

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.3
(975 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
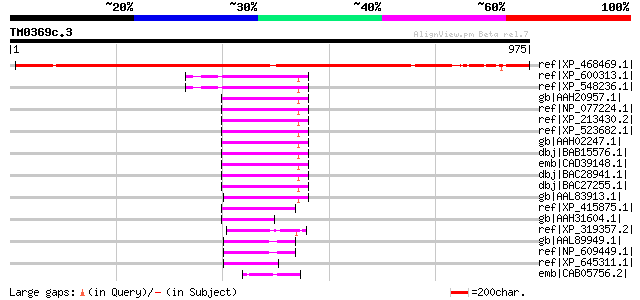
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_468469.1| unknown protein [Oryza sativa (japonica cultiva... 734 0.0
ref|XP_600313.1| PREDICTED: similar to hypothetical protein FLJ1... 100 2e-19
ref|XP_548236.1| PREDICTED: similar to hypothetical protein FLJ1... 100 3e-19
gb|AAH20957.1| FLJ10587 protein [Homo sapiens] gi|7022715|dbj|BA... 99 5e-19
ref|NP_077224.1| hypothetical protein LOC74133 [Mus musculus] gi... 99 5e-19
ref|XP_213430.2| PREDICTED: similar to RIKEN cDNA 1200011M11 [Ra... 99 5e-19
ref|XP_523682.1| PREDICTED: similar to hypothetical protein FLJ1... 99 5e-19
gb|AAH02247.1| 1200011M11Rik protein [Mus musculus] 99 5e-19
dbj|BAB15576.1| unnamed protein product [Homo sapiens] 99 5e-19
emb|CAD39148.1| hypothetical protein [Homo sapiens] gi|47777319|... 99 5e-19
dbj|BAC28941.1| unnamed protein product [Mus musculus] 99 5e-19
dbj|BAC27255.1| unnamed protein product [Mus musculus] 99 5e-19
gb|AAL83913.1| amplified in breast cancer 2 [Homo sapiens] 96 4e-18
ref|XP_415875.1| PREDICTED: similar to RIKEN cDNA 1200011M11 [Ga... 95 1e-17
gb|AAH31604.1| Unknown (protein for IMAGE:5178217) [Homo sapiens] 81 2e-13
ref|XP_319357.2| ENSANGP00000012357 [Anopheles gambiae str. PEST... 79 7e-13
gb|AAL89949.1| SD08609p [Drosophila melanogaster] 71 2e-10
ref|NP_609449.1| CG6729-PA [Drosophila melanogaster] gi|7297754|... 70 4e-10
ref|XP_645311.1| hypothetical protein DDB0217022 [Dictyostelium ... 59 8e-07
emb|CAB05756.2| Hypothetical protein K04B12.3 [Caenorhabditis el... 56 7e-06
>ref|XP_468469.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|48717085|dbj|BAD22858.1| unknown protein [Oryza sativa
(japonica cultivar-group)] gi|48716313|dbj|BAD22926.1|
unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 1154
Score = 734 bits (1895), Expect = 0.0
Identities = 435/977 (44%), Positives = 605/977 (61%), Gaps = 55/977 (5%)
Query: 12 VCHVIIYVQEGSRFDTRVLRNFRVLQAAKHAMAPFAKSQTMPPLPSR-SHSSLSSRPTSS 70
VCHVII++QEG RFDT++L+ FR+LQ++KHA+APF KS P +PS+ + S+ ++PT
Sbjct: 118 VCHVIIFLQEGFRFDTQILKKFRLLQSSKHAIAPFVKSLVAPAVPSKVARSNTPTKPTHR 177
Query: 71 ANNSSPGRGGGGNVSRNGSAMSLMSGLGSYASLFPGQCIPVTLFVFVDDFSNLSNSSTNG 130
A++ SP GG R+ SA+SLMSG GS+ + PG CIPV LFVF DD ++ + T+
Sbjct: 178 ASSISPPARRGG---RHPSAISLMSGTGSHPCMLPGLCIPVVLFVFEDDITDAPGAPTSP 234
Query: 131 EESSDVSSLSQSNSMSSAAKANLPAKVSGSVVMLARPASRSEGGFRKKLQASLEAQIRFL 190
++++D SS +Q+++ K N+ +K S SVVMLARPA RS+G F KKL +S+E QIRFL
Sbjct: 235 DDTNDTSS-NQASNTDGLPKPNMTSKGSSSVVMLARPAIRSDGTFSKKLHSSVEGQIRFL 293
Query: 191 IKKCHTLSGSEITHSGVRTGGTSTSAPLFSLDASRAVVLLDRFSNLKGESLEFATGLVED 250
+KKC TL G E H R + PLFSLD SR V LLDR + K E L+ GL ED
Sbjct: 294 LKKCRTLVGLEPGHIVSRGVSNVSHLPLFSLDTSRVVALLDRSISKKREPLDIIAGLFED 353
Query: 251 VLNGKATSDSLLLESHGQSANKEDLISVKEFIYRQSDILRGRGG-IVNSNSGSAAGVGMV 309
L K++ D LE++ A ED+ +K+FI+RQSD LRGRGG N+ +G +GVGMV
Sbjct: 354 SLTSKSSLDVSSLENNCHPATHEDVQFIKDFIFRQSDGLRGRGGHSSNTTAGPVSGVGMV 413
Query: 310 AVAAAAAAASAASGKTFTTPDLPNFEIWSSSSHHILSRILCAKGGCLDELEIIKRKPRPR 369
A AAAAAAASAASGK + PDLP F+ W S S ILS + + G L + +K P
Sbjct: 414 AAAAAAAAASAASGKQMSAPDLPTFDTWLSISSSILSALFSGEDG-LSSSQNMKASPTHT 472
Query: 370 NTVSSSTEGPLKSANPLDVAVSWLQCGRGLNTKFSTLWCQRAIPAAKEIYLKDLPDCYPT 429
