = 22/41 (53%), Positives = 31/41 (74%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRA 344
E+EEEE+ EE E++EEEEEEEEEEEN++ +T P++
Sbjct: 895 EEEEEEEEEEEEDEEEEEEEEEEEEEENIQSSPPRLTKPQS 935
Score = 38.5 bits (88), Expect = 0.40
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 5/53 (9%)
Query: 296 EWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
E + +E+EEEED EE +EEEEEEEE+EE E E SP
Sbjct: 880 ELSKESSEEEEEEEDEEE-----EEEEEEEEEDEEEEEEEEEEEEEENIQSSP 927
Score = 36.6 bits (83), Expect = 1.5
Identities = 19/32 (59%), Positives = 24/32 (74%), Gaps = 2/32 (6%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P+E+EEEE+ EE E++EEE EEEE NVE
Sbjct: 1167 PEEEEEEEEEEEE--EEEEEEGEEEEGGGNVE 1196
Score = 34.7 bits (78), Expect = 5.7
Identities = 18/44 (40%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Query: 295 KEWTARMPQ-EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
KE +AR+ E+EEEED E ED + ++E++ E+ EV E
Sbjct: 1216 KEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKE 1259
Score = 34.3 bits (77), Expect = 7.5
Identities = 15/26 (57%), Positives = 21/26 (80%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEE 330
++ E+D +P E++EEEEEEEEEEE
Sbjct: 1158 EKPEDDLIKPEEEEEEEEEEEEEEEE 1183
>gb|AAF00095.1| histone acetyltransferase MORF [Homo sapiens]
gi|20521021|dbj|BAA20837.2| KIAA0383 [Homo sapiens]
Length = 1781
Score = 48.5 bits (114), Expect = 4e-04
Identities = 25/55 (45%), Positives = 36/55 (65%), Gaps = 2/55 (3%)
Query: 289 RIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
++ +L KE + +E++EEE+ EE E+ EEEEEEEEEEE E + +PPR
Sbjct: 768 QLLELSKESSEEEEEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEENIQS--SPPR 820
Score = 48.1 bits (113), Expect = 5e-04
Identities = 22/41 (53%), Positives = 31/41 (74%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRA 344
E+EEEE+ EE E++EEEEEEEEEEEN++ +T P++
Sbjct: 786 EEEEEEEEEEEEDEEEEEEEEEEEEEENIQSSPPRLTKPQS 826
Score = 38.5 bits (88), Expect = 0.40
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 5/53 (9%)
Query: 296 EWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
E + +E+EEEED EE +EEEEEEEE+EE E E SP
Sbjct: 771 ELSKESSEEEEEEEDEEE-----EEEEEEEEEDEEEEEEEEEEEEEENIQSSP 818
Score = 36.6 bits (83), Expect = 1.5
Identities = 19/32 (59%), Positives = 24/32 (74%), Gaps = 2/32 (6%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P+E+EEEE+ EE E++EEE EEEE NVE
Sbjct: 1058 PEEEEEEEEEEEE--EEEEEEGEEEEGGGNVE 1087
Score = 34.7 bits (78), Expect = 5.7
Identities = 18/44 (40%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Query: 295 KEWTARMPQ-EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
KE +AR+ E+EEEED E ED + ++E++ E+ EV E
Sbjct: 1107 KEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKE 1150
Score = 34.3 bits (77), Expect = 7.5
Identities = 15/26 (57%), Positives = 21/26 (80%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEE 330
++ E+D +P E++EEEEEEEEEEE
Sbjct: 1049 EKPEDDLIKPEEEEEEEEEEEEEEEE 1074
>ref|XP_647315.1| hypothetical protein DDB0202150 [Dictyostelium discoideum]
gi|60475692|gb|EAL73627.1| hypothetical protein
DDB0202150 [Dictyostelium discoideum]
Length = 1354
Score = 48.5 bits (114), Expect = 4e-04
Identities = 24/39 (61%), Positives = 29/39 (73%)
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
M +E+EEEE+ EE E++EEEEEEEEEEE E NE V
Sbjct: 1174 MEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEGNEEV 1212
Score = 47.8 bits (112), Expect = 7e-04
