

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.2
(856 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
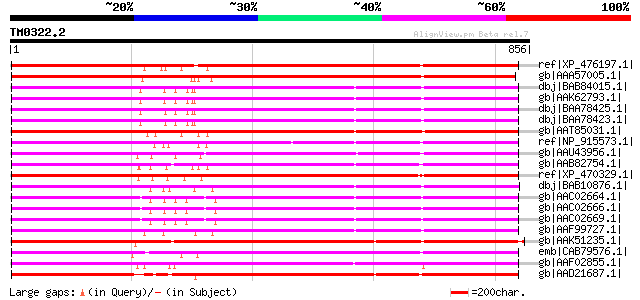
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_476197.1| putative polyprotein [Oryza sativa (japonica cu... 749 0.0
gb|AAA57005.1| copia-like retrotransposon Hopscotch polyprotein ... 748 0.0
dbj|BAB84015.1| polyprotein [Arabidopsis thaliana] gi|14475941|g... 746 0.0
gb|AAK62793.1| polyprotein, putative [Arabidopsis thaliana] 745 0.0
dbj|BAA78425.1| polyprotein [Arabidopsis thaliana] 744 0.0
dbj|BAA78423.1| polyprotein [Arabidopsis thaliana] 744 0.0
gb|AAT85031.1| putative polyprotein [Oryza sativa (japonica cult... 736 0.0
ref|NP_915573.1| putative gag/pol polyprotein [Oryza sativa (jap... 733 0.0
gb|AAU43956.1| unknown protein [Oryza sativa (japonica cultivar-... 730 0.0
gb|AAB82754.1| retrofit [Oryza longistaminata] gi|7444451|pir||T... 721 0.0
ref|XP_470329.1| putative copia-like retrotransposon protein [Or... 714 0.0
dbj|BAB10876.1| polyprotein [Arabidopsis thaliana] 709 0.0
gb|AAC02664.1| polyprotein [Arabidopsis thaliana] 707 0.0
gb|AAC02666.1| polyprotein [Arabidopsis thaliana] 704 0.0
gb|AAC02669.1| polyprotein [Arabidopsis thaliana] 703 0.0
gb|AAF99727.1| F17L21.7 [Arabidopsis thaliana] 701 0.0
gb|AAK51235.1| polyprotein [Arabidopsis thaliana] 687 0.0
emb|CAB79576.1| putative protein [Arabidopsis thaliana] gi|32692... 681 0.0
gb|AAF02855.1| Similar to retrotransposon proteins [Arabidopsis ... 679 0.0
gb|AAD21687.1| Strong similarity to gi|3600044 T12H20.12 proteas... 675 0.0
>ref|XP_476197.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46981313|gb|AAT07631.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|46981245|gb|AAT07563.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1431
Score = 749 bits (1933), Expect = 0.0
Identities = 406/910 (44%), Positives = 560/910 (60%), Gaps = 85/910 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+++ W++ LK KS+ + F F K++E Q ++K+ +QTD GGE+ L +F +GI H
Sbjct: 530 YSKFVWIYFLKHKSEVFQKFHEFQKLVERQFDRKILSMQTDWGGEYLKLHTFFNEIGISH 589
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CPHTH QNG VERKHRHIVE+GL+LLAHA + L++WD AFL YLINRLP+ V+
Sbjct: 590 HVSCPHTHQQNGAVERKHRHIVEVGLSLLAHASMPLKFWDEAFLAATYLINRLPSKVIKF 649
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
++P +LF+ + D K LR FGCAC+P LRPYN HKL SK+CVFLG+S HK +KCLD
Sbjct: 650 DTPLERLFKQTPDYKSLRTFGCACWPNLRPYNTHKLQFRSKQCVFLGFSNIHKGFKCLDV 709
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSV-----------------HSSIP 225
TGR+Y+S+DV F+E FP+ L A + + + + H +
Sbjct: 710 STGRVYVSRDVTFDEQVFPFANLHPNAGARLRAEISLLPPTLINEKTSDQGGEEHHDHLF 769
Query: 226 LFSFPQLTTLS-SPAPHSPNTTLP---------------ESSSPAS-------------- 255
S P T +S + +P + N+ +P ES+S ++
Sbjct: 770 NISMPNATDISCAESPRNVNSDIPGAFGRVHGANGDLAGESASDSASVQAQLQRQASGSA 829
Query: 256 ------QSSTPILP-EPTSPVPQSSPAPISPALS----------SPAPQSSSTDSSSSVS 298
Q + I P TSP + AP SPA + S P SS+ S+ +S
Sbjct: 830 TQGESEQQRSGIQPARATSPAASPTAAPPSPARAVASSSGGAQQSHQPGSSAPSESAQLS 889
Query: 299 PIVNPHNTHSMVTRAKDGIVKPRLVP------TLLLSQAEPTSAKLALKDPKWYSAMQDE 352
++ P TR + GI K ++ + S EP + AL D W AM E
Sbjct: 890 EVIRPK------TRLQSGIRKEKIYTDGTVRYSCFTSSGEPQTLHEALGDKNWKEAMDSE 943
Query: 353 FNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKE 412
+ AL N+TW LVP + I CKWV+++K ADGS++RYKAR+VAKG+ Q G DY++
Sbjct: 944 YQALMKNKTWHLVPSKKGQNIIDCKWVYKVKRKADGSLDRYKARVVAKGFKQRYGIDYED 1003
Query: 413 TFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSLVC 472
TF+PVVK T+R ILS+A+S+GW + QLDV NAFL+G LEE+V+M QPPG+E KD VC
Sbjct: 1004 TFNPVVKAATIRTILSIAISRGWTLRQLDVQNAFLHGVLEEDVFMRQPPGYEQKD-GYVC 1062
Query: 473 RLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIIT 532
+L+KALYGLKQAPRAW+ RL L + GF +S + SLF ++++LVYVDDII+
Sbjct: 1063 KLDKALYGLKQAPRAWYSRLSTKLHELGFKSSKSDTSLFFYSKGDVAMFMLVYVDDIIVA 1122
Query: 533 GNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMA 592
+S+ L+ LN EF+LK LG L YFLG++V+ +++ ++LTQ KY D+L+R +M+
Sbjct: 1123 SSSIDATNALLKNLNQEFALKDLGRLHYFLGIEVKEVNN-GIVLTQEKYAMDVLKRVNMS 1181
Query: 593 YAKGISTPMISNQKLTKHGANYMA--DPSLYRSIVGALQYVTITRPEISYSVNKVCQFLS 650
K ++TP+ ++KL+ H N D + YRS+VGALQY+T+TRP++S+SVNKVCQ+L
Sbjct: 1182 DCKAVNTPLSISEKLSAHEGNPFGPEDSTRYRSLVGALQYLTLTRPDLSFSVNKVCQYLH 1241
Query: 651 QPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSL 710
P +HWAA KRILRYLK TV GL + S L + AF DADWA DD ST G
Sbjct: 1242 APTTKHWAAAKRILRYLKHTVKLGLKI---SKSNSLLVSAFSDADWAGCLDDRHSTGGFA 1298
Query: 711 VFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHI-PHKTPLILCD 769
VF+GPNLVSW ++KQ+ V+RSSTEAEY+ LAN AE++W+Q+LL EL I K + CD
Sbjct: 1299 VFIGPNLVSWSARKQATVSRSSTEAEYKALANVTAEIMWIQTLLHELGIQAPKIAKVWCD 1358
Query: 770 NMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRF 829
N+ +T NPV H RTKH+E+D FVRE+V K L ++++ + DQ D TK LS +
