

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
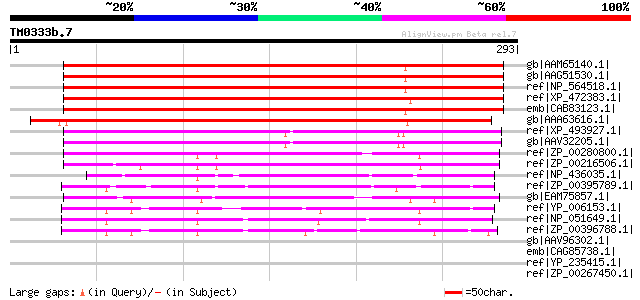
Query= TM0333b.7
(293 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM65140.1| dessication-related protein, putative [Arabidopsi... 299 6e-80
gb|AAG51530.1| dessication-related protein, putative; 70055-7184... 298 1e-79
ref|NP_564518.1| expressed protein [Arabidopsis thaliana] 298 1e-79
ref|XP_472383.1| OSJNBb0118P14.11 [Oryza sativa (japonica cultiv... 275 1e-72
emb|CAB83123.1| putative protein [Arabidopsis thaliana] gi|20147... 270 3e-71
gb|AAA63616.1| dessication-related protein [Craterostigma planta... 265 9e-70
ref|XP_493927.1| similar to Arabidopsis thaliana hypothetical pr... 205 1e-51
gb|AAV32205.1| hypothetical protein [Oryza sativa (japonica cult... 205 1e-51
ref|ZP_00280800.1| hypothetical protein Bcep02004480 [Burkholder... 105 2e-21
ref|ZP_00216506.1| hypothetical protein Bcepa03002342 [Burkholde... 92 2e-17
ref|NP_436035.1| hypothetical protein SMa1445 [Sinorhizobium mel... 91 3e-17
ref|ZP_00395789.1| hypothetical protein DgeoDRAFT_1759 [Deinococ... 89 1e-16
gb|EAM75857.1| Twin-arginine translocation pathway signal [Kineo... 84 4e-15
ref|YP_006153.1| dessication-related protein pcc13-62 precursor ... 70 7e-11
ref|NP_051649.1| dessication-associated protein [Deinococcus rad... 65 2e-09
ref|ZP_00396788.1| Twin-arginine translocation pathway signal [D... 64 7e-09
gb|AAV96302.1| oxidoreductase, FAD-binding [Silicibacter pomeroy... 35 1.9
emb|CAG85738.1| unnamed protein product [Debaryomyces hansenii C... 35 1.9
ref|YP_235415.1| hypothetical protein Psyr_2337 [Pseudomonas syr... 34 5.7
ref|ZP_00267450.1| COG1092: Predicted SAM-dependent methyltransf... 33 7.4
>gb|AAM65140.1| dessication-related protein, putative [Arabidopsis thaliana]
Length = 315
Score = 299 bits (766), Expect = 6e-80
Identities = 155/267 (58%), Positives = 192/267 (71%), Gaps = 13/267 (4%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
D L++F LNLE+LEAEF+ +G LG+GLD P L GGP P+G Q A LDP+ RDI Q
Sbjct: 43 DRKLLEFPLNLEYLEAEFFLFGALGFGLDKVAPNLTMGGPSPIGAQKANLDPLTRDIILQ 102
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
FA+Q+ GHLRAI + VKGF RP L++SK+AFA+V++ AFG PPF+PYANS NYL+A+
Sbjct: 103 FAWQEVGHLRAIKKTVKGFARPQLDLSKKAFAKVMDKAFGVKFVPPFNPYANSYNYLIAS 162
Query: 152 YMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTV 211
Y++PYVGLTGYVG NPKLQ +R LV+GLLG +SGQDA+IR +LY R V PY TV
Sbjct: 163 YLVPYVGLTGYVGANPKLQCPASRKLVAGLLGVESGQDAVIRGMLYARAAHIVYPYGVTV 222
Query: 212 YEVTHRLSLLRNDLGK-------------KQAEGKVVGQVFGLNMYSLTYPRSPEEILRI 258
T ++S LRN LGK AEG+V+G V N SL++ R+PEEILRI
Sbjct: 223 AAFTDKISDLRNKLGKAGVKDEGLIVPKFMGAEGQVIGNVLVGNELSLSFDRTPEEILRI 282
Query: 259 VYGSGDEHVPGGFFPKGGNGNIAKSYL 285
VYGSG+E VPGGF+PKG +G IAKSYL
Sbjct: 283 VYGSGNESVPGGFYPKGADGEIAKSYL 309
>gb|AAG51530.1| dessication-related protein, putative; 70055-71849 [Arabidopsis
thaliana] gi|25357137|pir||B96520 hypothetical protein
