

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0348.12
(180 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
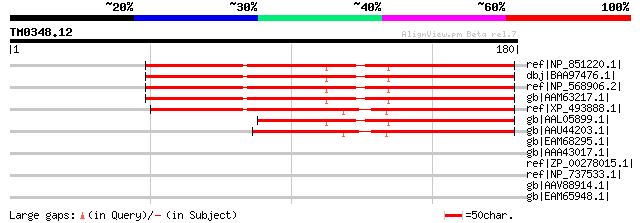
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_851220.1| expressed protein [Arabidopsis thaliana] 120 1e-26
dbj|BAA97476.1| unnamed protein product [Arabidopsis thaliana] 120 1e-26
ref|NP_568906.2| expressed protein [Arabidopsis thaliana] 120 1e-26
gb|AAM63217.1| unknown [Arabidopsis thaliana] 118 9e-26
ref|XP_493888.1| unknown protein [Oryza sativa] gi|14719338|gb|A... 106 3e-22
gb|AAL05899.1| AT5g59400/f2o15_60 [Arabidopsis thaliana] gi|1508... 86 5e-16
gb|AAU44203.1| unknown protein [Oryza sativa (japonica cultivar-... 79 7e-14
gb|EAM68295.1| DedA:TRAP C4-dicarboxylate transport system perm... 35 0.95
gb|AAA43017.1| envelope protein 34 1.6
ref|ZP_00278015.1| COG4177: ABC-type branched-chain amino acid t... 34 2.1
ref|NP_737533.1| hypothetical protein CE0923 [Corynebacterium ef... 33 3.6
gb|AAV88914.1| predicted membrane protein [Zymomonas mobilis sub... 33 3.6
gb|EAM65948.1| TRAP C4-dicarboxylate transport system permease ... 32 8.0
>ref|NP_851220.1| expressed protein [Arabidopsis thaliana]
Length = 299
Score = 120 bits (302), Expect = 1e-26
Identities = 67/136 (49%), Positives = 85/136 (62%), Gaps = 9/136 (6%)
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGI 108
W GS+S V YPRCS+ RQSTYADAE + L W + FGS AC+ P Y VG+
Sbjct: 106 WYGSKSVVKYPRCSLLRQSTYADAEDDASQVLLLA-TVWIMIFLFGSSACVLPTIYGVGL 164
Query: 109 AYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGFVIGYPLASAPVKVIQGLWRN 163
Y + F SGL + S L S+ + N I+ VLG GYP+AS+ V+V++GLWRN
Sbjct: 165 VYGGDPFDSGLVYSSQ---LSSSVPILSKFNGILLSVLGPAFGYPIASSAVRVLKGLWRN 221
Query: 164 DLVALRGACPNCGEEL 179
DL AL+G CPNCGEE+
Sbjct: 222 DLTALKGDCPNCGEEV 237
>dbj|BAA97476.1| unnamed protein product [Arabidopsis thaliana]
Length = 282
Score = 120 bits (302), Expect = 1e-26
Identities = 67/136 (49%), Positives = 85/136 (62%), Gaps = 9/136 (6%)
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGI 108
W GS+S V YPRCS+ RQSTYADAE + L W + FGS AC+ P Y VG+
Sbjct: 106 WYGSKSVVKYPRCSLLRQSTYADAEDDASQVLLLA-TVWIMIFLFGSSACVLPTIYGVGL 164
Query: 109 AYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGFVIGYPLASAPVKVIQGLWRN 163
Y + F SGL + S L S+ + N I+ VLG GYP+AS+ V+V++GLWRN
Sbjct: 165 VYGGDPFDSGLVYSSQ---LSSSVPILSKFNGILLSVLGPAFGYPIASSAVRVLKGLWRN 221
Query: 164 DLVALRGACPNCGEEL 179
DL AL+G CPNCGEE+
Sbjct: 222 DLTALKGDCPNCGEEV 237
>ref|NP_568906.2| expressed protein [Arabidopsis thaliana]
Length = 301
Score = 120 bits (302), Expect = 1e-26
Identities = 67/136 (49%), Positives = 85/136 (62%), Gaps = 9/136 (6%)
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGI 108
W GS+S V YPRCS+ RQSTYADAE + L W + FGS AC+ P Y VG+
Sbjct: 106 WYGSKSVVKYPRCSLLRQSTYADAEDDASQVLLLA-TVWIMIFLFGSSACVLPTIYGVGL 164
Query: 109 AYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGFVIGYPLASAPVKVIQGLWRN 163
Y + F SGL + S L S+ + N I+ VLG GYP+AS+ V+V++GLWRN
Sbjct: 165 VYGGDPFDSGLVYSSQ---LSSSVPILSKFNGILLSVLGPAFGYPIASSAVRVLKGLWRN 221
Query: 164 DLVALRGACPNCGEEL 179
DL AL+G CPNCGEE+