Identities = 22/35 (62%), Positives = 29/35 (82%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EEEE+ EE E++EEEEEEEEEE N EVV++
Sbjct: 1181 EEEEEEEEEEEEEEEEEEEEEEEEEEEGNEEVVDD 1215
Score = 46.6 bits (109), Expect = 0.001
Identities = 23/37 (62%), Positives = 28/37 (75%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
+E+EEEE+ EE E++EEEEEEEEEEE E EVV
Sbjct: 1177 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEGNEEVV 1213
Score = 42.4 bits (98), Expect = 0.027
Identities = 21/35 (60%), Positives = 25/35 (71%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E EEEE+ EE E++EEEEEEEEEEE E E
Sbjct: 1172 EEMEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 1206
Score = 41.6 bits (96), Expect = 0.047
Identities = 20/35 (57%), Positives = 25/35 (71%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+ EEE+ EE E++EEEEEEEEEEE E E
Sbjct: 1171 EEEMEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 1205
Score = 41.6 bits (96), Expect = 0.047
Identities = 20/35 (57%), Positives = 25/35 (71%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EE E+ EE E++EEEEEEEEEEE E E
Sbjct: 1169 EEEEEMEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 1203
Score = 41.6 bits (96), Expect = 0.047
Identities = 20/35 (57%), Positives = 25/35 (71%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+E EE+ EE E++EEEEEEEEEEE E E
Sbjct: 1170 EEEEMEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 1204
Score = 39.7 bits (91), Expect = 0.18
Identities = 20/46 (43%), Positives = 29/46 (62%)
Query: 288 ERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
E ++ +E +E+EEEE+ EE E++EEEEEEE EE V+
Sbjct: 1169 EEEEEMEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEGNEEVVD 1214
Score = 38.9 bits (89), Expect = 0.30
Identities = 19/57 (33%), Positives = 35/57 (61%)
Query: 280 PVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVN 336
P+ PL + + ++ ++ +E+EEEE+ E E++E+EEE EE+EE + + N
Sbjct: 592 PIKPLDINFSLFKKQRKEEKKVIKEEEEEEEEHEEEEEEEEQEEEGEEKEELLTIEN 648
Score = 38.1 bits (87), Expect = 0.52
Identities = 19/34 (55%), Positives = 23/34 (66%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+D EEE+ E E++EEEEEEEEEEE E E
Sbjct: 1166 DDFEEEEEMEEEEEEEEEEEEEEEEEEEEEEEEE 1199
Score = 37.4 bits (85), Expect = 0.88
Identities = 18/33 (54%), Positives = 23/33 (69%)
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
D++ E+ EE E++EEEEEEEEEEE E E
Sbjct: 1165 DDDFEEEEEMEEEEEEEEEEEEEEEEEEEEEEE 1197
Score = 37.0 bits (84), Expect = 1.2
Identities = 18/34 (52%), Positives = 23/34 (66%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+D+ EE+ E E++EEEEEEEEEEE E E
Sbjct: 1165 DDDFEEEEEMEEEEEEEEEEEEEEEEEEEEEEEE 1198
Score = 36.6 bits (83), Expect = 1.5
Identities = 19/37 (51%), Positives = 23/37 (61%)
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
M ED ++D EE ++EEEEEEEEEEE E E
Sbjct: 1158 MRNEDYTDDDFEEEEEMEEEEEEEEEEEEEEEEEEEE 1194
Score = 36.2 bits (82), Expect = 2.0
Identities = 17/37 (45%), Positives = 25/37 (66%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
+E+EEEE EE E++EEE EE+EE +E E++
Sbjct: 617 EEEEEEEHEEEEEEEEQEEEGEEKEELLTIENFKEMI 653
>dbj|BAA85450.1| S-locus protein 1 [Brassica rapa]
Length = 232
Score = 48.5 bits (114), Expect = 4e-04
Identities = 25/44 (56%), Positives = 32/44 (71%), Gaps = 8/44 (18%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE--------NVEVVNEV 338
+++EEEE+ EE AGE++EEEEEEEEEEE V+VVN V
Sbjct: 171 EDEEEEEEEEEEAGEEEEEEEEEEEEEEEEEKEEAVEVDVVNNV 214