Sbjct: 1359 NIGAKYMTANPVFHARTKHIEVDYHFVRERVARKFLQVEYISTKDQVADGFTKTLSVRQL 1418
Query: 830 EDLRTKLNVT 839
E R LN+T
Sbjct: 1419 EMFRNNLNLT 1428
>gb|AAA57005.1| copia-like retrotransposon Hopscotch polyprotein [Zea mays]
gi|7444442|pir||T02087 gag/pol polyprotein - maize
retrotransposon Hopscotch
Length = 1439
Score = 748 bits (1931), Expect = 0.0
Identities = 412/887 (46%), Positives = 547/887 (61%), Gaps = 61/887 (6%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+++TW++ L+ KSD Y +F F ++E +K+ Q+D GGE++ L +F +GI H
Sbjct: 543 YSKFTWIYLLRHKSDVYKSFCEFQHLVERMFGRKIIAFQSDWGGEYEKLNAHFKTIGIHH 602
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ +CPHTH QNG ERKHRHIVE+GLALLA + + L+YWDHAFL VYLINR P+ + +
Sbjct: 603 QVSCPHTHQQNGAAERKHRHIVEVGLALLAQSSMPLKYWDHAFLAAVYLINRTPSKTIAH 662
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
++P KL + D LRIFGCAC+P LRPYN HKL S CVFLGYS HK +KCLD
Sbjct: 663 DTPLHKLTGATPDYSSLRIFGCACWPNLRPYNQHKLQFRSTRCVFLGYSNMHKGFKCLDI 722
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSV------------------GSVHSSI 224
TGR+YIS+DV+F+E FP+ L A S V SS
Sbjct: 723 STGRIYISRDVVFDEHVFPFASLNKNAGVKYTSEVLLLPHDSCGNNMLTDHANNLPGSSS 782
Query: 225 PL-FSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPIS-PALS 282
PL F +S P S NT + +S ++ S P P+S VP +SPAP A +
Sbjct: 783 PLPFLAQHFLQGNSEVPTSNNTAMALPASGPNEVSVPPALVPSSLVPAASPAPTGVSANA 842
Query: 283 SPAPQSSSTDSSSSVS-------PIVNP----------HNTHSMV--------TRAKDGI 317
PAP++ S S V+ P +P H T TR + GI
Sbjct: 843 EPAPEADSLSSGPPVATESVTGVPDADPLLQAPGSSVAHQTPDSAPLSAAAPRTRLQHGI 902
Query: 318 VKPRLVP--TLLLSQA-----EPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSH 370
KP+ T+ A EP+S AL DP+W +AM+ EF AL N TWTLVP
Sbjct: 903 SKPKQFTDGTVRYGNAAARITEPSSVSEALADPQWRAAMEAEFQALQKNNTWTLVPPDRT 962
Query: 371 RKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLA 430
R I CKWVF++K NADGS++R KARLVAKG+ Q G DY +TFSPVVK T+R++LSLA
Sbjct: 963 RNLIDCKWVFKVKYNADGSIDRLKARLVAKGFKQQYGIDYDDTFSPVVKHSTIRLVLSLA 1022
Query: 431 VSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGF-EAKDRSLVCRLNKALYGLKQAPRAWF 489
VS+ W + QLDV NAFL+G LEE VYM QPPGF + + C L K+LYGLKQ PRAW+
Sbjct: 1023 VSQKWSLRQLDVQNAFLHGILEETVYMKQPPGFADTTHPNYHCHLQKSLYGLKQRPRAWY 1082
Query: 490 ERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAE 549
RL L GF S + SLF + ++Y+LVYVDDIIITG+S + +++AKL +
Sbjct: 1083 SRLSEKLQSLGFVPSKADVSLFIYNAHSTAIYILVYVDDIIITGSSPHAIDNVLAKLKDD 1142
Query: 550 FSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTK 609
F++K LG L YFLG++V H LLL Q KY +DLL+R M K + TP+ +++KL+
Sbjct: 1143 FAIKDLGDLHYFLGIEV-HRKGDGLLLCQEKYARDLLKRVGMECCKPVHTPVATSEKLSA 1201
Query: 610 HGANYMA--DPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYL 667
++ + + YRS+VGALQY+T+TRP++SY++N+VCQFL P + HW AVKRILR +
Sbjct: 1202 SAGTLLSPEETTKYRSVVGALQYLTLTRPDLSYAINRVCQFLHAPTDLHWTAVKRILRNI 1261
Query: 668 KGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSL 727
+ T+ GL ++P L L AF DADWA PDD +ST G +FLGPNL+SW SKKQS
Sbjct: 1262 QHTIGLGLTIRP---SLSLMLSAFSDADWAGCPDDRKSTGGYALFLGPNLISWNSKKQST 1318
Query: 728 VARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHNPVLHTRT 786
V+RSSTEAEY+ +AN AE++W+QSLL EL I P + CDN+ L+ P+ + RT
Sbjct: 1319 VSRSSTEAEYKAMANATAEVIWLQSLLHELGIRLTGIPRLWCDNLGATYLSSKPIFNART 1378
Query: 787 KHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLR 833
KH+E+D FVR++VL+K L I+ + + DQ D TKAL+ R + R
Sbjct: 1379 KHIEVDFHFVRDRVLSKKLDIRLISTNDQVADGFTKALTIGRLNEFR 1425
>dbj|BAB84015.1| polyprotein [Arabidopsis thaliana] gi|14475941|gb|AAK62788.1|
polyprotein, putative [Arabidopsis thaliana]
Length = 1466
Score = 746 bits (1927), Expect = 0.0
Identities = 430/915 (46%), Positives = 555/915 (59%), Gaps = 84/915 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS TF+ F ++E + ++ +D GGEF L YF+ GI H
Sbjct: 552 FTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISH 611
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL+HA I YW +AF VYLINRLPT +L
Sbjct: 612 LTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQL 671
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESPF KLF S + LR+FGCACYP+LRPYN HKL S++CVFLGYS T +Y CL
Sbjct: 672 ESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHL 731
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQ-----LSSVGSVHSSIPLFSFPQLTTLSS 237
T RLYIS+ V F+E FP+ S Q S V S H+++P + P L S
Sbjct: 732 QTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRT-PVLPAPSC 790
Query: 238 PAPHSPNTTLPESSSP-------------ASQSSTPILPEPTSP---VPQ---------- 271
PH T S+P + SS P PEPT+P PQ
Sbjct: 791 SDPHHAATPPSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQT 850
Query: 272 ---------------SSPAPISPALSSPAPQSSSTD------SSSSVSP----------- 299
SP+ ++ +LS+PA SSS+ SSSS SP
Sbjct: 851 QTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPP 910