T2J15.11 [imported] - Arabidopsis thaliana
Length = 302
Score = 298 bits (763), Expect = 1e-79
Identities = 155/267 (58%), Positives = 191/267 (71%), Gaps = 13/267 (4%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
D L++F LNLE+LEAEF+ +G LG GLD P L GGP P+G Q A LDP+ RDI Q
Sbjct: 30 DRKLLEFPLNLEYLEAEFFLFGALGLGLDKVAPNLTMGGPSPIGAQKANLDPLTRDIILQ 89
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
FA+Q+ GHLRAI + VKGF RP L++SK+AFA+V++ AFG PPF+PYANS NYL+A+
Sbjct: 90 FAWQEVGHLRAIKKTVKGFARPQLDLSKKAFAKVMDKAFGVKFVPPFNPYANSYNYLIAS 149
Query: 152 YMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTV 211
Y++PYVGLTGYVG NPKLQ +R LV+GLLG +SGQDA+IR +LY R V PY TV
Sbjct: 150 YLVPYVGLTGYVGANPKLQCPASRKLVAGLLGVESGQDAVIRGMLYARAAHIVYPYGVTV 209
Query: 212 YEVTHRLSLLRNDLGK-------------KQAEGKVVGQVFGLNMYSLTYPRSPEEILRI 258
T ++S LRN LGK AEG+V+G V N SL++ R+PEEILRI
Sbjct: 210 AAFTDKISDLRNKLGKAGVKDEGLIVPKFMGAEGQVIGNVLVGNELSLSFDRTPEEILRI 269
Query: 259 VYGSGDEHVPGGFFPKGGNGNIAKSYL 285
VYGSG+E VPGGF+PKG +G IAKSYL
Sbjct: 270 VYGSGNESVPGGFYPKGADGEIAKSYL 296
>ref|NP_564518.1| expressed protein [Arabidopsis thaliana]
Length = 315
Score = 298 bits (763), Expect = 1e-79
Identities = 155/267 (58%), Positives = 191/267 (71%), Gaps = 13/267 (4%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
D L++F LNLE+LEAEF+ +G LG GLD P L GGP P+G Q A LDP+ RDI Q
Sbjct: 43 DRKLLEFPLNLEYLEAEFFLFGALGLGLDKVAPNLTMGGPSPIGAQKANLDPLTRDIILQ 102
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
FA+Q+ GHLRAI + VKGF RP L++SK+AFA+V++ AFG PPF+PYANS NYL+A+
Sbjct: 103 FAWQEVGHLRAIKKTVKGFARPQLDLSKKAFAKVMDKAFGVKFVPPFNPYANSYNYLIAS 162
Query: 152 YMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTV 211
Y++PYVGLTGYVG NPKLQ +R LV+GLLG +SGQDA+IR +LY R V PY TV
Sbjct: 163 YLVPYVGLTGYVGANPKLQCPASRKLVAGLLGVESGQDAVIRGMLYARAAHIVYPYGVTV 222
Query: 212 YEVTHRLSLLRNDLGK-------------KQAEGKVVGQVFGLNMYSLTYPRSPEEILRI 258
T ++S LRN LGK AEG+V+G V N SL++ R+PEEILRI
Sbjct: 223 AAFTDKISDLRNKLGKAGVKDEGLIVPKFMGAEGQVIGNVLVGNELSLSFDRTPEEILRI 282
Query: 259 VYGSGDEHVPGGFFPKGGNGNIAKSYL 285
VYGSG+E VPGGF+PKG +G IAKSYL
Sbjct: 283 VYGSGNESVPGGFYPKGADGEIAKSYL 309
>ref|XP_472383.1| OSJNBb0118P14.11 [Oryza sativa (japonica cultivar-group)]
gi|38569173|emb|CAE05363.3| OJ000315_02.8 [Oryza sativa
(japonica cultivar-group)] gi|38346151|emb|CAD40673.2|
OSJNBb0118P14.11 [Oryza sativa (japonica
cultivar-group)]
Length = 323
Score = 275 bits (702), Expect = 1e-72
Identities = 139/267 (52%), Positives = 182/267 (68%), Gaps = 13/267 (4%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
DVDL++F LNLE+LEAEF+ + LG+GLD D L GGP PVG Q A L P +RDI +Q
Sbjct: 56 DVDLLEFPLNLEYLEAEFFCWSALGYGLDGIDASLTGGGPAPVGAQTAALTPFVRDIATQ 115
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
F +Q+ GHLR I + VKGFPRPLL+IS F +++ +A L+PPF+PY NSLN+LLA+
Sbjct: 116 FCYQEVGHLREIKQNVKGFPRPLLDISAANFGKIVETAMNTTLDPPFNPYENSLNFLLAS 175
Query: 152 YMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTV 211
