

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.4
(197 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
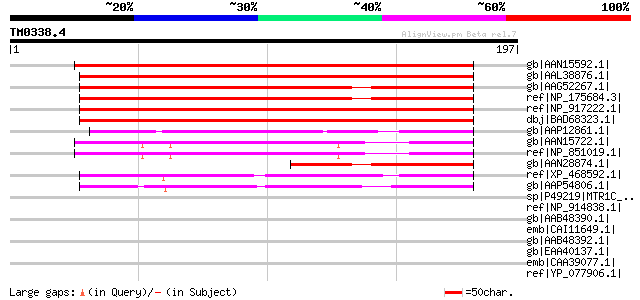
Score E
Sequences producing significant alignments: (bits) Value
gb|AAN15592.1| unknown protein [Arabidopsis thaliana] gi|2046645... 234 1e-60
gb|AAL38876.1| unknown protein [Arabidopsis thaliana] gi|2698389... 219 4e-56
gb|AAG52267.1| unknown protein; 18223-15857 [Arabidopsis thalian... 164 1e-39
ref|NP_175684.3| hydrolase, alpha/beta fold family protein [Arab... 164 1e-39
ref|NP_917222.1| P0038D11.18 [Oryza sativa (japonica cultivar-gr... 161 1e-38
dbj|BAD68323.1| alpha/beta hydrolase-like [Oryza sativa (japonic... 161 1e-38
gb|AAP12861.1| At2g40095 [Arabidopsis thaliana] gi|26449647|dbj|... 105 7e-22
gb|AAN15722.1| putative protein [Arabidopsis thaliana] gi|225309... 97 2e-19
ref|NP_851019.1| expressed protein [Arabidopsis thaliana] 97 2e-19
gb|AAN28874.1| At1g52750/F14G24_2 [Arabidopsis thaliana] gi|1931... 75 1e-12
ref|XP_468592.1| Unknown protein [Oryza sativa (japonica cultiva... 74 3e-12
gb|AAP54806.1| unknown protein [Oryza sativa (japonica cultivar-... 51 2e-05
sp|P49219|MTR1C_XENLA Melatonin receptor type 1C (MEL-1C-R) gi|5... 36 0.54
ref|NP_914838.1| P0413C03.14 [Oryza sativa (japonica cultivar-gr... 36 0.54
gb|AAB48390.1| Mel-1c(a) melatonin receptor [Xenopus laevis] gi|... 36 0.54
emb|CAI11649.1| novel protein similar to human and mouse androge... 36 0.70
gb|AAB48392.1| Mel-1c(b) melatonin receptor [Xenopus laevis] gi|... 36 0.70
gb|EAA40137.1| GLP_80_57878_54231 [Giardia lamblia ATCC 50803] 34 2.0
emb|CAA39077.1| glutamine-dependent carbamyl phosphate synthetas... 34 2.7
ref|YP_077906.1| Acyltransferase 3 [Bacillus licheniformis ATCC ... 34 2.7
>gb|AAN15592.1| unknown protein [Arabidopsis thaliana] gi|20466450|gb|AAM20542.1|
unknown protein [Arabidopsis thaliana]
gi|15220097|ref|NP_178144.1| hydrolase, alpha/beta fold
family protein [Arabidopsis thaliana]
gi|12324976|gb|AAG52432.1| unknown protein; 13661-11359
[Arabidopsis thaliana] gi|25406649|pir||C96834 unknown
protein F5I6.3 [imported] - Arabidopsis thaliana
Length = 647
Score = 234 bits (596), Expect = 1e-60
Identities = 111/155 (71%), Positives = 134/155 (85%)
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSL 85
+ L VSSLPV+V + D+LVP L+S+FTC+ CYG +H RYAFKSSLTDIPLVSL+RS
Sbjct: 26 VSLLVSSLPVLVAIGDVLVPTFLLSSFTCLTCYGFKEHLSRYAFKSSLTDIPLVSLVRSF 85
Query: 86 IILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRL 145
+++CVYS+ DAP LSHGPYLGTV+LCS++S+VLLSVK CVFT NS++ +AS+S RQRL
Sbjct: 86 LVICVYSLSDAPALSHGPYLGTVSLCSVVSVVLLSVKACVFTANSQLNDQASSSPSRQRL 145
Query: 146 HLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
HLKKSWGMPVLFLSSVVF+LGH+VVA RTSCRARR
Sbjct: 146 HLKKSWGMPVLFLSSVVFALGHMVVAYRTSCRARR 180
