

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0323a.10
(175 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
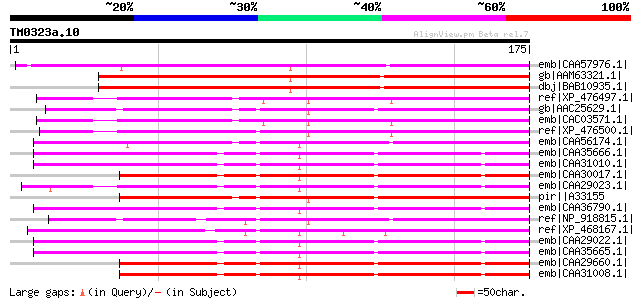
Score E
Sequences producing significant alignments: (bits) Value
emb|CAA57976.1| sts14 [Solanum tuberosum] gi|11177146|gb|AAG3215... 164 1e-39
gb|AAM63321.1| sts14 [Arabidopsis thaliana] 159 2e-38
dbj|BAB10935.1| unnamed protein product [Arabidopsis thaliana] g... 159 3e-38
ref|XP_476497.1| PR-1 type pathogenesis-related protein PR-1a [O... 125 7e-28
gb|AAC25629.1| pathogenesis related protein-1 [Zea mays] gi|7442... 125 7e-28
emb|CAC03571.1| PR1a protein [Oryza sativa (japonica cultivar-gr... 124 9e-28
ref|XP_476500.1| pathogenesis-related protein 1 [Oryza sativa (j... 122 4e-27
emb|CAA56174.1| PR-1 [Medicago truncatula] gi|2500715|sp|Q40374|... 122 6e-27
emb|CAA35666.1| unnamed protein product [Nicotiana tabacum] gi|1... 122 6e-27
emb|CAA31010.1| PR1c preprotein [Nicotiana tabacum] 122 6e-27
emb|CAA30017.1| unnamed protein product [Nicotiana tabacum] 121 7e-27
emb|CAA29023.1| PR-1c protein [Nicotiana tabacum] 121 9e-27
pir||A33155 pathogenesis-related protein 1 - maize gi|228409|prf... 121 9e-27
emb|CAA36790.1| unnamed protein product [Nicotiana tabacum] gi|1... 120 2e-26
ref|NP_918815.1| putative pathogenesis-related protein precursor... 120 2e-26
ref|XP_468167.1| putative pathogenesis related protein-1 [Oryza ... 119 4e-26
emb|CAA29022.1| PR-1b protein [Nicotiana tabacum] 118 6e-26
emb|CAA35665.1| unnamed protein product [Nicotiana tabacum] gi|4... 118 6e-26
emb|CAA29660.1| PR1a precursor (AA -30 to -1) [Nicotiana tabacum... 118 8e-26
emb|CAA31008.1| PR1a preprotein [Nicotiana tabacum] 118 8e-26
>emb|CAA57976.1| sts14 [Solanum tuberosum] gi|11177146|gb|AAG32153.1|
pistil-specific; similar to PR-1 proteins, Swiss-Prot
Accession Number P11670 [Solanum tuberosum]
gi|2500717|sp|Q41495|ST14_SOLTU STS14 protein precursor
gi|2129995|pir||S65052 pistil-specific protein sts14
precursor - potato gi|1589691|prf||2211417A sts14 gene
Length = 214
Score = 164 bits (415), Expect = 1e-39
Identities = 90/180 (50%), Positives = 106/180 (58%), Gaps = 9/180 (5%)
Query: 3 IFMQPFFSLALALFLLSAEAATVLDSVVPQLSAE------AKEFLDAHNWARAEVGVEPL 56
+F Q F L A L A TV P SA A+EFLDAHN AR+EVGV PL
Sbjct: 37 LFFQ-FLLLTTASSLTHISAQTVPPPPPPPTSAATPPSRAAQEFLDAHNKARSEVGVGPL 95
Query: 57 EWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQ-DLGGMGWTPSVAVQDWVNEKEF 115
WS LAK+ L RYQRD+ C AN S+ KYG NQ G TP +AV WV EK+F
Sbjct: 96 TWSPMLAKETSLLVRYQRDKQNCSFANLSNGKYGGNQLWASGTVVTPRMAVDSWVAEKKF 155
Query: 116 YSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
Y+Y NSC C YTQ+VW+KS ++GCAQ TC LT+CFY+PPGN+ GE PY