Query: 300 ----IVN-----PHNTHSMVTRAKDGIVKPRLVPTL---LLSQAEPTSAKLALKDPKWYS 347
IVN P NTHSM TRAK GI+KP +L L +++EP +A ALKD +W +
Sbjct: 911 PLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRN 970
Query: 348 AMQDEFNALHSNQTWTLV-PLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVE 406
AM E NA N TW LV P PSH +GC+W+F K N+DGS+NRYKARLVAKGY+Q
Sbjct: 971 AMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRP 1030
Query: 407 GFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK 466
G DY ETFSPV+K ++RI+L +AV + W I QLDVNNAFL G L ++VYM QPPGF K
Sbjct: 1031 GLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDK 1090
Query: 467 DR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVY 525
DR + VC+L KALYGLKQAPRAW+ L+ LL GF S + SLF L VY+LVY
Sbjct: 1091 DRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVY 1150
Query: 526 VDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDL 585
VDDI+ITGN +L + + L+ FS+K L YFLG++ + + L L+Q +YI DL
Sbjct: 1151 VDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRV-PTGLHLSQRRYILDL 1209
Query: 586 LERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKV 645
L R +M AK ++TPM + KL+ + + DP+ YR IVG+LQY+ TRP+ISY+VN++
Sbjct: 1210 LARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRL 1269
Query: 646 CQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRS 705
QF+ P EEH A+KRILRYL GT + G+ LK LSL A+ DADWA D DD+ S
Sbjct: 1270 SQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKK---GNTLSLHAYSDADWAGDKDDYVS 1326
Query: 706 TSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTP 764
T+G +V+LG + +SW SKKQ V RSSTEAEYR++ANT +E+ W+ SLL EL I + P
Sbjct: 1327 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPP 1386
Query: 765 LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKAL 824
+I CDN+ L NPV H+R KH+ +D F+R +V + +L + HV + DQ D LTK L
Sbjct: 1387 VIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1446
Query: 825 SPLRFEDLRTKLNVT 839
S F++ +K+ VT
Sbjct: 1447 SRTAFQNFASKIGVT 1461
>gb|AAK62793.1| polyprotein, putative [Arabidopsis thaliana]
Length = 1466
Score = 745 bits (1923), Expect = 0.0
Identities = 429/915 (46%), Positives = 554/915 (59%), Gaps = 84/915 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS TF+ F ++E + ++ +D GGEF L YF+ GI H
Sbjct: 552 FTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISH 611
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL+HA I YW +AF VYLINRLPT +L
Sbjct: 612 LTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQL 671
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESPF KLF S + LR+FGCACYP+LRPYN HKL S++CVFLGYS T +Y CL
Sbjct: 672 ESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHL 731
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQ-----LSSVGSVHSSIPLFSFPQLTTLSS 237
T RLYIS+ V F+E FP+ S Q S V S H+++P + P L S
Sbjct: 732 QTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRT-PVLPAPSC 790
Query: 238 PAPHSPNTTLPESSSP-------------ASQSSTPILPEPTSP---VPQ---------- 271
PH T S+P + SS P PEPT+P PQ
Sbjct: 791 SDPHHAATPPSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQT 850
Query: 272 ---------------SSPAPISPALSSPAPQSSSTD------SSSSVSP----------- 299
SP+ ++ +LS+PA SSS+ SSSS SP
Sbjct: 851 QTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPP 910
Query: 300 ----IVN-----PHNTHSMVTRAKDGIVKPRLVPTL---LLSQAEPTSAKLALKDPKWYS 347
IVN P NTHSM TRAK GI+KP +L L +++EP +A ALKD +W +
Sbjct: 911 PLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRN 970
Query: 348 AMQDEFNALHSNQTWTLV-PLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVE 406
AM E NA N TW LV P PSH +GC+W+F K N+DGS+NRYKAR VAKGY+Q
Sbjct: 971 AMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARFVAKGYNQRP 1030
Query: 407 GFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK 466
G DY ETFSPV+K ++RI+L +AV + W I QLDVNNAFL G L ++VYM QPPGF K
Sbjct: 1031 GLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDK 1090
Query: 467 DR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVY 525
DR + VC+L KALYGLKQAPRAW+ L+ LL GF S + SLF L VY+LVY
Sbjct: 1091 DRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVY 1150
Query: 526 VDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDL 585
VDDI+ITGN +L + + L+ FS+K L YFLG++ + + L L+Q +YI DL
Sbjct: 1151 VDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRV-PTGLHLSQRRYILDL 1209
Query: 586 LERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKV 645
L R +M AK ++TPM + KL+ + + DP+ YR IVG+LQY+ TRP+ISY+VN++
Sbjct: 1210 LARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRL 1269
Query: 646 CQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRS 705
QF+ P EEH A+KRILRYL GT + G+ LK LSL A+ DADWA D DD+ S
Sbjct: 1270 SQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKK---GNTLSLHAYSDADWAGDKDDYVS 1326
Query: 706 TSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTP 764
T+G +V+LG + +SW SKKQ V RSSTEAEYR++ANT +E+ W+ SLL EL I + P