Y++PYVGLTGYVG NP+L + R LV+GLLG +S QDA+IR LLYE +V Y V
Sbjct: 176 YIIPYVGLTGYVGANPRLLTPQARKLVAGLLGVESAQDAVIRALLYEHGLSRVASYGVGV 235
Query: 212 YEVTHRLSLLRNDLGKKQA-------------EGKVVGQVFGLNMYSLTYPRSPEEILRI 258
E+T +S LRN LG+K EG+ VG + + +SL Y R+PEEIL +
Sbjct: 236 AELTAHISELRNVLGRKGVKDEGLVVAPGQGPEGQTVGNIIAGDRFSLAYDRTPEEILGV 295
Query: 259 VYGSGDEHVPGGFFPKGGNGNIAKSYL 285
VYGSGD GGFFP+G +G IA++++
Sbjct: 296 VYGSGDPAKAGGFFPQGADGRIARAFI 322
>emb|CAB83123.1| putative protein [Arabidopsis thaliana] gi|20147275|gb|AAM10351.1|
AT3g62730/F26K9_160 [Arabidopsis thaliana]
gi|15294284|gb|AAK95319.1| AT3g62730/F26K9_160
[Arabidopsis thaliana] gi|15228845|ref|NP_191832.1|
expressed protein [Arabidopsis thaliana]
gi|11282971|pir||T48062 hypothetical protein F26K9.160 -
Arabidopsis thaliana
Length = 317
Score = 270 bits (691), Expect = 3e-71
Identities = 140/268 (52%), Positives = 182/268 (67%), Gaps = 14/268 (5%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
DVD V F +NLEF EAEF+ G G GLD + LA+GGPPP+G + A LDPI I +
Sbjct: 34 DVDRVHFAMNLEFTEAEFFLKGATGKGLDAYNATLAKGGPPPIGAKKANLDPITNRIIEE 93
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
F +Q+ GHLRAIT+ G PRPL+N+++E FA ++ A G+ NP FDPYANSLNYLLA+
Sbjct: 94 FGYQEIGHLRAITDMTGGIPRPLINLTRENFAVFMDRAVGRKSNPRFDPYANSLNYLLAS 153
Query: 152 YMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQW-T 210
Y +PYVGLTGYVGT P L + LV+GLLG +SGQDA+IRTLLYER+ +V+ Y T
Sbjct: 154 YYIPYVGLTGYVGTIPYLVYFNIKKLVAGLLGVESGQDAVIRTLLYERQNEKVEEYGGVT 213
Query: 211 VYEVTHRLSLLRNDLGK-------------KQAEGKVVGQVFGLNMYSLTYPRSPEEILR 257
V E+T+ +S LRN+LG AE + + + YSL+Y R+ +EILR
Sbjct: 214 VAELTNEISNLRNELGMCGIKDEGLCVPLWLGAENRTTSNILSADPYSLSYDRTAQEILR 273
Query: 258 IVYGSGDEHVPGGFFPKGGNGNIAKSYL 285
++YG+GDEH PGGF+P G NG IA+ +L
Sbjct: 274 VMYGTGDEHRPGGFWPCGANGRIARMFL 301
>gb|AAA63616.1| dessication-related protein [Craterostigma plantagineum]
gi|320600|pir||E45509 desiccation-related protein (clone
PCC13-62) - Craterostigma plantagineum
gi|118926|sp|P22242|DRPE_CRAPL Desiccation-related
protein PCC13-62 precursor gi|227781|prf||1710351E
abscisic acid responsive protein E
Length = 313
Score = 265 bits (678), Expect = 9e-70
Identities = 141/290 (48%), Positives = 188/290 (64%), Gaps = 24/290 (8%)
Query: 13 SSLVLSLFIPIFCSF---------EDVV--DVDLVQFHLNLEFLEAEFYFYGTLGWGLDH 61
S+ ++S F+ + CS +D+ DV L++F LNLE LEAEF+ + G G+D
Sbjct: 9 SAALVSFFLALICSCSYAAWHHEKDDIPKSDVSLLEFPLNLELLEAEFFAWAAFGKGIDE 68
Query: 62 ADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEA 121
+P+LA+GGP P+G Q A L P +RDI +QFA+Q+ GH+RAI V+GFPRPLL++S ++
Sbjct: 69 LEPELAKGGPSPIGVQKANLSPFIRDIIAQFAYQEFGHVRAIQSSVEGFPRPLLDLSAKS 128
Query: 122 FAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGL 181
FA V++SAFGK L PPFDPYAN +NYLLA Y++PYVGLTGYVG NPKL+ +R LV+GL
Sbjct: 129 FATVMDSAFGKTLKPPFDPYANDINYLLACYVVPYVGLTGYVGANPKLESPVSRKLVAGL 188
Query: 182 LGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLGKK------------- 228