Sbjct: 222 DLTALKGDCPNCGEEV 237
>gb|AAM63217.1| unknown [Arabidopsis thaliana]
Length = 299
Score = 118 bits (295), Expect = 9e-26
Identities = 65/136 (47%), Positives = 84/136 (60%), Gaps = 9/136 (6%)
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGI 108
W GS+S V YPRCS+ RQSTYADAE + L W + FGS AC+ P Y VG+
Sbjct: 106 WYGSKSVVKYPRCSLLRQSTYADAEDDASQVLLLA-TVWIMIFLFGSSACVLPTIYGVGL 164
Query: 109 AYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGFVIGYPLASAPVKVIQGLWRN 163
Y + F SGL + + L S+ + N I+ VLG GYP+AS+ V+ ++GLWRN
Sbjct: 165 VYGGDPFDSGLVYSNQ---LSSSVPILSKFNGILLSVLGPAFGYPIASSAVRALKGLWRN 221
Query: 164 DLVALRGACPNCGEEL 179
DL AL+G CPNCGEE+
Sbjct: 222 DLTALKGDCPNCGEEV 237
>ref|XP_493888.1| unknown protein [Oryza sativa] gi|14719338|gb|AAK73156.1| unknown
protein [Oryza sativa]
Length = 272
Score = 106 bits (264), Expect = 3e-22
Identities = 61/132 (46%), Positives = 87/132 (65%), Gaps = 8/132 (6%)
Query: 51 GSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGIAY 110
GS S V YPRCS++RQSTYADAE + L W L L FG+ A L P + + +
Sbjct: 83 GSPSVVKYPRCSLKRQSTYADAEEDKSMFMALS-SIWMLLLLFGTSAFLVPSLCILSLTF 141
Query: 111 QNAFGSG-LSHGSHNPGLGFSII--VNNIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVA 167
+AFG+ L +G+ + F +I VN+++ I LG++IGYP++SA V +QGL N+LVA
Sbjct: 142 GDAFGARYLLYGAKS----FDVITRVNDMVLIGLGYLIGYPISSASVGALQGLLTNNLVA 197
Query: 168 LRGACPNCGEEL 179
L+G+CPNCGE++
Sbjct: 198 LKGSCPNCGEQV 209
>gb|AAL05899.1| AT5g59400/f2o15_60 [Arabidopsis thaliana]
gi|15081620|gb|AAK82465.1| AT5g59400/f2o15_60
[Arabidopsis thaliana]
Length = 157
Score = 85.9 bits (211), Expect = 5e-16
Identities = 46/96 (47%), Positives = 61/96 (62%), Gaps = 8/96 (8%)
Query: 89 LFLAFGSLACLGPLSYAVGIAYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGF 143
+ FGS AC+ P Y VG+ Y + F SGL + S L S+ + N I+ VLG
Sbjct: 1 MIFLFGSSACVLPTIYGVGLVYGGDPFDSGLVYSSQ---LSSSVPILSKFNGILLSVLGP 57
Query: 144 VIGYPLASAPVKVIQGLWRNDLVALRGACPNCGEEL 179
GYP+AS+ V+V++GLWRNDL AL+G CPNCGEE+
Sbjct: 58 AFGYPIASSAVRVLKGLWRNDLTALKGDCPNCGEEV 93
>gb|AAU44203.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 163
Score = 78.6 bits (192), Expect = 7e-14
Identities = 42/96 (43%), Positives = 65/96 (66%), Gaps = 7/96 (7%)
Query: 87 WALFLAFGSLACLGPLSYAVGIAYQNAFGSG-LSHGSHNPGLGFSII--VNNIIFIVLGF 143
W L L FG+ A L P + + + +AFG+ L +G+ + F +I VN+++ I LG+
Sbjct: 9 WMLLLLFGTSAFLVPSLCILSLTFGDAFGARYLLYGAKS----FDVITRVNDMVLIGLGY 64
Query: 144 VIGYPLASAPVKVIQGLWRNDLVALRGACPNCGEEL 179
+IGYP++SA V +QGL N+LVAL+G+CPNCGE++
Sbjct: 65 LIGYPISSASVGALQGLLTNNLVALKGSCPNCGEQV 100
>gb|EAM68295.1| DedA:TRAP C4-dicarboxylate transport system permease DctM subunit
[Jannaschia sp. CCS1]
Length = 431
Score = 35.0 bits (79), Expect = 0.95
Identities = 23/73 (31%), Positives = 38/73 (51%), Gaps = 18/73 (24%)
Query: 91 LAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLA 150
++F LAC+ L + GI +G+SII+ +++++ LG G LA
Sbjct: 1 MSFEVLACIATLFFLAGIGTP---------------IGYSIILASLVYLGLG---GMDLA 42
Query: 151 SAPVKVIQGLWRN 163
A K+IQGL+R+
Sbjct: 43 LAGEKIIQGLYRS 55
>gb|AAA43017.1| envelope protein
Length = 595
Score = 34.3 bits (77), Expect = 1.6
