

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0383.4
(221 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
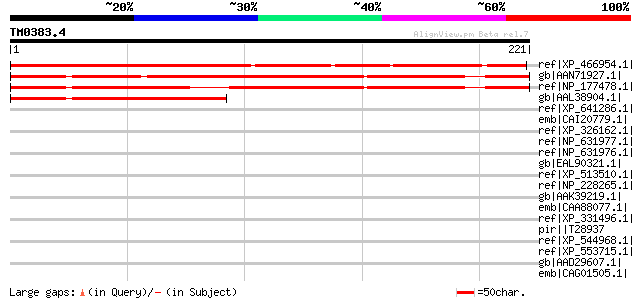
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_466954.1| unknown protein [Oryza sativa (japonica cultiva... 261 1e-68
gb|AAN71927.1| unknown protein [Arabidopsis thaliana] 256 3e-67
ref|NP_177478.1| expressed protein [Arabidopsis thaliana] gi|254... 223 2e-57
gb|AAL38904.1| unknown protein [Arabidopsis thaliana] 171 1e-41
ref|XP_641286.1| hypothetical protein DDB0206422 [Dictyostelium ... 42 0.010
emb|CAI20779.1| novel protein [Danio rerio] 41 0.028
ref|XP_326162.1| hypothetical protein [Neurospora crassa] gi|289... 40 0.048
ref|NP_631977.1| nexilin isoform b [Rattus norvegicus] gi|400352... 39 0.11
ref|NP_631976.1| nexilin isoform s [Rattus norvegicus] gi|400351... 39 0.11
gb|EAL90321.1| hypothetical protein Afu1g09920 [Aspergillus fumi... 39 0.14
ref|XP_513510.1| PREDICTED: similar to nexilin; nF actin binding... 39 0.14
ref|NP_228265.1| ribosomal protein L1 [Thermotoga maritima MSB8]... 38 0.18
gb|AAK39219.1| Hypothetical protein C52B9.8 [Caenorhabditis eleg... 38 0.24
emb|CAA88077.1| Hypothetical protein AH6.3 [Caenorhabditis elega... 38 0.24
ref|XP_331496.1| hypothetical protein [Neurospora crassa] gi|289... 38 0.24
pir||T28937 hypothetical protein C52B9.8 - Caenorhabditis elegans 38 0.24
ref|XP_544968.1| PREDICTED: similar to AF-4 [Canis familiaris] 37 0.31
ref|XP_553715.1| ENSANGP00000027587 [Anopheles gambiae str. PEST... 37 0.31
gb|AAD29607.1| unknown [Homo sapiens] 37 0.31
emb|CAG01505.1| unnamed protein product [Tetraodon nigroviridis] 37 0.31
>ref|XP_466954.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|49388711|dbj|BAD25892.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 215
Score = 261 bits (666), Expect = 1e-68
Identities = 145/220 (65%), Positives = 167/220 (75%), Gaps = 5/220 (2%)
Query: 1 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 60
M+GREVREYTNLSDPKD+KWGKGKDKIDDED+TFQRMVAKMQEVAGERGGYLHGRGALDS
Sbjct: 1 MSGREVREYTNLSDPKDRKWGKGKDKIDDEDITFQRMVAKMQEVAGERGGYLHGRGALDS 60
Query: 61 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIR 120
DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFK+AA L DS+PA VP LRVEPKPKS IR
Sbjct: 61 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKKAAILADSTPA-VPAALRVEPKPKSDIR 119
Query: 121 QQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSL 180
QQDLLK +V IKPKR + S S A +D + + ++ + QS SG +K Q
Sbjct: 120 QQDLLKNIVGIKPKRTK-VSSPSQPAENDKPKQSPEDSVNKV-SSPQSQSGSRKESSQRD 177
Query: 181 SGSTKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDD 220
+ + L ++KP+ N+ G LLGLAY SSD++
Sbjct: 178 GAVSFEKPLLKPVEPRESKPQ--NATGSLLGLAYESSDEE 215
>gb|AAN71927.1| unknown protein [Arabidopsis thaliana]
