

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0335b.5
(1648 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
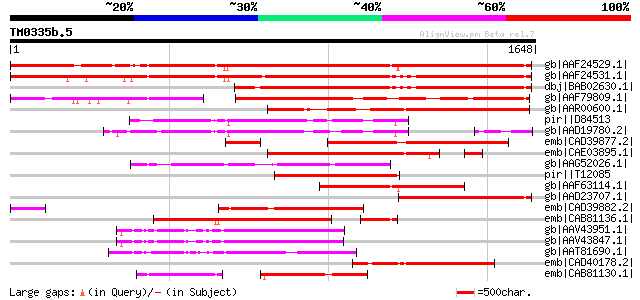
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF24529.1| F7F22.15 [Arabidopsis thaliana] 1537 0.0
gb|AAF24531.1| F7F22.17 [Arabidopsis thaliana] 1471 0.0
dbj|BAB02630.1| retroelement pol polyprotein-like [Arabidopsis t... 1161 0.0
gb|AAF79809.1| T32E20.9 [Arabidopsis thaliana] 1053 0.0
gb|AAR00600.1| putative reverse transcriptase [Oryza sativa (jap... 809 0.0
pir||D84513 probable retroelement pol polyprotein [imported] - A... 724 0.0
gb|AAD19780.2| hypothetical protein [Arabidopsis thaliana] 681 0.0
emb|CAD39877.2| OSJNBb0058J09.16 [Oryza sativa (japonica cultiva... 647 0.0
emb|CAE03895.1| OSJNBb0026I12.3 [Oryza sativa (japonica cultivar... 638 0.0
gb|AAG52026.1| polyprotein, putative; 77260-80472 [Arabidopsis t... 618 e-175
pir||T12085 reverse transcriptase homolog - fava bean (fragment)... 596 e-168
gb|AAF63114.1| Hypothetical protein [Arabidopsis thaliana] gi|25... 575 e-162
gb|AAD23707.1| putative retroelement pol polyprotein [Arabidopsi... 545 e-153
emb|CAD39882.2| OSJNBb0067G11.5 [Oryza sativa (japonica cultivar... 545 e-153
emb|CAB81136.1| putative athila transposon protein [Arabidopsis ... 535 e-150
gb|AAV43951.1| putative polyprotein [Oryza sativa (japonica cult... 523 e-146
gb|AAV43847.1| putative polyprotein [Oryza sativa (japonica cult... 518 e-145
gb|AAT81690.1| putative reverse transcriptase [Oryza sativa (jap... 514 e-144
emb|CAD40178.2| OSJNBa0061A09.17 [Oryza sativa (japonica cultiva... 461 e-128
emb|CAB81130.1| AT4g07600 [Arabidopsis thaliana] gi|5724765|gb|A... 456 e-126
>gb|AAF24529.1| F7F22.15 [Arabidopsis thaliana]
Length = 1862
Score = 1537 bits (3980), Expect = 0.0
Identities = 825/1718 (48%), Positives = 1108/1718 (64%), Gaps = 128/1718 (7%)
Query: 2 EDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAE 61
EDP H+++F R CN ++ VS + +L LFPFSL D A W + P S+TTW+D +
Sbjct: 178 EDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLPHDSITTWDDCKK 237
Query: 62 KFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYD 121
F ++FF + +L+N+I F+Q T E+ EAWE FK +C H T+A ++ Y
Sbjct: 238 AFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFKGYTNQCSHHGFTKASLLSTLYR 297
Query: 122 GLLYSSRFGLDAASSGEFDALPPQAGYDLIEKMAMRAMN-SEN-ERQIRRSSFEVETYDK 179
G+L R LD AS+G F + G++L+E +A N +EN +R +R ++ + + K
Sbjct: 298 GVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSNGNYNENCDRTVRGTADSDDKHRK 357
Query: 180 LI-ASNKKLSEEVAEMQKHIQDTKSIGAKVTSLECKFCGESHDSNQCTVND----DVSKV 234
I A N KL + KH+ D Q V D + +V
Sbjct: 358 EIKALNDKLDRILLSQHKHVHFLV------------------DDEQYEVQDGEGNQLEEV 399
Query: 235 RALTPGRGAPTSEQG-PLDQPQIVVKTVGKKSLEELIERFIDHTSSN--YKSHDMAIKSL 291
+ +G + P + ++ + ++ + S N + ++ A K
Sbjct: 400 SYINNNQGGYKGYNNFKTNNPNLSYRSTNVANPQDQVYPPQQQQSQNKPFVPYNQATKQT 459
Query: 292 ESQLGQLAKQMSEN------HQGKF---SSETLTLTEQENTAVVSTRSGRVLHEPKEKVE 342
G+ + E H GK E T+TE + G L K++ +
Sbjct: 460 SQLPGKAVQNPKEYAHAITLHSGKALPTREEPKTVTEDSED-----QDGEDLSLKKDQAD 514
Query: 343 GEKNEEAEVERDEGVME------ERAREPTRTRVVNTHTPELRKIPFPKALVKKNLDKQF 396
+ E D + + + PT V+ +P PK + KN +K
Sbjct: 515 KPLDLSLEQPLDLSLQQSLDPPLDSFTRPTTRPVIPAASPTA-----PKPVAVKNKEK-- 567
Query: 397 SKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETILMTEECSAILQRK 456
+ + IP +AL +P KF+KD++ +R + EV ++++ ECSAI+Q+K
Sbjct: 568 ---------VELRIPLVDALALIPDSHKFLKDLIVER--IQEVQGMVVLSHECSAIIQKK 616
Query: 457 M-PKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLGEVTPTMISLQMA 515
+ PKK DPGSFT+P + L LCDLGAS++LM L + KRL + ISL +A
Sbjct: 617 IIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGASVSLMPLSVAKRLGFTQYKSCNISLILA 676
Query: 516 DRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLATGEAEIKVAKGT 575
DRS++ P+G++E++ +++ P DFV+L+M+E+ K PLILGRPFLAT A I V KG
Sbjct: 677 DRSVRIPHGLLENLPIRIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGK 736
Query: 576 LTLKVGED-EVLFNIFDSLKHRADE-EVFRCEVVDELVCEEFVRISIKDPLETTIMEG-- 631
+ L +G+D + F++ D++K E ++F E +D+L E ++ +D L + + +
Sbjct: 737 IDLNLGKDFRMTFDVKDAMKKPTIEGQLFWIEEMDQLADELLEELAEEDHLNSALTKSGE 796
Query: 632 ---LEIEDQNLERELDSLYHEVNATLSQLESVSSLATKSIWKEE---------------- 672
L +E ++ LDS H+ E ++ AT+ + E
Sbjct: 797 DGFLHLETLGYQKLLDS--HKAMEESEPFEELNGPATEVMVMSEEGSTRVQPALSRTYSS 854
Query: 673 ----LTRDE-EIPI-----------EEKSELKSLPSSLKYAYLEEGENKPVILNSVLTPL 716
L+ DE PI K +LK LP L+YA+L PVI+N+ L
Sbjct: 855 NHSTLSTDEPREPIISTSDDWSELKAPKVDLKPLPKGLRYAFLGPNSTYPVIINAELNSD 914
Query: 717 KEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVR 776
+ LL LR ++ A+G+++ DIKGISP++C H+I LE ++P RRLNP++K+VV+
Sbjct: 915 EVNLLLSELRKYRRAIGYSLSDIKGISPSLCNHRIHLENESYSSIEPHRRLNPNLKEVVK 974
Query: 777 KEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRR 836
KEI+KLLDAGVIYPISDS WVSPV VPKK G+ VV NE +ELIPTR +T R+CIDYR+
Sbjct: 975 KEILKLLDAGVIYPISDSTWVSPVHCVPKKDGMIVVKNEKDELIPTRTITGHRMCIDYRK 1034
Query: 837 LNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFA 896
LN+ +RKDHFPLPFIDQML+RLA H YYCFLDGYSG+ QI + P DQEKT FTCPYG FA
Sbjct: 1035 LNAASRKDHFPLPFIDQMLERLANHPYYCFLDGYSGFFQIPIHPNDQEKTTFTCPYGTFA 1094
Query: 897 YKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQ 956
YKRMPFGLCNAPATFQRCM +IFSDLI+ +E+FMDDFSV+GP+F +CL NL VL RC+
Sbjct: 1095 YKRMPFGLCNAPATFQRCMTSIFSDLIKEMVEVFMDDFSVYGPSFSSCLLNLGRVLTRCE 1154
Query: 957 ETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAG 1016
ETNLVLNWEKCHFMV++GIVLGHK+SEKGIEVD+ K+EV+ +L PP +K IRSFLGHAG
Sbjct: 1155 ETNLVLNWEKCHFMVKEGIVLGHKISEKGIEVDKGKVEVMMQLQPPKTVKDIRSFLGHAG 1214
Query: 1017 FYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEI 1076
FYRRFIKDFSK+A+P+T LL KE F FD++CL++F++IK++LV+APV+ AP+W PFEI
Sbjct: 1215 FYRRFIKDFSKIARPLTRLLCKETEFKFDDDCLKSFQTIKDALVSAPVVRAPNWDYPFEI 1274
Query: 1077 MCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYIL 1136
MCDASD A+GAVL QK ++ L+VIYYASR L++AQ Y TTEKELL VVFA EKFR Y++
Sbjct: 1275 MCDASDYAVGAVLGQKIDKKLHVIYYASRTLDDAQGRYATTEKELLAVVFAFEKFRSYLV 1334
Query: 1137 GFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGG 1196
G KV V+TDHAALRHL+AK+D+KPRL+RW+LLLQEFD+EI+D++G +N ADHLSR+
Sbjct: 1335 GSKVTVYTDHAALRHLYAKKDTKPRLLRWILLLQEFDMEIVDKKGIENGAADHLSRMR-- 1392
Query: 1197 ACSPIPIQEEFSDEKLLAV-------STKE---------PLPWYVHFANFRVAGLIPHDL 1240
P+ I + +E+L+ V S KE PWY N+ G+ P +L
Sbjct: 1393 IEEPLLIDDSMPEEQLMVVEFFGKSYSGKEFHQLNAVEGESPWYADHVNYLACGVEPPNL 1452
Query: 1241 TWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCHGSSYGGHFSGER 1300
T ++KKF D Y WD+P+L+ +C D + RRC++E + E IL HCHGS+YGGHF+ +
Sbjct: 1453 TSYERKKFFRDIHHYYWDEPYLYTLCKDKIYRRCVSEDEVEGILLHCHGSAYGGHFATFK 1512
Query: 1301 TAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGIDFM 1360
T +K+LQ+GF+WPT+ ++++ FV CD CQR GNISRRNEMP ILE+E+FDVWGIDFM
Sbjct: 1513 TVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKGNISRRNEMPQNPILEVEIFDVWGIDFM 1572
Query: 1361 GPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGT 1420
GPFP S+G YILVAVDYVSKWVEA A TND+KVV+ K IF RFGVPR +ISDGG
Sbjct: 1573 GPFPSSYGNKYILVAVDYVSKWVEAIASPTNDAKVVLKLFKTIIFPRFGVPRVVISDGGK 1632
Query: 1421 HFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDD 1480
HF N+ FE+LL+K+GVKHKV+TPY+PQTSGQVEISNRE+K ILEK V +RKDWS KLDD
Sbjct: 1633 HFINKVFENLLKKHGVKHKVATPYNPQTSGQVEISNREIKTILEKTVGITRKDWSAKLDD 1692
Query: 1481 ALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWKVASEKRLLQLNE 1540
ALWAYRT FKTPIGT+PF+L++GK+CHLPVELE+KA WA++ LNFD K A EKRL+QL++
Sbjct: 1693 ALWAYRTTFKTPIGTTPFNLLYGKSCHLPVELEYKAMWAVKLLNFDIKTAEEKRLIQLSD 1752
Query: 1541 LDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVLLFNSRLRLFPGKLKSRWSGPF 1600
LDE RL AYES+ IYKE+TK +HD+KI+ ++F G VLLFNSRL+LFPGKLKSRWSGPF
Sbjct: 1753 LDEIRLEAYESSKIYKERTKLFHDKKIITKDFQVGDQVLLFNSRLKLFPGKLKSRWSGPF 1812
Query: 1601 VVKRVFPHGAVEVENPETKNIFTVNGQRLKVYQGGEVL 1638
+ V P+GAV + FTVNGQRLK Y ++L
Sbjct: 1813 CITEVRPYGAVTLTG--KSGDFTVNGQRLKKYLADQIL 1848
>gb|AAF24531.1| F7F22.17 [Arabidopsis thaliana]
Length = 1799
Score = 1471 bits (3807), Expect = 0.0
Identities = 814/1790 (45%), Positives = 1110/1790 (61%), Gaps = 200/1790 (11%)
Query: 2 EDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAE 61
EDP H+++F R CN ++ VS + +L LFPFSL D A W + P S+TTW+D +
Sbjct: 43 EDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLPHDSITTWDDCKK 102
Query: 62 KFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYD 121