L ++GQDA+IR LLYER +V+PY TV E T+++S LRN LG K
Sbjct: 189 LAVEAGQDAIIRALLYERATDKVEPYGITVAEFTNKISELRNKLGDKGVKDLGLIVEPEL 248
Query: 229 QAEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNG 278
AEGK+ G V + SL +PR+PE L + P F PK G
Sbjct: 249 GAEGKISGNVLAGDKNSLAFPRTPERCLGSCTAAAMRPSPAAFIPKAPTG 298
>ref|XP_493927.1| similar to Arabidopsis thaliana hypothetical protein (T48062)
[Oryza sativa]
Length = 328
Score = 205 bits (521), Expect = 1e-51
Identities = 117/294 (39%), Positives = 171/294 (57%), Gaps = 42/294 (14%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
D+ +QF LN +F+EAE++ +G LG G+D D L+ GGPPP G + A LD ++ ++
Sbjct: 32 DMAHIQFLLNAKFVEAEWFLHGALGRGIDFIDGALSGGGPPPTGARKATLDFRATEVAAE 91
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
+Q+ GH+RAIT+ + GFPRP +++S FA V++ A L+PPFDPYA+S+N+LLA+
Sbjct: 92 LGYQEVGHIRAITQSMGGFPRPAIDLSDAVFAAVMDDAMATRLDPPFDPYASSVNFLLAS 151
Query: 152 YMLPYVG--------------------------LTGYVGTNPKLQGVRTRNLVSGLLGAK 185
Y+LP++ L+G+ G + V L + +L +
Sbjct: 152 YILPHITASAALLPHGLLLQAGKPAKKPHRIRFLSGH-GGGETTESVADVQLQASMLAVE 210
Query: 186 SGQDAMIRTLLYERRYLQVKPYQW-TVYEVTHRLSLLRN--------DLG------KKQA 230
+GQDA+IR +LYER V PY+ TV E T R+S RN D G ++ A
Sbjct: 211 AGQDAVIRMMLYERADEVVAPYKGRTVAEFTRRISEWRNAASRCGAKDEGVKVLDRRQGA 270
Query: 231 EGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKSY 284
E + V + G SL + R+P E+LRI+YGSG+E VPGGF P+GGNG IAK +
Sbjct: 271 ERRTVSNILGAGDDSLGFARTPAEVLRILYGSGNEQVPGGFLPRGGNGTIAKGF 324
>gb|AAV32205.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|52353576|gb|AAU44142.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 361
Score = 205 bits (521), Expect = 1e-51
Identities = 117/294 (39%), Positives = 171/294 (57%), Gaps = 42/294 (14%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
D+ +QF LN +F+EAE++ +G LG G+D D L+ GGPPP G + A LD ++ ++
Sbjct: 65 DMAHIQFLLNAKFVEAEWFLHGALGRGIDFIDGALSGGGPPPTGARKATLDFRATEVAAE 124
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
+Q+ GH+RAIT+ + GFPRP +++S FA V++ A L+PPFDPYA+S+N+LLA+
Sbjct: 125 LGYQEVGHIRAITQSMGGFPRPAIDLSDAVFAAVMDDAMATRLDPPFDPYASSVNFLLAS 184
Query: 152 YMLPYVG--------------------------LTGYVGTNPKLQGVRTRNLVSGLLGAK 185
Y+LP++ L+G+ G + V L + +L +
Sbjct: 185 YILPHITASAALLPHGLLLQAGKPAKKPHRIRFLSGH-GGGETTESVADVQLQASMLAVE 243
Query: 186 SGQDAMIRTLLYERRYLQVKPYQW-TVYEVTHRLSLLRN--------DLG------KKQA 230
+GQDA+IR +LYER V PY+ TV E T R+S RN D G ++ A
Sbjct: 244 AGQDAVIRMMLYERADEVVAPYKGRTVAEFTRRISEWRNAASRCGAKDEGVKVLDRRQGA 303
Query: 231 EGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKSY 284
E + V + G SL + R+P E+LRI+YGSG+E VPGGF P+GGNG IAK +
Sbjct: 304 ERRTVSNILGAGDDSLGFARTPAEVLRILYGSGNEQVPGGFLPRGGNGTIAKGF 357
>ref|ZP_00280800.1| hypothetical protein Bcep02004480 [Burkholderia fungorum LB400]
Length = 291
Score = 105 bits (261), Expect = 2e-21
Identities = 73/266 (27%), Positives = 122/266 (45%), Gaps = 19/266 (7%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHA-DPKLAQGGPPPVGGQYAYLDPILRDIFS 90