Sbjct: 1327 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPP 1386
Query: 765 LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKAL 824
+I CDN+ L NPV H+R KH+ +D F+R +V + +L + HV + DQ D LTK L
Sbjct: 1387 VIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1446
Query: 825 SPLRFEDLRTKLNVT 839
S F++ +K+ VT
Sbjct: 1447 SRTAFQNFASKIGVT 1461
>dbj|BAA78425.1| polyprotein [Arabidopsis thaliana]
Length = 1447
Score = 744 bits (1922), Expect = 0.0
Identities = 428/915 (46%), Positives = 554/915 (59%), Gaps = 84/915 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS TF+ F ++E + ++ +D GGEF L YF+ GI H
Sbjct: 533 FTRYTWLYPLKQKSQVKETFITFKNLVENRFQTRIGTFYSDNGGEFVALREYFSQHGISH 592
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL+HA I YW +AF VYLINRLPT +L
Sbjct: 593 LTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQL 652
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESP KLF S + LR+FGCACYP+LRPYN HKL S++CVFLGYS T +Y CL
Sbjct: 653 ESPCQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHL 712
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQ-----LSSVGSVHSSIPLFSFPQLTTLSS 237
T R+YIS+ V F+E FP+ S Q S V S H+++P + P L S
Sbjct: 713 QTSRIYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRT-PVLPAPSC 771
Query: 238 PAPHSPNTTLPESSSPAS-------------QSSTPILPEPTSP---VPQ---------- 271
PH T S+P+ SS P PEPT+P PQ
Sbjct: 772 SDPHHAATPPSSPSAPSRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQT 831
Query: 272 ---------------SSPAPISPALSSPAPQSSSTD------SSSSVSP----------- 299
SP+ ++ +LS+PA SSS+ SSSS SP
Sbjct: 832 QTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPP 891
Query: 300 ----IVN-----PHNTHSMVTRAKDGIVKPRLVPTL---LLSQAEPTSAKLALKDPKWYS 347
IVN P NTHSM TRAK GI+KP L +L L +++EP +A ALKD +W +
Sbjct: 892 PLAQIVNNNNQAPINTHSMGTRAKAGIIKPNLKYSLAVSLAAESEPRTAIQALKDERWRN 951
Query: 348 AMQDEFNALHSNQTWTLV-PLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVE 406
AM E NA N TW LV P PSH +GC+W+F K N+DGS+NRYKARLVAKGY+Q
Sbjct: 952 AMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRP 1011
Query: 407 GFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK 466
G DY ETFSPV+K ++RI+L +AV + W I QLDVNNAFL G L ++VYM QPPGF K
Sbjct: 1012 GLDYVETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDK 1071
Query: 467 DR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVY 525
DR + VC+L KALYGLKQAPRAW+ L+ LL GF S + SLF L VY+LVY
Sbjct: 1072 DRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVY 1131
Query: 526 VDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDL 585
VDDI+ITGN +L + + L+ FS+K L YFLG++ + + L L+Q +YI DL
Sbjct: 1132 VDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRV-PTGLHLSQRRYILDL 1190
Query: 586 LERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKV 645
L R +M AK ++TPM + KL+ + + DP+ YR IVG+LQY+ TRP+ISY+VN++
Sbjct: 1191 LARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRL 1250
Query: 646 CQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRS 705
QF+ P EEH A+KRILRYL GT + G+ LK LSL A+ DADW D DD+ S
Sbjct: 1251 SQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKK---GNTLSLHAYSDADWTGDKDDYVS 1307
Query: 706 TSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTP 764
T+G +V+LG + +SW SKKQ V RSSTEAEYR++ANT +E+ W+ SLL EL I + P
Sbjct: 1308 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPP 1367
Query: 765 LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKAL 824
+I CDN+ L NPV H+R KH+ +D F+R +V + +L + HV + DQ D LTK L
Sbjct: 1368 VIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1427
Query: 825 SPLRFEDLRTKLNVT 839
S F++ +K+ VT
Sbjct: 1428 SRTAFQNFASKIGVT 1442
>dbj|BAA78423.1| polyprotein [Arabidopsis thaliana]
Length = 1431
Score = 744 bits (1922), Expect = 0.0
Identities = 429/915 (46%), Positives = 553/915 (59%), Gaps = 84/915 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS TF+ F ++E + ++ +D GGEF L YF+ GI H
Sbjct: 517 FTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISH 576
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL+HA I YW +AF VYLINRLPT +L
Sbjct: 577 LTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQL 636
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESPF KLF S + LR+FGCACYP+LRPYN HKL S++CVFLGYS T +Y CL
Sbjct: 637 ESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHL 696
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQ-----LSSVGSVHSSIPLFSFPQLTTLSS 237
T RLYIS+ V F+E FP+ S Q S V S H+++P + P L S
Sbjct: 697 QTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRT-PGLPAPSG 755
Query: 238 PAPHSPNTTLPESSSPAS-------------QSSTPILPEPTSP---VPQ---------- 271
PH T S+P SS P PEPT+P PQ
Sbjct: 756 SDPHHAATPPSSPSAPFRNSQVSSSNRESYFSSSFPSSPEPTAPRQNGPQPKTQPTQTQT 815
Query: 272 ---------------SSPAPISPALSSPAPQSSSTD------SSSSVSP----------- 299
SP+ ++ +LS+PA SSS+ SSSS SP