++ + + P +N + A+S L+ +GLN KFS+ WCQR +PAAKE+YLKDLP YPT
Sbjct: 473 SSFPKNDQLPSAGSNAIQTALSCLEGNKGLNVKFSSSWCQRILPAAKEVYLKDLPAFYPT 532
Query: 430 SQHEDHLDKALHAFHSMVKGPSVQVFARKLEEECTSIWKSGRQLCDAISLTGKPCMHQRH 489
S HE L KAL +FHSMVKGP+VQVF++KL++EC +IW+SGRQ CDA+SLTG+PC HQRH
Sbjct: 533 SMHEVQLQKALRSFHSMVKGPAVQVFSKKLKDECQAIWESGRQQCDAVSLTGRPCKHQRH 592
Query: 490 DVETSNSNIGASPPSHK--PHSSGYFFLHACACGRSRQLRPDPFDFESADA--SCFSDCD 545
G S PS HSSGY FLHACACGRSR+LR DPFDFE+A+ +CFS+C+
Sbjct: 593 ---------GKSSPSDAALQHSSGYVFLHACACGRSRRLRDDPFDFEAANMTFNCFSNCE 643
Query: 546 KLLPSVKLPEIEVAGPVQSSAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEK 605
LLP++ LP AG S+W L+R+G A+YY+ +KGLLQ+GFC+ EK LL+WT+ L K
Sbjct: 644 DLLPTLVLPRETNAGAFPVSSWRLVRLGGARYYKPTKGLLQAGFCSKEKYLLRWTISLGK 703
Query: 606 QKRPNGSTETIVKQSSVIR-EPKVEHI-ADTKKTGVRKPHPAVQNGG-GDQRTSLGISKD 662
+ +G+ T S+ +P+ I A K+ V + +++ + R +
Sbjct: 704 GQGKHGTHATNKPFSTASNADPQAPPIVAGEVKSAVTQVTAEIKSMKLENSRKQPEVESM 763
Query: 663 DDKKISFGRGFPIFKMRKPFSEVVAGSAAVDSGFPPLQQRKLPASGSEKGMKLSKPSNQN 722
++ I+FG+G P F M+KPF+EVVAG A DS FP LQQ++ G+ K + ++Q
Sbjct: 764 NNSSINFGKGLPNFTMKKPFAEVVAGHTARDSEFPALQQKRPLKPGNWKDERQVSGADQT 823
Query: 723 GERVNAAVDYEISQKSQDILLTEGSLDGSGNNSCRDDDPFLRIGSNVVPVYLSGGERSKP 782
R + A+ ++ ++ +GS PFL+IGSN+VP+ + G E +
Sbjct: 824 NGRGHPALSQGPIADNESEKVSRDKSNGSAGGK-----PFLQIGSNIVPMVV-GKETKEV 877
Query: 783 HSSLKHVIVYIGFEHECPRGHRFLLNARHLNELGSSYSLLEESCISSSMEPASINQAYHS 842
+ S++ +VY+GFEHEC GHRFLL+ +HL E+ SSY E S +++ E
Sbjct: 878 NQSIQQFMVYVGFEHECSYGHRFLLSEKHLKEIDSSYLQFERSNLNNEAE---------- 927
Query: 843 RVSKNASWAKVHQSSNEIPPASTNKERDADKSNEMMPNGDLNSDGLLYTSIPLKEQNLTS 902
SK+ S K+ Q+++ + A+T K N M + NS L + + L
Sbjct: 928 --SKHGS-QKLPQNASRL--AATMDVTSGGKLNRPMDSSGRNSQQQLLKP-RVDAETLQP 981
Query: 903 GNALAKTQNLMKDSGGDLQ----AIGMGGDELAFSMLNRNLPIYMMCPHCRNSRNMKDTP 958
+ L+ QN K G+L + GG+ AFS+LNRNLPIYM CPHC++S + K
Sbjct: 982 SHWLSDPQNERK---GELSLHYVTLDDGGE--AFSLLNRNLPIYMHCPHCKSS-DRKGNQ 1035
Query: 959 KVKFASGISQLKRIFLV 975
K A+ +SQL+RIF+V
Sbjct: 1036 DAKVAAAVSQLQRIFIV 1052
>ref|XP_600313.1| PREDICTED: similar to hypothetical protein FLJ10587 [Bos taurus]
Length = 825
Score = 100 bits (249), Expect = 2e-19
Identities = 66/237 (27%), Positives = 107/237 (44%), Gaps = 24/237 (10%)
Query: 330 DLPNFEIWSSSSHHILSRILCAKGGCLDELEIIKRKPRPRNTVSSSTEGPLKSANPLDVA 389
+LP ++ W S++ + E+ I ++ P + T L S L+
Sbjct: 355 ELPTYQKWISAASKLY------------EVAIDGKEEDPASPTGELTSKILSSIKVLEGF 402
Query: 390 VSWLQCGRGLNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKG 449
+ ++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +G
Sbjct: 403 LD-------IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARG 455
Query: 450 PSVQVFARKLEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPH 508
P+ +A +L E+C W +G QLC+ SLT + C+H+ H + S A P H
Sbjct: 456 PAFHKYAMQLHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYH 515
Query: 509 SSGYFFLHACACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
+S AC CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 516 NSRARSTGACNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 572
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 678 SSWSLVKLGPAKSYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 728
>ref|XP_548236.1| PREDICTED: similar to hypothetical protein FLJ10587 [Canis
familiaris]
