

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0378b.13
(282 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
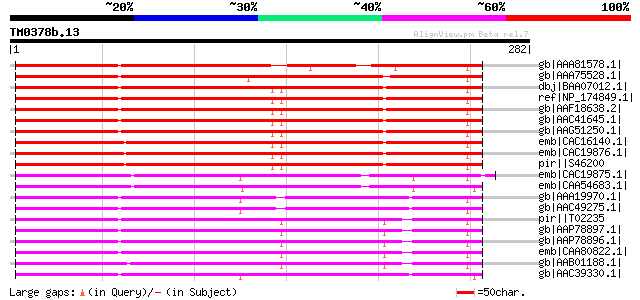
Sequences producing significant alignments: (bits) Value
gb|AAA81578.1| acetyl-CoA carboxylase gi|7438097|pir||T07081 ace... 213 5e-54
gb|AAA75528.1| acetyl CoA carboxylase gi|7438099|pir||T07084 ace... 210 3e-53
dbj|BAA07012.1| acetyl-CoA carboxylase [Arabidopsis thaliana] 200 4e-50
ref|NP_174849.1| acetyl-CoA carboxylase 1 (ACC1) [Arabidopsis th... 200 4e-50
gb|AAF18638.2| F5J5.19 [Arabidopsis thaliana] gi|25293894|pir||D... 200 4e-50
gb|AAC41645.1| acetyl-CoA carboxylase gi|11869927|gb|AAG40563.1|... 200 4e-50
gb|AAG51250.1| acetyl-CoA carboxylase, putative, 5' partial; 1-7... 200 4e-50
emb|CAC16140.1| acetyl coa carboxylase [Brassica napus] 197 2e-49
emb|CAC19876.1| acetyl-CoA carboxylase [Brassica napus] 197 2e-49
pir||S46200 acetyl-CoA carboxylase (EC 6.4.1.2) - rape (fragments) 197 2e-49
emb|CAC19875.1| acetyl-CoA carboxylase [Brassica napus] 182 7e-45
emb|CAA54683.1| acetyl-CoA carboxylase [Brassica napus] gi|74380... 177 2e-43
gb|AAA19970.1| cytosolic acetyl-CoA carboxylase [Triticum aestiv... 170 4e-41
gb|AAC49275.1| acetyl-CoA carboxylase gi|1588584|prf||2208491A A... 170 4e-41
pir||T02235 acetyl-CoA carboxylase (EC 6.4.1.2) - maize gi|85473... 162 1e-38
gb|AAP78897.1| acetyl-coenzyme A carboxylase ACC1B [Zea mays] 162 1e-38
gb|AAP78896.1| acetyl-coenzyme A carboxylase ACC1A [Zea mays] 162 1e-38
emb|CAA80822.1| acetyl CoA carboxylase [Zea mays] gi|7438094|pir... 162 1e-38
gb|AAB01188.1| acetyl CoA carboxylase gi|7438096|pir||T02750 ace... 160 5e-38
gb|AAC39330.1| acetyl-coenzyme A carboxylase [Triticum aestivum]... 159 8e-38
>gb|AAA81578.1| acetyl-CoA carboxylase gi|7438097|pir||T07081 acetyl-CoA
carboxylase (EC 6.4.1.2) B - soybean (fragment)
Length = 1978
Score = 213 bits (542), Expect = 5e-54
Identities = 147/277 (53%), Positives = 167/277 (60%), Gaps = 40/277 (14%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY EEEAAGT LLIDGR LLQNDHDP KL L +LV+ S
Sbjct: 642 LMQLDGNSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLVAETPSKLLRYLVADD-SHV 700
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS II F +SEGQ MQA ELIARLDLDDPSAVRKAEPF
Sbjct: 701 DADTPYAEVEVMKMCMPLLSPASGIIHFTMSEGQAMQAGELIARLDLDDPSAVRKAEPFT 760
Query: 124 SWALLLQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LC-------------KV*SIVYS 170
+L + V Q A S N R + + + + S
Sbjct: 761 GSFPVLGPPTAISGKVHQK--------CAASLNAARMILAGYEHNIDEVVVQSLLNCLDS 812
Query: 171 PEFPFLQW*ECFAVVATRLCKVLATHLPIDLKN*FPSR-----GFQAPKFLTF-AKLLKG 224
PE PFLQW EC AV+ATR LP DLKN S+ G + + + F AKLLKG
Sbjct: 813 PELPFLQWQECLAVLATR--------LPKDLKNELESKYKEFEGISSSQIVDFPAKLLKG 864
Query: 225 NLEAHISSCPDKEKGAQERLK*P----D*SCEGGRET 257
LEAH+SSCPDKEKGAQERL P S EGGRE+
Sbjct: 865 ILEAHLSSCPDKEKGAQERLVEPLLSLVKSYEGGRES 901
>gb|AAA75528.1| acetyl CoA carboxylase gi|7438099|pir||T07084 acetyl-CoA
carboxylase (EC 6.4.1.2) A - soybean
Length = 2261
Score = 210 bits (535), Expect = 3e-53
Identities = 142/264 (53%), Positives = 168/264 (62%), Gaps = 14/264 (5%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY E EAAGT LLIDGR LLQNDHDP KL L +LV+ S