Sbjct: 156 YNYENNSCTGDD-KCGVYTQIVWKKSIELGCAQRTCYEGPATLTVCFYNPPGNVIGEKPY 214
>gb|AAM63321.1| sts14 [Arabidopsis thaliana]
Length = 185
Score = 159 bits (403), Expect = 2e-38
Identities = 78/147 (53%), Positives = 101/147 (68%), Gaps = 3/147 (2%)
Query: 31 PQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
P +SA AK F DAHN ARA VGV PL WS+ L A ARYQR++ C+ A+ + KYG
Sbjct: 40 PTISAAAKAFTDAHNKARAMVGVPPLVWSQTLEAAASRLARYQRNQKKCEFASLNPGKYG 99
Query: 91 SNQ--DLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQ 148
+NQ G + TPS+AV+ WV EK FY+Y++++C A + C Y QVVWR SK++GCAQ
Sbjct: 100 ANQLWAKGLVAVTPSLAVETWVKEKPFYNYKSDTCAA-NHTCGVYKQVVWRNSKELGCAQ 158
Query: 149 LTCVVDKLILTICFYDPPGNINGESPY 175
TC + +LTICFY+PPGNI G+ PY
Sbjct: 159 ATCTKESTVLTICFYNPPGNIIGQKPY 185
>dbj|BAB10935.1| unnamed protein product [Arabidopsis thaliana]
gi|20259800|gb|AAM13247.1| unknown protein [Arabidopsis
thaliana] gi|15240015|ref|NP_201460.1| allergen
V5/Tpx-1-related family protein [Arabidopsis thaliana]
gi|14423500|gb|AAK62432.1| Unknown protein [Arabidopsis
thaliana]
Length = 185
Score = 159 bits (402), Expect = 3e-38
Identities = 77/147 (52%), Positives = 101/147 (68%), Gaps = 3/147 (2%)
Query: 31 PQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
P +SA AK F DAHN ARA VGV PL WS+ L A ARYQR++ C+ A+ + KYG
Sbjct: 40 PTISAAAKAFTDAHNKARAMVGVPPLVWSQTLEAAASRLARYQRNQKKCEFASLNPGKYG 99
Query: 91 SNQ--DLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQ 148
+NQ G + TPS+AV+ WV EK FY+Y++++C A + C Y QVVWR SK++GCAQ
Sbjct: 100 ANQLWAKGLVAVTPSLAVETWVKEKPFYNYKSDTCAA-NHTCGVYKQVVWRNSKELGCAQ 158
Query: 149 LTCVVDKLILTICFYDPPGNINGESPY 175
TC + +LTICFY+PPGN+ G+ PY
Sbjct: 159 ATCTKESTVLTICFYNPPGNVIGQKPY 185
>ref|XP_476497.1| PR-1 type pathogenesis-related protein PR-1a [Oryza sativa
(japonica cultivar-group)] gi|50509799|dbj|BAD31924.1|
PR-1 type pathogenesis-related protein PR-1a [Oryza
sativa (japonica cultivar-group)]
gi|34395126|dbj|BAC84842.1| PR-1 type
pathogenesis-related protein PR-1a [Oryza sativa
(japonica cultivar-group)]
Length = 168
Score = 125 bits (313), Expect = 7e-28
Identities = 72/170 (42%), Positives = 97/170 (56%), Gaps = 14/170 (8%)
Query: 10 SLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLF 69
S L + +A AAT +S A++F+D HN ARA+VGV P+ W + +A A +
Sbjct: 9 SCCLLVLAAAAMAATAQNS--------AQDFVDPHNAARADVGVGPVSWDDTVAAYAESY 60
Query: 70 ARYQRDRLGCKLANF-SHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAP 126
A ++ CKL + S KYG N G G WT + AV WV+EK++Y + +NSC AP
Sbjct: 61 AAQRQG--DCKLEHSDSGGKYGENIFWGSAGGDWTAASAVSAWVSEKQWYDHGSNSCSAP 118
Query: 127 S-FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +GCA++ C D + C Y PPGN G+SPY
Sbjct: 119 EGSSCGHYTQVVWRDSTAIGCARVVCDGDLGVFITCNYSPPGNFVGQSPY 168