D +++ F LNLE+LE++FY Y T G GL + + G G Q + DP+++ +
Sbjct: 30 DAEILNFALNLEYLESQFYTYATTGAGLAASMTAGVGTMGAVIPGQQVPFQDPVVKAYAN 89
Query: 91 QFAFQQAGHLRAITEEV--KGFPRPLLNIS----KEAFAEVINSAFGKPLNPPFDPYANS 144
+ A + H+ + + P ++I AF+ +A P PF+PYAN
Sbjct: 90 EIANDEREHVTFLRSALGSAAVAMPAIDIGGTDPNGAFSNAARAAGLVPAGTPFNPYAND 149
Query: 145 LNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQV 204
N+LL AY+ VG+T Y G +P + +G+L A++ ++RT+LY +
Sbjct: 150 NNFLLGAYIFEDVGVTAYKGASPLITNKTFLEAAAGILAAEAYHAGLVRTVLYSKGIDMT 209
Query: 205 KPYQWTVYEVTHRLSLLRNDLGKKQAEGKVV-------GQVFGLNMYSLTYPRSPEEILR 257
+ + +S RN L + + + + L+ L + RS +L
Sbjct: 210 -----GLVTAANAISAARNSLDHNGHDDQGITGASAGTSNIVPLDSNGLAFSRSYSNVLN 264
Query: 258 IVYGSGDEHVPGGFFPKGGNGNIAKS 283
IVY + GGFFP G NG++ S
Sbjct: 265 IVYLTSSAATKGGFFPNGVNGSLNMS 290
>ref|ZP_00216506.1| hypothetical protein Bcepa03002342 [Burkholderia cepacia R18194]
Length = 286
Score = 92.0 bits (227), Expect = 2e-17
Identities = 72/262 (27%), Positives = 119/262 (44%), Gaps = 11/262 (4%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPV--GGQYAYLDPILRDIF 89
D +++ F LNLE+LE++FY Y T G GL A G P V G Q + DP+++
Sbjct: 25 DAEILNFALNLEYLESQFYTYATTGSGLP-AGMVTGVGTPGAVIPGQQVPFQDPVVQAYA 83
Query: 90 SQFAFQQAGHLRAITEEV--KGFPRPLLNIS----KEAFAEVINSAFGKPLNPPFDPYAN 143
++ A + H+ + + +P ++I AF+ +A F+PYA+
Sbjct: 84 NEIAKDEREHVTFLRTALGSAAVAQPAIDIGGTDPNGAFSVAARAAGLVGSGVAFNPYAS 143
Query: 144 SLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQ 203
N+LL A++ VG+T Y G +P + +G+L A++ ++RT+L+ +
Sbjct: 144 DNNFLLGAFIFEDVGVTAYKGASPLITNKTFLEAAAGILAAEAYHAGLVRTVLFAKGVDM 203
Query: 204 VKPYQWTVYEVTHRLSLLRNDLGKKQAEGKVVG--QVFGLNMYSLTYPRSPEEILRIVYG 261
R SL + + G G + L+ L Y R +L IVY
Sbjct: 204 TSLVNAANAISAARASLDQVGNDDQGITGSTPGSSNIVPLDSNGLAYSRGYGNVLNIVYL 263
Query: 262 SGDEHVPGGFFPKGGNGNIAKS 283
+ + GGFFP G NG++ S
Sbjct: 264 TSTAAMKGGFFPNGVNGSLNMS 285
>ref|NP_436035.1| hypothetical protein SMa1445 [Sinorhizobium meliloti 1021]
gi|14523914|gb|AAK65447.1| Conserved hypothetical
protein [Sinorhizobium meliloti 1021]
gi|25505187|pir||E95360 hypothetical protein SMa1445
[imported] - Sinorhizobium meliloti (strain 1021)
magaplasmid pSymA
Length = 234
Score = 91.3 bits (225), Expect = 3e-17
Identities = 69/239 (28%), Positives = 111/239 (45%), Gaps = 11/239 (4%)
Query: 45 LEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAIT 104
+EAE+Y GT G G+D +D + GP G Q ++ P + + + A + H+R
Sbjct: 1 MEAEYYLRGTTGKGIDDSDAG-SDAGPVTGGKQVSFDTPAIGEFMQEVAENELAHVRFYR 59
Query: 105 EEV--KGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGY 162
+ + + PRP ++ FA V SA L FDP+ N N++L + VG+T Y
Sbjct: 60 KTLADQAVPRPAIDFDA-GFAAVAKSA---GLGEDFDPFGNETNFVLGGMLFEDVGVTAY 115
Query: 163 VGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLR 222
G L+ +G+L ++ M R+ LY + K Q V + ++
Sbjct: 116 AGAATVLKNKDFLAAAAGILAVEAYHMGMARSTLYRKGEEAWKAAQ-AVSDARDKIDGPE 174
Query: 223 NDLGKKQAEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHV-PGGFFPKGGNGNI 280