Length = 991
Score = 100 bits (248), Expect = 3e-19
Identities = 66/237 (27%), Positives = 107/237 (44%), Gaps = 24/237 (10%)
Query: 330 DLPNFEIWSSSSHHILSRILCAKGGCLDELEIIKRKPRPRNTVSSSTEGPLKSANPLDVA 389
+LP ++ W S++ + E+ I ++ P + T L S L+
Sbjct: 441 ELPTYQKWISAASKLY------------EVAIDGKEEDPGSPTGELTSKILSSIKVLEGF 488
Query: 390 VSWLQCGRGLNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKG 449
+ ++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +G
Sbjct: 489 LD-------IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARG 541
Query: 450 PSVQVFARKLEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPH 508
P+ +A +L E+C W +G QLC+ SLT + C+H+ H + S A P H
Sbjct: 542 PAFHKYAMQLHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYH 601
Query: 509 SSGYFFLHACACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
+S AC CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 602 NSRARSTGACNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 658
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 764 SSWSLVKLGPAKSYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 814
>gb|AAH20957.1| FLJ10587 protein [Homo sapiens] gi|7022715|dbj|BAA91699.1| unnamed
protein product [Homo sapiens]
Length = 642
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 142 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 201
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 202 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 261
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 262 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 309
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 415 SSWSLVKLGPAKSYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 465
>ref|NP_077224.1| hypothetical protein LOC74133 [Mus musculus]
gi|56206524|emb|CAI25313.1| novel protein [Mus musculus]
gi|18043597|gb|AAH20005.1| RIKEN cDNA 1200011M11 [Mus
musculus] gi|26334424|dbj|BAB23498.2| unnamed protein
product [Mus musculus]
Length = 991
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 491 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 550
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 551 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 610
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 611 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 658
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 764 SSWSLVKLGPAKSYNFHTGLDQQGFVPGTNYLMPWDIVIRTRAEDEGDLDT 814
>ref|XP_213430.2| PREDICTED: similar to RIKEN cDNA 1200011M11 [Rattus norvegicus]
Length = 991
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 491 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 550
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 551 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 610
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 611 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 658
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 764 SSWSLVKLGPAKSYNFHTGLDQQGFVPGTNYLMPWDIVIRTRAEDEGDLDT 814
>ref|XP_523682.1| PREDICTED: similar to hypothetical protein FLJ10587 [Pan
troglodytes]
Length = 991
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 491 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 550
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 551 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 610