>gb|AAC25629.1| pathogenesis related protein-1 [Zea mays] gi|7442179|pir||T02054
pathogenesis related protein-1 - maize
Length = 163
Score = 125 bits (313), Expect = 7e-28
Identities = 72/165 (43%), Positives = 97/165 (58%), Gaps = 8/165 (4%)
Query: 13 LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARY 72
LA L A AA V+ Q S + +++D HN ARA+VGV P+ W + +A A +A
Sbjct: 5 LACLLALAMAAIVVAPCTAQNSPQ--DYVDPHNAARADVGVGPVSWDDTVAAYAQSYAAQ 62
Query: 73 QRDRLGCKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPSFPC 130
++ CKL + S YG N G G W+ S AV WV+EK++Y + TNSC A C
Sbjct: 63 RQG--DCKLIH-SGGPYGENLFWGSAGADWSASDAVGSWVSEKQYYDHDTNSC-AEGQVC 118
Query: 131 WHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S +GCA++ C + + IC Y+PPGN+ GESPY
Sbjct: 119 GHYTQVVWRDSTAIGCARVVCDNNAGVFIICSYNPPGNVVGESPY 163
>emb|CAC03571.1| PR1a protein [Oryza sativa (japonica cultivar-group)]
gi|12005673|gb|AAG44566.1| acidic PR-1 type
pathogenesis-related protein PR-1a [Oryza sativa subsp.
japonica] gi|11277195|pir||JC7330 acidic
pathogenesis-related protein 1a precursor - rice
Length = 168
Score = 124 bits (312), Expect = 9e-28
Identities = 72/170 (42%), Positives = 97/170 (56%), Gaps = 14/170 (8%)
Query: 10 SLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLF 69
S L + +A AAT +S A++F+D HN ARA+VGV P+ W + +A A +
Sbjct: 9 SCCLLVLAAAAMAATAQNS--------AQDFVDPHNAARADVGVGPVSWDDTVAAYAESY 60
Query: 70 ARYQRDRLGCKLANF-SHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAP 126
A ++ CKL + S KYG N G G WT + AV WV+EK++Y + +NSC AP
Sbjct: 61 AAQRQG--DCKLEHSDSGGKYGENIFWGSPGGDWTAASAVSAWVSEKQWYDHGSNSCSAP 118
Query: 127 S-FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +GCA++ C D + C Y PPGN G+SPY
Sbjct: 119 EGSSCGHYTQVVWRDSTAIGCARVVCDGDLGVFITCNYSPPGNFVGQSPY 168
>ref|XP_476500.1| pathogenesis-related protein 1 [Oryza sativa (japonica
cultivar-group)] gi|21304633|gb|AAM45439.1|
pathogenesis-related protein 1 [Oryza sativa]
gi|34395061|dbj|BAC84723.1| pathogenesis-related protein
1 [Oryza sativa (japonica cultivar-group)]
Length = 165
Score = 122 bits (306), Expect = 4e-27
Identities = 67/168 (39%), Positives = 97/168 (56%), Gaps = 12/168 (7%)
Query: 11 LALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFA 70
L A+ +A AAT +S A++++DAHN AR++VGV P+ W + +A A +A
Sbjct: 7 LVAAVLAAAAMAATAQNS--------AQDYVDAHNAARSDVGVGPVSWDDTVAAYAESYA 58
Query: 71 RYQRDRLGCKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPS- 127
++ + ++ S KYG N G G WT + AV WV EK++Y + +NSC AP+
Sbjct: 59 AQRQGDCALEHSD-SGGKYGENIFWGSAGGDWTAASAVSSWVAEKQWYDHDSNSCSAPAG 117
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVW S +GCA++ C + C Y PPGN++GESPY