Length = 208
Score = 256 bits (654), Expect = 3e-67
Identities = 143/221 (64%), Positives = 165/221 (73%), Gaps = 13/221 (5%)
Query: 1 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 60
MAGREV+EYTNLSDPKDKK GKGK IDDEDVTFQRMVAKMQEVAGERGGYLHGRG DS
Sbjct: 1 MAGREVKEYTNLSDPKDKKSGKGK--IDDEDVTFQRMVAKMQEVAGERGGYLHGRG--DS 56
Query: 61 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIR 120
DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFK+AA+ DSSPAS+PLPLRVEPKPKSGIR
Sbjct: 57 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKKAATQADSSPASIPLPLRVEPKPKSGIR 116
Query: 121 QQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSL 180
QQDLL+KVVE+KPKRP+ + S S S V ++R P ++ Q + +++
Sbjct: 117 QQDLLRKVVEVKPKRPKISTPSSPSLSPPVR-SDRGPTEAKVHRDKQKEEAMPVLKKPGA 175
Query: 181 SGSTKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE 221
+ S+G N+ GLLGLAY SSD+++
Sbjct: 176 DRNDGKATESNGQG--------QNAPKGLLGLAYDSSDEED 208
>ref|NP_177478.1| expressed protein [Arabidopsis thaliana] gi|25406262|pir||A96760
hypothetical protein T9L24.52 [imported] - Arabidopsis
thaliana gi|11120800|gb|AAG30980.1| hypothetical protein
[Arabidopsis thaliana]
Length = 194
Score = 223 bits (569), Expect = 2e-57
Identities = 129/221 (58%), Positives = 150/221 (67%), Gaps = 27/221 (12%)
Query: 1 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 60
MAGREV+EYTNLSDPKDKK GKGK IDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS
Sbjct: 1 MAGREVKEYTNLSDPKDKKSGKGK--IDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 58
Query: 61 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIR 120
DDLLYLKEQMEAEEDAE A+ DSSPAS+PLPLRVEPKPKSGIR
Sbjct: 59 DDLLYLKEQMEAEEDAE----------------PATQADSSPASIPLPLRVEPKPKSGIR 102
Query: 121 QQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSL 180
QQDLL+KVVE+KPKRP+ + S S S V ++R P ++ Q + +++
Sbjct: 103 QQDLLRKVVEVKPKRPKISTPSSPSLSPPVR-SDRGPTEAKVHRDKQKEEAMPVLKKPGA 161
Query: 181 SGSTKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE 221
+ S+G N+ GLLGLAY SSD+++
Sbjct: 162 DRNDGKATESNGQG--------QNAPKGLLGLAYDSSDEED 194
>gb|AAL38904.1| unknown protein [Arabidopsis thaliana]
Length = 130
Score = 171 bits (434), Expect = 1e-41
Identities = 88/92 (95%), Positives = 89/92 (96%), Gaps = 2/92 (2%)
Query: 1 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 60
MAGREV+EYTNLSDPKDKK GKGK IDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS
Sbjct: 1 MAGREVKEYTNLSDPKDKKSGKGK--IDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 58
Query: 61 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFK 92
DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFK
Sbjct: 59 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFK 90
>ref|XP_641286.1| hypothetical protein DDB0206422 [Dictyostelium discoideum]