Sbjct: 642 LMQLDGNSHVIYAEGEAAGTRLLIDGRTCLLQNDHDPSKLVAETPCKLLRYLVADD-SHV 700
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF- 122
+ PYAEVEV+KMCMPLLSPAS II F +SEGQ MQA ELIARLDLDDPSAVRKAEPF
Sbjct: 701 DADTPYAEVEVMKMCMPLLSPASGIIHFTMSEGQAMQAGELIARLDLDDPSAVRKAEPFT 760
Query: 123 *SWALL---LQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LCK-V*SIVYSPEFPFLQW 178
S+ +L VK ++ + R L N+ + + + + + SPE PFLQW
Sbjct: 761 GSFPVLGPPTAISVKVHQKCAASLNAARMILSGYEHNIDEVVVQSLLNCLDSPELPFLQW 820
Query: 179 *ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTF-AKLLKGNLEAHISSCPDKE 237
EC AV+ATRL K L L + G + + + F AKLLKG +EAH+SSCPDKE
Sbjct: 821 QECLAVLATRLPKELKNELESKYQE---FEGISSSQIVDFPAKLLKGIIEAHLSSCPDKE 877
Query: 238 KGAQERLK*P----D*SCEGGRET 257
KGAQERL P S EGGRE+
Sbjct: 878 KGAQERLVEPLLSLVKSYEGGRES 901
>dbj|BAA07012.1| acetyl-CoA carboxylase [Arabidopsis thaliana]
Length = 2254
Score = 200 bits (508), Expect = 4e-50
Identities = 139/262 (53%), Positives = 161/262 (61%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +L+S S
Sbjct: 640 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLMRYLISDN-SNI 698
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA LDLDDPSAVRKAEPF
Sbjct: 699 DADTPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIANLDLDDPSAVRKAEPFH 758
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 759 GSFPRLGLPTAISGRVHQRCAATLNAARMILAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 818
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+SSC +KE+G
Sbjct: 819 ECFAVLATRLPKNLRNMLESKYRE-FESISRNSLTTDFPAKLLKGILEAHLSSCDEKERG 877
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 878 ALERLIEPLMSLAKSYEGGRES 899
>ref|NP_174849.1| acetyl-CoA carboxylase 1 (ACC1) [Arabidopsis thaliana]
Length = 2247
Score = 200 bits (508), Expect = 4e-50
Identities = 139/262 (53%), Positives = 161/262 (61%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +L+S S
Sbjct: 640 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLMRYLISDN-SNI 698
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA LDLDDPSAVRKAEPF
Sbjct: 699 DADTPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIANLDLDDPSAVRKAEPFH 758
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 759 GSFPRLGLPTAISGRVHQRCAATLNAARMILAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 818
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+SSC +KE+G
Sbjct: 819 ECFAVLATRLPKNLRNMLESKYRE-FESISRNSLTTDFPAKLLKGILEAHLSSCDEKERG 877
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 878 ALERLIEPLMSLAKSYEGGRES 899
>gb|AAF18638.2| F5J5.19 [Arabidopsis thaliana] gi|25293894|pir||D86483 protein
F5J5.19 [imported] - Arabidopsis thaliana
Length = 2257
Score = 200 bits (508), Expect = 4e-50
Identities = 139/262 (53%), Positives = 161/262 (61%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +L+S S
Sbjct: 640 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLMRYLISDN-SNI 698
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA LDLDDPSAVRKAEPF
Sbjct: 699 DADTPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIANLDLDDPSAVRKAEPFH 758
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 759 GSFPRLGLPTAISGRVHQRCAATLNAARMILAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 818