Sbjct: 118 SSCGHYTQVVWSNSTAIGCARVVCDNSLGVFITCNYSPPGNVDGESPY 165
>emb|CAA56174.1| PR-1 [Medicago truncatula] gi|2500715|sp|Q40374|PR1_MEDTR
Pathogenesis-related protein PR-1 precursor
gi|629627|pir||S47171 gene PR-1 protein - barrel medic
Length = 173
Score = 122 bits (305), Expect = 6e-27
Identities = 73/172 (42%), Positives = 96/172 (55%), Gaps = 9/172 (5%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAK----EFLDAHNWARAEVGVEPLEWSEALAK 64
FS AL L LL+ + + + S +++ +FL N ARA VG+ PL W + L
Sbjct: 6 FSFALFLLLLTFISHVSASYIPNKKSFKSRSFKNQFLIPQNIARAAVGLRPLVWDDKLTH 65
Query: 65 DAGLFARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSC 123
A +A +R+ C L + S+ YG N G G+GW P+ AV WV+EK+FY+Y NSC
Sbjct: 66 YAQWYANQRRN--DCALEH-SNGPYGENIFWGSGVGWNPAQAVSAWVDEKQFYNYWHNSC 122
Query: 124 VAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
V C HYTQVVW + +VGCA + C DK C YDPPGN GE PY
Sbjct: 123 VDGEM-CGHYTQVVWGSTTKVGCASVVCSDDKGTFMTCNYDPPGNYYGERPY 173
>emb|CAA35666.1| unnamed protein product [Nicotiana tabacum]
gi|130828|sp|P09042|PR1C_TOBAC Pathogenesis-related
protein 1C precursor (PR-1C)
Length = 168
Score = 122 bits (305), Expect = 6e-27
Identities = 72/168 (42%), Positives = 101/168 (59%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S +++LDAHN ARA+VGVEPL W + +A A
Sbjct: 6 FSQMSSFFLVSTLLLFLIISHSCHAQNSQQDYLDAHNTARADVGVEPLTWDDQVAAYAQN 65
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+A + C L + SH +YG N G G T + AV+ WVNEK++Y++ +N+C A
Sbjct: 66 YA--SQLAADCNLVH-SHGQYGENLAWGSGDFLTAAKAVEMWVNEKQYYAHDSNTC-AQG 121
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA++ C I++ C YDPPGN+ G+SPY
Sbjct: 122 QVCGHYTQVVWRNSVRVGCARVQCNNGGYIVS-CNYDPPGNVIGKSPY 168
>emb|CAA31010.1| PR1c preprotein [Nicotiana tabacum]
Length = 163
Score = 122 bits (305), Expect = 6e-27
Identities = 72/168 (42%), Positives = 101/168 (59%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S +++LDAHN ARA+VGVEPL W + +A A
Sbjct: 1 FSQMSSFFLVSTLLLFLIISHSCHAQNSQQDYLDAHNTARADVGVEPLTWDDQVAAYAQN 60
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+A + C L + SH +YG N G G T + AV+ WVNEK++Y++ +N+C A
Sbjct: 61 YA--SQLAADCNLVH-SHGQYGENLAWGSGDFLTAAKAVEMWVNEKQYYAHDSNTC-AQG 116
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA++ C I++ C YDPPGN+ G+SPY
Sbjct: 117 QVCGHYTQVVWRNSVRVGCARVQCNNGGYIVS-CNYDPPGNVIGKSPY 163
>emb|CAA30017.1| unnamed protein product [Nicotiana tabacum]
Length = 168
Score = 121 bits (304), Expect = 7e-27
Identities = 67/139 (48%), Positives = 89/139 (63%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
+++LDAHN ARA+VGVEPL W + +A A +A + C L + SH +YG N G
Sbjct: 35 QDYLDAHNTARADVGVEPLTWDDQVAAYAQNYA--SQLAADCNLVH-SHGQYGENLAEGS 91