Sbjct: 816 QTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPP 875
Query: 300 ----IVN-----PHNTHSMVTRAKDGIVKPRLVPTL---LLSQAEPTSAKLALKDPKWYS 347
IVN P NTHSM TRAK GI+KP +L L +++EP +A ALKD +W +
Sbjct: 876 PLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRN 935
Query: 348 AMQDEFNALHSNQTWTLV-PLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVE 406
AM E NA N TW LV P PSH +GC+W+F K N+DGS+NRYKAR VAKGY+Q
Sbjct: 936 AMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARFVAKGYNQRP 995
Query: 407 GFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK 466
G DY ETFSPV+K ++RI+L +AV + W I QLDVNNAFL G L ++VYM QPPGF K
Sbjct: 996 GLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDK 1055
Query: 467 DR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVY 525
DR + VC+L KALYGLKQAPRAW+ L+ LL GF S + SLF L VY+LVY
Sbjct: 1056 DRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVY 1115
Query: 526 VDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDL 585
VDDI+ITGN +L + + L+ FS+K L YFLG++ + + L L+Q +YI DL
Sbjct: 1116 VDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRV-PTGLHLSQRRYILDL 1174
Query: 586 LERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKV 645
L R +M AK ++TPM + KL+ + + DP+ YR IVG+LQY+ TRP+ISY+VN++
Sbjct: 1175 LARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRL 1234
Query: 646 CQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRS 705
QF+ P EEH A+KRILRYL GT + G+ LK LSL A+ DADWA D DD+ S
Sbjct: 1235 SQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKK---GNTLSLHAYSDADWAGDKDDYVS 1291
Query: 706 TSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTP 764
T+G +V+LG + +SW SKKQ V RSSTEAEYR++ANT +E+ W+ SLL EL I + P
Sbjct: 1292 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPP 1351
Query: 765 LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKAL 824
+I CDN+ L NPV H+R KH+ +D F+R +V + +L + HV + DQ D LTK L
Sbjct: 1352 VIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1411
Query: 825 SPLRFEDLRTKLNVT 839
S F++ +K+ VT
Sbjct: 1412 SRTAFQNFASKIGVT 1426
>gb|AAT85031.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1437
Score = 736 bits (1900), Expect = 0.0
Identities = 403/892 (45%), Positives = 550/892 (61%), Gaps = 61/892 (6%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+++TW++ LK KS+ + F F ++E N+K+ +QTD GGE++ L +F +GI H
Sbjct: 541 YSKFTWIYLLKYKSEVFDKFHEFQSLVERLFNRKIVAMQTDWGGEYQKLHSFFNKVGITH 600
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CPHTH QNG ERKHRHIVE+GLALLA++ + L++W AFL+ VYLINR P+ VL +
Sbjct: 601 HVSCPHTHQQNGSAERKHRHIVEVGLALLAYSSMPLKFWGEAFLSAVYLINRTPSRVLHD 660
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLDP 183
SP +L D LR+FGCAC+P LRPYN HKL S C FLGYST HK +KCLDP
Sbjct: 661 VSPLERLLGHKPDYNALRVFGCACWPNLRPYNKHKLQFRSTTCTFLGYSTLHKGFKCLDP 720
Query: 184 -TGRLYISKDVLFNEFRFPYKEL---FGCASSPQLSSVGSVHSSIP--------LFSFPQ 231
TGR+YIS+DV+F+E +FP+ +L G +++ V + +S+P + + P+
Sbjct: 721 STGRVYISRDVVFDETQFPFTKLHPNVGAKLRAEIALVPELAASLPRGLQQISSVINTPE 780
Query: 232 LTTLSSP----------APHSPNTTLPE---SSSPASQSSTPILPEPTSPVPQSSPAPIS 278
+S+ P + P+ +++PA S +P + EP SP +S A S
Sbjct: 781 NANVSNENMQQDSTYDNEPETETDGAPDTVSANAPAESSGSPPINEPASPFGESDSATAS 840
Query: 279 PALS--------------SPAPQSSSTDSSSSVSPIVNPHNTHSMV--------TRAKDG 316
PA + S AP+ S++ + I +PH ++ TR + G
Sbjct: 841 PASAPVNSAPHPDAAASGSSAPRGSTSQGGTPSVAIDDPHPATTVTGQEAQRPRTRLQSG 900
Query: 317 IVKPRLVPT------LLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSH 370
I K ++ +L S EP + + AL++ W AM E+ AL N TW LVP
Sbjct: 901 IRKEKVYTDGTVKWGMLTSTGEPENLQDALQNNNWKCAMDAEYMALIKNNTWHLVPPQQG 960
Query: 371 RKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLA 430
R I CKWV++IK DGS++RYKARLVAKG+ Q G DY++TFSPVVK T+RIILS+A
Sbjct: 961 RNVIDCKWVYKIKRKQDGSLDRYKARLVAKGFKQRYGIDYEDTFSPVVKAATIRIILSIA 1020
Query: 431 VSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFE-AKDRSLVCRLNKALYGLKQAPRAWF 489
VS+GW + QLDV NAFL+G LEEEVYM QPPG+E VC+L+KALYGLKQAPRAW+
Sbjct: 1021 VSRGWCLRQLDVQNAFLHGVLEEEVYMKQPPGYENPSTPDYVCKLDKALYGLKQAPRAWY 1080
Query: 490 ERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAE 549
RL L GF S + SLF ++++L+YVDDII+ + + L+ L E
Sbjct: 1081 SRLSGKLHDLGFKGSKADTSLFFYNKGSLTIFLLIYVDDIIVVSSRKEAVSALLQDLQKE 1140
Query: 550 FSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTK 609
F+LK LG L YFLG++V + +L++Q KY DLL+R +M+ K ++TP+ +++KL
Sbjct: 1141 FALKDLGDLHYFLGIEVTKI-PGGILMSQEKYASDLLKRVNMSDCKSVATPLSASEKLIA 1199
Query: 610 HGANYMA--DPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYL 667
+ D + YRSIVGALQY+T+TR +I++SVNKVCQFL P EHWAAVKRILRY+
Sbjct: 1200 GKGTILGPNDATQYRSIVGALQYLTLTRLDIAFSVNKVCQFLHNPTTEHWAAVKRILRYI 1259
Query: 668 KGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSL 727
K GL + S + + + DADWA DD RST G V+LG NLVSW +KKQ+
Sbjct: 1260 KQCTGLGLRICKSS---SMIVSGYSDADWAGCLDDRRSTGGFAVYLGDNLVSWNAKKQAT 1316