+ Q +GK + ++ + R+P+E+LRIVY S E GGF+P G NG I
Sbjct: 175 DKDQGLQVDGK--ANIVPSTPDAIAFTRTPQEVLRIVYISDKEGASKGGFYPNGMNGKI 231
>ref|ZP_00395789.1| hypothetical protein DgeoDRAFT_1759 [Deinococcus geothermalis DSM
11300] gi|66782690|gb|EAL83653.1| hypothetical protein
DgeoDRAFT_1759 [Deinococcus geothermalis DSM 11300]
Length = 307
Score = 89.4 bits (220), Expect = 1e-16
Identities = 79/266 (29%), Positives = 127/266 (47%), Gaps = 27/266 (10%)
Query: 31 VDVDLVQFHLNLEFLEAEFYFYGT--------LGWGLDHADPKLAQGGPPPVGGQYAYLD 82
+D +++ F LNLE+LEA FY +G G A+ +L G G Q+ D
Sbjct: 52 IDAEVLNFALNLEYLEAAFYLAAVGRVDELRAIGGG---AEIRLPAGLDRMRGMQFK--D 106
Query: 83 PILRDIFSQFAFQQAGHLRAITEEV--KGFPRPLLNISKEAFAEVINSAFGKPLNPPFDP 140
++ + A + H++ + + PRP+L+++ AF +A G + F+P
Sbjct: 107 SNVQALARDIAEDELAHVKFLHGALGKAAAPRPVLDLAG-AFDAAGQAASGGKIKG-FNP 164
Query: 141 YANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERR 200
YAN L +L A++ VG+T Y G + +G+L ++ IRT+LY++R
Sbjct: 165 YANDLFFLHGAFIFEDVGVTAYNGAATLITNPAYLQAAAGILAVEAYHGGAIRTMLYQQR 224
Query: 201 YLQVKPYQWTVYEVTHRLSLLR------NDLGKKQAEGKVVGQVFGLNMYSLTYPRSPEE 254
+ + V +V +S LR DLG A G +V V + + +PRS E
Sbjct: 225 QVSAAAGLY-VGQVVQAISNLRAKVGGGKDLGLSDAHGGMV--VAPADQNGVAFPRSTRE 281
Query: 255 ILRIVYGSGDEHVPGGFFPKGGNGNI 280
+L IVY + H GGF+P G NG+I
Sbjct: 282 VLNIVYLAPGAH-KGGFYPNGLNGSI 306
>gb|EAM75857.1| Twin-arginine translocation pathway signal [Kineococcus
radiotolerans SRS30216]
Length = 359
Score = 84.3 bits (207), Expect = 4e-15
Identities = 71/267 (26%), Positives = 121/267 (44%), Gaps = 29/267 (10%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQG----GPPPVGGQYAYLDPILRD 87
+V ++ F LNLE+LEAEFY + G GL A +A G G G + + +R
Sbjct: 107 EVSVLNFALNLEYLEAEFYCFAAYGHGLAEA---MATGTGTMGGVTGGHRVPFKSKAMRY 163
Query: 88 IFSQFAFQQAGHLRAITEEVKG--FPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSL 145
+ A + H++ + + RP +++ + +F +A FDP+++
Sbjct: 164 YAEEIANDEIAHVKFLRSALGAGAVSRPAIDL-QSSFTGAAVAAGVIEQGQTFDPFSSEE 222
Query: 146 NYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVK 205
+LL A++ VG+T Y G P + +G+L ++ ++RTLLY+
Sbjct: 223 FFLLGAFLFEDVGVTAYKGAAPLITNKTYLEAAAGILAVEAYHSGIVRTLLYQN------ 276
Query: 206 PYQWTVYEVTHRLSLLRNDLGKKQA---EGKVVGQVFGLNMY-----SLTYPRSPEEILR 257
+ T+ +S R+ L A +G G N+ ++ + R+P+E+L
Sbjct: 277 ----GLAAPTNLISAARDSLDGSAASKDQGITTGASGRANLVAADKDAIAFSRTPQEVLN 332
Query: 258 IVYGSGDEHV-PGGFFPKGGNGNIAKS 283
IVY + V GGF+P G NG I S
Sbjct: 333 IVYLTAGAGVSKGGFYPNGLNGEIKTS 359
>ref|YP_006153.1| dessication-related protein pcc13-62 precursor [Thermus
thermophilus HB27] gi|46198090|gb|AAS82500.1|
dessication-related protein pcc13-62 precursor [Thermus
thermophilus HB27]
Length = 292
Score = 70.1 bits (170), Expect = 7e-11
Identities = 71/265 (26%), Positives = 121/265 (44%), Gaps = 32/265 (12%)
Query: 31 VDVDLVQFHLNLEFLEAEFYFYGT-----LGWGLDHADPKLAQG--GPPPVGGQYAYLDP 83
+DV ++ F LNLE+LE FY T L +A L G G PV G L