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G T + AV+ WVNEK++Y + +N+C A C HYTQVVWR S +VGCA++ C
Sbjct: 92 GDFMTAAKAVEMWVNEKQYYDHDSNTC-AQGQVCGHYTQVVWRNSVRVGCARVQCNNGGY 150
Query: 157 ILTICFYDPPGNINGESPY 175
++T C YDPPGN GESPY
Sbjct: 151 VVT-CNYDPPGNYRGESPY 168
>emb|CAA29023.1| PR-1c protein [Nicotiana tabacum]
Length = 161
Score = 121 bits (303), Expect = 9e-27
Identities = 74/173 (42%), Positives = 105/173 (59%), Gaps = 15/173 (8%)
Query: 5 MQPFFSLA-LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALA 63
M FF ++ L LFL+ + + +S +++LDAHN ARA+VGVEPL W + +A
Sbjct: 2 MSSFFLVSTLLLFLIISHSCHAQNS--------QQDYLDAHNTARADVGVEPLTWDDQVA 53
Query: 64 KDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNS 122
A +A + C L + SH +YG N G G T + AV+ WVNEK++Y++ +N+
Sbjct: 54 AYAQNYA--SQLAADCNLVH-SHGQYGENLAWGSGDFLTAAKAVEMWVNEKQYYAHDSNT 110
Query: 123 CVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C A C HYTQVVWR S +VGCA++ C I++ C YDPPGN+ G+SPY
Sbjct: 111 C-AQGQVCGHYTQVVWRNSVRVGCARVQCNNGGYIVS-CNYDPPGNVIGKSPY 161
>pir||A33155 pathogenesis-related protein 1 - maize gi|228409|prf||1803521A
pathogenesis-related protein 1
Length = 140
Score = 121 bits (303), Expect = 9e-27
Identities = 63/140 (45%), Positives = 87/140 (62%), Gaps = 6/140 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGG 97
++++D HN ARA+VGV P+ W + +A A +A ++ CKL + S YG N G
Sbjct: 5 QDYVDPHNAARADVGVGPVSWDDTVAAYAQSYAAQRQG--DCKLIH-SGGPYGENLFWGS 61
Query: 98 MG--WTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDK 155
G W+ S AV WV+EK++Y + TNSC A C HYTQVVWR S +GCA++ C +
Sbjct: 62 AGADWSASDAVGSWVSEKQYYDHDTNSC-AEGQVCGHYTQVVWRDSTAIGCARVVCDNNA 120
Query: 156 LILTICFYDPPGNINGESPY 175
+ IC Y+PPGN+ GESPY
Sbjct: 121 GVFIICSYNPPGNVVGESPY 140
>emb|CAA36790.1| unnamed protein product [Nicotiana tabacum] gi|100370|pir||S10205
pathogenesis-related protein 1 - common tobacco
Length = 184
Score = 120 bits (301), Expect = 2e-26
Identities = 71/168 (42%), Positives = 99/168 (58%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S+ +++LDAHN ARA+VGVEPL W + +A A
Sbjct: 6 FSQMPSFFLVSTLLLFLIISLSCGAQNSPQDYLDAHNTARADVGVEPLTWDDQVAAYAQN 65
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+A + C L + SH +YG N G G T + AV+ WVNEK++Y + +N+C A
Sbjct: 66 YA--SQLAADCMLVH-SHGQYGENLAWGSGDFMTAAKAVEMWVNEKQYYDHDSNTC-AQG 121
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA+ C +++ C YDPPGN G+SPY
Sbjct: 122 QVCGHYTQVVWRNSVRVGCARAQCNSGGYVVS-CNYDPPGNFVGQSPY 168
>ref|NP_918815.1| putative pathogenesis-related protein precursor [Oryza sativa
(japonica cultivar-group)] gi|22535624|dbj|BAC10798.1|
putative pathogenesis-related protein [Oryza sativa
(japonica cultivar-group)] gi|18461277|dbj|BAB84473.1|
putative pathogenesis-related protein [Oryza sativa