>gb|AAL38876.1| unknown protein [Arabidopsis thaliana] gi|26983890|gb|AAN86197.1|
unknown protein [Arabidopsis thaliana]
gi|15218212|ref|NP_173002.1| hydrolase, alpha/beta fold
family protein [Arabidopsis thaliana]
gi|5103847|gb|AAD39677.1| Contains PF|00561 alpha/beta
hydrolase fold. [Arabidopsis thaliana]
Length = 648
Score = 219 bits (557), Expect = 4e-56
Identities = 102/153 (66%), Positives = 128/153 (82%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
L VSSLP++V + D+LVP L+S+FTC+ CYG +H RY FK SLTDIP+VS++RS ++
Sbjct: 28 LLVSSLPLLVAIGDVLVPSFLLSSFTCVTCYGAKEHLRRYGFKRSLTDIPIVSVVRSFLV 87
Query: 88 LCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
+C+Y + D P LSHGPYLGTV+LCS++S++LLSVK C+FTVNS++ EAS S R+RLHL
Sbjct: 88 ICIYLLSDVPALSHGPYLGTVSLCSVVSVLLLSVKACLFTVNSQLNNEASFSPSRKRLHL 147
Query: 148 KKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
KKSWGMPVLFLSSVVF+LGH VVA RTSCRARR
Sbjct: 148 KKSWGMPVLFLSSVVFALGHTVVAYRTSCRARR 180
>gb|AAG52267.1| unknown protein; 18223-15857 [Arabidopsis thaliana]
gi|25405595|pir||E96568 unknown protein, 18223-15857
[imported] - Arabidopsis thaliana
Length = 614
Score = 164 bits (415), Expect = 1e-39
Identities = 82/153 (53%), Positives = 110/153 (71%), Gaps = 7/153 (4%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
L SSLPV++ V D++VPC+++S+ TC+ C+ +HF +Y+FK+SL D+PLVSL+RSL+I
Sbjct: 11 LLASSLPVLITVADVVVPCLIVSSITCLTCHSPAQHFRQYSFKTSLIDVPLVSLIRSLVI 70
Query: 88 LCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
+C+ +C+ L++G YL TV L S I+LL VKTCVFTVNS +E + + HL
Sbjct: 71 ICLSWLCEDTRLAYGLYLETVMLSSFGGILLLLVKTCVFTVNSTLE-------EAKVYHL 123
Query: 148 KKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
K SWGMPVL LSS VF L H+VVA R SC ARR
Sbjct: 124 KISWGMPVLLLSSAVFGLAHVVVAYRKSCGARR 156
>ref|NP_175684.3| hydrolase, alpha/beta fold family protein [Arabidopsis thaliana]
Length = 633
Score = 164 bits (415), Expect = 1e-39
Identities = 82/153 (53%), Positives = 110/153 (71%), Gaps = 7/153 (4%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
L SSLPV++ V D++VPC+++S+ TC+ C+ +HF +Y+FK+SL D+PLVSL+RSL+I
Sbjct: 30 LLASSLPVLITVADVVVPCLIVSSITCLTCHSPAQHFRQYSFKTSLIDVPLVSLIRSLVI 89
Query: 88 LCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
+C+ +C+ L++G YL TV L S I+LL VKTCVFTVNS +E + + HL
Sbjct: 90 ICLSWLCEDTRLAYGLYLETVMLSSFGGILLLLVKTCVFTVNSTLE-------EAKVYHL 142
Query: 148 KKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
K SWGMPVL LSS VF L H+VVA R SC ARR
Sbjct: 143 KISWGMPVLLLSSAVFGLAHVVVAYRKSCGARR 175
>ref|NP_917222.1| P0038D11.18 [Oryza sativa (japonica cultivar-group)]
Length = 629
Score = 161 bits (407), Expect = 1e-38
Identities = 82/153 (53%), Positives = 105/153 (68%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
L ++S P +V D+ V L C+RC+G+ H RY F+SSL DIPLVS+ RS++I