F ++FF + +L+N+I F+Q T E+ EAWE FK +CP H T+A ++ Y
Sbjct: 103 VFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFKGYTNQCPHHGFTKASLLSTLYR 162
Query: 122 GLLYSSRFGLDAASSGEFDALPPQAGYDLIEKMAMRA--MNSENERQIRRSSFEVETYDK 179
G+L R LD AS+G F + G++L+E +A N + +R +R ++ + + K
Sbjct: 163 GVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSDGNYNEDCDRTVRGTADSDDKHRK 222
Query: 180 ------------LIASNKKLSEEVAEMQKHIQDTKSIGAKVTSLECKFCGESHDSNQCTV 227
L++ +K + V + Q +QD + + S G N
Sbjct: 223 EIKALNDKLDRILLSQHKHVHFLVDDEQYEVQDGEGNQLEEVSYINNNQGGYKGYNYFKT 282
Query: 228 NDDVSKVRALT-----------------------------------------PGRGAPTS 246
N+ R+ P AP
Sbjct: 283 NNPNLSYRSTNVANPQDQVYPPQQQQSQNKPFVPYNQGFVPKQQFQGNYQQQPPGFAPPQ 342
Query: 247 EQGPL----DQPQIVVKTV-GKKS----LEELIERFIDHTSSNYKSHDMAIKSLESQLGQ 297
QGP D Q++ + + G+ S + + I + +Y ++ +++L++++
Sbjct: 343 HQGPAAPDADMKQMLQQLLHGQASCSMEMAKKISELHNKLDCSYNDLNVKMETLDTKVRY 402
Query: 298 LAKQMSENHQGKFSSET---LTLTEQENTAVVSTRSGRVL---HEPK---EKVEGEKNEE 348
L + + K +S+ +E ++ RSG+ L EPK E E + E+
Sbjct: 403 LEGHSTSSSATKQTSQLPGKAVQNPKEYAHAITLRSGKALPTREEPKTVTEDSEDQDGED 462
Query: 349 AEVERDEGVME---------------------ERAREPTRTRVVNTHTPELRK------- 380
+E+D+ + PT V+ +P K
Sbjct: 463 LSLEKDQADKPLDLSLEQPLDLSLQQSLDPPLDSFTRPTTRPVIPAASPTAPKPVAVKNK 522
Query: 381 ------------IPFPKALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKD 428
+PFP K DK + F + K++ + IP +AL +P KF+KD
Sbjct: 523 EKVFVPPPYKSQLPFPGRHKKALADKYRAMFAKNIKEVELRIPLVDALALIPDSHKFLKD 582
Query: 429 ILSKRRKLSEVDETILMTEECSAILQRKM-PKKRRDPGSFTIPVEIEGLTMVEALCDLGA 487
++ +R + EV ++++ ECSAI+Q+K+ PKK DPGSFT+P + L LCDLGA
Sbjct: 583 LIVER--IQEVQGMVVLSHECSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGA 640
Query: 488 SINLMSLRMFKRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDM 547
S++LM L + KRL + ISL +ADRS++ P+G++E++ +++ P DFV+L+M
Sbjct: 641 SVSLMPLSVAKRLGFTQYKSCNISLILADRSVRIPHGLLENLPIRIGAVEIPTDFVVLEM 700
Query: 548 EEDSKVPLILGRPFLATGEAEIKVAKGTLTLKVGED-EVLFNIFDSLKHRADE-EVFRCE 605
+E+ K PLILGRPFLAT A I V KG + L +G+D + F++ D++K E ++F E
Sbjct: 701 DEEPKDPLILGRPFLATAGAMIDVKKGKIDLNLGKDFRMTFDVKDAMKKPTIEGQLFWIE 760
Query: 606 VVDELVCEEFVRISIKDPLETTIMEG-----LEIEDQNLERELDSLYHEVNATLSQLESV 660
+D+L E ++ +D L + + + L +E ++ LDS H+ E +
Sbjct: 761 EMDQLADELLEELAEEDHLNSALTKSGEDGFLHLETLGYQKLLDS--HKAMEESEPFEEL 818
Query: 661 SSLATKSIWKEE--------------------LTRDEE----IPIEE--------KSELK 688
+ AT+ + E L+ DE IP + K +LK
Sbjct: 819 NGPATEVMVMSEEGSTRVQPALSRTYSSNHSTLSTDEPREPIIPTSDDWSELKAPKVDLK 878
Query: 689 SLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAICM 748
LP L+YA+L PVI+N+ L + LL L+ ++ A+G+++ DIKGISP++C
Sbjct: 879 PLPKGLRYAFLGPNSTYPVIINAELNSDEVNLLLSELKKYRRAIGYSLSDIKGISPSLCN 938
Query: 749 HKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGG 808
H+I LE ++PQRRLNP++K+VV+KEI+KLLDAGVIYPISD + VPKK G
Sbjct: 939 HRIHLENESYSSIEPQRRLNPNLKEVVKKEILKLLDAGVIYPISD------MHCVPKKDG 992
Query: 809 ITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
+TVV NE +ELIPTR +T R+CIDYR+LN+ +RKDHFPLPFIDQML+RLA H YYCFLD
Sbjct: 993 MTVVKNEKDELIPTRTITGHRMCIDYRKLNAASRKDHFPLPFIDQMLERLANHPYYCFLD 1052
Query: 869 GYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIE 928
GY+G+ QI + P DQEKT FTCPYG FAYKRMPFGLCNAPATFQRCM +IFSDLIE +E
Sbjct: 1053 GYNGFFQIPIHPNDQEKTTFTCPYGTFAYKRMPFGLCNAPATFQRCMTSIFSDLIEEMVE 1112
Query: 929 IFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEV 988
+FMDDFSV+GP+F +CL NL VL RC+ETNLVLNWEKCHFMV++GIVLGHK+SEKGIEV
Sbjct: 1113 VFMDDFSVYGPSFSSCLLNLGRVLTRCEETNLVLNWEKCHFMVKEGIVLGHKISEKGIEV 1172
Query: 989 DRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENC 1048
D+ K+EV+ +L PP +K IRSFLGHAGFYRRFIKDFSK+A+P+T LL KE F FD++C
Sbjct: 1173 DKGKVEVMMQLQPPKTVKDIRSFLGHAGFYRRFIKDFSKIARPLTRLLCKETEFKFDDDC 1232
Query: 1049 LRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLN 1108
L++F++IK++LV+APV+ AP+W PFEIMCDASD A+GAVL QK ++ L+VIYYASR L+
Sbjct: 1233 LKSFQTIKDALVSAPVVRAPNWDYPFEIMCDASDYAVGAVLGQKIDKKLHVIYYASRTLD 1292
Query: 1109 EAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLL 1168
+AQ Y TTEKELL VVFA EKFR Y++G KV V+TDHAALRHL+AK+D+KPRL+RW+LL
Sbjct: 1293 DAQGRYATTEKELLVVVFAFEKFRSYLVGSKVTVYTDHAALRHLYAKKDTKPRLLRWILL 1352
Query: 1169 LQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFA 1228
LQEFD+EI+D++G +N ADHLSR+ P+ I + +E+L+ V F
Sbjct: 1353 LQEFDMEIVDKKGIENGAADHLSRMR--IEEPLLIDDSMPEEQLMV----------VEFF 1400
Query: 1229 NFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCH 1288
+G K H + + P+ RC++E + E IL HCH
Sbjct: 1401 GKSYSG------------KEFHQLNAVEGESPW-----------RCVSEDEVEGILLHCH 1437
Query: 1289 GSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILE 1348
GS+YGGHF+ +T +K+LQ+GF+WPT+ ++++ FV CD CQR GNISRRNEMP ILE
Sbjct: 1438 GSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKGNISRRNEMPQNPILE 1497
Query: 1349 IELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRF 1408
+E+FDVWGIDFMGPFP S+G YILVAVDYVSKWVEA A TND+KVV+ K IF RF
Sbjct: 1498 VEIFDVWGIDFMGPFPSSYGNKYILVAVDYVSKWVEAIASPTNDAKVVLKLFKTIIFPRF 1557
Query: 1409 GVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVN 1468
GVPR +ISDGG HF N+ FE+LL+K+GVKHKV+TPY+PQTSGQVEISNRE+K ILEK V
Sbjct: 1558 GVPRVVISDGGKHFINKVFENLLKKHGVKHKVATPYNPQTSGQVEISNREIKTILEKTVG 1617
Query: 1469 SSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWK 1528
+RKDWS KLDDALWAYRTAFKTPIGT+PF+L++GK+CHLPVELE+KA WA++ LNFD K
Sbjct: 1618 ITRKDWSAKLDDALWAYRTAFKTPIGTTPFNLLYGKSCHLPVELEYKAMWAVKLLNFDIK 1677
Query: 1529 VASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVLLFNSRLRLF 1588
A EKRL+QL++LDE RL AYES+ IYKE+TK +HD+KI+ ++F G VLLFNSRL+LF
Sbjct: 1678 TAEEKRLIQLSDLDEIRLEAYESSKIYKERTKLFHDKKIITKDFQFGVQVLLFNSRLKLF 1737
Query: 1589 PGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVYQGGEVL 1638
PGKLKSRWSGPF + V P+GAV + FTVNGQRLK Y ++L
Sbjct: 1738 PGKLKSRWSGPFCITEVRPYGAVTLAG--KSGDFTVNGQRLKKYLADQIL 1785
>dbj|BAB02630.1| retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 897
Score = 1161 bits (3004), Expect = 0.0
Identities = 568/932 (60%), Positives = 708/932 (75%), Gaps = 49/932 (5%)
Query: 707 VILNSVLTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRR 766
+I+N+ L + L LR ++ A+G+++ DIKGISP++C H+I LE ++PQRR
Sbjct: 1 MIINAELNSDEVNLQLSELRKYRRAIGYSLSDIKGISPSLCNHRIHLENESYSSIEPQRR 60
Query: 767 LNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVT 826
LNP+ DAGVIYPISDS WVS V VPKKGG+TVV NE +ELIPTR +T
Sbjct: 61 LNPNF------------DAGVIYPISDSTWVSLVYCVPKKGGMTVVKNEKDELIPTRTIT 108
Query: 827 KWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKT 886
R+CIDYR+LN+ +RKDHFPLPFIDQML+RLA H YYCFLDGYSG+ QI + P DQEKT
Sbjct: 109 GHRMCIDYRKLNAASRKDHFPLPFIDQMLERLANHPYYCFLDGYSGFFQIPIHPNDQEKT 168
Query: 887 AFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLG 946
FTCPYG FAYKRMPFGLCNAPATFQRCM +IFSDLIE +E+FMDDFS +GP+F +CL
Sbjct: 169 TFTCPYGTFAYKRMPFGLCNAPATFQRCMTSIFSDLIEEMVEVFMDDFSGYGPSFSSCLL 228
Query: 947 NLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIK 1006
NL VL RC+ETNLVLNWEKCHFMV++GIVLGHK+SEKGIEVD+ K+EV+ +L PP +K
Sbjct: 229 NLGRVLTRCEETNLVLNWEKCHFMVKEGIVLGHKISEKGIEVDKGKVEVMMQLQPPKTVK 288
Query: 1007 GIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIV 1066
IRSFLGHAGFYRRFIKDFSK+ +P+T LL KE F FDE+CL++F++IK++LV+APV+
Sbjct: 289 EIRSFLGHAGFYRRFIKDFSKIVRPLTRLLCKETEFEFDEDCLKSFQTIKDALVSAPVVR 348
Query: 1067 APDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVF 1126
AP+W PFEIMCDASD +GAVL QK ++ L+VIYYASR L++AQ Y TTEKELL VVF
Sbjct: 349 APNWDYPFEIMCDASDYVVGAVLGQKIDKKLHVIYYASRTLDDAQGRYATTEKELLAVVF 408
Query: 1127 ACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSV 1186
A EKFR Y++G KV V+TDHAALRHL+AK+D+KPRL+RW+LLLQEFD+EI+D++G +N
Sbjct: 409 AFEKFRSYLVGSKVTVYTDHAALRHLYAKKDTKPRLLRWILLLQEFDMEIVDKKGIENGA 468
Query: 1187 ADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFANFRVAGLIPHDLTWQQKK 1246
ADHL R+ P+PI + +E+L+ V F +G
Sbjct: 469 ADHLLRMR--IEEPLPIDDSMPEEQLMV----------VEFFGKSYSG------------ 504
Query: 1247 KFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCHGSSYGGHFSGERTAAKVL 1306
K H + P+ RC++E + E IL HCH S+YGGHF+ +T +K+L
Sbjct: 505 KEFHQLNDVEGESPW-----------RCVSEDEVEGILLHCHCSAYGGHFATFKTVSKIL 553
Query: 1307 QSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGIDFMGPFPPS 1366
Q+GF+WPT+ ++++ FV CD CQR NISR NEMP I+E+E+FDVWGIDFMG FP S
Sbjct: 554 QAGFWWPTMFKDAQEFVSKCDSCQRKDNISRINEMPQNPIVEVEIFDVWGIDFMGLFPSS 613