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 611 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 658
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 764 SSWSLVKLGPAKSYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 814
>gb|AAH02247.1| 1200011M11Rik protein [Mus musculus]
Length = 271
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 34 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 93
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 94 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 153
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 154 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 201
>dbj|BAB15576.1| unnamed protein product [Homo sapiens]
Length = 991
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 491 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 550
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 551 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 610
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 611 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 658
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 764 SSWSLVKLGPAKSYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 814
>emb|CAD39148.1| hypothetical protein [Homo sapiens] gi|47777319|ref|NP_060619.4|
hypothetical protein LOC55181 [Homo sapiens]
Length = 991
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 491 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 550
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 551 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 610
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 611 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 658
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 764 SSWSLVKLGPAKSYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 814
>dbj|BAC28941.1| unnamed protein product [Mus musculus]
Length = 642
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 142 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 201
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 202 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 261
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 262 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 309
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 415 SSWSLVKLGPAKSYNFHTGLDQQGFVPGTNYLMPWDIVIRTRAEDEGDLDT 465
>dbj|BAC27255.1| unnamed protein product [Mus musculus]
Length = 980
Score = 99.4 bits (246), Expect = 5e-19
Identities = 55/168 (32%), Positives = 84/168 (49%), Gaps = 5/168 (2%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 480 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 539
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + S A P H+S A
Sbjct: 540 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGA 599
Query: 518 CACGRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
C CGR + R DPFD ++A+ + C L + P E + P
Sbjct: 600 CNCGRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 647
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 753 SSWSLVKLGPAKSYNFHTGLDQQGFVPGTNYLMPWDIVIRTRAEDEGDLDT 803
>gb|AAL83913.1| amplified in breast cancer 2 [Homo sapiens]
Length = 1023
Score = 96.3 bits (238), Expect = 4e-18
Identities = 54/165 (32%), Positives = 80/165 (47%), Gaps = 5/165 (3%)