Sbjct: 8 LLMASAPALVAAGDVAVALWLEVRLGCLRCHGLRGHLERYGFRSSLVDIPLVSIARSVVI 67
Query: 88 LCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
CVY + DA LSHGPYLGT T CS+ S+++L +K V++ I E S SL +L L
Sbjct: 68 TCVYLMSDASGLSHGPYLGTATCCSLASLLILLIKASVYSPAQEIGPELSPSLADHKLSL 127
Query: 148 KKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
KK GMPVLFLSS+VF+LGH+VVA RTSCRARR
Sbjct: 128 KKLSGMPVLFLSSLVFALGHVVVAYRTSCRARR 160
>dbj|BAD68323.1| alpha/beta hydrolase-like [Oryza sativa (japonica cultivar-group)]
gi|55296846|dbj|BAD68190.1| alpha/beta hydrolase-like
[Oryza sativa (japonica cultivar-group)]
Length = 650
Score = 161 bits (407), Expect = 1e-38
Identities = 82/153 (53%), Positives = 105/153 (68%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
L ++S P +V D+ V L C+RC+G+ H RY F+SSL DIPLVS+ RS++I
Sbjct: 29 LLMASAPALVAAGDVAVALWLEVRLGCLRCHGLRGHLERYGFRSSLVDIPLVSIARSVVI 88
Query: 88 LCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
CVY + DA LSHGPYLGT T CS+ S+++L +K V++ I E S SL +L L
Sbjct: 89 TCVYLMSDASGLSHGPYLGTATCCSLASLLILLIKASVYSPAQEIGPELSPSLADHKLSL 148
Query: 148 KKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
KK GMPVLFLSS+VF+LGH+VVA RTSCRARR
Sbjct: 149 KKLSGMPVLFLSSLVFALGHVVVAYRTSCRARR 181
>gb|AAP12861.1| At2g40095 [Arabidopsis thaliana] gi|26449647|dbj|BAC41948.1|
unknown protein [Arabidopsis thaliana]
gi|30688139|ref|NP_850325.1| expressed protein
[Arabidopsis thaliana]
Length = 209
Score = 105 bits (262), Expect = 7e-22
Identities = 59/149 (39%), Positives = 82/149 (54%), Gaps = 11/149 (7%)
Query: 32 SLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLIILCVY 91
S P+ + V D L+P L+ F+ ++ H Y F+ SL DIPL+S++RS +ILCVY
Sbjct: 39 SAPIFLAVADALLPSALLHRFSSPAT--LSSHLTNYDFRHSLIDIPLISIIRSAVILCVY 96
Query: 92 SICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKKSW 151
+CD P LS GPYL +CSI S++ +S+K F + + R
Sbjct: 97 GLCDGPKLSRGPYLTITMICSISSLIYVSLK-AAFVFGEPAIGDGGGNYFRA-------- 147
Query: 152 GMPVLFLSSVVFSLGHIVVACRTSCRARR 180
LFL S V ++GHIVVA RTSCR R+
Sbjct: 148 AEVALFLCSSVLAIGHIVVAYRTSCRERK 176
>gb|AAN15722.1| putative protein [Arabidopsis thaliana] gi|22530950|gb|AAM96979.1|
putative protein [Arabidopsis thaliana]
gi|30694339|ref|NP_191147.2| expressed protein
[Arabidopsis thaliana]
Length = 215
Score = 97.4 bits (241), Expect = 2e-19
Identities = 60/167 (35%), Positives = 83/167 (48%), Gaps = 29/167 (17%)
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLIS------NFTCIRCYGIT--KHFGRYAFKSSLTDIP 77
L + S P+ + V D ++P ++S N + T + Y F+ SL DIP
Sbjct: 34 LSFIIFSAPIFLAVADAVLPSAILSSSSPSLNRLSPSSFPATIYSYLSNYDFRYSLIDIP 93
Query: 78 LVSLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVF----TVNSRIE 133
L+S++RS IILCVY +CD P LS GPYL +CSI S++ +S K + +
Sbjct: 94 LISIIRSAIILCVYGLCDGPKLSRGPYLTITMICSISSLIYVSFKAAIVFGEPVIGGYFR 153