gi|60469321|gb|EAL67315.1| hypothetical protein
DDB0206422 [Dictyostelium discoideum]
Length = 1221
Score = 42.4 bits (98), Expect = 0.010
Identities = 43/215 (20%), Positives = 81/215 (37%), Gaps = 13/215 (6%)
Query: 6 VREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDSDDLLY 65
+ + + + PK K+W K + + + E + G G SDD
Sbjct: 82 LENHPHKNQPKSKQWQKAYN------AAIKYLKNSESEPDSDSDGDYSG-----SDDSNI 130
Query: 66 LKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLL 125
+E ++E++ +T K++ + ++S++++S +S + S L
Sbjct: 131 ESASSSSESESEKV--KTSKKSEKSTSGSSSIINNSSSSSKSSDSKKKSSSSSSSSSSSL 188
Query: 126 KKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTK 185
++ + K+ S K ++ SS V + +K S K E+ L T
Sbjct: 189 SSNLKSESKKSTSSLKSKSNGSSKVKASKPDKTPEPIKTESSSTKSEGKTEKAKLKKETV 248
Query: 186 VEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDD 220
LSS S + K NS G + SSD +
Sbjct: 249 SPTLSSSSSSSSKKKSSTNSSNGKRKIKSISSDSE 283
>emb|CAI20779.1| novel protein [Danio rerio]
Length = 476
Score = 40.8 bits (94), Expect = 0.028
Identities = 38/155 (24%), Positives = 68/155 (43%), Gaps = 26/155 (16%)
Query: 76 AERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLR---------------VEPKPKSGIR 120
AER L E+ +F A VD + S+P+P+R V P+S
Sbjct: 4 AERTLPELERNSFLHCDMAERGVDVNTLSIPVPMRRGGSETCLDLDSDLTVRDGPRSRFG 63
Query: 121 QQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTN--RKPDHGRLKE--------TGQSLS 170
L +K++++K + ++ Q + + + + N K GR+K+ T ++S
Sbjct: 64 LDSLQQKILKVKEQLRIEQNLQDENVAEYLKLINCADKQQSGRIKQVFEKKNQKTAHNIS 123
Query: 171 GVKKVEEQSLSGSTKVEDLSSGSHDADAKPKINNS 205
++K EQ K D+S+G+ A K K N++
Sbjct: 124 QLQKKLEQ-YQRKMKESDISNGTKHATPKDKDNDN 157
>ref|XP_326162.1| hypothetical protein [Neurospora crassa] gi|28924178|gb|EAA33333.1|
hypothetical protein [Neurospora crassa]
Length = 419
Score = 40.0 bits (92), Expect = 0.048
Identities = 40/157 (25%), Positives = 68/157 (42%), Gaps = 26/157 (16%)
Query: 57 ALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPK 116
A+D+D++LY+K ++A+ + ERL + A AA + AAS V + A+ V + K
Sbjct: 111 AIDADEILYVKPPVDAKAEKERLKKEKAAAAAAAAQGAASAVQGAAAT------VVDRTK 164
Query: 117 SGIRQQDLLKKVVEIK----------------PKRPRSESKQSTSASSDVSVTNRKPDHG 160
+ +++K VE+K PK+ + E K + + + P
Sbjct: 165 EAVAA--VVEKAVEVKDQVVQAATDAAPAAAAPKKEKKEKKVNENRRPKATPPPPAPLSP 222
Query: 161 RLKE--TGQSLSGVKKVEEQSLSGSTKVEDLSSGSHD 195
L + G L +K E SL ST + G+ D
Sbjct: 223 GLIDLRVGHILKAIKHPEADSLFVSTIAVGDAPGTED 259
>ref|NP_631977.1| nexilin isoform b [Rattus norvegicus] gi|4003521|gb|AAC95129.1|
s-nexilin [Rattus norvegicus]
Length = 606
Score = 38.9 bits (89), Expect = 0.11
Identities = 35/129 (27%), Positives = 61/129 (47%), Gaps = 6/129 (4%)
Query: 67 KEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKP-KSGIRQQDLL 125
+E+ +AEE+A R + EK+AFA +R+ L D SP + P K I + LL