Query: 1367 FGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRA 1426
+G YILVA+DYVSKWVEA A+ TND+KVV+ K IF RFGVPR +ISDGG HF N+
Sbjct: 614 YGNKYILVAIDYVSKWVEAIAIPTNDAKVVLKLFKTIIFPRFGVPRVVISDGGKHFINKV 673
Query: 1427 FESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYR 1486
FE+LL+K+GVKHKV+TPYHPQTSGQVEIS+RE+K ILEK V +RKDWS KLDDALWAYR
Sbjct: 674 FENLLKKHGVKHKVATPYHPQTSGQVEISDREIKTILEKTVGITRKDWSAKLDDALWAYR 733
Query: 1487 TAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRL 1546
TAFKTPIGT+PF++++GK+CHLPVELE+KA WA++ LNFD K A EKRL+QL++LDE RL
Sbjct: 734 TAFKTPIGTTPFNILYGKSCHLPVELEYKAMWAVKLLNFDIKTAEEKRLIQLSDLDEIRL 793
Query: 1547 RAYESASIYKEKTKKWHDRKILNREFVSGQLVLLFNSRLRLFPGKLKSRWSGPFVVKRVF 1606
AYES+ IYKE+TK +HD+KI+ ++F G VLLFNSRL+LFPGKLKSRWSGPF + +V
Sbjct: 794 EAYESSKIYKERTKLFHDKKIITKDFQVGDQVLLFNSRLKLFPGKLKSRWSGPFCITKVR 853
Query: 1607 PHGAVEVENPETKNIFTVNGQRLKVYQGGEVL 1638
P+GAV + FTVNGQRLK Y ++L
Sbjct: 854 PYGAVTLAG--KSGDFTVNGQRLKKYLADQIL 883
>gb|AAF79809.1| T32E20.9 [Arabidopsis thaliana]
Length = 1586
Score = 1053 bits (2722), Expect = 0.0
Identities = 524/925 (56%), Positives = 651/925 (69%), Gaps = 115/925 (12%)
Query: 708 ILNSVLTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRL 767
I N++ P E + + A+G+++DDIKGISP +C H+I LE ++PQRRL
Sbjct: 757 ITNTMKKPTIEGNIFWIEEMDMKAIGYSLDDIKGISPTLCTHRIHLENESYSSIEPQRRL 816
Query: 768 NPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTK 827
NP++K+VV+KEI+KLLDAGVIYPISDS WVSPV VPKKGG+TVV N +ELIPTR +T
Sbjct: 817 NPNLKEVVKKEILKLLDAGVIYPISDSTWVSPVHCVPKKGGMTVVKNSKDELIPTRTITG 876
Query: 828 WRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTA 887
R+CI+YR+LN +RK+HFPLPFID ML+RLA H YYCFLD YSG+ QI + P DQ KT
Sbjct: 877 HRMCIEYRKLNVASRKEHFPLPFIDHMLERLANHPYYCFLDSYSGFFQIPIHPNDQGKTT 936
Query: 888 FTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGN 947
FTCPYG FAYKRMPFGLCNAPATFQRCM +IFSDLIE +E+FMDDFSV+G +F +CL N
Sbjct: 937 FTCPYGTFAYKRMPFGLCNAPATFQRCMTSIFSDLIEEMVEVFMDDFSVYGSSFSSCLLN 996
Query: 948 LALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKG 1007
L VLKRC+ETNLVLNWEKCHFMVR+GIVLG K+SE+GIEVD+AKI+V+ +L PP +K
Sbjct: 997 LCRVLKRCEETNLVLNWEKCHFMVREGIVLGRKISEEGIEVDKAKIDVMMQLQPPKTVKD 1056
Query: 1008 IRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVA 1067
IRSFLGHAGFYR FIKDFSKLA+P+T LL KE F FD+ CL AF+ IKE+L+TAP++ A
Sbjct: 1057 IRSFLGHAGFYRIFIKDFSKLARPLTRLLCKETEFAFDDECLTAFKLIKEALITAPIVQA 1116
Query: 1068 PDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFA 1127
P+W PFEI+ +++AQ Y TTEKELL VVFA
Sbjct: 1117 PNWDFPFEII----------------------------TMDDAQVRYATTEKELLAVVFA 1148
Query: 1128 CEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVA 1187
EKFR Y++G KV ++TDHAALRH++AK+D+KPRL+RW+LLLQEFD+EI+D++G +N
Sbjct: 1149 FEKFRSYLVGSKVTIYTDHAALRHIYAKKDTKPRLLRWILLLQEFDMEIVDKKGIEN--- 1205
Query: 1188 DHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFANFRVAGLIPHDLTWQQKKK 1247
E LPWY N+ V+G P +L+ +KKK
Sbjct: 1206 -------------------------------EKLPWYADHVNYLVSGEEPPNLSSYEKKK 1234
Query: 1248 FLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCHGSSYGGHFSGERTAAKVLQ 1307
F D + WD+P+L+ +C D + R C++E + E IL HCHG +YGGHF+ +T +K+LQ
Sbjct: 1235 FFKDINHFYWDEPYLYTLCKDKIYRTCVSEDEIEGILLHCHGFAYGGHFATFKTMSKILQ 1294
Query: 1308 SGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGIDFMGPFPPSF 1367
+ E+E FDVWGIDFMGPFP S+
Sbjct: 1295 A---------------------------------------EVENFDVWGIDFMGPFPSSY 1315
Query: 1368 GC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAF 1427
G YILVA+DYVSKWVEA A TND++VV+ K IF RFGVPR +ISDGG HF N+ F
Sbjct: 1316 GNKYILVAIDYVSKWVEAIASHTNDARVVLKLFKTIIFPRFGVPRIVISDGGKHFINKGF 1375
Query: 1428 ESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRT 1487
E+LL+K+GVKHK VEISNRE+K ILEK V S+RKDWS KL+D LWAYRT
Sbjct: 1376 ENLLKKHGVKHK------------VEISNREIKAILEKTVGSTRKDWSAKLNDTLWAYRT 1423
Query: 1488 AFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLR 1547
AFKTPIGT+PF+L++GK+CHLPVELE+KA WA++ LNFD K A EKRL+QLN+L++ RL
Sbjct: 1424 AFKTPIGTTPFNLLYGKSCHLPVELEYKAMWAVKLLNFDIKTAEEKRLIQLNDLNKIRLE 1483
Query: 1548 AYESASIYKEKTKKWHDRKILNREFVSGQLVLLFNSRLRLFPGKLKSRWSGPFVVKRVFP 1607
AYES+ IYKE+TK +HD+KI++R+F G VLLFNSRLRLFPGKLKSRWSGPF V V P
Sbjct: 1484 AYESSKIYKERTKSFHDKKIVSRDFKVGDQVLLFNSRLRLFPGKLKSRWSGPFSVTAVRP 1543
Query: 1608 HGAVEVENPETKNIFTVNGQRLKVY 1632
+GA+ + FTVNGQRLK Y
Sbjct: 1544 YGAITLAG--KNGDFTVNGQRLKKY 1566
Score = 265 bits (677), Expect = 1e-68
Identities = 204/707 (28%), Positives = 327/707 (45%), Gaps = 121/707 (17%)
Query: 2 EDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAE 61
EDP H+++F R C+ ++ VS + +L LFPFSL D A +W S PQGS+T+W D +
Sbjct: 92 EDPLDHLDEFDRLCSLTKINRVSEDGFKLRLFPFSLGDKAHQWEKSLPQGSITSWNDCKK 151
Query: 62 KFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYD 121
F +FF S +L+NDI FTQ+ +E YEAWE FK +CP H +
Sbjct: 152 AFLAKFFSNSRTARLRNDISGFTQTNNETFYEAWERFKGYQTQCPHHEML---------- 201
Query: 122 GLLYSSRFGLDAASSGEFDALPPQAGYDLIEKMAMR--AMNSENERQIRRSSFEVETYDK 179
LD AS+G F + G++++E +A N + +R IR SS E + +
Sbjct: 202 ---------LDTASNGNFLNKDVEDGWEVVENLAQSDGNYNEDYDRSIRTSSDSDEKHRR 252
Query: 180 -LIASNKKLSEEVAEMQKHI-----------QDTKSIGAKVTS----------------- 210
+ A N KL + + QKHI QD +++ + S
Sbjct: 253 EMKAMNDKLDKLLLMQQKHIHFLGDDETLQVQDGETLQLEEVSYVQNQGGYNKGFNNFKQ 312
Query: 211 ----LECKFCGESHDSNQCTVNDDVSKVRALTP---GRGAPTSEQ------------GPL 251
L + ++ +Q + ++ + P G+G +Q G
Sbjct: 313 NHPNLSYRSTNVANRQDQVYPSQQQNQPKPFVPYNQGQGYVPKQQYQGNYQQQLPPPGFT 372
Query: 252 DQPQIVVKTVGKKSLEELIERFID----------------HTSSNYKSHDMAIK--SLES 293
Q Q T L+ ++++ + H + +D+ IK +L S
Sbjct: 373 QQQQQPALTTPDSDLKNMLQQILQGQATGAMDLSKRMAEIHNKVDCSYNDINIKVEALTS 432
Query: 294 QLGQLAKQMSENHQGKFS--SETLTLTEQENTAVVSTRSGRVL---HEPKEKVEGEKNEE 348
++ + Q KF+ S +E ++ RSG+ L P + E +++
Sbjct: 433 KIRYIEGQTGSTAAPKFTGPSGKSMSNSKEYAHAITLRSGKELPTKESPNQNTEDSLDQD 492
Query: 349 AEVERDEGVMEERA------REPTRTRV------------------VNTHTPELRKIPFP 384
E G E+A +PTR V P +PFP
Sbjct: 493 GEDFCQNGNSAEKAIEEPILHQPTRPLAPAASPLVEKPAAAKTKENVFIPPPYKPPLPFP 552
Query: 385 KALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETIL 444
K + K + + K L + +P + L +P K++KD++++R + EV ++
Sbjct: 553 GRFKKVMIQKYKALLEKQLKNLEVTMPLVDCLALIPDSNKYVKDMITER--IKEVQGMVV 610
Query: 445 MTEECSAILQRK-MPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLG 503
++ ECSAI+Q+K +PKK DPGSFT+P + L + LCDLGAS++LM L + K+L
Sbjct: 611 LSHECSAIIQQKIIPKKLGDPGSFTLPCALGPLAFSKCLCDLGASVSLMPLPVAKKLGFN 670
Query: 504 EVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLA 563
+ P ISL +ADRS++ +G++ED+ V + P DFV+L+M+E+ K PLILGRPFLA
Sbjct: 671 KYKPCNISLILADRSVRISHGLLEDLPVMIGVVEVPTDFVVLEMDEEPKDPLILGRPFLA 730
Query: 564 TGEAEIKVAKGTLTLKVGED-EVLFNIFDSLKHRADE-EVFRCEVVD 608
A I V KG + L +G D ++ F+I +++K E +F E +D
Sbjct: 731 RARAIIDVKKGKIDLNLGRDLKMTFDITNTMKKPTIEGNIFWIEEMD 777
>gb|AAR00600.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|50901488|ref|XP_463177.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 743
Score = 809 bits (2090), Expect = 0.0
Identities = 405/825 (49%), Positives = 538/825 (65%), Gaps = 105/825 (12%)
Query: 809 ITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
+TVVAN NELIP R VT WR+CIDYR+LN T++DHFPLPFID+ML+R+A H ++CFLD
Sbjct: 1 MTVVANAQNELIPQRTVTGWRMCIDYRKLNKATKEDHFPLPFIDEMLERMANHSFFCFLD 60
Query: 869 GYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIE 928
GYSGY+QI + PEDQ KT FTCPYG +AY+RM FGLCNAPA+FQRCM +IFSD+IE
Sbjct: 61 GYSGYHQIPIHPEDQSKTTFTCPYGTYAYRRMSFGLCNAPASFQRCMMSIFSDMIE---- 116
Query: 929 IFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEV 988
D VF +F V G
Sbjct: 117 ---DIMEVFMDDFS---------------------------------VYG---------- 130
Query: 989 DRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENC 1048
+ +++ PP NIKGIRSFLGHAGFYRRFIKDFS +A+P+TNLL K+APF D+ C
Sbjct: 131 -----KTFDQIRPPVNIKGIRSFLGHAGFYRRFIKDFSTIARPLTNLLAKDAPFESDDAC 185
Query: 1049 LRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLN 1108
L++FE +K++LV+AP+I PDW+L FEIMCDASD A+GA
Sbjct: 186 LKSFEVLKKALVSAPIIQPPDWTLSFEIMCDASDFAVGA--------------------- 224
Query: 1109 EAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLL 1168
FR Y++ KVI++TDHAAL++ K+D+KPRL+RW+LL
Sbjct: 225 ----------------------FRSYLVRAKVIIYTDHAALKYPLTKKDAKPRLLRWILL 262
Query: 1169 LQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFA 1228