Query: 134 AEASASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
E A LFL S + ++GHIVVA RTSCR RR
Sbjct: 154 TEEMA-----------------LFLCSWILAIGHIVVAYRTSCRERR 183
>ref|NP_851019.1| expressed protein [Arabidopsis thaliana]
Length = 198
Score = 97.4 bits (241), Expect = 2e-19
Identities = 60/167 (35%), Positives = 83/167 (48%), Gaps = 29/167 (17%)
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLIS------NFTCIRCYGIT--KHFGRYAFKSSLTDIP 77
L + S P+ + V D ++P ++S N + T + Y F+ SL DIP
Sbjct: 34 LSFIIFSAPIFLAVADAVLPSAILSSSSPSLNRLSPSSFPATIYSYLSNYDFRYSLIDIP 93
Query: 78 LVSLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVF----TVNSRIE 133
L+S++RS IILCVY +CD P LS GPYL +CSI S++ +S K + +
Sbjct: 94 LISIIRSAIILCVYGLCDGPKLSRGPYLTITMICSISSLIYVSFKAAIVFGEPVIGGYFR 153
Query: 134 AEASASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
E A LFL S + ++GHIVVA RTSCR RR
Sbjct: 154 TEEMA-----------------LFLCSWILAIGHIVVAYRTSCRERR 183
>gb|AAN28874.1| At1g52750/F14G24_2 [Arabidopsis thaliana]
gi|19310418|gb|AAL84946.1| At1g52750/F14G24_2
[Arabidopsis thaliana]
Length = 523
Score = 74.7 bits (182), Expect = 1e-12
Identities = 42/71 (59%), Positives = 47/71 (66%), Gaps = 7/71 (9%)
Query: 110 LCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIV 169
L S I+LL VKTCVFTVNS +E + + HLK SWGMPVL LSS VF L H+V
Sbjct: 2 LSSFGGILLLLVKTCVFTVNSTLE-------EAKVYHLKISWGMPVLLLSSAVFGLAHVV 54
Query: 170 VACRTSCRARR 180
VA R SC ARR
Sbjct: 55 VAYRKSCGARR 65
>ref|XP_468592.1| Unknown protein [Oryza sativa (japonica cultivar-group)]
gi|25446696|gb|AAN74843.1| Unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 212
Score = 73.6 bits (179), Expect = 3e-12
Identities = 51/155 (32%), Positives = 75/155 (47%), Gaps = 12/155 (7%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCY--GITKHFGRYAFKSSLTDIPLVSLMRSL 85
L V + P+++VV+DLL+P L+SNF + + + F+SSL D+P VS RSL
Sbjct: 39 LLVCAPPLLIVVLDLLLPPALLSNFHRAANHPTSLIDQARGFHFRSSLVDLPAVSAARSL 98
Query: 86 IILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRL 145
+ILC Y+ C YL CS+ S+ + K V + A K Q +
Sbjct: 99 LILCAYTACG----GGAAYLWVAVACSVGSVCYVVAKAAVVFGAAPDRAVLGLQGKGQLV 154
Query: 146 HLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
+ +FL S+ + HI +A R SCR RR
Sbjct: 155 ------AVEAMFLMSLALAAAHIAMAYRASCRERR 183
>gb|AAP54806.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37536434|ref|NP_922519.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|18057098|gb|AAL58121.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 219
Score = 51.2 bits (121), Expect = 2e-05
Identities = 48/163 (29%), Positives = 69/163 (41%), Gaps = 26/163 (15%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYG----------ITKHFGRYAFKSSLTDIP 77
L V + PV+VV++DL +P L+S +R G + + F+SSL D+
Sbjct: 48 LLVCAPPVLVVILDLALPPALLS--ARLRGGGGGDDASFVAAVVAQARAFDFRSSLVDLL 105