Identities = 20/51 (39%), Positives = 32/51 (62%), Gaps = 7/51 (13%)
Query: 114 FGSGL-SH-GSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVIQGLWR 162
FG L SH + NPGLG SII ++F+++ Y L ++P K+++ LW+
Sbjct: 344 FGKDLWSHIANWNPGLGASIIKYIVVFLLI-----YLLLTSPPKILRALWK 389
>ref|ZP_00278015.1| COG4177: ABC-type branched-chain amino acid transport system,
permease component [Burkholderia fungorum LB400]
Length = 590
Score = 33.9 bits (76), Expect = 2.1
Identities = 31/86 (36%), Positives = 37/86 (42%), Gaps = 12/86 (13%)
Query: 86 AW--ALFLAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGF 143
AW ALF FG+L LG GI N FG L G H I I +VLG
Sbjct: 118 AWGLALFYLFGNLEMLGKYDGINGIPVLNVFGINLESGRH--------IYYLIWVVVLGA 169
Query: 144 VIGYP--LASAPVKVIQGLWRNDLVA 167
VI L S P + I+ L ++A
Sbjct: 170 VISVQNLLNSRPGRAIRALRGGGMMA 195
>ref|NP_737533.1| hypothetical protein CE0923 [Corynebacterium efficiens YS-314]
gi|23492761|dbj|BAC17733.1| conserved hypothetical
protein [Corynebacterium efficiens YS-314]
Length = 149
Score = 33.1 bits (74), Expect = 3.6
Identities = 32/104 (30%), Positives = 46/104 (43%), Gaps = 26/104 (25%)
Query: 79 LWFLH*QAWAL--FLAFGSLACLGPLSYAVGIA-YQNA------FGSGLSHGSHNPGLGF 129
+W L W ++ FG LAC+ ++ GIA ++ A FG + +H G G
Sbjct: 20 IWLLFGGIWLALGYVLFGILACILIITIPAGIASFRMANYALWPFGRTVVRKTHG-GNGL 78
Query: 130 SIIVNNIIFIVLGF----------------VIGYPLASAPVKVI 157
S I N I FI+ G +IG PLA A +K+I
Sbjct: 79 SAISNFIWFIIAGLWLAIGHITTAAAQAITIIGIPLAIANIKMI 122
>gb|AAV88914.1| predicted membrane protein [Zymomonas mobilis subsp. mobilis ZM4]
gi|56551186|ref|YP_162025.1| predicted membrane protein
[Zymomonas mobilis subsp. mobilis ZM4]
Length = 418
Score = 33.1 bits (74), Expect = 3.6
Identities = 27/90 (30%), Positives = 39/90 (43%), Gaps = 2/90 (2%)
Query: 74 GRRIYLWFLH*QAWALFLAFGSLACLGPLS--YAVGIAYQNAFGSGLSHGSHNPGLGFSI 131
GR + +F H AW +AF A +G L+ +AV +++ G H + P I
Sbjct: 29 GRELNHFFPHQWAWGFLIAFSEAAMVGGLADWFAVTALFRHPLGLPFPHTALIPHAKDRI 88
Query: 132 IVNNIIFIVLGFVIGYPLASAPVKVIQGLW 161
VN FI F+ LA V GL+
Sbjct: 89 AVNLADFIETHFLRSIVLARRLKPVDAGLF 118
>gb|EAM65948.1| TRAP C4-dicarboxylate transport system permease DctM subunit
[Jannaschia sp. CCS1]
Length = 438
Score = 32.0 bits (71), Expect = 8.0
Identities = 21/58 (36%), Positives = 33/58 (56%), Gaps = 5/58 (8%)
Query: 90 FLAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFVIGY 147
+L FG+ A +G + GIA+ G LS S NP + +++V +FI+LGF I +
Sbjct: 290 WLVFGTNALIGVYNLMGGIAFA---GEVLSGVSDNPSIILAVMVG--VFILLGFFIDW 342
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.340 0.149 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 297,931,239
Number of Sequences: 2540612
Number of extensions: 11317557
Number of successful extensions: 32257
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 32236
Number of HSP's gapped (non-prelim): 13
length of query: 180
length of database: 863,360,394
effective HSP length: 120
effective length of query: 60
effective length of database: 558,486,954
effective search space: 33509217240
effective search space used: 33509217240
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0348.12