LQEFDLEI D++G +NSVADHLSRL+ PI + D+ L+ V ++ PWY +
Sbjct: 263 LQEFDLEIKDKKGVENSVADHLSRLQITNMPEQPINDFLRDDMLMTV--RDSNPWYANIV 320
Query: 1229 NFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCH 1288
N+ V IP + ++ ++++ ++WD+P+L+++CSDG++RRC+ + KI+ CH
Sbjct: 321 NYMVFKYIPPG---KNQRNLKYESRRHIWDEPYLYRVCSDGLLRRCVPTEEGLKIIERCH 377
Query: 1289 GSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILE 1348
S YGGH+ RT AK+ QSGF+WPT++ +S+ FV C CQR G I+ R+ MPL L+
Sbjct: 378 ASPYGGHYGVFRTQAKIWQSGFFWPTMYEDSKEFVRRCTSCQRQGGITARDAMPLTYNLQ 437
Query: 1349 IELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRF 1408
+E+FDVWGIDFMGPFP S C YILVAVDYVSKWV+A S D++ + IF +F
Sbjct: 438 VEIFDVWGIDFMGPFPKSRNCEYILVAVDYVSKWVKAMPCSATDARHAKKMFIEIIFPKF 497
Query: 1409 GVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVN 1468
PR +ISDGG+HF ++ F LL + G KH V+TPYHPQTSGQ E SN+++K IL+K VN
Sbjct: 498 DTPRMVISDGGSHFIDKTFRDLLREMGAKHNVATPYHPQTSGQAETSNKQIKNILQKTVN 557
Query: 1469 SSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWK 1528
W +L DAL AY A+KTPI SP+ +V+ K C LPVELEH+AYWAIR N D++
Sbjct: 558 KMGTGWKDRLPDALCAYWIAYKTPIRMSPYQIVYRKPCRLPVELEHRAYWAIRNWNMDFE 617
Query: 1529 VASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVLLFNSRLRLF 1588
A E R +Q+ EL+E+R +AY +A IYKE+TK+WHD++ ++F G VL+FNSR++LF
Sbjct: 618 GAGEWRKMQIAELEEWREKAYHNAKIYKERTKRWHDKRKKIKKFKPGDKVLMFNSRVKLF 677
Query: 1589 -PGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVY 1632
GKL+S+W GPF V HGA+ + + E NIF VNGQRLK++
Sbjct: 678 GHGKLRSKWEGPFDVIDTSSHGAITLRD-ELGNIFKVNGQRLKIF 721
>pir||D84513 probable retroelement pol polyprotein [imported] - Arabidopsis
thaliana
Length = 841
Score = 724 bits (1870), Expect = 0.0
Identities = 421/916 (45%), Positives = 548/916 (58%), Gaps = 144/916 (15%)
Query: 376 PELRKIPFPKALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRK 435
P + K+PFP K +K++S F E+ ++L P
Sbjct: 19 PYVPKLPFPGRQRKIQREKEYSLFDEIMRQLQRYSP------------------------ 54
Query: 436 LSEVDETILMTEECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLR 495
ECSAILQ +P KR DPGSF +P I T LCDLGA ++LM
Sbjct: 55 ------------ECSAILQNVIPVKREDPGSFVLPSRIGEYTFDRCLCDLGAGVSLMPFS 102
Query: 496 MFKRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPL 555
+ KRL TPT +SL + DRS+ P G+ EDV V+V F P DFVI++++E+ + L
Sbjct: 103 VAKRLGDTNFTPTKMSLVLGDRSISFPVGVAEDVQVRVGNFYIPTDFVIIELDEEPRHRL 162
Query: 556 ILGRPFLATGEAEIKVAKGTLTLKVGEDEVLFNIFDSLKHRADE-EVFRCEVVDELVCEE 614
ILGRPFL A I V K + L++G+ FN+ + E + F +++DELV E
Sbjct: 163 ILGRPFLNIVAALIDVRKSKINLRIGDIVQEFNMERIMSKPTTECQTFWVDIMDELVNEL 222
Query: 615 FVRISIKDPLETTIMEGLEIEDQNLERELDSLYHEVNATLSQLESVSSLATKSIWKEELT 674
++ +DPL+T + + E E L E +++ SS TK + EL
Sbjct: 223 LAELNTEDPLQTVLTKE--------ESEFGYL-GEATTRFARILDSSSPMTKVVAFAELG 273
Query: 675 RDEEIPIEE-------------------KSELKSLPSSLKYAYLEEGENKPVILNSVLTP 715
+E +E+ K ELK LP+ L+ A+L VI+N L
Sbjct: 274 DNE---VEKALVVSSPKDCDDWSELNAPKMELKPLPARLRVAFLGPNSTYLVIINPELNN 330
Query: 716 LKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVV 775
++ LL LR ++ ALG+++DDI GISP +CMH I LE V+ QRRLN +++DVV
Sbjct: 331 VESALLLCELRKYRKALGYSLDDITGISPTLCMHMIHLEGESITSVEHQRRLNSNLRDVV 390
Query: 776 RKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYR 835
+KEI+KLLDAG+IYPISDS WVSPV VVPKKGG+ V+ NE NELIPTR VT R+CIDYR
Sbjct: 391 KKEIMKLLDAGIIYPISDSTWVSPVHVVPKKGGVIVIKNEKNELIPTRTVTGHRMCIDYR 450
Query: 836 RLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVF 895
+LNS TRKD+FPL FIDQML+RL+ YYCFLDGY G+ QI + P+DQEKT FTCPYG F
Sbjct: 451 KLNSATRKDNFPLSFIDQMLERLSNQPYYCFLDGYLGFFQILIHPDDQEKTTFTCPYGTF 510
Query: 896 AYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRC 955
AY+RMPFGLCNAPATFQ CM IFSD+IE +E+F+DDF LVL+RC
Sbjct: 511 AYRRMPFGLCNAPATFQHCMKYIFSDMIEDFMEVFIDDF---------------LVLQRC 555
Query: 956 QETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHA 1015
++ +LVLNWEK HFMVRDGIVLGHK+SEKG+EVDRAKIE++ FLGHA
Sbjct: 556 EDKHLVLNWEKSHFMVRDGIVLGHKISEKGVEVDRAKIEIMR-------------FLGHA 602
Query: 1016 GFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFE 1075
GFYRRFIKDFSK+A+P T LL KE FD+ CL+AF +KESLV+A ++ P+W LPFE
Sbjct: 603 GFYRRFIKDFSKIARPCTQLLCKEQKSEFDDTCLQAFNIVKESLVSALIVQPPEWELPFE 662
Query: 1076 IMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYI 1135
++CDASD +GAVL Q+K++ L+ IYYASR L+ AQ
Sbjct: 663 VICDASDYVVGAVLGQRKDKKLHAIYYASRTLDGAQ------------------------ 698
Query: 1136 LGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRL-- 1193
VIVHT HAALR+L +K+D+KPRL+RW+LLLQ+FDLEI D++G +N VAD+LSRL
Sbjct: 699 ----VIVHTVHAALRYLLSKKDAKPRLLRWILLLQDFDLEIKDKKGIENEVADNLSRLRV 754
Query: 1194 --EGGACSPIP------IQEEFSDE---------KLLAVSTKEP-LPWYVHFANFRVAGL 1235
E +P I++ + DE +L+ + T E LPWY FAN+ +
Sbjct: 755 QEEVLMRDILPGENLASIEDCYMDEVGRLRVSTLELMTLHTGESNLPWYADFANYLSCEV 814
Query: 1236 IPHDLTWQQKKKFLHD 1251
P D T KKK L +
Sbjct: 815 PPPDFTGYFKKKLLKE 830
>gb|AAD19780.2| hypothetical protein [Arabidopsis thaliana]
Length = 1048
Score = 681 bits (1757), Expect = 0.0
Identities = 417/1006 (41%), Positives = 571/1006 (56%), Gaps = 163/1006 (16%)
Query: 294 QLGQLAKQMSENHQGKFSSETLTLTEQENTAVVSTRSGRVLH--------EPKEKVEGEK 345
Q+ LA+ ++N + + S + +N +SGR L+ + K K +
Sbjct: 12 QILSLARVGNQNSRSENSGGRAP--KHQNPEKGGHKSGRELNPILKKEKAKEKSKAASDD 69
Query: 346 NEEAEVERDEGVMEERAREPTRTRVVNTHTPELRKIPFPKALVKKNLDKQFSKFLEVFKK 405
EE ++ +EE+A++ P + K+PFP K +K++S F E+ ++
Sbjct: 70 LEEDNTGKESDTLEEKAKKTVLP-------PYVPKLPFPGRQRKIQREKEYSLFDEIMRQ 122
Query: 406 LHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETILMTEECSAILQRKMPKKRRDPG 465
L + +PF + ++ + IY K +KDIL+ +R L E +L++ ECSAILQ R
Sbjct: 123 LQVKLPFLDLVQNVSIYRKHLKDILTNKRTLEEGH--VLISHECSAILQNVALFSRM--- 177
Query: 466 SFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLGEVTPTMISLQMADRSLKTPYGI 525
I T LCDLGA ++LM + KRL TPT +SL + DRS+ P G+
Sbjct: 178 -------IGEYTFDRCLCDLGAGVSLMPFSVAKRLGDTNFTPTKMSLVLGDRSISFPVGV 230
Query: 526 IEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLATGEAEIKVAKGTLTLKVGEDEV 585
EDV V+V F P DFVI++++E+ + LILGRPFL A I V K + L++G+
Sbjct: 231 AEDVQVRVGNFYIPTDFVIIELDEEPRHRLILGRPFLNIVAALIDVRKSKINLRIGDIVQ 290
Query: 586 LFNIFDSLKHRADE-EVFRCEVVDELVCEEFVRISIKDPLETTIMEGLEIEDQNLERELD 644
FN+ + E + F +++DELV E ++ +DPL+T + + E E
Sbjct: 291 EFNMERIMSKPTTECQTFWVDIMDELVNELLAELNTEDPLQTVLTKE--------ESEFG 342
Query: 645 SLYHEVNATLSQLESVSSLATKSIWKEELTRDEEIPIEE-------------------KS 685
L E +++ SS TK + EL +E +E+ K
Sbjct: 343 YL-GEATTRFARILDSSSPMTKVVAFAELGDNE---VEKALVVSSPKDCDDWSELNAPKM 398
Query: 686 ELKSLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWTIDDIKGISPA 745
ELK LP+ L+ A+L VI+N L ++ LL LR ++ ALG+++DDI GISP
Sbjct: 399 ELKPLPARLRVAFLGPNSTYLVIINPELNNVESALLLCELRKYRKALGYSLDDITGISPT 458
Query: 746 ICMHKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPK 805
+CMH I LE V+ QRRLN +++DVV+KEI+KLLDAG+IYPISDS WVSPV VVPK
Sbjct: 459 LCMHMIHLEGESITSVEHQRRLNSNLRDVVKKEIMKLLDAGIIYPISDSTWVSPVHVVPK 518
Query: 806 KGGITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYC 865
KGG+ V+ NE NELIPTR VT R+CIDYR+LNS TRKD+FPL FIDQML+RL+ YYC
Sbjct: 519 KGGVIVIKNEKNELIPTRTVTGHRMCIDYRKLNSATRKDNFPLSFIDQMLERLSNQPYYC 578
Query: 866 FLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIET 925
FLDGY G+ QI + P+DQEKT FTCPYG FAY+RMPFGLCNAPATFQ CM IFSD+IE
Sbjct: 579 FLDGYLGFFQILIHPDDQEKTTFTCPYGTFAYRRMPFGLCNAPATFQHCMKYIFSDMIE- 637
Query: 926 CIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKG 985
DF ++RC++ +LVLNWEK HFMVRDGIVLGHK+SEKG
Sbjct: 638 -------DF-----------------MERCEDKHLVLNWEKSHFMVRDGIVLGHKISEKG 673
Query: 986 IEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFD 1045
+EVDRAKIEV+ + PP ++K +E FD
Sbjct: 674 VEVDRAKIEVMMSMQPPNSVKD-----------------------------HEEQKSEFD 704
Query: 1046 ENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASR 1105
+ CL+AF +KESLV+A ++ P+W LPFE++CDASD +GAVL Q+K++ L+ IYYASR
Sbjct: 705 DTCLQAFNIVKESLVSALIVQPPEWELPFEVICDASDYVVGAVLGQRKDKKLHAIYYASR 764
Query: 1106 VLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRW 1165
L+ AQ VIVHT HAALR+L +K+D+KPRL+RW
Sbjct: 765 TLDGAQ----------------------------VIVHTVHAALRYLLSKKDAKPRLLRW 796
Query: 1166 VLLLQEFDLEIIDRRGKDNSVADHLSRL----EGGACSPIP------IQEEFSDE----- 1210
+LLLQ+FDLEI D++G +N VAD+LSRL E +P I++ + DE
Sbjct: 797 ILLLQDFDLEIKDKKGIENEVADNLSRLRVQEEVLMRDILPGENLASIEDCYMDEVGRLR 856
Query: 1211 ----KLLAVSTKEP-LPWYVHFANFRVAGLIPHDLTWQQKKKFLHD 1251
+L+ + T E LPWY FAN+ + P D T KKK L +
