

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.8
(61 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
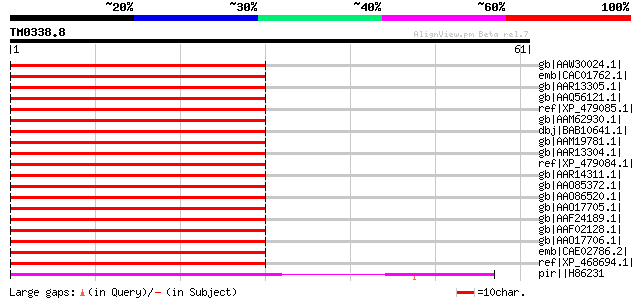
Score E
Sequences producing significant alignments: (bits) Value
gb|AAW30024.1| At5g15630 [Arabidopsis thaliana] gi|30685446|ref|... 67 2e-10
emb|CAC01762.1| putative phytochelatin synthetase [Arabidopsis t... 67 2e-10
gb|AAR13305.1| phytochelatin synthetase-like protein [Phaseolus ... 65 6e-10
gb|AAQ56121.1| BRITTLE CULM1 [Oryza sativa (indica cultivar-grou... 64 8e-10
ref|XP_479085.1| putative phytochelatin synthetase [Oryza sativa... 64 1e-09
gb|AAM62930.1| phytochelatin synthetase-like protein [Arabidopsi... 60 2e-08
dbj|BAB10641.1| phytochelatin synthetase [Arabidopsis thaliana] ... 60 2e-08
gb|AAM19781.1| AT5g60920/MSL3_40 [Arabidopsis thaliana] gi|18424... 60 2e-08
gb|AAR13304.1| phytochelatin synthetase-like protein [Phaseolus ... 60 2e-08
ref|XP_479084.1| putative phytochelatin synthetase [Oryza sativa... 58 8e-08
gb|AAR14311.1| phytochelatin synthetase [Triticum aestivum] 57 2e-07
gb|AAO85372.1| phytochelatin synthetase [Hordeum vulgare subsp. ... 57 2e-07
gb|AAO86520.1| phytochelatin synthetase [Triticum monococcum] 57 2e-07
gb|AAO17705.1| phytochelatin synthetase-like protein 1 [Sorghum ... 57 2e-07
gb|AAF24189.1| phytochelatin synthetase-like protein [Zea mays] 57 2e-07
gb|AAF02128.1| unknown protein [Arabidopsis thaliana] gi|2645213... 56 3e-07
gb|AAO17706.1| phytochelatin synthetase-like protein 2 [Sorghum ... 56 3e-07
emb|CAE02786.2| OSJNBa0011L07.10 [Oryza sativa (japonica cultiva... 55 4e-07
ref|XP_468694.1| putative phytochelatin synthetase [Oryza sativa... 55 5e-07
pir||H86231 hypothetical protein [imported] - Arabidopsis thalia... 55 5e-07
>gb|AAW30024.1| At5g15630 [Arabidopsis thaliana] gi|30685446|ref|NP_197067.2|
phytochelatin synthetase family protein / COBRA cell
expansion protein COBL4 [Arabidopsis thaliana]
gi|48525349|gb|AAT44976.1| At5g15630 [Arabidopsis
thaliana] gi|34222647|sp|Q9LFW3|CBL4_ARATH COBRA-like
protein 4 precursor
Length = 431
Score = 66.6 bits (161), Expect = 2e-10
Identities = 26/30 (86%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD+ TF+FKQGWAFPRKVYFNGDECM+PPP
Sbjct: 372 KDQKTFTFKQGWAFPRKVYFNGDECMLPPP 401
>emb|CAC01762.1| putative phytochelatin synthetase [Arabidopsis thaliana]
gi|11358657|pir||T51392 probable phytochelatin
synthetase - Arabidopsis thaliana
Length = 395
Score = 66.6 bits (161), Expect = 2e-10
Identities = 26/30 (86%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD+ TF+FKQGWAFPRKVYFNGDECM+PPP
Sbjct: 336 KDQKTFTFKQGWAFPRKVYFNGDECMLPPP 365
>gb|AAR13305.1| phytochelatin synthetase-like protein [Phaseolus vulgaris]
Length = 463
Score = 64.7 bits (156), Expect = 6e-10
Identities = 25/30 (83%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+TF+ KQGWAFPRKVYFNG+ECM+PPP
Sbjct: 403 KDKDTFTLKQGWAFPRKVYFNGEECMLPPP 432
>gb|AAQ56121.1| BRITTLE CULM1 [Oryza sativa (indica cultivar-group)]
gi|34014146|gb|AAQ56120.1| BRITTLE CULM1 [Oryza sativa
(japonica cultivar-group)] gi|50916593|ref|XP_468693.1|
putative phytochelatin synthetase [Oryza sativa
(japonica cultivar-group)] gi|41469147|gb|AAS07098.1|