(japonica cultivar-group)]
Length = 167
Score = 120 bits (301), Expect = 2e-26
Identities = 72/165 (43%), Positives = 95/165 (56%), Gaps = 10/165 (6%)
Query: 14 ALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQ 73
+LF+L+ AAT+ Q S + ++L N AR+ VGV P+ WS L G Y
Sbjct: 10 SLFVLAVVAATMFHCSDAQNSPQ--DYLSPQNAARSAVGVGPMSWSTKLQ---GFAEDYA 64
Query: 74 RDRLG-CKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPSFPC 130
R R G C+L + S YG N G G WT + AV+ WV+EK++Y+Y +NSC A C
Sbjct: 65 RQRKGDCRLQH-SGGPYGENIFWGSAGADWTAADAVRSWVDEKKYYNYASNSCAAGKV-C 122
Query: 131 WHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S VGCA++ C ++ I IC Y+P GNI G PY
Sbjct: 123 GHYTQVVWRDSTNVGCARVRCDANRGIFIICNYEPRGNIVGRRPY 167
>ref|XP_468167.1| putative pathogenesis related protein-1 [Oryza sativa (japonica
cultivar-group)] gi|47497162|dbj|BAD19210.1| putative
pathogenesis related protein-1 [Oryza sativa (japonica
cultivar-group)]
Length = 178
Score = 119 bits (298), Expect = 4e-26
Identities = 74/176 (42%), Positives = 92/176 (52%), Gaps = 11/176 (6%)
Query: 7 PFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDA 66
PF ++ + LL A S + + A FLDAHN AR +VGV PL W E LA A
Sbjct: 7 PFAAVTAVILLLHGAADAKSSSSSGKTKSLASGFLDAHNAARRQVGVPPLRWDERLASYA 66
Query: 67 GLFARYQRDRLG----CKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVN-EKEFYSYRT 120
ARY R G C L + SH YG N G G+GW P+ V WV+ E+ Y +
Sbjct: 67 ---ARYAAARSGAGGGCALVH-SHGPYGENLFHGSGVGWAPADVVAAWVSRERALYDAAS 122
Query: 121 NSCVA-PSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
NSC + C HYTQVVWR++ VGCA TC + +C Y+PPGN G PY
Sbjct: 123 NSCRGGDAAACGHYTQVVWRRTTAVGCALATCAGGRGTYGVCSYNPPGNYVGVRPY 178
>emb|CAA29022.1| PR-1b protein [Nicotiana tabacum]
Length = 164
Score = 118 bits (296), Expect = 6e-26
Identities = 69/168 (41%), Positives = 98/168 (58%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S +++LDAHN ARA+VGVEPL W +A A
Sbjct: 2 FSQMPSFFLVSTLLLFLIISHSSHAQNSQQDYLDAHNTARADVGVEPLTWDNGVAAYAQN 61
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+ + C L + SH +YG N G G T + AV+ WV+EK++Y + +N+C A
Sbjct: 62 YV--SQLAADCNLVH-SHGQYGENLAQGSGDFMTAAKAVEMWVDEKQYYDHDSNTC-AQG 117
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA++ C +++ C YDPPGN+ G+SPY
Sbjct: 118 QVCGHYTQVVWRNSVRVGCARVKCNNGGYVVS-CNYDPPGNVIGQSPY 164
>emb|CAA35665.1| unnamed protein product [Nicotiana tabacum]
gi|456200|emb|CAA27183.1| PR-1b precursor; (aa -30-138)
[Nicotiana tabacum] gi|130827|sp|P07053|PR1B_TOBAC
Pathogenesis-related protein 1B precursor (PR-1B)
gi|218306|dbj|BAA14221.1| PR1b protein precursor
[Nicotiana tabacum] gi|224881|prf||1203245A protein
1b,pathogenesis related
Length = 168
Score = 118 bits (296), Expect = 6e-26