Sbjct: 857 VSTLELMTLHTGESNLPWYADFANYLSCEVPPPDFTGYFKKKLLKE 902
Score = 133 bits (335), Expect = 4e-29
Identities = 74/181 (40%), Positives = 105/181 (57%), Gaps = 43/181 (23%)
Query: 1460 KRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWA 1519
K++L++V+ W T +KTP+GT+PF+L++GK+CHLPVELE+KA WA
Sbjct: 897 KKLLKEVLEERPPSW------------TTYKTPVGTTPFNLIYGKSCHLPVELEYKALWA 944
Query: 1520 IRKLNFDWKVASE-KRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLV 1578
+ +N+D K A E +RLLQLNELDE + G V
Sbjct: 945 TKLMNYDIKPAGERRRLLQLNELDE-----------------------------IQGDKV 975
Query: 1579 LLFNSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVYQGGEVL 1638
LLFNS+L++FPGKL+S+WSGPFV+ + P+G+ + + K F+ NGQRLK Y L
Sbjct: 976 LLFNSKLKIFPGKLRSKWSGPFVIMEMRPNGSAILWDNNGKP-FSTNGQRLKPYLAHSTL 1034
Query: 1639 K 1639
+
Sbjct: 1035 E 1035
>emb|CAD39877.2| OSJNBb0058J09.16 [Oryza sativa (japonica cultivar-group)]
gi|50922333|ref|XP_471527.1| OSJNBb0058J09.16 [Oryza
sativa (japonica cultivar-group)]
Length = 1045
Score = 647 bits (1670), Expect = 0.0
Identities = 308/567 (54%), Positives = 406/567 (71%), Gaps = 23/567 (4%)
Query: 998 KLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKE 1057
+L PP NIKGI SFLGHAGFYRRFIKDFS +A+P+TNLL K+APF F++ CLR+F+ +K+
Sbjct: 500 RLPPPVNIKGICSFLGHAGFYRRFIKDFSTIARPLTNLLAKDAPFEFNDACLRSFKILKK 559
Query: 1058 SLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTT 1117
+LV+AP+I PDW+LPFEIMCDASD A+GAVL Q K++ + I YAS+ L AQ NY TT
Sbjct: 560 ALVSAPIIQPPDWTLPFEIMCDASDFAVGAVLGQTKDKKHHAICYASKTLTGAQLNYATT 619
Query: 1118 EKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEII 1177
EKELL VVFA +KFR Y++G KVI++TDHAAL++L K+D+KPRL+RW+LLLQEFD+EI
Sbjct: 620 EKELLTVVFAIDKFRSYLVGAKVIIYTDHAALKYLLTKKDAKPRLLRWILLLQEFDIEIK 679
Query: 1178 DRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFANFRVAGLIP 1237
D++G +N VADHLSRL PI + D+ L+ IP
Sbjct: 680 DKKGVENYVADHLSRLHITNMQEQPINDLLRDDMLMTC--------------------IP 719
Query: 1238 HDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCHGSSYGGHFS 1297
+ ++K +++ ++WD+P+L+++CSDG++RRC+ + KI+ CH S YGGH+
Sbjct: 720 PG---ENQRKLKYESHRHIWDEPYLYRVCSDGLLRRCVPTDEGLKIIERCHASPYGGHYG 776
Query: 1298 GERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGI 1357
RT AK+ QSGF+WPT++ +S+ F+ C CQR G I+ R+ MPL L++E+FDVWGI
Sbjct: 777 AFRTQAKIWQSGFFWPTMYEDSKEFIRRCTSCQRQGGITARDAMPLTYNLQVEIFDVWGI 836
Query: 1358 DFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISD 1417
DFMGPFP S C YILVAVDYVSKWVEA S D++ + IF RFG R +ISD
Sbjct: 837 DFMGPFPKSRNCEYILVAVDYVSKWVEAMPCSAADARHAKKMFTETIFPRFGTRRMVISD 896
Query: 1418 GGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRK 1477
G HF ++ F LL + KH V+TPYHPQTSGQ E SN+++K IL+K VN +W +
Sbjct: 897 RGFHFIDKTFRDLLREMEAKHNVATPYHPQTSGQAETSNKQIKNILQKTVNKMGTEWKDR 956
Query: 1478 LDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWKVASEKRLLQ 1537
L DALWAYRTA+KTPIG SP+ +V+GK+C LPVELEH+AYWAIR N D++ A E R +Q
Sbjct: 957 LQDALWAYRTAYKTPIGMSPYQIVYGKSCRLPVELEHRAYWAIRNSNMDFEGAGEWRKMQ 1016
Query: 1538 LNELDEFRLRAYESASIYKEKTKKWHD 1564
+ EL+E+R AY +A IYKE+TK+WHD
Sbjct: 1017 IAELEEWREMAYHNAKIYKERTKRWHD 1043
Score = 96.7 bits (239), Expect = 6e-18
Identities = 46/110 (41%), Positives = 73/110 (65%)
Query: 678 EIPIEEKSELKSLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWTID 737
E P + ELK LP L+YA+L PVI++ L+ + ++L VL H+S LG+++
Sbjct: 358 EPPPQPPLELKPLPLGLRYAFLHNDREAPVIISDKLSKDETQRLPTVLEKHRSVLGYSLQ 417
Query: 738 DIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGV 787
D++GI+ A+C H+I ++ P +PQRRLN M++VV+KE++KLL AG+
Sbjct: 418 DLRGINLALCTHRIPIDPKSTPSREPQRRLNNEMQEVVKKEVLKLLHAGI 467
>emb|CAE03895.1| OSJNBb0026I12.3 [Oryza sativa (japonica cultivar-group)]
gi|50921891|ref|XP_471306.1| OSJNBb0026I12.3 [Oryza
sativa (japonica cultivar-group)]
Length = 589
Score = 638 bits (1646), Expect = 0.0
Identities = 308/548 (56%), Positives = 411/548 (74%), Gaps = 14/548 (2%)
Query: 809 ITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
+TVVAN NNELIP R VT WR+CIDY++LN T+KD+FPLP ID+ML+RLA H +CFLD
Sbjct: 1 MTVVANVNNELIPQRTVTGWRMCIDYQKLNKATKKDNFPLPVIDEMLERLANHSLFCFLD 60
Query: 869 GYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIE 928
GYSGY+QI + P+DQ KT FTCPYG +AY+RM FGLCNAPA+FQRCM +IFSD+IE +E
Sbjct: 61 GYSGYHQIPIHPKDQCKTTFTCPYGTYAYRRMSFGLCNAPASFQRCMMSIFSDMIEAIME 120
Query: 929 IFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEV 988
+FMDDFS++G FD CL NL V +RCQE +LVLNWEKCHFMV +GIVLGH+VSE+GIEV
Sbjct: 121 VFMDDFSLYGKTFDHCLQNLDKVFQRCQEKDLVLNWEKCHFMVHEGIVLGHRVSERGIEV 180
Query: 989 DRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENC 1048
DRAKIEVI++L PP NIKGIR+FLGHAGF+RRFIK+FS +A+P+TNLL K+APF FD+ C
Sbjct: 181 DRAKIEVIDQLPPPVNIKGIRNFLGHAGFHRRFIKEFSTIARPLTNLLAKDAPFEFDDVC 240
Query: 1049 LRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLN 1108
L++F+++K++LV+A +I PD LPFEIMCDASD +GAVL Q K++ L+ I YAS L
Sbjct: 241 LKSFKTLKKALVSALIIQPPDGMLPFEIMCDASDFIVGAVLGQTKDKKLHAICYASETLA 300
Query: 1109 EAQRNYTTTEKELLGVVFACEKF-RPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVL 1167
AQ NY TTEKELL VVFA +KF R ++G KVIV+T+HAAL++L K+D+KPRL+RW+
Sbjct: 301 GAQSNYATTEKELLAVVFAIDKFRRSDLVGAKVIVYTEHAALKYLLTKKDAKPRLLRWIP 360
Query: 1168 LLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHF 1227
LLQEFDLEI D++G +NSVADHLSRL+ PI D+ L+AV + PWY +
Sbjct: 361 LLQEFDLEIKDKKGVENSVADHLSRLQLTNMQEPPINYFLQDDMLMAVRNSD--PWYANI 418
Query: 1228 ANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHC 1287
N+ V+ +P + ++K +++ ++WD+P+L+++CS+G++RRC+ + + KI+ C
Sbjct: 419 VNYMVSKYVPQG---ENQRKLKYESHCHIWDEPYLYRVCSNGLLRRCVPKEEGFKIIERC 475
Query: 1288 HGSSYGGHFSGERTAAKVLQSGFYWPTL-------HRNSRAFVESCDRCQRTGNISRRNE 1340
H + Y GH+ RT AK+ QS F+WPT+ N++ + + Q G N+
Sbjct: 476 HAAPYRGHYGAFRTQAKIWQSRFFWPTMTFRELLRKMNAKHNIATPYHPQTNGQAEMSNK 535
Query: 1341 MPLKNILE 1348
+KNIL+
Sbjct: 536 Q-IKNILQ 542
Score = 72.0 bits (175), Expect = 2e-10
Identities = 30/57 (52%), Positives = 39/57 (67%)
Query: 1427 FESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALW 1483
F LL K KH ++TPYHPQT+GQ E+SN+++K IL+K V+ W KL DALW
Sbjct: 505 FRELLRKMNAKHNIATPYHPQTNGQAEMSNKQIKNILQKTVHEMGTGWKDKLPDALW 561
>gb|AAG52026.1| polyprotein, putative; 77260-80472 [Arabidopsis thaliana]
gi|25405064|pir||B96492 probable polyprotein, 77260-80472
[imported] - Arabidopsis thaliana
Length = 884
Score = 618 bits (1593), Expect = e-175
Identities = 350/820 (42%), Positives = 492/820 (59%), Gaps = 122/820 (14%)
Query: 380 KIPFPKALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEV 439
K+PFPK+ K + ++ + KL + +P +A++ P+ +F+K I++K +
Sbjct: 160 KVPFPKSPRKSKQELDDARCKVMMDKLIVEMPLIDAVKSSPMIRQFVKRIVTK----DML 215
Query: 440 DETILMT--EECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMF 497
ET++MT + S I+Q K+P+K D
Sbjct: 216 TETVVMTMSTQVSDIIQNKIPQKLPDR--------------------------------- 242
Query: 498 KRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLIL 557
++ PT I+L +ADRS++ GII DV V+V P D V+L E++ K PLIL
Sbjct: 243 ------DLEPTQITLVLADRSVRRSDGIICDVPVQVGTSYIPTDLVVLSYEKEPKDPLIL 296
Query: 558 GRPFLATGEAEIKVAKGTLTLKVGEDEVLFNIFDSLKHRA-DEEVFRCEVVDELVCEEFV 616
GRPFL T A I V KG + L VG+ + F+I +K D + F + + L E +
Sbjct: 297 GRPFLVTTGAIIDVRKGIIGLNVGDLTMQFDINKVVKKPTIDGKTFYLDTISFLADEFLM 356
Query: 617 RISIKDPLETTIMEGLEIEDQNLERELDSLYHEVNATLSQ--LESVSSLATKSIWKEELT 674
++++ PL+ ++ ++ + + LD + H + + + + S AT W E
Sbjct: 357 KMTLAVPLKHALIPSIDKVPKGYGKLLDRIEHVMQLVAQEELIGTSSQAATIGDWSLEKA 416
Query: 675 RDEEIPIEEKSELKSLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGW 734
K +LK LP L+YA+L E VI+N+ L ++ LL LR ++ ALG+
Sbjct: 417 --------PKVDLKPLPPGLRYAFLGENSTYHVIVNASLNKVELTLLLSKLRKYRKALGY 468
Query: 735 TIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDS 794
++DDI GISP +CMH+I L++ KP V+ +RRLNP++ K+ +K
Sbjct: 469 SLDDITGISPDLCMHRIHLKDESKPSVEHRRRLNPNL-----KDAVK------------- 510
Query: 795 EWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQM 854
+++ K +LN+ TRK+HFPL FIDQ+
Sbjct: 511 ----------------------------KEIMK--------KLNAATRKNHFPLQFIDQL 534
Query: 855 LDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRC 914
L+RL+ H+YYC LDGYSG+ QI + P+DQEKT FTCPYG FAY RMPFGLCNAPA F+RC
Sbjct: 535 LERLSNHKYYCVLDGYSGFFQIPIHPDDQEKTMFTCPYGTFAYSRMPFGLCNAPAIFERC 594
Query: 915 MFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDG 974
M +IF+D+IE IE+FMDDFSV+G +F+ACL NL VL RC+E NLVLNWEKCHFMV++G