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+SSC +KE+G
Sbjct: 819 ECFAVLATRLPKNLRNMLESKYRE-FESISRNSLTTDFPAKLLKGILEAHLSSCDEKERG 877
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 878 ALERLIEPLMSLAKSYEGGRES 899
>gb|AAC41645.1| acetyl-CoA carboxylase gi|11869927|gb|AAG40563.1| acetyl-CoA
carboxylase 1 [Arabidopsis thaliana]
gi|1090217|prf||2018327A Ac-CoA carboxylase
Length = 2254
Score = 200 bits (508), Expect = 4e-50
Identities = 139/262 (53%), Positives = 161/262 (61%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +L+S S
Sbjct: 640 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLMRYLISDN-SNI 698
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA LDLDDPSAVRKAEPF
Sbjct: 699 DADTPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIANLDLDDPSAVRKAEPFH 758
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 759 GSFPRLGLPTAISGRVHQRCAATLNAARMILAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 818
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+SSC +KE+G
Sbjct: 819 ECFAVLATRLPKNLRNMLESKYRE-FESISRNSLTTDFPAKLLKGILEAHLSSCDEKERG 877
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 878 ALERLIEPLMSLAKSYEGGRES 899
>gb|AAG51250.1| acetyl-CoA carboxylase, putative, 5' partial; 1-7710 [Arabidopsis
thaliana]
Length = 1865
Score = 200 bits (508), Expect = 4e-50
Identities = 139/262 (53%), Positives = 161/262 (61%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +L+S S
Sbjct: 258 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLMRYLISDN-SNI 316
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA LDLDDPSAVRKAEPF
Sbjct: 317 DADTPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIANLDLDDPSAVRKAEPFH 376
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 377 GSFPRLGLPTAISGRVHQRCAATLNAARMILAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 436
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+SSC +KE+G
Sbjct: 437 ECFAVLATRLPKNLRNMLESKYRE-FESISRNSLTTDFPAKLLKGILEAHLSSCDEKERG 495
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 496 ALERLIEPLMSLAKSYEGGRES 517
>emb|CAC16140.1| acetyl coa carboxylase [Brassica napus]
Length = 796
Score = 197 bits (502), Expect = 2e-49
Identities = 138/262 (52%), Positives = 161/262 (60%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +LVS S
Sbjct: 320 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLLRYLVSDNSSI- 378
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA+LDLDDPSAVRKAEPF
Sbjct: 379 DADMPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIAKLDLDDPSAVRKAEPFH 438
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 439 GGFPRLGLPTAISGKVHQRCAATLNAARMVLAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 498
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+ SC +K++G
Sbjct: 499 ECFAVLATRLPKDLRMMLESKYRE-FESISRNSLTADFPAKLLKGILEAHLLSCDEKDRG 557
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 558 ALERLIEPLMSLAKSYEGGRES 579
>emb|CAC19876.1| acetyl-CoA carboxylase [Brassica napus]
Length = 1798
Score = 197 bits (502), Expect = 2e-49
Identities = 138/262 (52%), Positives = 161/262 (60%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +LVS S
Sbjct: 637 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLLRYLVSDNSSI- 695