Identities = 69/168 (41%), Positives = 98/168 (58%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S +++LDAHN ARA+VGVEPL W +A A
Sbjct: 6 FSQMPSFFLVSTLLLFLIISHSSHAQNSQQDYLDAHNTARADVGVEPLTWDNGVAAYAQN 65
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+ + C L + SH +YG N G G T + AV+ WV+EK++Y + +N+C A
Sbjct: 66 YV--SQLAADCNLVH-SHGQYGENLAQGSGDFMTAAKAVEMWVDEKQYYDHDSNTC-AQG 121
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA++ C +++ C YDPPGN+ G+SPY
Sbjct: 122 QVCGHYTQVVWRNSVRVGCARVKCNNGGYVVS-CNYDPPGNVIGQSPY 168
>emb|CAA29660.1| PR1a precursor (AA -30 to -1) [Nicotiana tabacum]
gi|19936|emb|CAA29392.1| PR-1a precursor (AA -30 to 138)
[Nicotiana tabacum] gi|19934|emb|CAA31233.1| unnamed
protein product [Nicotiana tabacum]
gi|130826|sp|P08299|PR1A_TOBAC Pathogenesis-related
protein 1A precursor (PR-1A)
Length = 168
Score = 118 bits (295), Expect = 8e-26
Identities = 65/139 (46%), Positives = 89/139 (63%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
+++LDAHN ARA+VGVEPL W + +A A +A + C L + SH +YG N G
Sbjct: 35 QDYLDAHNTARADVGVEPLTWDDQVAAYAQNYA--SQLAADCNLVH-SHGQYGENLAEGS 91
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G T + AV+ WV+EK++Y + +N+C A C HYTQVVWR S +VGCA++ C
Sbjct: 92 GDFMTAAKAVEMWVDEKQYYDHDSNTC-AQGQVCGHYTQVVWRNSVRVGCARVQCNNGGY 150
Query: 157 ILTICFYDPPGNINGESPY 175
+++ C YDPPGN GESPY
Sbjct: 151 VVS-CNYDPPGNYRGESPY 168
>emb|CAA31008.1| PR1a preprotein [Nicotiana tabacum]
Length = 165
Score = 118 bits (295), Expect = 8e-26
Identities = 65/139 (46%), Positives = 89/139 (63%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
+++LDAHN ARA+VGVEPL W + +A A +A + C L + SH +YG N G
Sbjct: 32 QDYLDAHNTARADVGVEPLTWDDQVAAYAQNYA--SQLAADCNLVH-SHGQYGENLAEGS 88
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G T + AV+ WV+EK++Y + +N+C A C HYTQVVWR S +VGCA++ C
Sbjct: 89 GDFMTAAKAVEMWVDEKQYYDHDSNTC-AQGQVCGHYTQVVWRNSVRVGCARVQCNNGGY 147
Query: 157 ILTICFYDPPGNINGESPY 175
+++ C YDPPGN GESPY
Sbjct: 148 VVS-CNYDPPGNYRGESPY 165
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 309,455,623
Number of Sequences: 2540612
Number of extensions: 11985666
Number of successful extensions: 24134
Number of sequences better than 10.0: 530
Number of HSP's better than 10.0 without gapping: 322
Number of HSP's successfully gapped in prelim test: 208
Number of HSP's that attempted gapping in prelim test: 23197
Number of HSP's gapped (non-prelim): 550
length of query: 175
length of database: 863,360,394
effective HSP length: 119
effective length of query: 56
effective length of database: 561,027,566
effective search space: 31417543696
effective search space used: 31417543696
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0323a.10