Sbjct: 254 EEKRKAEEEARRRMEE-EKKAFAEARRSMVLDDDSPEIYKAVSQESLTPGKLEINFEQLL 312
Query: 126 KKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEE---QSLSG 182
++ +E + +R E +Q + + G+ +E +S ++ EE SG
Sbjct: 313 RQKMEEERRRTEEERRQKLEMEKQ-EFEQLRQEMGKEEEENESFGLSREYEELIKLKRSG 371
Query: 183 STKVEDLSS 191
S + ++L S
Sbjct: 372 SIQAKNLKS 380
>ref|NP_631976.1| nexilin isoform s [Rattus norvegicus] gi|4003519|gb|AAC95128.1|
F-actin binding protein b-Nexilin [Rattus norvegicus]
Length = 656
Score = 38.9 bits (89), Expect = 0.11
Identities = 35/129 (27%), Positives = 61/129 (47%), Gaps = 6/129 (4%)
Query: 67 KEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKP-KSGIRQQDLL 125
+E+ +AEE+A R + EK+AFA +R+ L D SP + P K I + LL
Sbjct: 304 EEKRKAEEEARRRMEE-EKKAFAEARRSMVLDDDSPEIYKAVSQESLTPGKLEINFEQLL 362
Query: 126 KKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEE---QSLSG 182
++ +E + +R E +Q + + G+ +E +S ++ EE SG
Sbjct: 363 RQKMEEERRRTEEERRQKLEMEKQ-EFEQLRQEMGKEEEENESFGLSREYEELIKLKRSG 421
Query: 183 STKVEDLSS 191
S + ++L S
Sbjct: 422 SIQAKNLKS 430
>gb|EAL90321.1| hypothetical protein Afu1g09920 [Aspergillus fumigatus Af293]
Length = 247
Score = 38.5 bits (88), Expect = 0.14
Identities = 56/233 (24%), Positives = 96/233 (41%), Gaps = 30/233 (12%)
Query: 16 KDKKWGKGKDKIDDE-----------------DVTFQRMVAKMQEVAGERGGYLHGRGAL 58
+D +W + + ++++E +V Q +AK QE E+ + AL
Sbjct: 17 RDDEWRRAQQELEEERRRKAELGRQEGGKSLYEVLQQNKMAK-QEAFEEKMKLKNQFRAL 75
Query: 59 DSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKR------AASLVDSSPASVPLPLRVE 112
D D++ +L ME+ E +++ F+R A L +SS VP +
Sbjct: 76 DEDEVEFLDSIMESTRAQEAAVKKETAEQLELFRRQREEAEKALLENSSTDVVPSNDGED 135
Query: 113 PKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGV 172
K S R++D K+ + I K+ +S S+ + SS P + GQ + V
Sbjct: 136 WKIPSRKRRRDKSKEFL-IPGKKRKSTSEGAAERSSPSESKAEAPKVQSPSKDGQIPAAV 194
Query: 173 KKVEEQSLSGSTKVEDLSSGSHDADAKPKI-----NNSGGGLLGLAYASSDDD 220
+ + ++ +TK +SS S AK + S LGLA SSD +
Sbjct: 195 EVPQVAEVTKTTKSLGISSTSATQGAKASLPAAPAAKSSTLSLGLAGYSSDSE 247
>ref|XP_513510.1| PREDICTED: similar to nexilin; nF actin binding protein [Pan
troglodytes]
Length = 382
Score = 38.5 bits (88), Expect = 0.14
Identities = 43/194 (22%), Positives = 82/194 (42%), Gaps = 18/194 (9%)
Query: 22 KGKDKIDDEDVTFQRMVAKM-------QEVAGERGGYLHGRGALDSDDLLYLK---EQME 71
+ + ++++E F+ +M Q+ A GY G+ L +++ + E+ +
Sbjct: 98 EARKRLEEEKRAFEEARRQMVNEDEENQDTAKIFKGYRPGKLKLSFEEMERQRREDEKRK 157
Query: 72 AEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKP-KSGIRQQDLLKKVVE 130
AEE+A + + EK+AFA +R + D SP + P K I ++LLK+ +E
Sbjct: 158 AEEEARKRIEE-EKKAFAEARRNMVVDDDSPEMYKTISQEFLTPGKLEINFEELLKQKME 216
Query: 131 IKPKRPRSESKQSTSASSDVSVTNRKPDHG------RLKETGQSLSGVKKVEEQSLSGST 184
+ +R E K DV V + +K + ++ ++ EEQ
Sbjct: 217 EEKRRTEEERKHKLEMEDDVDVRPARKSEAPFTHKVNMKARFEQMAKAREEEEQRRIEEQ 276