Query: 402 KFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARKLEE 461
KFS CQ+A+P A Y +LP YP H++ L +A + +GP+ +A +L E
Sbjct: 494 KFSENRCQKALPMAHSAYQSNLPHNYPMPVHKNQLAQARRVYSQHARGPAFHKYAMQLHE 553
Query: 462 ECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHACAC 520
+C W +G QLC+ SLT + C+H+ H + S A P H+S AC C
Sbjct: 554 DCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGEKPEADRNPPVLYHNSRARSTGACNC 613
Query: 521 GRSRQLRPDPFDFESADASCF----SDCDKLLPSVKLPEIEVAGP 561
GR + R DPFD ++A+ + C L + P E + P
Sbjct: 614 GRKQAPRDDPFDIKAANYDFYQLLEEKCCGKLDHINFPVFEPSTP 658
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 764 SSWSLVKLGPAKSYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 814
>ref|XP_415875.1| PREDICTED: similar to RIKEN cDNA 1200011M11 [Gallus gallus]
Length = 885
Score = 95.1 bits (235), Expect = 1e-17
Identities = 49/140 (35%), Positives = 76/140 (54%), Gaps = 1/140 (0%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 384 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 443
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGAS-PPSHKPHSSGYFFLHA 517
L E+C W +G QLC+ SLT + C+H+ H + + A P H+S A
Sbjct: 444 LNEDCYKFWSNGHQLCEERSLTDQHCVHKFHLLPKAGEKPEADRNPPILYHNSRARSTGA 503
Query: 518 CACGRSRQLRPDPFDFESAD 537
C CGR + R DPFD ++A+
Sbjct: 504 CNCGRKQAPRDDPFDIKAAN 523
Score = 38.9 bits (89), Expect = 0.84
Identities = 17/51 (33%), Positives = 26/51 (50%)
Query: 565 SAWSLLRIGSAKYYESSKGLLQSGFCATEKNLLKWTVYLEKQKRPNGSTET 615
S+WSL+++G AK Y GL Q GF L+ W + + + G +T
Sbjct: 658 SSWSLVKLGPAKAYNFHTGLDQQGFIPGTNYLMPWDIVIRTRAEDEGDLDT 708
>gb|AAH31604.1| Unknown (protein for IMAGE:5178217) [Homo sapiens]
Length = 584
Score = 80.9 bits (198), Expect = 2e-13
Identities = 36/98 (36%), Positives = 57/98 (57%)
Query: 399 LNTKFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARK 458
++TKFS CQ+A+P A Y +LP Y + H++ L +AL + +GP+ +A +
Sbjct: 487 IDTKFSENRCQKALPMAHSAYQSNLPHNYTMTVHKNQLAQALRVYSQHARGPAFHKYAMQ 546
Query: 459 LEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNS 496
L E+C W +G QLC+ SLT + C+H+ H + S S
Sbjct: 547 LHEDCYKFWSNGHQLCEERSLTDQHCVHKFHSLPKSGS 584
>ref|XP_319357.2| ENSANGP00000012357 [Anopheles gambiae str. PEST]
gi|55236386|gb|EAA13828.2| ENSANGP00000012357 [Anopheles
gambiae str. PEST]
Length = 879
Score = 79.0 bits (193), Expect = 7e-13
Identities = 49/157 (31%), Positives = 78/157 (49%), Gaps = 19/157 (12%)
Query: 408 CQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARKLEEECTSIW 467
C++ + A Y LP Y S HE +AL+ +GP ++ +KL+E C +IW
Sbjct: 341 CEQGLDLATLKYKDMLPPHYSASFHETKYHEALNLLLEYARGPELKPSIQKLKEYCDAIW 400
Query: 468 KSGRQLCDAISLTGKPCMHQRHDVETSNSNIGASPPSHKPHSSGYFFLHACACGRSRQLR 527
+SG+Q C+ SL G PC+ +H +I + HSSG ++ +C CGR++ R
Sbjct: 401 QSGKQQCEYPSLRGNPCVLGKH------KSIDLA-----EHSSGVIYVSSCNCGRTQGHR 449
Query: 528 PDPFDFESAD-------ASCFSDCDKLLPSVKLPEIE 557
DP+ A+ A S+C++ L V+ P E
Sbjct: 450 EDPYTIRQANYEFYQMIAKSCSNCNR-LERVQFPVFE 485
Score = 35.8 bits (81), Expect = 7.1
Identities = 17/39 (43%), Positives = 26/39 (66%), Gaps = 7/39 (17%)
Query: 778 ERSKPHSS--LKH-----VIVYIGFEHECPRGHRFLLNA 809
++ PHSS +H + ++IG E+EC RGHRF++NA
Sbjct: 707 KKGNPHSSKSFEHNKTFTLKIFIGVEYECLRGHRFIMNA 745
>gb|AAL89949.1| SD08609p [Drosophila melanogaster]
Length = 944
Score = 70.9 bits (172), Expect = 2e-10
Identities = 39/136 (28%), Positives = 58/136 (41%), Gaps = 13/136 (9%)