Query: 78 LVSLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEAS 137
VS R+L+IL Y C YL V + S+ + K + R A A
Sbjct: 106 AVSAARALLILGAYMACGG---GGAAYLWVVATSAAGSVSYVLAKAAAAVLPRRGVAPAP 162
Query: 138 ASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
+ G + L SV + H+ VA RTSCR RR
Sbjct: 163 -----------EGKGPEPMLLLSVALAAAHLAVAYRTSCRERR 194
>sp|P49219|MTR1C_XENLA Melatonin receptor type 1C (MEL-1C-R) gi|505667|gb|AAA70166.1|
Mel-1c receptor subtype
Length = 420
Score = 36.2 bits (82), Expect = 0.54
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 19 SCTTRSKLDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGIT---KHFGRYAFKSSLTD 75
SCT + SS + VVVV +VP +++ F +R + + KH R FK LT
Sbjct: 181 SCTFAQTVS---SSYTITVVVVHFIVPLSVVT-FCYLRIWVLVIQVKHRVRQDFKQKLTQ 236
Query: 76 IPLVSLMRSLIILCVYSICDAP 97
L + + ++ ++++C AP
Sbjct: 237 TDLRNFLTMFVVFVLFAVCWAP 258
>ref|NP_914838.1| P0413C03.14 [Oryza sativa (japonica cultivar-group)]
Length = 592
Score = 36.2 bits (82), Expect = 0.54
Identities = 29/94 (30%), Positives = 41/94 (42%), Gaps = 15/94 (15%)
Query: 82 MRSLIILCVYSICDA--------PTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIE 133
M LI +C+ +C P L GP +G +C SI+ L T+
Sbjct: 95 MIRLIEICLALLCSRRGRQAWPPPLLERGPRVGVARICRRCSILDL-------TIVGLDP 147
Query: 134 AEASASLKRQRLHLKKSWGMPVLFLSSVVFSLGH 167
AE +L+ Q+ H K+ P L SVV+ LGH
Sbjct: 148 AENETTLQAQQPHYLKTCCKPSLDHGSVVYLLGH 181
>gb|AAB48390.1| Mel-1c(a) melatonin receptor [Xenopus laevis]
gi|1857129|gb|AAB48389.1| Mel-1c(a) melatonin receptor
[Xenopus laevis]
Length = 354
Score = 36.2 bits (82), Expect = 0.54
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 19 SCTTRSKLDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGIT---KHFGRYAFKSSLTD 75
SCT + SS + VVVV +VP +++ F +R + + KH R FK LT
Sbjct: 181 SCTFAQTVS---SSYTITVVVVHFIVPLSVVT-FCYLRIWVLVIQVKHRVRQDFKQKLTQ 236
Query: 76 IPLVSLMRSLIILCVYSICDAP 97
L + + ++ ++++C AP
Sbjct: 237 TDLRNFLTMFVVFVLFAVCWAP 258
>emb|CAI11649.1| novel protein similar to human and mouse androgen-induced 1 (AIG1)
[Danio rerio]
Length = 227
Score = 35.8 bits (81), Expect = 0.70
Identities = 27/92 (29%), Positives = 47/92 (50%), Gaps = 5/92 (5%)
Query: 81 LMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSV--KTCVFTVNSRIEAEASA 138
L+ ILC Y D P +H Y G+ + + +V+ +V CV T S + + SA
Sbjct: 14 LLSYFSILCNYKAIDMP--AHQTYGGSWKFLTFIDLVIQAVFFGVCVLTDLSSLLTKGSA 71
Query: 139 SLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVV 170
S++++R L+K G+ ++ + F +G VV
Sbjct: 72 SMEQER-QLRKLIGLRDWMMAVLAFPVGVFVV 102
>gb|AAB48392.1| Mel-1c(b) melatonin receptor [Xenopus laevis]
gi|1857133|gb|AAB48391.1| Mel-1c(b) melatonin receptor
[Xenopus laevis]
Length = 354
Score = 35.8 bits (81), Expect = 0.70
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 19 SCTTRSKLDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGIT---KHFGRYAFKSSLTD 75