Query: 185 KVEDLSSGSHDADA 198
K+ + + DA
Sbjct: 277 KLLRMQFEQREIDA 290
>ref|NP_228265.1| ribosomal protein L1 [Thermotoga maritima MSB8]
gi|48185|emb|CAA77860.1| ribosomal protein L1
[Thermotoga maritima] gi|4980961|gb|AAD35538.1|
ribosomal protein L1 [Thermotoga maritima MSB8]
gi|71061|pir||R5HG1T ribosomal protein L1 - Thermotoga
maritima (strain MSB8) gi|132757|sp|P29393|RL1_THEMA 50S
ribosomal protein L1
Length = 233
Score = 38.1 bits (87), Expect = 0.18
Identities = 35/124 (28%), Positives = 57/124 (45%), Gaps = 12/124 (9%)
Query: 67 KEQMEAEED---AERLLRRTEKRAFAAFKRAASLVDSSPASVPL-----PLRVEPKPKSG 118
KE +EA D AE L+ + EK F F A + D L P + P PKSG
Sbjct: 85 KEALEAGADYVGAEDLVEKIEKEGFLDFDVAIATPDMMRIIGRLGKILGPRGLMPSPKSG 144
Query: 119 IRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQ 178
Q++ + V E K+ R E + + + + V R D+ +LKE ++ +K++ +
Sbjct: 145 TVTQEVAEAVKEF--KKGRIEVRTDKTGNIHIPVGKRSFDNEKLKE--NIIAAIKQIMQM 200
Query: 179 SLSG 182
+G
Sbjct: 201 KPAG 204
>gb|AAK39219.1| Hypothetical protein C52B9.8 [Caenorhabditis elegans]
gi|17551114|ref|NP_508736.1| brahma (XE918)
[Caenorhabditis elegans]
Length = 1336
Score = 37.7 bits (86), Expect = 0.24
Identities = 46/210 (21%), Positives = 85/210 (39%), Gaps = 30/210 (14%)
Query: 26 KIDD---EDVTFQ--RMVAKMQEVAGERGGYLHGRGALDSDDLLYLKEQMEAEED----- 75
K+DD +TF+ R+ K+ G+ H R DSDD KE+ +ED
Sbjct: 1007 KVDDVPAPPLTFKVPRLTIKL---GGDEKKRKHNRSESDSDDNSLKKEKKHRKEDHPKEK 1063
Query: 76 -AERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPK 134
E+ + ++++ + K +++ + + EP K +K+ E PK
Sbjct: 1064 EKEKKKEKEQEKSTDSEKDLKRKIEAPIEKIRIKFSGEPSEK--------YRKINEEPPK 1115
Query: 135 RP-----RSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTKVEDL 189
+ R E K S D + +K H +++ +S KK + + K+ +
Sbjct: 1116 KEHRDKGRKEEKSHKHRSDDDDSSPKKKKH---RDSDESSEKKKKKHKHDSDSALKLREG 1172
Query: 190 SSGSHDADAKPKINNSGGGLLGLAYASSDD 219
S S D + P G G ++ A+++D
Sbjct: 1173 SPLSVDKEKSPMKIRIGQGQPSISLAANED 1202
>emb|CAA88077.1| Hypothetical protein AH6.3 [Caenorhabditis elegans]
gi|1176643|sp|Q09202|YP23_CAEEL Hypothetical protein
AH6.3 in chromosome II gi|17531187|ref|NP_496041.1|
putative nuclear protein, with a coiled coil-4 domain
(2J805) [Caenorhabditis elegans]
Length = 230
Score = 37.7 bits (86), Expect = 0.24
Identities = 35/118 (29%), Positives = 52/118 (43%), Gaps = 13/118 (11%)
Query: 92 KRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVE---IKPKRPRS----ESKQST 144
K AS SP + P +P P+S + Q+ E + PK P+S E+K+
Sbjct: 29 KPTASTESVSPLNEP-----KPPPQSAVPQKPAAPAAEEKAPVDPKDPKSKDVDEAKKPD 83
Query: 145 SASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLS-GSTKVEDLSSGSHDADAKPK 201
A+S S ++KP+ G K S KK EE+ +S + D + DAK K
Sbjct: 84 PANSKKSNKSKKPEKGSKKSKKSEKSKKKKTEEKVMSEDKPERPDKLTEEEKPDAKEK 141