Sbjct: 44 LDVAILNFALNLEYLEGLFYLAATGRISELNQVGGNAQIVLPPGFNGTSPVPG----LTG 99
Query: 84 ILRDIFSQFAFQQAGHLRAITEEV--KGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPY 141
L D+ + A + H+ + + + + RP++++ A + F+P+
Sbjct: 100 DLLDLADEIADDEKAHVLFLRQALGSQAVSRPVIDLYNSFNA----------IQSGFNPF 149
Query: 142 ANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLV---SGLLGAKSGQDAMIRTLLYE 198
+ +++ + A++ VG+T Y G P + +N++ +G+L A++ IR L E
Sbjct: 150 NDPVSFFVGAFVFEDVGVTAYNGAAPLI--TDKQNVLAPAAGILAAEAYHAGAIRRHLIE 207
Query: 199 RRYLQVKPYQWTVYEVTHRLSLLRNDLGKKQAEGKVV---GQVFGLNMYSLTYPRSPEEI 255
R V TV ++ + +S RN L EG V + + + R+ + +
Sbjct: 208 IRTQTVPGTGLTVEQLANAISNARNSLSGGGDEGLTVMGTPNNVAADPNGVAFSRTTDGV 267
Query: 256 LRIVYGSGDEHVPGGFFPKGGNGNI 280
L+IVY + + PGGFFP+G NG I
Sbjct: 268 LKIVYLNAQKQ-PGGFFPQGLNGQI 291
>ref|NP_051649.1| dessication-associated protein [Deinococcus radiodurans R1]
gi|6460847|gb|AAF12551.1|AE001826_20
dessication-associated protein [Deinococcus radiodurans]
gi|7471800|pir||B75631 dessication-associated protein -
Deinococcus radiodurans (strain R1)
Length = 337
Score = 65.5 bits (158), Expect = 2e-09
Identities = 71/273 (26%), Positives = 114/273 (41%), Gaps = 27/273 (9%)
Query: 31 VDVDLVQFHLNLEFLEAEFYFYGT------LGWGLDHADPKLAQGGPPPVGGQYAYLDPI 84
+D + F LNLE+LEA FY G D + L G G L
Sbjct: 61 LDATIFNFALNLEYLEAAFYLAAVGRLNELTAAGGDASKVTLPSGVTGMGGTAVPGLTGD 120
Query: 85 LRDIFSQFAFQQAGHLRAITEEV--KGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYA 142
LR + + A + H++ I + +P L++S A ++ G N F+PYA
Sbjct: 121 LRAMMEEIADDELAHVKVIRSVLGSAAVAQPRLDLSASFLAAGSLASNGAITN--FNPYA 178
Query: 143 NSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRT-RNL--VSGLLGAKSGQDAMIRTLLYER 199
N L +L A++ VG+T Y G L G + NL +G+L ++ IRT L+ R
Sbjct: 179 NPLFFLHGAFVFEDVGVTAYKGAARLLVGDKPGGNLENAAGILAVEAYHAGSIRTQLFMR 238
Query: 200 RYLQVKPYQWTVYEVTHRLSLLRNDLGKKQAEGKVV------------GQVFGLNMYSLT 247
R Q TV +V +S LR+ + + + + + +
Sbjct: 239 RTEQAAA-GLTVEQVVQAISNLRDSVDGADDRDQGITANGNAGVLARDANIIPTDSNGIA 297
Query: 248 YPRSPEEILRIVY-GSGDEHVPGGFFPKGGNGN 279
+ R+P ++ IV+ + + GGFFP G G+
Sbjct: 298 FSRTPRQVANIVFLDTTGKAARGGFFPDGLTGD 330
>ref|ZP_00396788.1| Twin-arginine translocation pathway signal [Deinococcus
geothermalis DSM 11300] gi|66781740|gb|EAL82707.1|
Twin-arginine translocation pathway signal [Deinococcus
geothermalis DSM 11300]
Length = 311
Score = 63.5 bits (153), Expect = 7e-09
Identities = 77/274 (28%), Positives = 120/274 (43%), Gaps = 31/274 (11%)
Query: 31 VDVDLVQFHLNLEFLEAEFYFYGT------LGWGLDHADPKLAQG--GPPPVGGQYAYLD 82
+D + F LNLE+LEA FY T G D + L G G PV G L
Sbjct: 43 LDAAIFNFALNLEYLEAAFYLAATGRLGELTAVGGDASKVILPSGFTGSSPVPG----LT 98
Query: 83 PILRDIFSQFAFQQAGHLRAITEEV--KGFPRPLLNISKEAFAEVINSAFGKPLNPPFDP 140
L ++ A + H++ I + P+P L++S A ++ GK N F+P
Sbjct: 99 GDLLARANEIADDEKAHVKVIRAVLGNAAVPQPRLDLSASFVAAGKAASGGKIDN--FNP 156
Query: 141 YANSLNYLLAAYMLPYVGLTGYVGTNPKL----QGVRTRNLVSGLLGAKSGQDAMIRTLL 196
+AN L +L A++ VG++ Y G L G N +G+L ++ IR+ L