Query: 728 VARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLIL-CDNMSTVALTHNPVLHTRT 786
V+RSSTEAEY+ LAN AE++WVQ+LLQEL+I L CDNM L+ NPV H RT
Sbjct: 1317 VSRSSTEAEYKALANATAEIMWVQTLLQELNIVSPAMAQLWCDNMGAKYLSFNPVFHART 1376
Query: 787 KHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
KH+E+D FVRE+V K L + +V + DQ D TKAL + E+ + LN+
Sbjct: 1377 KHIEVDYHFVRERVARKLLQVDYVSTNDQVADGFTKALPVKQLENFKYNLNL 1428
>ref|NP_915573.1| putative gag/pol polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1449
Score = 733 bits (1892), Expect = 0.0
Identities = 411/911 (45%), Positives = 547/911 (59%), Gaps = 82/911 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
++++TW++ +K KS+ F F ++E N+K+ +Q+D GGE++ L +FT +GI H
Sbjct: 542 FSKFTWIYLIKFKSEVIQKFHEFQALVERLFNRKILAMQSDWGGEYEKLNSFFTKIGITH 601
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CPHTH QNG ERKHRHIVEMGLALLA++ + L++WD AF + VYLINR P+ +L
Sbjct: 602 HVSCPHTHQQNGSAERKHRHIVEMGLALLAYSSMPLKFWDEAFTSAVYLINRTPSRLLKY 661
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLDP 183
+P +LF+ + D L +FGCAC+P+LRPYNA KL SK+C FLGYS HK +KCLDP
Sbjct: 662 ATPLERLFKQTPDYNILCVFGCACWPHLRPYNARKLQFRSKQCTFLGYSPLHKGFKCLDP 721
Query: 184 -TGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSS----- 237
TGR+YIS+DV+F E FP+ L A + S + + S + + P + +S+
Sbjct: 722 STGRVYISRDVIFYENVFPFASLHPNAGAKLRSEILLLPSHLDIPIIPGVQQMSTTVANV 781
Query: 238 --------------------------------------PAPHSPNTTLPESSS------- 252
PA SP+T+ SSS
Sbjct: 782 PTNPANQISGDELSLQENLAANNAAPDGADAATAATKDPAADSPHTSDAASSSVLQRRVE 841
Query: 253 -PASQSSTP-----ILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNT 306
S +P L SPV SP SP SS + + S++ +S +
Sbjct: 842 ATVESSCSPDRLHVELTPAHSPVAMQSPDATSPTASSDSQEPSTSLDASVAGEVDTQQQQ 901
Query: 307 HSMV----TRAKDGIVKPRLV---------PTLLLSQAEPTSAKLALKDPKWYSAMQDEF 353
H ++ TR + GI++ V T L EP + ALKD W AM EF
Sbjct: 902 HQVISRPATRLQSGIIRKEKVYTDGTVKYRTTFLTVTGEPENLDDALKDNNWKKAMDAEF 961
Query: 354 NALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKET 413
AL +N+TW LVP R I CKWV++IK DGS++RYKARLVAKG+ Q G DY++T
Sbjct: 962 MALQNNKTWHLVPPQKGRNVIDCKWVYKIKRKQDGSLDRYKARLVAKGFKQRYGIDYEDT 1021
Query: 414 FSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSL--- 470
FSPVVK T+RIILS+AVSKGW + QLDV NAFL+G LEEEVYM QPPG+E D SL
Sbjct: 1022 FSPVVKAATIRIILSIAVSKGWCMRQLDVQNAFLHGVLEEEVYMKQPPGYE--DSSLPGY 1079
Query: 471 VCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDII 530
VC+L+KALYGLKQAPRAW+ RL L GF S + SLF ++VL+YVDDII
Sbjct: 1080 VCKLDKALYGLKQAPRAWYSRLSKKLYDMGFQGSKADTSLFFYNKGDLIIFVLIYVDDII 1139
Query: 531 ITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERAS 590
+T + + L+ L EF+LK LG L YFLG++V + ++LTQ KY+ DLL R +
Sbjct: 1140 VTSSRQEAVPALLQDLQKEFALKDLGDLHYFLGIEVTKV-SGGIVLTQDKYVNDLLRRVN 1198
Query: 591 MAYAKGISTPMISNQKLTKHGANYMA--DPSLYRSIVGALQYVTITRPEISYSVNKVCQF 648
M K +STP+ +++KL+ + + + D + YRS+VGALQY+T+TRP+I++ VNKVCQF
Sbjct: 1199 MFDCKPVSTPLSTSEKLSINEGDPLGPNDATHYRSVVGALQYLTLTRPDIAFPVNKVCQF 1258
Query: 649 LSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSG 708
L P HWAAVKRILRYLK GL L S + + + A DADWA DD RST G
Sbjct: 1259 LHAPTTHHWAAVKRILRYLKQCTRLGLKL---SKSRSMLVSACSDADWAGSLDDRRSTGG 1315
Query: 709 SLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLIL- 767
VFLG NLVSW ++KQ+ V+RSSTE+EY+ LAN AE++WVQ+LL EL + + L
Sbjct: 1316 FAVFLGDNLVSWSARKQATVSRSSTESEYKALANATAEIMWVQTLLAELQVESPSMAKLW 1375
Query: 768 CDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPL 827
CDN+ L+ NPV H RTKH+E+D FVRE+V K L I VP+ DQ D TKALS
Sbjct: 1376 CDNLGAKYLSANPVFHARTKHIEVDYHFVRERVSRKLLEIDFVPTGDQVADGFTKALSVR 1435
Query: 828 RFEDLRTKLNV 838
+ E+ + LN+
Sbjct: 1436 QLENFKYNLNL 1446
>gb|AAU43956.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|52353503|gb|AAU44069.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1447
Score = 730 bits (1884), Expect = 0.0
Identities = 405/909 (44%), Positives = 546/909 (59%), Gaps = 81/909 (8%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+++TW++ +K KS+ F F ++E N+K+ +Q+D GGE++ L +FT +GI H
Sbjct: 543 YSKFTWLYLIKHKSEVIQKFHEFQALVERLFNRKIIAMQSDWGGEYEKLHSFFTKIGITH 602
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CPHTH QNG ERKHRHIVE+GL LLA++ + L++WD AF VYLINR PT +L
Sbjct: 603 HVSCPHTHQQNGSAERKHRHIVEVGLTLLAYSSMPLKFWDEAFQAAVYLINRTPTKLLQF 662
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLDP 183
+P LF + D LR+FGCAC+P+LRPYN HKL SK+C FLGYST HK +KCLDP
Sbjct: 663 LTPLEHLFNQTPDYSSLRVFGCACWPHLRPYNTHKLQFRSKQCTFLGYSTLHKGFKCLDP 722
Query: 184 -TGRLYISKDVLFNEFRFPYKELFGCA-------------------SSPQLSSVGSVHSS 223
TGR+YIS+DV+F+E FP+ +L A S+P + ++
Sbjct: 723 STGRVYISRDVIFDETNFPFAKLHTNAGARLRSEILLLPSHLLNPTSNPGEQQLDDNMAN 782
Query: 224 IPLFSFPQL--------------------------TTLSSPAPHSPNTTLPESSSPASQS 257
IP+ Q+ + SP+ + +S P + +
Sbjct: 783 IPVNPANQIFGSNVQPVSAENDEADDSSGATENLAEQIHQDTAASPSASDTAASQPGAAT 842