putative phytochelatin synthetase [Oryza sativa
(japonica cultivar-group)]
Length = 468
Score = 64.3 bits (155), Expect = 8e-10
Identities = 25/30 (83%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD NTF+F QGWAFPRK+YFNGDEC MPPP
Sbjct: 406 KDYNTFTFSQGWAFPRKIYFNGDECKMPPP 435
>ref|XP_479085.1| putative phytochelatin synthetase [Oryza sativa (japonica
cultivar-group)] gi|34394570|dbj|BAC83873.1| putative
phytochelatin synthetase [Oryza sativa (japonica
cultivar-group)]
Length = 451
Score = 63.9 bits (154), Expect = 1e-09
Identities = 25/30 (83%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F+QGWAFPRKVYFNGDEC MPPP
Sbjct: 389 KDARTFTFRQGWAFPRKVYFNGDECQMPPP 418
>gb|AAM62930.1| phytochelatin synthetase-like protein [Arabidopsis thaliana]
Length = 456
Score = 60.1 bits (144), Expect = 2e-08
Identities = 21/30 (70%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD++TF+F++GWAFPR++YFNGD C+MPPP
Sbjct: 394 KDQSTFTFEKGWAFPRRIYFNGDNCVMPPP 423
>dbj|BAB10641.1| phytochelatin synthetase [Arabidopsis thaliana]
gi|3559805|emb|CAA07251.1| putative phytochelatin
synthetase [Arabidopsis thaliana]
gi|25456127|pir||T52038 probable phytochelatin
synthetase [imported] - Arabidopsis thaliana
Length = 362
Score = 60.1 bits (144), Expect = 2e-08
Identities = 21/30 (70%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD++TF+F++GWAFPR++YFNGD C+MPPP
Sbjct: 300 KDQSTFTFEKGWAFPRRIYFNGDNCVMPPP 329
>gb|AAM19781.1| AT5g60920/MSL3_40 [Arabidopsis thaliana]
gi|18424412|ref|NP_568930.1| phytochelatin synthetase,
putative / COBRA cell expansion protein COB, putative
[Arabidopsis thaliana] gi|14193657|gb|AAK56072.1|
putative glycosylphosphatidylinositol-anchored protein
[Arabidopsis thaliana] gi|34222620|sp|Q94KT8|COBR_ARATH
COBRA protein precursor (Cell expansion protein)
gi|24796996|gb|AAN64510.1| At5g60920/MSL3_40
[Arabidopsis thaliana]
Length = 456
Score = 60.1 bits (144), Expect = 2e-08
Identities = 21/30 (70%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD++TF+F++GWAFPR++YFNGD C+MPPP
Sbjct: 394 KDQSTFTFEKGWAFPRRIYFNGDNCVMPPP 423
>gb|AAR13304.1| phytochelatin synthetase-like protein [Phaseolus vulgaris]
Length = 448
Score = 59.7 bits (143), Expect = 2e-08
Identities = 22/30 (73%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK TF+F +GWAFPR++YFNGD C+MPPP
Sbjct: 385 KDKATFTFDKGWAFPRRIYFNGDNCVMPPP 414
>ref|XP_479084.1| putative phytochelatin synthetase [Oryza sativa (japonica
cultivar-group)] gi|34394569|dbj|BAC83872.1| putative
phytochelatin synthetase [Oryza sativa (japonica
cultivar-group)]
Length = 446
Score = 57.8 bits (138), Expect = 8e-08
Identities = 21/30 (70%), Positives = 27/30 (90%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD +F+F++GWAFPR+VYFNGD C+MPPP
Sbjct: 382 KDPKSFTFEKGWAFPRRVYFNGDNCVMPPP 411
>gb|AAR14311.1| phytochelatin synthetase [Triticum aestivum]
Length = 456
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 391 KDSETFTFQKGWAFPRRVYFNGDNCVMPSP 420
>gb|AAO85372.1| phytochelatin synthetase [Hordeum vulgare subsp. vulgare]
Length = 112
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 78 KDSETFTFQKGWAFPRRVYFNGDNCVMPSP 107
>gb|AAO86520.1| phytochelatin synthetase [Triticum monococcum]
Length = 457