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA+LDLDDPSAVRKAEPF
Sbjct: 696 DADMPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIAKLDLDDPSAVRKAEPFH 755
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 756 GGFPRLGLPTAISGKVHQRCAATLNAARMVLAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 815
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+ SC +K++G
Sbjct: 816 ECFAVLATRLPKDLRMMLESKYRE-FESISRNSLTADFPAKLLKGILEAHLLSCDEKDRG 874
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 875 ALERLIEPLMSLAKSYEGGRES 896
>pir||S46200 acetyl-CoA carboxylase (EC 6.4.1.2) - rape (fragments)
Length = 1561
Score = 197 bits (502), Expect = 2e-49
Identities = 138/262 (52%), Positives = 161/262 (60%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +LVS S
Sbjct: 320 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLLRYLVSDNSSI- 378
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA+LDLDDPSAVRKAEPF
Sbjct: 379 DADMPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIAKLDLDDPSAVRKAEPFH 438
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 439 GGFPRLGLPTAISGKVHQRCAATLNAARMVLAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 498
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+ SC +K++G
Sbjct: 499 ECFAVLATRLPKDLRMMLESKYRE-FESISRNSLTADFPAKLLKGILEAHLLSCDEKDRG 557
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 558 ALERLIEPLMSLAKSYEGGRES 579
>emb|CAC19875.1| acetyl-CoA carboxylase [Brassica napus]
Length = 2321
Score = 182 bits (463), Expect = 7e-45
Identities = 129/272 (47%), Positives = 160/272 (58%), Gaps = 18/272 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY +EE +GT LLIDG+ LLQN+HDP KL L +LVS S
Sbjct: 736 LMQLDGKSHVIYAQEETSGTRLLIDGKTCLLQNEHDPSKLMAETPCKLVRYLVSDDSSID 795
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ M ELIA LDL DPS VRKAEPF
Sbjct: 796 ADT-PYAEVEVMKMCMPLLSPASGVIHFKMSEGQVMLPGELIANLDLTDPSTVRKAEPFH 854
Query: 124 S----WALLLQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + K ++ R L + + + S + SPE PFLQW
Sbjct: 855 GRFPRLGLPTEISAKVHKRCAATLNAARMILAGYEHQVDEVVQDLVSCLDSPELPFLQWQ 914
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTF---AKLLKGNLEAHISSCPDK 236
ECFAV+ATRL K L I L++ + + LT AKLLKG L+AH++SC +
Sbjct: 915 ECFAVLATRLPK----DLRIMLESKYMEYECISRNSLTADFPAKLLKGILKAHVASCDEN 970
Query: 237 EKGAQERLK*P----D*SCEGGRETCTYNCSI 264
E+GA ERL P S EGGRE ++ C+I
Sbjct: 971 ERGALERLIEPLMSLANSYEGGRE--SHACAI 1000
>emb|CAA54683.1| acetyl-CoA carboxylase [Brassica napus] gi|7438090|pir||T07920
probable acetyl-CoA carboxylase (EC 6.4.1.2) - rape
Length = 2304
Score = 177 bits (450), Expect = 2e-43
Identities = 126/265 (47%), Positives = 153/265 (57%), Gaps = 16/265 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY +EE +GT LLIDG+ LLQN+HDP KL L +LVS S
Sbjct: 736 LMQLDGKSHVIYAQEETSGTRLLIDGKTCLLQNEHDPSKLMAETPCKLLRYLVSDDSSID 795
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+ EGQ M ELIA LDL DPS VRKAEPF
Sbjct: 796 ADT-PYAEVEVMKMCMPLLSPASGVIHFKMCEGQVMLPGELIANLDLADPSTVRKAEPFH 854
Query: 124 SW----ALLLQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + K ++ R L + + S + SPE PFLQW
Sbjct: 855 GGFPRLGLPTEISAKVHQRCAATLDAARMILAGYEHQVDEVVQDFVSCLDSPELPFLQWQ 914
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTF---AKLLKGNLEAHISSCPDK 236