Sbjct: 595 MMSIFTDMIENFIEVFMDDFSVYGSSFEACLENLRKVLARCEEKNLVLNWEKCHFMVQEG 654
Query: 975 IVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTN 1034
IVLGHKVS GIEV++AKIEV+ L ++ RF+KDFSK+A+P+T
Sbjct: 655 IVLGHKVSGAGIEVNKAKIEVMTSLQALDSVN------------LRFVKDFSKIARPLTA 702
Query: 1035 LLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKE 1094
LL K+ F F+ C AF IK +LV+AP++ DW+LPFEIMCDA D A+GAVL Q+K
Sbjct: 703 LLCKDVKFDFNSECHDAFNQIKNALVSAPIVQPLDWNLPFEIMCDAGDYAVGAVLGQRKG 762
Query: 1095 RVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFA 1154
L+ I+YASR L++AQRN TTEKELL VVFA +KFR Y++G KVIVHTDH+AL++L
Sbjct: 763 NKLHAIHYASRTLDDAQRNNATTEKELLAVVFAFDKFRSYLVGSKVIVHTDHSALKYLMQ 822
Query: 1155 KQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLE 1194
K+D+KPRL+RW+LLLQEFD+E+ DR+G +N VADHLSR++
Sbjct: 823 KKDAKPRLLRWILLLQEFDIEVRDRKGIENGVADHLSRIK 862
>pir||T12085 reverse transcriptase homolog - fava bean (fragment)
gi|2522228|dbj|BAA22787.1| reverse transcriptase-like
protein [Vicia faba]
Length = 407
Score = 596 bits (1536), Expect = e-168
Identities = 282/397 (71%), Positives = 342/397 (86%), Gaps = 5/397 (1%)
Query: 830 VCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFT 889
VCIDYRRLN TRKDHFPLPFIDQML+RLA H+YYCFLDGYSGYNQI V PEDQEKT FT
Sbjct: 1 VCIDYRRLNLATRKDHFPLPFIDQMLERLADHEYYCFLDGYSGYNQIAVVPEDQEKTTFT 60
Query: 890 CPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLA 949
CP+G+F+Y+R+PFGLCNAPATFQRCM +IF+D++E +E+FMDDFSVFG +FD CL NLA
Sbjct: 61 CPFGIFSYRRIPFGLCNAPATFQRCMQSIFADMLEKYMEVFMDDFSVFGKSFDNCLSNLA 120
Query: 950 LVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIR 1009
LVL+RCQE+NL+LNWEKCHFMVR+GIVLGHK+S KGIEVD+AKIEVI KL PPTN KGIR
Sbjct: 121 LVLERCQESNLILNWEKCHFMVREGIVLGHKISYKGIEVDQAKIEVISKLHPPTNEKGIR 180
Query: 1010 SFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPD 1069
SFLGH GFYRRFI+DFSK+ KP+T LL K+ F FD+ C+ AFE++K LV+AP++VAPD
Sbjct: 181 SFLGHVGFYRRFIRDFSKIEKPLTTLLVKDKDFVFDKECVVAFETLKGKLVSAPIVVAPD 240
Query: 1070 WSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACE 1129
W LPFEIMCDASD+A+GAVL Q++E+ L+VIYYAS VLN AQ NY TTEKELL +V+A +
Sbjct: 241 WYLPFEIMCDASDIAVGAVLGQRREKRLHVIYYASHVLNPAQMNYATTEKELLVMVYAFD 300
Query: 1130 KFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADH 1189
KFR Y+LG KVIV+TDHAAL++LF+KQDSKPRL+RW+LLLQEFD+EI D+RG +N+VADH
Sbjct: 301 KFRQYMLGSKVIVYTDHAALKYLFSKQDSKPRLLRWILLLQEFDVEIRDKRGCENTVADH 360
Query: 1190 LSRLEGGACSPI--PIQEEFSDEKLLAVSTKEPLPWY 1224
LSR+ + + PI++EF+DE +L V +PW+
Sbjct: 361 LSRMSHIEETKVKQPIKDEFADEHILVVI---GVPWF 394
>gb|AAF63114.1| Hypothetical protein [Arabidopsis thaliana] gi|25405147|pir||B96502
hypothetical protein F28H19.8 [imported] - Arabidopsis
thaliana
Length = 640
Score = 575 bits (1482), Expect = e-162
Identities = 280/471 (59%), Positives = 352/471 (74%), Gaps = 17/471 (3%)
Query: 973 DGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPM 1032
+GIVLGHK+S KGIEVD+AKI+V+ +L P +K IRSFLGHAGFYRRFIKDFSK+A+P+
Sbjct: 170 EGIVLGHKISGKGIEVDKAKIDVMVRLQPLKTVKDIRSFLGHAGFYRRFIKDFSKIARPL 229
Query: 1033 TNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQK 1092
T LL KEA F FDE+CL+AF SIKE+LV AP++ AP+W+ PFEIMCDASD A+GAVL Q+
Sbjct: 230 TRLLCKEAEFDFDEDCLKAFHSIKEALVLAPIVQAPNWNHPFEIMCDASDYAVGAVLGQR 289
Query: 1093 KERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHL 1152
++ L+VIYYASR L++ Q Y TTEKELL VVFA EKFR Y++G KV V+TDHAAL+H+
Sbjct: 290 IDKKLHVIYYASRTLDDTQSRYATTEKELLVVVFAFEKFRSYLVGSKVTVYTDHAALKHI 349
Query: 1153 FAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKL 1212
+AK+D+KPRL+RW+L LQEFD+EI+D+RG +N VADHLSR+ +PI + +E+L
Sbjct: 350 YAKKDTKPRLLRWILFLQEFDMEIVDKRGSENGVADHLSRMR--MTEAVPIDDSMPEEQL 407
Query: 1213 LAVST--------------KEPLPWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWD 1258
LAV T LPWYV N+ AG++P +L+ QKKKF D Y WD
Sbjct: 408 LAVGTSFDKFKIETMCATRSGKLPWYVDLVNYLTAGIVPSELSSYQKKKFFRDINHYYWD 467
Query: 1259 DPFLFKICSDGVIRRCITEVDFEKILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRN 1318
+P L+K DG+ RRCI E + + +L CH S YGGHF+ +T KVLQ+G +WP++ ++
Sbjct: 468 EPVLYKKGVDGLFRRCIAEEEVQGVLELCHNSPYGGHFATFKTVQKVLQAGLWWPSMFKD 527
Query: 1319 SRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGIDFMGPF-PPSFGC*YILVAVD 1377
+ + CDRCQR GNISRRNEMP ILE+E+FDVWGIDFMGPF P S+G YILVAVD
Sbjct: 528 AHEHITRCDRCQRVGNISRRNEMPQNPILEVEVFDVWGIDFMGPFEPASYGNKYILVAVD 587
Query: 1378 YVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFE 1428
YVSKWVEA A TNDSKVV+ K IF RFG+PR +ISDGG+HF N+ FE
Sbjct: 588 YVSKWVEAIASPTNDSKVVLKLFKTIIFPRFGIPRVVISDGGSHFINKVFE 638
>gb|AAD23707.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25411232|pir||G84476 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 466
Score = 545 bits (1403), Expect = e-153
Identities = 255/417 (61%), Positives = 319/417 (76%), Gaps = 2/417 (0%)
Query: 1222 PWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFE 1281
PWY N+ G+ P +LT ++K F D Y WD+P+L+ +C D + R ++E + E
Sbjct: 38 PWYADHVNYLACGIEPPNLTSYERKNFFRDIHHYYWDEPYLYTLCKDKIYRSYVSEDEVE 97
Query: 1282 KILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEM 1341
IL HCH S+YGGHF+ +T +K+LQ+GF+WPT+ ++++ FV CD CQR GNISRRNEM
Sbjct: 98 GILLHCHDSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKGNISRRNEM 157
Query: 1342 PLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLK 1401
P ILE+++FDVWGIDFMGPFP S+G YILV VDYVSKWVEA A TND+KVV+ K
Sbjct: 158 PQNPILEVDIFDVWGIDFMGPFPSSYGNKYILVVVDYVSKWVEAIASPTNDAKVVLKLFK 217
Query: 1402 KNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKR 1461
IF RFGV +ISDGG HF N+ FE+LL+K+GVKHKV+TPYHPQTSGQVEISNRE+K
Sbjct: 218 TIIFPRFGVSWVVISDGGKHFINKVFENLLKKHGVKHKVATPYHPQTSGQVEISNREIKT 277
Query: 1462 ILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIR 1521
ILEK V +RKDWS KLDDALWAY+TAFKTPIGT+PF+L+ K+CHL VELE+KA WA++
Sbjct: 278 ILEKTVGITRKDWSTKLDDALWAYKTAFKTPIGTTPFNLLCVKSCHLHVELEYKAMWAVK 337
Query: 1522 KLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVLLF 1581
LNFD K A EKRL+QL+ELDE RL AYES+ IYKE+TK +HD+KI+ ++F G VLLF
Sbjct: 338 LLNFDIKTAEEKRLIQLSELDEIRLEAYESSKIYKERTKLFHDKKIITKDFQVGDQVLLF 397
Query: 1582 NSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVYQGGEVL 1638
NSRL++FPGKLKSRWSGPF + V P+GAV + +FTVNGQRLK Y ++L
Sbjct: 398 NSRLKIFPGKLKSRWSGPFCITEVRPYGAVTLAG--KSGVFTVNGQRLKKYLANQIL 452
>emb|CAD39882.2| OSJNBb0067G11.5 [Oryza sativa (japonica cultivar-group)]
gi|50922247|ref|XP_471484.1| OSJNBb0067G11.5 [Oryza
sativa (japonica cultivar-group)]
Length = 791
Score = 545 bits (1403), Expect = e-153
Identities = 264/458 (57%), Positives = 342/458 (74%), Gaps = 17/458 (3%)
Query: 654 LSQLESVSSL-ATKSIWKEELTRDEEIPIEEKSELKSLPSSLKYAYLEEGENKPVILNSV 712
L ++ES + A E ++E+P K E L+Y +L PVI++
Sbjct: 350 LQEIESAEEVTAVPPHESPEALLEDEVPDFIKEEADP---GLRYDFLHNDREAPVIISDK 406
Query: 713 LTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMK 772
L+ + + LL VL H+S LG+++ D++GI+PA+C H+IL++ P +PQR LN +M+
Sbjct: 407 LSEDETQHLLTVLEKHRSILGYSLQDLRGINPALCTHRILIDPESTPSREPQRWLNNAMR 466
Query: 773 DVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCI 832
+VV+KE++KLL AG+IYPI SEWVSPVQVVPKKGG+TVV +WR+CI
Sbjct: 467 EVVKKEVLKLLHAGIIYPIPYSEWVSPVQVVPKKGGMTVVY-------------RWRMCI 513
Query: 833 DYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPY 892
DYR+LN T+KDHFPLPFID+ML+RLA H ++CFLDGYSGY+QI + PEDQ KT FTCPY
Sbjct: 514 DYRKLNKATKKDHFPLPFIDEMLERLANHSFFCFLDGYSGYHQIPIHPEDQSKTTFTCPY 573
Query: 893 GVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVL 952
G +AY+RM FGLCN PA+FQRCM +IFSD+IE +E+FMDDFSV+G CL NL VL
Sbjct: 574 GTYAYRRMSFGLCNTPASFQRCMMSIFSDMIEDIMEVFMDDFSVYGKTLGHCLQNLDKVL 633
Query: 953 KRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFL 1012
+RCQE +LVLN EKCHFMVR+GIVLGH+VSE+G+EVDRAKI+VI++L PP NIKGI SF
Sbjct: 634 QRCQEKDLVLNREKCHFMVREGIVLGHRVSERGVEVDRAKIDVIDQLPPPMNIKGIHSFF 693
Query: 1013 GHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSL 1072
GHAGFYRRFIKDFS +A+P+TNLL K+AP FD+ CL++FE +K++LV+AP+I PDW+L
Sbjct: 694 GHAGFYRRFIKDFSTIARPLTNLLAKDAPIEFDDVCLKSFEILKKALVSAPIIQPPDWTL 753
Query: 1073 PFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEA 1110
PFEIMCDASD A+GAVL Q K++ + I YAS+ L A
Sbjct: 754 PFEIMCDASDFAVGAVLGQTKDKKHHAICYASKTLTGA 791
Score = 70.1 bits (170), Expect = 6e-10
Identities = 33/109 (30%), Positives = 60/109 (54%)
Query: 2 EDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAE 61