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 392 KDSETFTFQKGWAFPRRVYFNGDNCVMPSP 421
>gb|AAO17705.1| phytochelatin synthetase-like protein 1 [Sorghum bicolor]
Length = 425
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 386 KDSRTFTFEKGWAFPRRVYFNGDNCVMPAP 415
>gb|AAF24189.1| phytochelatin synthetase-like protein [Zea mays]
Length = 407
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 297 KDSRTFTFEKGWAFPRRVYFNGDNCVMPSP 326
>gb|AAF02128.1| unknown protein [Arabidopsis thaliana] gi|26452135|dbj|BAC43156.1|
GPI-anchored protein [Arabidopsis thaliana]
gi|28951027|gb|AAO63437.1| At3g02210 [Arabidopsis
thaliana] gi|15232863|ref|NP_186870.1| phytochelatin
synthetase family protein / COBRA cell expansion protein
COBL3 [Arabidopsis thaliana]
gi|34222657|sp|Q9SRT7|CBL1_ARATH COBRA-like protein 1
precursor
Length = 452
Score = 55.8 bits (133), Expect = 3e-07
Identities = 19/30 (63%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+ + F+F++GWAFPR++YFNGD C+MPPP
Sbjct: 389 KEASAFTFEKGWAFPRRIYFNGDNCVMPPP 418
>gb|AAO17706.1| phytochelatin synthetase-like protein 2 [Sorghum bicolor]
Length = 449
Score = 55.8 bits (133), Expect = 3e-07
Identities = 21/30 (70%), Positives = 25/30 (83%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F +GWAFPR+VYFNGD C+MP P
Sbjct: 385 KDSRTFTFDKGWAFPRRVYFNGDNCVMPSP 414
>emb|CAE02786.2| OSJNBa0011L07.10 [Oryza sativa (japonica cultivar-group)]
gi|50927043|ref|XP_473354.1| OSJNBa0011L07.10 [Oryza
sativa (japonica cultivar-group)]
Length = 439
Score = 55.5 bits (132), Expect = 4e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+ F+F GWAFPR+VYF+G EC+MPPP
Sbjct: 375 KDKSDFTFSGGWAFPRRVYFDGHECVMPPP 404
>ref|XP_468694.1| putative phytochelatin synthetase [Oryza sativa (japonica
cultivar-group)] gi|41469124|gb|AAS07075.1| putative
phytochelatin synthetase [Oryza sativa (japonica
cultivar-group)]
Length = 458
Score = 55.1 bits (131), Expect = 5e-07
Identities = 19/30 (63%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD++TF+F +GWAFPR++YFNG+ C+MP P
Sbjct: 384 KDRSTFTFDKGWAFPRRIYFNGESCVMPSP 413
>pir||H86231 hypothetical protein [imported] - Arabidopsis thaliana
gi|2160169|gb|AAB60732.1| F21M12.17 gene product
[Arabidopsis thaliana]
Length = 454
Score = 55.1 bits (131), Expect = 5e-07
Identities = 28/61 (45%), Positives = 35/61 (56%), Gaps = 16/61 (26%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTHFCQILLLQA*FP----SRHSSSHYSS 56
KD F+F++GWAFPR++ FNGDEC+MP P FP S HSSS S+
Sbjct: 399 KDMGNFTFREGWAFPRRILFNGDECVMPSPDD------------FPRLPKSAHSSSSSSA 446
Query: 57 F 57
F
Sbjct: 447 F 447
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.335 0.145 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 121,990,658
Number of Sequences: 2540612
Number of extensions: 3862585
Number of successful extensions: 9777
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 9739
Number of HSP's gapped (non-prelim): 38
length of query: 61
length of database: 863,360,394
effective HSP length: 37
effective length of query: 24
effective length of database: 769,357,750
effective search space: 18464586000
effective search space used: 18464586000
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0338.8