Query: 258 STPILPEPTSPVPQS-----SPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTR 312
S ++ P SP + S +P P+ ++ P S+ S S P T VT+
Sbjct: 843 SLDVVHFPASPDVMTHHSADSSSPSQPSHATATPASNDDVPSPSHISASEPFATGEEVTQ 902
Query: 313 A------------------KDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFN 354
A DG VK + T L EP + AL++ W AM E+
Sbjct: 903 APSRPTTRLQRGIRKEKVYTDGTVKYK--NTFLTVTGEPKNLTDALQNTNWKKAMDIEYE 960
Query: 355 ALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETF 414
AL +N+TW LVP R I CKWV++IK DGS++RYKARLVAKG+ Q G DY++TF
Sbjct: 961 ALMNNKTWHLVPPKQGRNVIDCKWVYKIKRKQDGSLDRYKARLVAKGFKQRYGIDYEDTF 1020
Query: 415 SPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKD-RSLVCR 473
SPVVK T+RI+LS+AVS+GW + QLDV NAFL+G LEEEVYM QPPG+E + VC+
Sbjct: 1021 SPVVKAATIRIVLSIAVSRGWCMRQLDVQNAFLHGFLEEEVYMKQPPGYEDESFPGYVCK 1080
Query: 474 LNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITG 533
L+KALYGLKQAPRAW+ RL L GF S + SLF G ++VL+YVDDII+T
Sbjct: 1081 LDKALYGLKQAPRAWYSRLSKKLYDLGFQGSKGDTSLFFYNKGGLIIFVLIYVDDIIVTS 1140
Query: 534 NSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAY 593
+ + L+ L EF+LK LG L YFLG++V + D+ ++LTQ KY DLL R +M
Sbjct: 1141 SRQEAVSALLQDLKKEFALKDLGDLHYFLGIEVNKVTDE-IILTQDKYACDLLRRVNMFD 1199
Query: 594 AKGISTPMISNQKLTKHGANYMA--DPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQ 651
K +STP+ +++KL+ H + + D + YRS+VGALQY+T+TRP+I++ VNKVCQFL
Sbjct: 1200 CKPVSTPLSTSEKLSAHEGDLLGPLDATNYRSVVGALQYLTLTRPDIAFPVNKVCQFLHA 1259
Query: 652 PLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLV 711
P HWAAVKRILRYLK GL L C + + + A+ DADWA DD RST G V
Sbjct: 1260 PTTVHWAAVKRILRYLKQCTKLGLKL--CK-SKSMLVSAYSDADWAGSLDDRRSTGGFAV 1316
Query: 712 FLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPL--ILCD 769
FLG NLVSWC++KQ+ V+RSSTE+EY+ LAN AE++WVQ+LL EL + P+ + CD
Sbjct: 1317 FLGDNLVSWCARKQATVSRSSTESEYKALANATAEIMWVQTLLTELQV-QSPPMAKLWCD 1375
Query: 770 NMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRF 829
N+ L+ NPV H RTKH+E+D FVRE+V K L I VP+ DQ D TKAL +
Sbjct: 1376 NLGAKYLSSNPVFHARTKHIEVDYHFVRERVSQKLLEIDFVPTGDQVADGFTKALPVRQL 1435
Query: 830 EDLRTKLNV 838
E+ + LN+
Sbjct: 1436 ENFKHNLNL 1444
>gb|AAB82754.1| retrofit [Oryza longistaminata] gi|7444451|pir||T10728 probable
gag/pol polyprotein - long-staminate rice retrotransposon
retrofit
Length = 1445
Score = 721 bits (1860), Expect = 0.0
Identities = 407/910 (44%), Positives = 547/910 (59%), Gaps = 84/910 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDG-GGEFKPLTPYFTNLGIV 62
++++TW++ LK KS+ + F F ++E ++K+ +QTD GG ++ L +F +G++
Sbjct: 542 FSKFTWIYLLKYKSEVFEKFKEFQALVERMFDRKIIAMQTDWRGGRYQKLNSFFAQIGLI 601
Query: 63 HRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLD 122
QNG ERKHRHIVE+GL+LL++A + L++WD AF+ YLINR+P+ +
Sbjct: 602 IMCHVLTLIRQNGSAERKHRHIVEVGLSLLSYASMPLKFWDEAFVAATYLINRIPSKTIQ 661
Query: 123 NESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD 182
N +P KLF D LR+FGCAC+P+LRPYN HKL SK+CVFLG+ST HK +KCLD
Sbjct: 662 NSTPLEKLFNQKPDYSSLRVFGCACWPHLRPYNTHKLQFRSKQCVFLGFSTHHKGFKCLD 721
Query: 183 -PTGRLYISKDVLFNEFRFPYKELFGCAS-----------SP----QLSSVGSVHSSIPL 226
+GR+YIS+DV+F+E FP+ L A SP +S G H P+
Sbjct: 722 VSSGRVYISRDVVFDENVFPFSTLHSNAGARLRSEILLLPSPLTNYNTASAGGTHVVAPV 781
Query: 227 FSFP----QLTTLSSPAPHSPNTTLPESSSPASQ---------------SSTPILPEPTS 267
+ P L + ++ N+ E Q +S P+L +PT+
Sbjct: 782 ANTPLPSDNLISNAADVTSGENSAAHEQEMENEQEIENVMHGNDVHGDAASGPVLDQPTA 841
Query: 268 PVPQSSPAPISPALSSPAPQSSSTDSSSSVSPI-----------------VNPHNTHSMV 310
SS AP A +S A +++D+ + + V P T +++
Sbjct: 842 ---DSSTAPDQGADTSDAVSGAASDAGGDTATLGAGAANSAAAGGEESQPVQPDVTGTVL 898
Query: 311 ----------TRAKDGIVKPRLVPT------LLLSQAEPTSAKLALKDPKWYSAMQDEFN 354
TR + GI K ++ S EP + K AL D W AM+ E+N
Sbjct: 899 ATVAPASRPHTRLRSGIRKEKVYTDGTVKYGCFSSTGEPQNDKEALGDKNWRDAMETEYN 958
Query: 355 ALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETF 414
AL N TW LVP + IGCKWV++IK ADG+++RYKARLVAKG+ Q G DY++TF
Sbjct: 959 ALIKNDTWHLVPYEKGQNIIGCKWVYKIKRKADGTLDRYKARLVAKGFKQRYGIDYEDTF 1018
Query: 415 SPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCR 473
SPVVK T+RIILS+AVS+GW + QLDV NAFL+G LEEEVYM QPPGFE+ + VC+
Sbjct: 1019 SPVVKAATIRIILSIAVSRGWSLRQLDVQNAFLHGFLEEEVYMQQPPGFESSSKPDYVCK 1078
Query: 474 LNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITG 533
L+KALYGLKQAPRAW+ RL L++ GF AS + SLF L G ++VLVYVDDII+
Sbjct: 1079 LDKALYGLKQAPRAWYSRLSKKLVELGFEASKADTSLFFLNKGGILMFVLVYVDDIIVAS 1138
Query: 534 NSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAY 593
++ L+ LN EF+LK LG L YFLG++V + ++LTQ KY DLL+R +M+
Sbjct: 1139 STEKATTALLKDLNKEFALKDLGDLHYFLGIEVTKV-SNGVILTQEKYANDLLKRVNMSN 1197
Query: 594 AKGISTPMISNQKLTKHGANYMA--DPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQ 651
K +STP+ ++KLT + + + D YRSIVGALQY+T+TRP+I+YSVNKVCQFL