ECFAV+ATRL K L I L++ + + LT AKLLKG LEAH++SC +
Sbjct: 915 ECFAVLATRLPK----DLRIMLESKYMEYECISRNSLTADFPAKLLKGILEAHVASCDET 970
Query: 237 EKGAQERLK*PD*SC----EGGRET 257
E+GA RL P S EGGRE+
Sbjct: 971 ERGALARLIEPLMSLAKCYEGGRES 995
>gb|AAA19970.1| cytosolic acetyl-CoA carboxylase [Triticum aestivum]
gi|2130099|pir||A57710 acetyl-CoA carboxylase (EC
6.4.1.2) - wheat
Length = 2257
Score = 170 bits (431), Expect = 4e-41
Identities = 120/266 (45%), Positives = 156/266 (58%), Gaps = 18/266 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY E EAAGT LLI+GR LLQ +HDP +L L +LV+ S
Sbjct: 629 LMQLDGNSHVIYAETEAAGTRLLINGRTCLLQKEHDPSRLLADTPCKLLRFLVADG-SHV 687
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
PYAEVEV+KMCMPLL PAS +I F + EGQ MQA +LIARLDLDDPS+VR+AEPF
Sbjct: 688 VADTPYAEVEVMKMCMPLLLPASGVIHFVMPEGQAMQASDLIARLDLDDPSSVRRAEPFH 747
Query: 124 S--------WALLLQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LCKV*SIVYSPEFPF 175
A+ + KF +V A ++ L N+ + + + + SPE PF
Sbjct: 748 GTFPKLGPPTAISGKVHQKFAASVNSAHMI----LAGYEHNINHVVQDLLNCLDSPELPF 803
Query: 176 LQW*ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPD 235
LQW E +V+ATRL K L L K + F+ K AKLL+G +EA+++ C +
Sbjct: 804 LQWQELMSVLATRLPKDLRNELDAKYKEYELNADFRKSKDFP-AKLLRGVIEANLAYCSE 862
Query: 236 KEKGAQERLK*P----D*SCEGGRET 257
K++ ERL P S EGGRE+
Sbjct: 863 KDRVTSERLVEPLMSLVKSYEGGRES 888
>gb|AAC49275.1| acetyl-CoA carboxylase gi|1588584|prf||2208491A Ac-CoA carboxylase
Length = 2260
Score = 170 bits (431), Expect = 4e-41
Identities = 120/266 (45%), Positives = 156/266 (58%), Gaps = 18/266 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY E EAAGT LLI+GR LLQ +HDP +L L +LV+ S
Sbjct: 632 LMQLDGNSHVIYAETEAAGTRLLINGRTCLLQKEHDPSRLLADTPCKLLRFLVADG-SHV 690
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
PYAEVEV+KMCMPLL PAS +I F + EGQ MQA +LIARLDLDDPS+VR+AEPF
Sbjct: 691 VADTPYAEVEVMKMCMPLLLPASGVIHFVMPEGQAMQASDLIARLDLDDPSSVRRAEPFH 750
Query: 124 S--------WALLLQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LCKV*SIVYSPEFPF 175
A+ + KF +V A ++ L N+ + + + + SPE PF
Sbjct: 751 GTFPKLGPPTAISGKVHQKFAASVNSAHMI----LAGYEHNINHVVQDLLNCLDSPELPF 806
Query: 176 LQW*ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPD 235
LQW E +V+ATRL K L L K + F+ K AKLL+G +EA+++ C +
Sbjct: 807 LQWQELMSVLATRLPKDLRNELDAKYKEYELNADFRKSKDFP-AKLLRGVIEANLAYCSE 865
Query: 236 KEKGAQERLK*P----D*SCEGGRET 257
K++ ERL P S EGGRE+
Sbjct: 866 KDRVTSERLVEPLMSLVKSYEGGRES 891
>pir||T02235 acetyl-CoA carboxylase (EC 6.4.1.2) - maize
gi|854731|gb|AAA80214.1| acetyl-coenzyme A carboxylase
Length = 2325
Score = 162 bits (410), Expect = 1e-38
Identities = 116/266 (43%), Positives = 152/266 (56%), Gaps = 18/266 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY EEEA GT L IDG+ LLQNDHDP KL L +LV+ +
Sbjct: 735 LMQLDGNSHVIYAEEEAGGTRLQIDGKTCLLQNDHDPSKLLAETPCKLLRFLVADG-AHV 793
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I +SEGQ +QA +LIARLDLDDPSAV++AEPF
Sbjct: 794 DADVPYAEVEVMKMCMPLLSPASGVIHCMMSEGQALQAGDLIARLDLDDPSAVKRAEPFD 853
Query: 124 SWALLLQFLVKFNRNVPQA*ILH----R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
++ V + V + R L N+ + + + +PE PFLQW
Sbjct: 854 GIFPQMELPVAVSSQVHKRYAASLNAARMVLAGYEHNINEVVQDLVCCLDNPELPFLQWD 913