Score = 44.7 bits (104), Expect = 0.006
Identities = 22/36 (61%), Positives = 25/36 (69%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
EDEEEE+ EE ++EEEEEEEEEEE E E V
Sbjct: 171 EDEEEEEEEEEEAGEEEEEEEEEEEEEEEEEKEEAV 206
>dbj|BAB47484.1| KIAA1855 protein [Homo sapiens]
Length = 1270
Score = 48.1 bits (113), Expect = 5e-04
Identities = 33/82 (40%), Positives = 40/82 (48%), Gaps = 4/82 (4%)
Query: 280 PVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV- 338
P PL + A +E E+EEEED EE E++EEEEEEEEEEE E V
Sbjct: 667 PQVPLAPVQPQANGKEEEEEEEEDEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEAAAAVA 726
Query: 339 ---VTPPRADPSPAWMEAAFGR 357
V PR+ S E + R
Sbjct: 727 LGEVLGPRSGSSSEGSERSTDR 748
Score = 45.4 bits (106), Expect = 0.003
Identities = 26/66 (39%), Positives = 40/66 (60%), Gaps = 2/66 (3%)
Query: 278 RLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV-- 335
++P+AP+ + + +E +E+E+EE+ EE E++EEEEEEEEEEE V
Sbjct: 668 QVPLAPVQPQANGKEEEEEEEEDEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEAAAAVAL 727
Query: 336 NEVVTP 341
EV+ P
Sbjct: 728 GEVLGP 733
Score = 38.5 bits (88), Expect = 0.40
Identities = 23/57 (40%), Positives = 30/57 (52%), Gaps = 2/57 (3%)
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRM 358
PQ + +EE+ EE ED+EEEEE+EEEEE E E + A A G +
Sbjct: 676 PQANGKEEEEEEE--EDEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEAAAAVALGEV 730
>ref|XP_573479.1| PREDICTED: similar to LRRGT00170 [Rattus norvegicus]
Length = 497
Score = 48.1 bits (113), Expect = 5e-04
Identities = 28/60 (46%), Positives = 36/60 (59%), Gaps = 6/60 (10%)
Query: 274 VIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
VI++ PL K+R Q R +E+EE+E+ EE GE+ EEEEEEEEEEE E
Sbjct: 134 VIQRTNTQLPLHPKQRQPQ------RRQEEEEEEKEEEEEEEGEEGEEEEEEEEEEEEEE 187
Score = 41.2 bits (95), Expect = 0.061
Identities = 18/27 (66%), Positives = 23/27 (84%)
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEE 330
E+EEEE+ EE ED+EE++EEEEEEE
Sbjct: 174 EEEEEEEEEEEEEEDEEEDDEEEEEEE 200
Score = 40.8 bits (94), Expect = 0.080
Identities = 20/39 (51%), Positives = 27/39 (68%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTP 341
+E+EEEE+ EE ED EEEEEEEEE+ + + V +P
Sbjct: 177 EEEEEEEEEEEDEEEDDEEEEEEEEEKRKHKKKHLVQSP 215
Score = 40.8 bits (94), Expect = 0.080
Identities = 17/28 (60%), Positives = 24/28 (85%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E+EEEE+ EE E++++EEEEEEEEE
Sbjct: 175 EEEEEEEEEEEEEDEEEDDEEEEEEEEE 202
Score = 40.4 bits (93), Expect = 0.10
Identities = 19/40 (47%), Positives = 26/40 (64%)
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPP 342
+E+EEEE+ EE E+ +EEEEEEEEE+ +V P
Sbjct: 176 EEEEEEEEEEEEDEEEDDEEEEEEEEEKRKHKKKHLVQSP 215
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 810,024,038
Number of Sequences: 2540612
Number of extensions: 38247436
Number of successful extensions: 607083
Number of sequences better than 10.0: 4159
Number of HSP's better than 10.0 without gapping: 3235
Number of HSP's successfully gapped in prelim test: 1033
Number of HSP's that attempted gapping in prelim test: 453519
Number of HSP's gapped (non-prelim): 54260
length of query: 422
length of database: 863,360,394
effective HSP length: 131
effective length of query: 291
effective length of database: 530,540,222
effective search space: 154387204602
effective search space used: 154387204602
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0388a.2