Sbjct: 157 FANELFFLHGAFVFEDVGVSAYKGAARFLVDDKAGGNLEN-AAGILAVEAYHAGEIRSEL 215
Query: 197 YERRYLQVKPYQWTVYEVTHRLSLLRNDL-GKKQAEGKVVGQVFGLNMY-----SLTYPR 250
Y RR + TV +V +S LR+ + G + + N+ + + R
Sbjct: 216 YRRRG-EAAAAGLTVEQVVQAISDLRDSVDGSSDDDQGISNMGASANIVLADGNGIAFSR 274
Query: 251 SPEEILRIVYGSGDEHVPGGFFPKG--GNGNIAK 282
+P ++ IV+ S GGFFP G +GN+ K
Sbjct: 275 TPRQVGNIVFLSAGA-TKGGFFPDGLSDDGNLGK 307
>gb|AAV96302.1| oxidoreductase, FAD-binding [Silicibacter pomeroyi DSS-3]
gi|56697899|ref|YP_168270.1| oxidoreductase, FAD-binding
[Silicibacter pomeroyi DSS-3]
Length = 470
Score = 35.4 bits (80), Expect = 1.9
Identities = 17/47 (36%), Positives = 24/47 (50%), Gaps = 2/47 (4%)
Query: 246 LTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKSYLHPNAPAP 292
L PRS EE+ R++ +G + VP P GG + + P PAP
Sbjct: 45 LALPRSTEEVARLIRAAGTKRVP--VLPYGGGTGLVGGQVMPEGPAP 89
>emb|CAG85738.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50417629|ref|XP_457712.1| unnamed protein product
[Debaryomyces hansenii]
Length = 304
Score = 35.4 bits (80), Expect = 1.9
Identities = 21/53 (39%), Positives = 29/53 (54%), Gaps = 2/53 (3%)
Query: 10 FLLSSLVLSLFI--PIFCSFEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGLD 60
FLLS L+ + I P + D +DVD++Q+ L LE LE FY W +D
Sbjct: 5 FLLSGLMAASTIAAPANIAKRDDIDVDILQYALTLEHLENAFYKKALSTWSVD 57
>ref|YP_235415.1| hypothetical protein Psyr_2337 [Pseudomonas syringae pv. syringae
B728a] gi|63256281|gb|AAY37377.1| conserved hypothetical
protein [Pseudomonas syringae pv. syringae B728a]
Length = 320
Score = 33.9 bits (76), Expect = 5.7
Identities = 21/70 (30%), Positives = 29/70 (41%), Gaps = 6/70 (8%)
Query: 177 LVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRN------DLGKKQA 230
L+ L+ + Q + TLL + RYL QW EV + N D GKKQ
Sbjct: 69 LLHALIDSPQWQQSQAHTLLLQHRYLPQSTTQWLTGEVVDEWVITENGLRYKVDFGKKQN 128
Query: 231 EGKVVGQVFG 240
G + +G
Sbjct: 129 SGLFLDMRYG 138
>ref|ZP_00267450.1| COG1092: Predicted SAM-dependent methyltransferases [Pseudomonas
fluorescens PfO-1]
Length = 313
Score = 33.5 bits (75), Expect = 7.4
Identities = 20/74 (27%), Positives = 36/74 (48%), Gaps = 6/74 (8%)
Query: 175 RNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLL------RNDLGKK 228
+ L+ + G++ Q + TLL + RYL +W + E ++++ R DLG+K
Sbjct: 67 KRLLLEITGSEPWQQSGAHTLLIQHRYLPQSTAEWLLGEEIDEMTIVEGGLKYRVDLGRK 126
Query: 229 QAEGKVVGQVFGLN 242
Q G + +G N
Sbjct: 127 QNTGLFLDMRYGRN 140
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 551,340,463
Number of Sequences: 2540612
Number of extensions: 25301800
Number of successful extensions: 48576
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 48533
Number of HSP's gapped (non-prelim): 28
length of query: 293
length of database: 863,360,394
effective HSP length: 127
effective length of query: 166
effective length of database: 540,702,670
effective search space: 89756643220
effective search space used: 89756643220
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0333b.7