Query: 180 ECFAVVATRLCKVLATHLPIDLK----N*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPD 235
E +V+ATRL + L + L K N + + P +KLL+ +E ++S +
Sbjct: 914 ELMSVLATRLPRNLKSELEDKYKEYKLNFYHGKNEDFP-----SKLLRDIIEENLSYGSE 968
Query: 236 KEKGAQERLK*P----D*SCEGGRET 257
KEK ERL P S EGGRE+
Sbjct: 969 KEKATNERLVEPLMNLLKSYEGGRES 994
>gb|AAP78897.1| acetyl-coenzyme A carboxylase ACC1B [Zea mays]
Length = 2325
Score = 162 bits (410), Expect = 1e-38
Identities = 116/266 (43%), Positives = 152/266 (56%), Gaps = 18/266 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY EEEA GT L IDG+ LLQNDHDP KL L +LV+ +
Sbjct: 735 LMQLDGNSHVIYAEEEAGGTRLQIDGKTCLLQNDHDPSKLLAETPCKLLRFLVADG-AHV 793
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I +SEGQ +QA +LIARLDLDDPSAV++AEPF
Sbjct: 794 DADVPYAEVEVMKMCMPLLSPASGVIHCMMSEGQALQAGDLIARLDLDDPSAVKRAEPFD 853
Query: 124 SWALLLQFLVKFNRNVPQA*ILH----R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
++ V + V + R L N+ + + + +PE PFLQW
Sbjct: 854 GIFPQMELPVAVSSQVHKRYAASLNAARMVLAGYEHNINEVVQDLVCCLDNPELPFLQWD 913
Query: 180 ECFAVVATRLCKVLATHLPIDLK----N*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPD 235
E +V+ATRL + L + L K N + + P +KLL+ +E ++S +
Sbjct: 914 ELMSVLATRLPRNLKSELEDKYKEYKLNFYHGKNEDFP-----SKLLRDIIEENLSYGSE 968
Query: 236 KEKGAQERLK*P----D*SCEGGRET 257
KEK ERL P S EGGRE+
Sbjct: 969 KEKATNERLVEPLMNLLKSYEGGRES 994
>gb|AAP78896.1| acetyl-coenzyme A carboxylase ACC1A [Zea mays]
Length = 2324
Score = 162 bits (410), Expect = 1e-38
Identities = 116/266 (43%), Positives = 152/266 (56%), Gaps = 18/266 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY EEEA GT L IDG+ LLQNDHDP KL L +LV+ +
Sbjct: 735 LMQLDGNSHVIYAEEEAGGTRLQIDGKTCLLQNDHDPSKLLAETPCKLLRFLVADG-AHV 793
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I +SEGQ +QA +LIARLDLDDPSAV++AEPF
Sbjct: 794 DADVPYAEVEVMKMCMPLLSPASGVIHCMMSEGQALQAGDLIARLDLDDPSAVKRAEPFD 853
Query: 124 SWALLLQFLVKFNRNVPQA*ILH----R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
++ V + V + R L N+ + + + +PE PFLQW
Sbjct: 854 GIFPQMELPVAVSSQVHKRYAASLNAARMVLAGYEHNINEVVQDLVCCLDNPELPFLQWD 913
Query: 180 ECFAVVATRLCKVLATHLPIDLK----N*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPD 235
E +V+ATRL + L + L K N + + P +KLL+ +E ++S +
Sbjct: 914 ELMSVLATRLPRNLKSELEDKYKEYKLNFYHGKNEDFP-----SKLLRDIIEENLSYGSE 968
Query: 236 KEKGAQERLK*P----D*SCEGGRET 257
KEK ERL P S EGGRE+
Sbjct: 969 KEKATNERLVEPLMNLLKSYEGGRES 994
>emb|CAA80822.1| acetyl CoA carboxylase [Zea mays] gi|7438094|pir||T02921 acetyl-CoA
carboxylase (EC 6.4.1.2) (clone A3) - maize (fragment)
Length = 1625
Score = 162 bits (410), Expect = 1e-38
Identities = 116/266 (43%), Positives = 152/266 (56%), Gaps = 18/266 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY EEEA GT L IDG+ LLQNDHDP KL L +LV+ +
Sbjct: 36 LMQLDGNSHVIYAEEEAGGTRLQIDGKTCLLQNDHDPSKLLAETPCKLLRFLVADG-AHV 94
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I +SEGQ +QA +LIARLDLDDPSAV++AEPF
Sbjct: 95 DADVPYAEVEVMKMCMPLLSPASGVIHCMMSEGQALQAGDLIARLDLDDPSAVKRAEPFD 154
Query: 124 SWALLLQFLVKFNRNVPQA*ILH----R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
++ V + V + R L N+ + + + +PE PFLQW
Sbjct: 155 GIFPQMELPVAVSSQVHKRYAASLNAARMVLAGYEHNINEVVQDLVCCLDNPELPFLQWD 214