E+P+ H+ F + C+ ++ + E + LFPFSL++ A++W S +WE L +
Sbjct: 51 ENPYHHLRDFEQVCSCLKIRGMRQETVWWKLFPFSLQERAKQWYTSTVGCVNGSWEKLRD 110
Query: 62 KFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNL 110
+F FFP + + L+ +I+ F Q +E++ AW F L++ P +L
Sbjct: 111 RFCLTFFPVTRITALRVEILNFKQIENESIGAAWSRFTNLVQSGPTLSL 159
>emb|CAB81136.1| putative athila transposon protein [Arabidopsis thaliana]
gi|25407395|pir||D85075 probable athila transposon
protein [imported] - Arabidopsis thaliana
Length = 724
Score = 535 bits (1377), Expect = e-150
Identities = 290/596 (48%), Positives = 386/596 (64%), Gaps = 38/596 (6%)
Query: 451 AILQRKM-PKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLGEVTPTM 509
AI Q+K+ PKK DPGSFT+P + L LCDLGA ++LM L + KRL +
Sbjct: 3 AITQKKIVPKKLIDPGSFTLPCSLGPLAFKRCLCDLGALVSLMPLSVAKRLGFTQYKSCN 62
Query: 510 ISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLATGEAEI 569
ISL +ADRS++ P+ + E++ +++ P DFV+L+M+E+ K PLILGRPFLAT A
Sbjct: 63 ISLILADRSVRIPHSLFENLPIRIGAVDIPTDFVVLEMDEEPKDPLILGRPFLATAGAMN 122
Query: 570 KVAKGTLTLKVGE-DEVLFNIFDSLKHRADE-EVFRCEVVDELVCEEFVRISIKDPLETT 627
V KG + L +G+ + F++ D++K + ++F E +D+L E + +D L +
Sbjct: 123 DVKKGKIDLNLGKYCRMTFDVKDAMKKPTIKGQLFWIEEIDQLADELLEERAEEDHLYSA 182
Query: 628 IMEG-----LEIEDQNLERELDS--------LYHEVNA---------------------- 652
+ + L +E ++ LDS + E+N
Sbjct: 183 LTKRGEDGFLHLETLGYQKLLDSHKAMEESEPFEELNGPETEVMVMSEEGSTQVQPAHSR 242
Query: 653 TLSQLESVSSLATKSIWKEELTRDEEIPIEEKSELKSLPSSLKYAYLEEGENKPVILNSV 712
T S S S+++ + D K +LKSLP L+Y + PVI+N
Sbjct: 243 TYSTNNSTSTISNSGELIIPTSDDWSELKAPKVDLKSLPKGLRYVFFGPNSTYPVIINVE 302
Query: 713 LTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMK 772
L + LL LR +K A+G+++ DIK ISP++C H+I LE ++PQRRLN ++K
Sbjct: 303 LNDNEVNLLLSELRKYKRAIGYSLSDIKRISPSLCNHRIHLENESYSSIEPQRRLNLNLK 362
Query: 773 DVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCI 832
+VV+KEI+KLLDAGVIYPISDS WV PV VPKKGG+TVV NE +ELIPTR +T RVCI
Sbjct: 363 EVVKKEILKLLDAGVIYPISDSTWVFPVHCVPKKGGMTVVKNEKDELIPTRTITGHRVCI 422
Query: 833 DYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPY 892
DYR+LN+ +RKDHFPLPF +QML+ LA H Y CFLDGYSG+ QI + P DQEKT FTCPY
Sbjct: 423 DYRKLNAASRKDHFPLPFTNQMLEGLANHLYNCFLDGYSGFFQIPIHPNDQEKTTFTCPY 482
Query: 893 GVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVL 952
G FAYKRMPFGLCNAP TFQRCM +IFSDLIE +E+FMDDFSV+GP+F +CL NL VL
Sbjct: 483 GTFAYKRMPFGLCNAPTTFQRCMTSIFSDLIEKMVEVFMDDFSVYGPSFSSCLLNLGRVL 542
Query: 953 KRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGI 1008
+ +ETNLVLNWEKC+FMV++GIVLGHK+SEKGIEVD+ KI+V+ +L PP +K I
Sbjct: 543 TKWEETNLVLNWEKCYFMVKEGIVLGHKISEKGIEVDKEKIKVMMQLQPPKTVKDI 598
Score = 136 bits (343), Expect = 5e-30
Identities = 68/116 (58%), Positives = 90/116 (76%), Gaps = 2/116 (1%)
Query: 1100 IYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSK 1159
IYYASR L+EAQ Y TTEKELL VVFA +KFR Y++ KV V+TDHAALRH++AK+D+K
Sbjct: 598 IYYASRTLDEAQGRYATTEKELLVVVFAFKKFRSYLVESKVTVYTDHAALRHMYAKKDTK 657
Query: 1160 PRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAV 1215
PRL+R +LLLQEFD+EI++++G +N ADHLSR+ PI I + +E+L+ V
Sbjct: 658 PRLLRGILLLQEFDMEIVEKKGIENGAADHLSRMR--IEEPILIDDSMPEEQLMVV 711
>gb|AAV43951.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 963
Score = 523 bits (1346), Expect = e-146
Identities = 315/734 (42%), Positives = 426/734 (57%), Gaps = 84/734 (11%)
Query: 335 HEPKEKVEGEKNEE-------AEVERDEGVMEERAREPTRTRVVNTHTPELRKIPFPKAL 387
+EP+ + K EE AE ++E+ E TP+ I F
Sbjct: 291 NEPQPEASVSKIEEPLTIEPLAEPSTFASSIDEKVPEQPSVENEEIQTPDCATILF---- 346
Query: 388 VKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEV--DETILM 445
K D+ + L F K +P P+ +F+++ + R+L+ + DE +
Sbjct: 347 -KDGFDEDYGNTLNYFSKRKPLVPLPPP---DPMELRFLRETI---RELTTIMSDEWLRE 399
Query: 446 TEECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRM-FKRLNLGE 504
E S ++ R + IP +EG V L NL+S F L+
Sbjct: 400 AELSSEVI-------RINTAPRIIPCHLEG-NDVSILYSPTVGANLVSGSFAFAYLSDKA 451
Query: 505 VTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLAT 564
VTPT + ++ +GI++DV V + +DF + ++++ +++G P
Sbjct: 452 VTPTNKFFKHPSGNIIKRFGIMQDVPVCFEDKEAILDFHVFEIQD---FDILIGLPI--- 505
Query: 565 GEAEIKVAKGTLTLKVGEDEVLFNI--FDSLKHRADEEVFRCEVVDELVCEEFVRISIKD 622
+++L N DSLK E F RI + D
Sbjct: 506 ------------------EQLLINTPRLDSLKITLGETEFSIPF-------SRARIVLTD 540
Query: 623 PLETTIMEGLEIEDQNLERELDSLYHEVNATLSQLESVSSLATKSIWKEELTRDE--EIP 680
PL E ++ E + HE L + E + KEE E ++P
Sbjct: 541 PLP---------EIESTEEVIAVPPHESPEALLEDEVPDFI------KEEADPGETLDLP 585
Query: 681 IEEKS-----ELKSLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWT 735
I E ELK LP SL YA+L PVI++ L + ++LL VL H S LG++
Sbjct: 586 IMEPPPWPPLELKPLPPSLHYAFLHHDREAPVIISDKLFEDETQRLLAVLEKHCSVLGYS 645
Query: 736 IDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSE 795
+ D+KGI+P +C+H+I ++ P +PQRRLN +M++VV+KE++KLL AG+IYP+ SE
Sbjct: 646 LQDLKGINPVLCIHRIPIDPESTPSREPQRRLNNAMREVVKKEVLKLLHAGIIYPVPYSE 705
Query: 796 WVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQML 855
WVSPVQVVPKKGGI VVAN NELIP R VT+WR+CIDYR+LN T+KDHFPLPFID+ML
Sbjct: 706 WVSPVQVVPKKGGIMVVANAQNELIPQRTVTEWRMCIDYRKLNKATKKDHFPLPFIDEML 765
Query: 856 DRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCM 915
+RLA H ++CFLDGYSGY+QI + PEDQ KT FTCPYG +AY+RM FGLCNAPA+FQRCM
Sbjct: 766 ERLANHSFFCFLDGYSGYHQIPIHPEDQSKTTFTCPYGTYAYRRMSFGLCNAPASFQRCM 825
Query: 916 FAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGI 975
+IFSD+IE +E+FMDDF V+G F L NL VL+RCQE +LVLNWEKCHFMVR+GI
Sbjct: 826 MSIFSDMIEYIMEVFMDDFLVYGKTFGHYLQNLDKVLQRCQEKDLVLNWEKCHFMVREGI 885
Query: 976 VLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNL 1035
VLGH+VSE+GIEVDRAKI+VI++L PP NIKGIRSFLGHA FYRRFI+D+S + +P+TNL
Sbjct: 886 VLGHRVSERGIEVDRAKIDVIDQLPPPVNIKGIRSFLGHASFYRRFIEDYSTIVRPLTNL 945
Query: 1036 LEKEAPFTFDENCL 1049
L K+APF FD+ CL
Sbjct: 946 LAKDAPFEFDDACL 959
>gb|AAV43847.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1005
Score = 518 bits (1335), Expect = e-145
Identities = 313/731 (42%), Positives = 424/731 (57%), Gaps = 84/731 (11%)
Query: 335 HEPKEKVEGEKNEE-------AEVERDEGVMEERAREPTRTRVVNTHTPELRKIPFPKAL 387
+EP+ + K EE AE ++E+ E TP+ I F
Sbjct: 296 NEPQPEASVSKIEEPLTIEPLAEPSTFASSIDEKVPEQPSVENEEIQTPDCATILF---- 351
Query: 388 VKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEV--DETILM 445
K D+ + L F K +P P+ +F+++ + R+L+ + DE +
Sbjct: 352 -KDGFDEDYGNTLNYFSKRKPLVPLPPP---DPMELRFLRETI---RELTTIMSDEWLRE 404
Query: 446 TEECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRM-FKRLNLGE 504
E S ++ R + IP +EG V L NL+S F L+
Sbjct: 405 AELSSEVI-------RINTAPRIIPCHLEG-NDVSILYSPTVGANLVSGSFAFAYLSDKA 456
Query: 505 VTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLAT 564
VTPT + ++ +GI++DV V + +DF + ++++ +++G P
Sbjct: 457 VTPTNKFFKHPSGNIIKRFGIMQDVPVCFEDKEAILDFHVFEIQD---FDILIGLPI--- 510
Query: 565 GEAEIKVAKGTLTLKVGEDEVLFNI--FDSLKHRADEEVFRCEVVDELVCEEFVRISIKD 622
+++L N DSLK E F RI + D
Sbjct: 511 ------------------EQLLINTPRLDSLKITLGETEFSIPF-------SRARIVLTD 545
Query: 623 PLETTIMEGLEIEDQNLERELDSLYHEVNATLSQLESVSSLATKSIWKEELTRDE--EIP 680
PL E ++ E + HE L + E + KEE E ++P
Sbjct: 546 PLP---------EIESTEEVIAVPPHESPEALLEDEVPDFI------KEEADPGETLDLP 590
Query: 681 IEEKS-----ELKSLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWT 735
I E ELK LP SL YA+L PVI++ L + ++LL VL H S LG++
Sbjct: 591 IMEPPPWPPLELKPLPPSLHYAFLHHDREAPVIISDKLFEDETQRLLAVLEKHCSVLGYS 650
Query: 736 IDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSE 795
+ D+KGI+P +C+H+I ++ P +PQRRLN +M++VV+KE++KLL AG+IYP+ SE
Sbjct: 651 LQDLKGINPVLCIHRIPIDPESTPSREPQRRLNNAMREVVKKEVLKLLHAGIIYPVPYSE 710
Query: 796 WVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQML 855
WVSPVQVVPKKGGI VVAN NELIP R VT+WR+CIDYR+LN T+KDHFPLPFID+ML
Sbjct: 711 WVSPVQVVPKKGGIMVVANAQNELIPQRTVTEWRMCIDYRKLNKATKKDHFPLPFIDEML 770
Query: 856 DRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCM 915
+RLA H ++CFLDGYSGY+QI + PEDQ KT FTCPYG +AY+RM FGLCNAPA+FQRCM
Sbjct: 771 ERLANHSFFCFLDGYSGYHQIPIHPEDQSKTTFTCPYGTYAYRRMSFGLCNAPASFQRCM 830
Query: 916 FAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGI 975
+IFSD+IE +E+FMDDF V+G F L NL VL+RCQE +LVLNWEKCHFMVR+GI
Sbjct: 831 MSIFSDMIEYIMEVFMDDFLVYGKTFGHYLQNLDKVLQRCQEKDLVLNWEKCHFMVREGI 890
Query: 976 VLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNL 1035
VLGH+VSE+GIEVDRAKI+VI++L PP NIKGIRSFLGHA FYRRFI+D+S + +P+TNL
Sbjct: 891 VLGHRVSERGIEVDRAKIDVIDQLPPPVNIKGIRSFLGHASFYRRFIEDYSTIVRPLTNL 950