Query: 402 KFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARKLEE 461
KF C+ + Y P+ Y ++ H L +A AF +GP Q KL
Sbjct: 393 KFWAHLCELGLKKGIAAYKNAAPENYGSATHRQLLAEATLAFEEEGRGPQAQAALAKLAS 452
Query: 462 ECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGASPPSHKPHSSGYFFLHACACG 521
C W+ GRQ C+ +SL PC + H+ H+SG + +C CG
Sbjct: 453 ICHKHWQDGRQQCEQLSLRSHPCTQPK-------------KVPHEKHNSGVIHISSCNCG 499
Query: 522 RSRQLRPDPFDFESAD 537
R++ R DPF+ A+
Sbjct: 500 RTQGRREDPFNLRQAN 515
>ref|NP_609449.1| CG6729-PA [Drosophila melanogaster] gi|7297754|gb|AAF53005.1|
CG6729-PA [Drosophila melanogaster]
Length = 944
Score = 69.7 bits (169), Expect = 4e-10
Identities = 39/136 (28%), Positives = 58/136 (41%), Gaps = 13/136 (9%)
Query: 402 KFSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARKLEE 461
KF C+ + Y P+ Y ++ H L +A AF +GP Q KL
Sbjct: 393 KFWAHLCELGLKKGIAAYKNAAPENYGSATHRQLLAEATLAFEEEGRGPQAQAALAKLAS 452
Query: 462 ECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGASPPSHKPHSSGYFFLHACACG 521
C W+ GRQ C+ +SL PC + H+ H+SG + +C CG
Sbjct: 453 ICHKHWQDGRQQCEQLSLRSHPCTLPK-------------KVPHEKHNSGVIHISSCNCG 499
Query: 522 RSRQLRPDPFDFESAD 537
R++ R DPF+ A+
Sbjct: 500 RTQGRREDPFNLRQAN 515
>ref|XP_645311.1| hypothetical protein DDB0217022 [Dictyostelium discoideum]
gi|42761571|gb|AAS45389.1| similar to Multicopy
Suppressor of STA10 - 11; Mss11p [Saccharomyces
cerevisiae] [Dictyostelium discoideum]
gi|60473441|gb|EAL71387.1| hypothetical protein
DDB0217022 [Dictyostelium discoideum]
Length = 1002
Score = 58.9 bits (141), Expect = 8e-07
Identities = 32/102 (31%), Positives = 51/102 (49%)
Query: 403 FSTLWCQRAIPAAKEIYLKDLPDCYPTSQHEDHLDKALHAFHSMVKGPSVQVFARKLEEE 462
FS C+ + E Y ++LP Y T H + KA+ GP+++ L E
Sbjct: 519 FSKKLCRMNNKQSMEQYQENLPISYGTKIHCSRVAKAIKTICFNASGPALEFSIDYLRME 578
Query: 463 CTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNSNIGASPPS 504
C IWK G++LC+A SL+G+ C + H+V ++ PP+
Sbjct: 579 CDQIWKRGKRLCEAESLSGRICTLKTHNVPPLPNSCMPPPPT 620
>emb|CAB05756.2| Hypothetical protein K04B12.3 [Caenorhabditis elegans]
Length = 891
Score = 55.8 bits (133), Expect = 7e-06
Identities = 27/111 (24%), Positives = 56/111 (50%), Gaps = 8/111 (7%)
Query: 437 DKALHAFHSMVKGPSVQVFARKLEEECTSIWKSGRQLCDAISLTGKPCMHQRHDVETSNS 496
++A H S+V G + + +L+ +C +W+S + C+++S+ G+ C+ + H
Sbjct: 407 NEATHYIDSVV-GVNSREALSQLQAQCNEMWQSDMRACESVSMMGRSCVRKIHPTFGDQ- 464
Query: 497 NIGASPPSHK--PHSSGYFFLHACACGRSRQLRPDPFDFESADASCFSDCD 545
+ P H+ H + + C CGR + +RP+PF + A++ + D
Sbjct: 465 ----TAPEHRWTAHDASNTMISTCVCGRKQLIRPEPFSVKEANSDFYDHPD 511
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.130 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,672,647,305
Number of Sequences: 2540612
Number of extensions: 73002335
Number of successful extensions: 204425
Number of sequences better than 10.0: 206
Number of HSP's better than 10.0 without gapping: 47
Number of HSP's successfully gapped in prelim test: 171
Number of HSP's that attempted gapping in prelim test: 203389
Number of HSP's gapped (non-prelim): 892
length of query: 975
length of database: 863,360,394
effective HSP length: 138
effective length of query: 837
effective length of database: 512,755,938
effective search space: 429176720106
effective search space used: 429176720106
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0369c.3