>ref|XP_331496.1| hypothetical protein [Neurospora crassa] gi|28926718|gb|EAA35685.1|
hypothetical protein [Neurospora crassa]
Length = 2300
Score = 37.7 bits (86), Expect = 0.24
Identities = 39/192 (20%), Positives = 87/192 (45%), Gaps = 22/192 (11%)
Query: 17 DKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS-------DDLLYLKEQ 69
D+ + +++++D ++R ++ E A L R ++++ D LL+++E+
Sbjct: 1872 DELMRRHQNEVEDLQTRYERQLSNTTEDAQRTEKNLLDRLSIETSKAQHLQDKLLHVEEK 1931
Query: 70 MEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEP---KPKSGIRQQDLLK 126
+E ++A R + K + A+ ++ S+P VP R+ P + + Q+ L
Sbjct: 1932 LEIAKEAARAAAQAAKASVASTPAGPAVSKSTPNDVPSSERISPQALRESIMVLQEQLQA 1991
Query: 127 KVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTKV 186
+ + I+ + E ++ ++ R + L+E L V+ + Q + G+
Sbjct: 1992 RELRIEELEQQLE---KADPEAETKISKRDDEIIWLRE----LLAVRHSDLQDIIGA--- 2041
Query: 187 EDLSSGSHDADA 198
LSS ++D DA
Sbjct: 2042 --LSSDNYDKDA 2051
>pir||T28937 hypothetical protein C52B9.8 - Caenorhabditis elegans
Length = 1257
Score = 37.7 bits (86), Expect = 0.24
Identities = 46/210 (21%), Positives = 85/210 (39%), Gaps = 30/210 (14%)
Query: 26 KIDD---EDVTFQ--RMVAKMQEVAGERGGYLHGRGALDSDDLLYLKEQMEAEED----- 75
K+DD +TF+ R+ K+ G+ H R DSDD KE+ +ED
Sbjct: 928 KVDDVPAPPLTFKVPRLTIKL---GGDEKKRKHNRSESDSDDNSLKKEKKHRKEDHPKEK 984
Query: 76 -AERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPK 134
E+ + ++++ + K +++ + + EP K +K+ E PK
Sbjct: 985 EKEKKKEKEQEKSTDSEKDLKRKIEAPIEKIRIKFSGEPSEK--------YRKINEEPPK 1036
Query: 135 RP-----RSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTKVEDL 189
+ R E K S D + +K H +++ +S KK + + K+ +
Sbjct: 1037 KEHRDKGRKEEKSHKHRSDDDDSSPKKKKH---RDSDESSEKKKKKHKHDSDSALKLREG 1093
Query: 190 SSGSHDADAKPKINNSGGGLLGLAYASSDD 219
S S D + P G G ++ A+++D
Sbjct: 1094 SPLSVDKEKSPMKIRIGQGQPSISLAANED 1123
>ref|XP_544968.1| PREDICTED: similar to AF-4 [Canis familiaris]
Length = 1541
Score = 37.4 bits (85), Expect = 0.31
Identities = 41/161 (25%), Positives = 63/161 (38%), Gaps = 17/161 (10%)
Query: 52 LHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKR------AFAAFKRAASLVDSSPASV 105
L RGA D+ L K + EAE+D + R EK + + K ++ P+S
Sbjct: 987 LEKRGA-DNSSKLVKKRKAEAEKDHDSKKARLEKETKLQSLSSSPHKESSKTKTPRPSSE 1045
Query: 106 PLPLRVEPKP--KSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRK----PDH 159
P + P P S K + +P+R S Q S S+ S N K P H
Sbjct: 1046 PSKKEMLPPPPVSSSSSSSQKPAKPAQKRPRREADPSSQDASQSASSSKGNHKDSSTPKH 1105
Query: 160 GRLKETGQSLSGVKKVEEQSLSGSTKVEDLSSGSHDADAKP 200
+++ G S E + SG + L + ++KP
Sbjct: 1106 RKVEGKGSGSS----TEHKGSSGDSAYPFLVPSLPNGNSKP 1142
>ref|XP_553715.1| ENSANGP00000027587 [Anopheles gambiae str. PEST]
gi|55236057|gb|EAL39215.1| ENSANGP00000027587 [Anopheles
gambiae str. PEST]
Length = 1446
Score = 37.4 bits (85), Expect = 0.31