SCT + SS + VVVV +VP +++ F +R + + KH R FK LT
Sbjct: 181 SCTFAQTVS---SSYTITVVVVHFIVPLSVVT-FCYLRIWVLVIQVKHRVRQDFKQKLTP 236
Query: 76 IPLVSLMRSLIILCVYSICDAP 97
L + + ++ ++++C AP
Sbjct: 237 TDLRNFLTMFVVFVLFAVCWAP 258
>gb|EAA40137.1| GLP_80_57878_54231 [Giardia lamblia ATCC 50803]
Length = 1215
Score = 34.3 bits (77), Expect = 2.0
Identities = 37/124 (29%), Positives = 58/124 (45%), Gaps = 31/124 (25%)
Query: 31 SSLPVVVVVVDL---LVPCVLIS-------NFTCIRCYGITKHFGRYAFKSSLTDIP--- 77
S PVVVV++ L L+P V +S + T + I +FG SSL+D+
Sbjct: 999 SYAPVVVVLLSLTSLLLPAVELSPGSSRSSSGTPVSVRAIGGYFGTGGSGSSLSDVARLQ 1058
Query: 78 -----LVSLMRSLIILCVYSICDAP------TLSHGPYLG-------TVTLCSILSIVLL 119
V+L S+I+ C+YSI AP +++ YLG + CS LS++
Sbjct: 1059 LVTNVCVALYGSIILRCMYSIIRAPFHLSGASVNATGYLGYILYDAAALLCCSTLSLIPF 1118
Query: 120 SVKT 123
++ T
Sbjct: 1119 TLLT 1122
>emb|CAA39077.1| glutamine-dependent carbamyl phosphate synthetase [Dictyostelium
discoideum]
Length = 1042
Score = 33.9 bits (76), Expect = 2.7
Identities = 22/82 (26%), Positives = 44/82 (52%), Gaps = 10/82 (12%)
Query: 104 YLGTVTLCSI-LSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKKSWGMPVLFLSSVV 162
+LG + C+I S+ S + C+ VN+R+ ++ + K+ G P+ F+S+ V
Sbjct: 264 HLGVIGECNIQYSLNPYSEEYCIIEVNARLSRSSALA--------SKATGYPLAFISAKV 315
Query: 163 FSLGHIVVACRTSCRARRSSCF 184
+LG+ + A R + + ++CF
Sbjct: 316 -ALGYDLAALRNTITKKTTACF 336
>ref|YP_077906.1| Acyltransferase 3 [Bacillus licheniformis ATCC 14580]
gi|52002326|gb|AAU22268.1| Acyltransferase 3 [Bacillus
licheniformis ATCC 14580]
Length = 335
Score = 33.9 bits (76), Expect = 2.7
Identities = 37/143 (25%), Positives = 61/143 (41%), Gaps = 12/143 (8%)
Query: 42 LLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLIILCVYSICDAPTLSH 101
LL+P + S + + +HF F S +D+ V + + YS+ D L+H
Sbjct: 68 LLLPYFIFSVLAYLFWVLVERHF---TFTGS-SDVDPVVPFKGIF----YSMPDDYKLTH 119
Query: 102 GPYLGTVTLCSILSIVLLSVKTC----VFTVNSRIEAEASASLKRQRLHLKKSWGMPVLF 157
P L +T I+ I+ VK +F V + + + A L Q L +K W + V
Sbjct: 120 NPALWFLTCLFIVEILFFIVKRWGKGRLFIVIAVLVSAALGFLNYQYLGIKLPWNIEVAL 179
Query: 158 LSSVVFSLGHIVVACRTSCRARR 180
+ V +S G ++ R RR
Sbjct: 180 TAIVFYSAGFLLKNNLKEMRDRR 202
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.343 0.149 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 266,626,581
Number of Sequences: 2540612
Number of extensions: 9122792
Number of successful extensions: 45951
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 45926
Number of HSP's gapped (non-prelim): 30
length of query: 197
length of database: 863,360,394
effective HSP length: 121
effective length of query: 76
effective length of database: 555,946,342
effective search space: 42251921992
effective search space used: 42251921992
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0338.4