Query: 180 ECFAVVATRLCKVLATHLPIDLK----N*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPD 235
E +V+ATRL + L + L K N + + P +KLL+ +E ++S +
Sbjct: 215 ELMSVLATRLPRNLKSELEDKYKEYKLNFYHGKNEDFP-----SKLLRDIIEENLSYGSE 269
Query: 236 KEKGAQERLK*P----D*SCEGGRET 257
KEK ERL P S EGGRE+
Sbjct: 270 KEKATNERLVEPLMNLLKSYEGGRES 295
>gb|AAB01188.1| acetyl CoA carboxylase gi|7438096|pir||T02750 acetyl-CoA
carboxylase (EC 6.4.1.2) - maize (fragment)
Length = 1685
Score = 160 bits (404), Expect = 5e-38
Identities = 115/266 (43%), Positives = 150/266 (56%), Gaps = 18/266 (6%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY EEEA GT L I+G+ LLQNDHDP KL L +LV+
Sbjct: 95 LMQLDGNSHVIYAEEEAGGTRLQINGKTCLLQNDHDPSKLLAETPCKLLRFLVADGAHVG 154
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
PYAEVEV+KMCMPLLSPAS +I +SEGQ +QA +LIARLDLDDPSAV++AEPF
Sbjct: 155 A-DVPYAEVEVMKMCMPLLSPASGVIHCMMSEGQALQAGDLIARLDLDDPSAVKRAEPFD 213
Query: 124 SWALLLQFLVKFNRNVPQA*ILH----R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L+ V + V + R L N+ + + + +PE PFLQW
Sbjct: 214 GMFPLMDLPVAASSQVHKRYAASLNAARMVLAGYEHNINEVVQDLVCCLDNPELPFLQWD 273
Query: 180 ECFAVVATRLCKVLATHLPIDLK----N*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPD 235
E +V+ATRL + L + L K N + + P +KLL+ +E +++ +
Sbjct: 274 ELMSVLATRLPRNLKSELEDKYKEYKLNFYHGKNKDFP-----SKLLRDIVEENLAYGSE 328
Query: 236 KEKGAQERLK*P----D*SCEGGRET 257
KEK ERL P S EGGRE+
Sbjct: 329 KEKATNERLVEPLMNLLKSYEGGRES 354
>gb|AAC39330.1| acetyl-coenzyme A carboxylase [Triticum aestivum]
gi|7438101|pir||T06161 acetyl-CoA carboxylase (EC
6.4.1.2) - wheat
Length = 2311
Score = 159 bits (402), Expect = 8e-38
Identities = 112/262 (42%), Positives = 153/262 (57%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDGN+HVIY EEEA GT LLIDG+ LLQNDHDP +L L +LV+ +
Sbjct: 735 LMQLDGNSHVIYAEEEAGGTRLLIDGKTCLLQNDHDPSRLLAETPCKLLRFLVADG-AHV 793
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
E PYAEVEV+KMCMPLLSPA+ +I +SEGQ MQA +LIARLDLDDPSAV++AEPF
Sbjct: 794 EADVPYAEVEVMKMCMPLLSPAAGVINVLLSEGQPMQAGDLIARLDLDDPSAVKRAEPFN 853
Query: 124 S----WALLLQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
+L + + ++ + R L + + + + S + +PE PFLQW
Sbjct: 854 GSFPEMSLPIAASGQVHKRCATSLNAARMVLAGYDHPINKVVQDLVSCLDAPELPFLQWE 913
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
E +V+ATRL ++L + L + G K +K+L+ +E +++ +KE
Sbjct: 914 ELMSVLATRLPRLLKSELEGKYSEYKLNVGHGKSKDFP-SKMLREIIEENLAHGSEKEIA 972
Query: 240 AQERLK*PD*SC----EGGRET 257
ERL P S EGGRE+
Sbjct: 973 TNERLVEPLMSLLKSYEGGRES 994
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.359 0.161 0.584
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 404,853,087
Number of Sequences: 2540612
Number of extensions: 14360157
Number of successful extensions: 49451
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 102
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 49131
Number of HSP's gapped (non-prelim): 178
length of query: 282
length of database: 863,360,394
effective HSP length: 126
effective length of query: 156
effective length of database: 543,243,282
effective search space: 84745951992
effective search space used: 84745951992
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0378b.13