Query: 1036 LEKEAPFTFDE 1046
L K+APF FD+
Sbjct: 951 LAKDAPFEFDD 961
>gb|AAT81690.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1017
Score = 514 bits (1325), Expect = e-144
Identities = 321/792 (40%), Positives = 450/792 (56%), Gaps = 101/792 (12%)
Query: 309 KFSSETLTLTEQ--ENTAVVSTRSGRVLHEPKEKVEGEKNEE--AEVERDEGVMEERARE 364
K SE T+ ++ ENT+ ++ + L K+E E AE ++E+ E
Sbjct: 315 KTPSEGKTILDRILENTSFMTQSNEPQLEASVSKIEEPLTIEPFAEPSTSASSIDEKVPE 374
Query: 365 PTRTRVVNTHTPELRKIPFPKALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAK 424
+ TP+ I F + D + L F K +P S P+
Sbjct: 375 QPSVKNEEIQTPDCATIMF-----RDGFDVDYGNTLNYFSKRKPPVPLSPP---DPMELG 426
Query: 425 FMKDILSKRRKLSEVDETILMTEECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCD 484
F+++ + R+L T +M++E + R + IP +EG V L
Sbjct: 427 FLRETV---REL-----TSIMSDEWQREAELSSEVIRINTAPRIIPCHLEG-NDVGILYS 477
Query: 485 LGASINLMSLRM-FKRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFV 543
NL+S+ F L+ VTPT + +R++ +GI++DV V + +DF
Sbjct: 478 PTIGANLISVSFAFAYLSDKAVTPTNKFFKHPNRNIIEGFGIVQDVPVFFEDREAILDFH 537
Query: 544 ILDMEEDSKVPLILGRPFLATGEAEIKVAK-GTLTLKVGEDEVLFNIFDSLKHRADEEVF 602
+ ++++ +++G P + I ++ G+L + +GE+E F RA
Sbjct: 538 VFEIQD---FDILIGLPI---EQLLINTSRLGSLKITLGENE-----FSIPFSRA----- 581
Query: 603 RCEVVDELVCEEFVRISIKDPLETTIMEGLEIEDQNLERELDSLYHEVNATLSQLESVSS 662
RI++ DPL T ++ E HE L + E
Sbjct: 582 --------------RITLTDPLPET---------ESAEEVTAVPPHESTKALLEDEVPDF 618
Query: 663 LATKSIWKEELTRDEEIPIEEKS-----ELKSLPSSLKYAYLEEGENKPVILNSVLTPLK 717
+ ++ E L ++PI E ELK LP L+YA+L PVI++ L+ +
Sbjct: 619 IKEEADPSETL----DLPIMEPPPRPPLELKPLPPGLRYAFLHNDREAPVIISDKLSEDE 674
Query: 718 EEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVRK 777
++LL VL H+S LG+++ D++GI+PA+C H+I ++ P +PQRRLN +M++VV+K
Sbjct: 675 TQRLLTVLEKHRSILGYSLQDLRGINPALCTHRIPIDLESTPSREPQRRLNNAMREVVKK 734
Query: 778 EIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRRL 837
E++KLL AG+IYP+ SEWVS VQV+PKKGG+TVVAN NELIP R VT WR+CIDYR+L
Sbjct: 735 EVLKLLHAGIIYPVPYSEWVSKVQVMPKKGGMTVVANSQNELIPQRTVTGWRMCIDYRKL 794
Query: 838 NSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAY 897
N T+KDHF LPF D+ML+ LA H ++CFLDG
Sbjct: 795 NKATKKDHFLLPFNDEMLEWLANHSFFCFLDG---------------------------- 826
Query: 898 KRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQE 957
M FGLCNAPA FQRCM +IFSD+IE +E+FMDDFSV+G F CL NL VL+RCQE
Sbjct: 827 --MSFGLCNAPAPFQRCMMSIFSDMIEDIMEVFMDDFSVYGKTFGHCLQNLDKVLQRCQE 884
Query: 958 TNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGF 1017
+LVLNWEKCHFMVR+GI+LGH+VSE+G EVDRAKI+VI++L P NIKGI SFLGHAGF
Sbjct: 885 KDLVLNWEKCHFMVREGIILGHRVSERGSEVDRAKIDVIDQLPPLVNIKGIHSFLGHAGF 944
Query: 1018 YRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIM 1077
YRRFIKDFS +A+P+TNLL K+APF FD+ CL++FE +K++LV AP+I PDW+LPFEIM
Sbjct: 945 YRRFIKDFSIIARPLTNLLAKDAPFEFDDACLKSFEILKKALVFAPIIQPPDWTLPFEIM 1004
Query: 1078 CDASDLALGAVL 1089
CDASD A+ AVL
Sbjct: 1005 CDASDFAVAAVL 1016
>emb|CAD40178.2| OSJNBa0061A09.17 [Oryza sativa (japonica cultivar-group)]
gi|50921885|ref|XP_471303.1| OSJNBa0061A09.17 [Oryza
sativa (japonica cultivar-group)]
Length = 430
Score = 461 bits (1186), Expect = e-128
Identities = 220/445 (49%), Positives = 303/445 (67%), Gaps = 16/445 (3%)
Query: 1077 MCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYIL 1136
MCDAS+ A+GAVL Q K+R + I YAS+ L AQ NY+TTEKELL VV
Sbjct: 1 MCDASNFAVGAVLGQTKDRKQHAIAYASKTLTGAQLNYSTTEKELLAVV----------- 49
Query: 1137 GFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGG 1196
G KV+V+TDHAAL++L K+D+KPRLIRW+L+LQEFD+EI +++G +NSVADHLSR++
Sbjct: 50 GAKVVVYTDHAALKYLLTKKDAKPRLIRWILILQEFDIEINNKKGIENSVADHLSRMQIT 109
Query: 1197 ACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYL 1256
+ I + D+ LL V+ +P WY NF VA +P + KK+ ++++ L
Sbjct: 110 NMQELSINDYLRDDMLLKVTDSDP--WYAPIVNFMVAWHVPPG---ENKKRLSYESRKRL 164
Query: 1257 WDDPFLFKICSDGVIRRCITEVDFEKILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLH 1316
WD P+L+++CSD ++RRC+ + +I+ C + YG H+ RT AK+ QSGF+WPT++
Sbjct: 165 WDAPYLYRVCSDSLLRRCVPADEGMEIIEKCRVAPYGSHYGAFRTHAKIWQSGFFWPTMY 224
Query: 1317 RNSRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAV 1376
+++ F+ C CQ+ G I+ ++ MPL L++ELFDVWGIDFMGPFP + C YILVAV
Sbjct: 225 GDTKEFIRRCTSCQKHGGITAQDAMPLTYNLQVELFDVWGIDFMGPFPKFYDCEYILVAV 284
Query: 1377 DYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGV 1436
DYVSKWVEA ++K + IF FG PR +ISDGG+HF ++ F++ L++ G
Sbjct: 285 DYVSKWVEAMPCRAANTKHAHRMFDEIIFPCFGTPRTVISDGGSHFIDKTFQNFLQELGA 344
Query: 1437 KHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTS 1496
+H + TPYHP+T GQ E N+++ IL+K VN K W KL +ALWAYRT + PIG S
Sbjct: 345 QHNIVTPYHPKTGGQAETRNKQIMNILQKTVNEMGKAWKNKLPNALWAYRTTYNMPIGMS 404
Query: 1497 PFHLVFGKACHLPVELEHKAYWAIR 1521
P+ LV+GK CHLPVELEHKA+WAIR
Sbjct: 405 PYQLVYGKTCHLPVELEHKAHWAIR 429
>emb|CAB81130.1| AT4g07600 [Arabidopsis thaliana] gi|5724765|gb|AAD48069.1| contains
similarity to Pfam family PF00078 -943 Reverse
transcriptase (RNA-dependent DNA polymerase); score 65.8,
E=9.4e-16, N=1; may be a pseudogene [Arabidopsis
thaliana] gi|25407387|pir||F85074 hypothetical protein
AT4g07600 [imported] - Arabidopsis thaliana
Length = 630
Score = 456 bits (1174), Expect = e-126
Identities = 222/341 (65%), Positives = 263/341 (77%), Gaps = 26/341 (7%)
Query: 788 IYPISDSEW-------VSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRRLNSV 840
I P SD +W VSPVQ +PKKGGITV+ NE +ELIP R +T R+CIDYR LN+
Sbjct: 309 IIPTSD-DWSKLKAPKVSPVQCIPKKGGITVIKNEKDELIPIRTITGLRMCIDYRNLNAA 367
Query: 841 TRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRM 900
+R DHFPLPF DQML+RLA H YYCFLDGYSG+ QI + P D EKT FTCPYG FAY+RM
Sbjct: 368 SRNDHFPLPFTDQMLERLANHPYYCFLDGYSGFFQIPIHPNDHEKTTFTCPYGTFAYERM 427
Query: 901 PFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNL 960
PFGLCNAPATFQRCM +IFSD+IE +E+FMDDFSV+GP+F +CL NL VL RC+ETNL
Sbjct: 428 PFGLCNAPATFQRCMTSIFSDIIEEMVEVFMDDFSVYGPSFSSCLLNLGRVLTRCEETNL 487
Query: 961 VLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRR 1020
VLNWEKCHFMV++GI+LGHK+SEKGIEVD+ FLGHAGFYRR
Sbjct: 488 VLNWEKCHFMVKEGIMLGHKISEKGIEVDKG------------------CFLGHAGFYRR 529
Query: 1021 FIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDA 1080
FIKDFSK+A+P+T LL KE F FDE+C ++F +IKE+LV+APV+ A +W PFEIMCDA
Sbjct: 530 FIKDFSKIARPLTRLLCKETEFEFDEDCPKSFHTIKEALVSAPVVRATNWDYPFEIMCDA 589
Query: 1081 SDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKEL 1121
D A+GAVL QK ++ L+VIYYASR L+EAQ Y TTEKEL
Sbjct: 590 LDYAVGAVLGQKIDKKLHVIYYASRTLDEAQGRYATTEKEL 630
Score = 149 bits (377), Expect = 6e-34
Identities = 95/276 (34%), Positives = 160/276 (57%), Gaps = 12/276 (4%)
Query: 399 FLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETILMTEECSAILQRKM- 457
F + K++ + IP +AL + KF+KD++ +R + EV ++++ ECSAI+Q+K+
Sbjct: 2 FAKNIKEVELRIPLVDALALILDTHKFLKDLIVER--IQEVQGMVVLSHECSAIIQKKIV 59
Query: 458 PKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLGEVTPTMISLQMADR 517
PKK DPGSFT+P + + LCDLGA ++ M L + KRL + ISL +ADR
Sbjct: 60 PKKLSDPGSFTLPCFLGTVAFNRCLCDLGALVSPMPLSIAKRLGFTQYKSCNISLILADR 119
Query: 518 SLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLATGEAEIKVAKGTLT 577
S++ +G+++++ +++ DFVIL+M+E+ K PLIL RPFLAT A I V KG +
Sbjct: 120 SVRISHGLLKNLPIRIGAAEISTDFVILEMDEEPKDPLILRRPFLATAGAMIDVKKGKID 179
Query: 578 LKVGED-EVLFNIFDSLKH-RADEEVFRCEVVDELVCEEFVRISIKDPLETTIMEG---- 631
L +G+D + F+I D++K + ++F E +D+L E ++ +D L + + +
Sbjct: 180 LNLGKDFRMTFDIKDAMKKPNIEGQLFWIEEMDQLADELLEELAQEDHLNSALTKTGQDG 239
Query: 632 -LEIEDQNLERELDSLYHEVNATLSQLESVSSLATK 666
L ++ ++ LDS H+ E ++ ATK
Sbjct: 240 FLHLKVLGYQKLLDS--HKAMEESKPFEELNGPATK 273
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,774,580,428
Number of Sequences: 2540612
Number of extensions: 119412828
Number of successful extensions: 395342
Number of sequences better than 10.0: 26456
Number of HSP's better than 10.0 without gapping: 4978
Number of HSP's successfully gapped in prelim test: 21492
Number of HSP's that attempted gapping in prelim test: 358943
Number of HSP's gapped (non-prelim): 50655
length of query: 1648
length of database: 863,360,394
effective HSP length: 142
effective length of query: 1506
effective length of database: 502,593,490
effective search space: 756905795940
effective search space used: 756905795940
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0335b.5