Identities = 37/141 (26%), Positives = 61/141 (43%), Gaps = 8/141 (5%)
Query: 53 HGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVE 112
HG L + D +++ + E E +K AFA F L + S L ++
Sbjct: 737 HGITYLPNTDKIFVLREGEISESGTYQELMDKKGAFAEFL-IQHLQEVSEEEEDLD-EIK 794
Query: 113 PKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHG-----RLKETGQ 167
+ ++ + ++LL ++ KR RSES T + D+S N D R E
Sbjct: 795 QQLENSVGGEELLNQLKRSNSKRSRSESTSETGSVKDISRQNSTTDSNTSLRRRTSEKAP 854
Query: 168 SLSGVKKVE-EQSLSGSTKVE 187
++ K +E E+S +GS K E
Sbjct: 855 EVAKTKLIETEKSETGSVKWE 875
>gb|AAD29607.1| unknown [Homo sapiens]
Length = 448
Score = 37.4 bits (85), Expect = 0.31
Identities = 44/184 (23%), Positives = 81/184 (43%), Gaps = 16/184 (8%)
Query: 22 KGKDKIDDEDVTFQRMVAKM-------QEVAGERGGYLHGRGALDSDDLLYLK---EQME 71
+ + ++++E F+ +M Q+ A GY G+ L +++ + E+ +
Sbjct: 41 EARKRLEEEKRAFEEARRQMVNEDEENQDTAKIFKGYRPGKLKLSFEEMERQRREDEKRK 100
Query: 72 AEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKP-KSGIRQQDLLKKVVE 130
AEE+A R + EK+AFA +R + D SP + P K I ++LLK+ +E
Sbjct: 101 AEEEARRRIEE-EKKAFAEARRNMVVDDDSPEMYKTISQEFLTPGKLEINFEELLKQKME 159
Query: 131 IKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEE---QSLSGSTKVE 187
+ +R E K + + G +E ++ K+ EE SGS + +
Sbjct: 160 EEKRRTEEERKHKLEMEKQ-EFEQLRQEMGEEEEENETFGLSKEYEELIKLKRSGSIQAK 218
Query: 188 DLSS 191
+L S
Sbjct: 219 NLKS 222
>emb|CAG01505.1| unnamed protein product [Tetraodon nigroviridis]
Length = 280
Score = 37.4 bits (85), Expect = 0.31
Identities = 40/170 (23%), Positives = 67/170 (38%), Gaps = 34/170 (20%)
Query: 60 SDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGI 119
+D L YL+++M+ +ED +LL+ A +++ P S+ LP R G+
Sbjct: 66 ADRLTYLEQRMQMQEDEIQLLK-------MALADVLKRLNTRPVSLALPPR-----HPGV 113
Query: 120 RQQDLLKKVVEI---------KPKRPRSESKQ----------STSASSDVSVTNRKPDHG 160
LKK + P PRS K T+ S S ++RKP+ G
Sbjct: 114 TSGASLKKSCTLPSSASARNYSPTPPRSGGKSPLGNVKDGPCKTTKSQPTSSSSRKPEEG 173
Query: 161 RLKETGQSLSGVKKVEEQSLSGSTKVEDLS---SGSHDADAKPKINNSGG 207
+ + G ++V ++ + LS S DA ++ GG
Sbjct: 174 SKSKEAAAAVGSRRVTHCKVTMQIYLSPLSRKTGSSEDAKLAARVPTRGG 223
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.307 0.127 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 372,896,497
Number of Sequences: 2540612
Number of extensions: 15413979
Number of successful extensions: 38187
Number of sequences better than 10.0: 340
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 321
Number of HSP's that attempted gapping in prelim test: 38051
Number of HSP's gapped (non-prelim): 427
length of query: 221
length of database: 863,360,394
effective HSP length: 123
effective length of query: 98
effective length of database: 550,865,118
effective search space: 53984781564
effective search space used: 53984781564
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0383.4