Sbjct: 1198 CKPVSTPLSVSEKLTLYEGSPLGPNDAIQYRSIVGALQYLTLTRPDIAYSVNKVCQFLHA 1257
Query: 652 PLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLI--AFCDADWAADPDDHRSTSGS 709
P HW AVKRILRYL GLH +H+ S + + DADWA DD +ST G
Sbjct: 1258 PTTSHWIAVKRILRYLNQCTSLGLH-----IHKSASTLVHGYSDADWAGSIDDRKSTGGF 1312
Query: 710 LVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPH-KTPLILC 768
VFLG NLVSW ++KQ V+RSSTEAEY+ +ANT AEL+WVQ+LL+EL I K I C
Sbjct: 1313 AVFLGSNLVSWSARKQPTVSRSSTEAEYKAVANTTAELIWVQTLLKELGIESPKAAKIWC 1372
Query: 769 DNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLR 828
DN+ L+ NPV H RTKH+E+D FVRE+V K L I VPS DQ D TKALS
Sbjct: 1373 DNLGAKYLSANPVFHARTKHIEVDYHFVRERVSQKLLEIDFVPSGDQVADGFTKALSACL 1432
Query: 829 FEDLRTKLNV 838
E+ + LN+
Sbjct: 1433 LENFKHNLNL 1442
>ref|XP_470329.1| putative copia-like retrotransposon protein [Oryza sativa (japonica
cultivar-group)] gi|40714683|gb|AAR88589.1| putative
copia-like retrotransposon protein [Oryza sativa
(japonica cultivar-group)]
Length = 1399
Score = 714 bits (1843), Expect = 0.0
Identities = 394/890 (44%), Positives = 539/890 (60%), Gaps = 59/890 (6%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+++TWV+ LK KS+ + F F ++E K+ VQTD GGE++ L +F +GI H
Sbjct: 505 YSKFTWVYLLKHKSEVFQKFQEFQTLVERFFGHKILAVQTDWGGEYQKLNTFFAKIGISH 564
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CP+ H QNG ERKHRH++E+ L LLAHA + +++WD A L YLINR P+ V++
Sbjct: 565 HVSCPYAHQQNGSAERKHRHLIEVALTLLAHASMPIKFWDEAVLAAAYLINRTPSKVINF 624
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
P +LF+ + LRIFGCA +P LRPYN HKL+ SK CVFLGYS HK +KCL+
Sbjct: 625 ACPLEQLFKEKPNYTALRIFGCAVWPNLRPYNKHKLAFRSKRCVFLGYSNLHKGFKCLEI 684
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASS--------------PQLSSVGSVHSSIPLFS 228
TGR+Y+S+DV F+E FP+ EL A + P LSS+G ++ L
Sbjct: 685 ATGRVYVSRDVTFDESIFPFSELHSNAGARLRAEISLLPPSLVPHLSSLGGEQNNHVLNY 744
Query: 229 FPQLT------------TLSSPAPHSPNTTLPESSSPASQSST---------PILPEPTS 267
P +T + + + E+++ A+ PE +S
Sbjct: 745 PPNVTDQFGEENAEIGEEIVANGEENAAAAADENAAAAANGGAQDDVHGVAYDASPEHSS 804
Query: 268 PVPQSSPAPISPALSSPAPQ----SSSTDSSSSVSP-IVNPHNTHSMVTRAK-------- 314
PV + A + +P + +S ++SS SP + + H VT +
Sbjct: 805 PVTDDAMASAAEQHGNPIQEEHLVQASPQTASSTSPSVASSAGVHDDVTTDQSDQTDQAM 864
Query: 315 -DGIVKPRLVPTLLLSQ-AEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRK 372
+ V P T L S + S + A+ + W AM E+ AL N+TW LVP R
Sbjct: 865 PEAAVAPIRPKTRLQSGIRKEKSLEEAVNNKHWKEAMDAEYMALIENKTWHLVPPQKGRN 924
Query: 373 AIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVS 432
I CKWV+++K ADGS++RYKARLVAKG+ Q G DY++TFSPVVK T+RI+LSLAVS
Sbjct: 925 VIDCKWVYKVKRKADGSLDRYKARLVAKGFKQRYGIDYEDTFSPVVKAATIRIVLSLAVS 984
Query: 433 KGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLNKALYGLKQAPRAWFER 491
+GW + QLDV NAFL+G LEEEVYM QPPG+E K + VC+L+KALYGLKQAPRAW+ R
Sbjct: 985 RGWSLRQLDVKNAFLHGVLEEEVYMKQPPGYEKKSMPNYVCKLDKALYGLKQAPRAWYSR 1044
Query: 492 LKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFS 551
L L + GF S + SLF S+++L+YVDDII+ + L+ +L+ +F+
Sbjct: 1045 LSTKLSELGFVPSKADTSLFFYKKGQVSIFLLIYVDDIIVASSVPDATSTLLQELSKDFA 1104
Query: 552 LKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHG 611
LK LG L YFLG++V + D L+L+Q KY DLL R M K +STP+ +++KL+ +
Sbjct: 1105 LKDLGDLHYFLGIEVHKVKD-GLMLSQEKYASDLLRRVGMYECKPVSTPLSTSEKLSVNE 1163
Query: 612 ANYMA--DPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKG 669
+ D + YRS+VGALQY+T+TRP+IS+S+NKVCQFL P HWAAVKRILRY+K
Sbjct: 1164 GTLLGPQDSTQYRSVVGALQYLTLTRPDISFSINKVCQFLHAPTTTHWAAVKRILRYVKY 1223
Query: 670 TVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVA 729
TVD G LK C + L + F DADWA PDD RST G VFLGPNLVSW ++KQ+ V+
Sbjct: 1224 TVDTG--LKFCR-NPSLLVSGFSDADWAGSPDDRRSTGGFAVFLGPNLVSWSARKQATVS 1280
Query: 730 RSSTEAEYRNLANTVAELLWVQSLLQELHIPH-KTPLILCDNMSTVALTHNPVLHTRTKH 788
RSS EAEY+ LAN AE++WVQ+LLQEL + + + CDN+ L+ NP+ H RTKH
Sbjct: 1281 RSSIEAEYKALANATAEIMWVQTLLQELGVESPRAAKLWCDNLGAKYLSANPIFHARTKH 1340
Query: 789 MELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
+E+D FVRE+V K L I ++ + DQ D TKA+ + E + LN+
Sbjct: 1341 IEVDFHFVRERVARKLLEIAYISTKDQVADGFTKAIPVRQMEMFKNNLNL 1390
>dbj|BAB10876.1| polyprotein [Arabidopsis thaliana]
Length = 1429
Score = 709 bits (1830), Expect = 0.0
Identities = 399/896 (44%), Positives = 526/896 (58%), Gaps = 64/896 (7%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS FV F ++E + +++ + +D GGEF L P+ GI H
Sbjct: 535 FTRYTWMYPLKQKSQVKDVFVAFKALVENRFQSRIRTLYSDNGGEFIGLRPFLAAHGISH 594
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GLALL HA + +W +AF T VYLINR+PT VL
Sbjct: 595 LTSPPHTPEHNGLAERKHRHIVETGLALLTHASLPKTFWTYAFATAVYLINRMPTEVLQG 654
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
SP+ KLFQ+S + LR+FGC CYP+LRPYN +KL S CVFLGYS T +Y CLD
Sbjct: 655 TSPYVKLFQMSPNYLKLRVFGCLCYPWLRPYNTNKLEARSTMCVFLGYSLTQSAYLCLDI 714
Query: 183 PTGRLYISKDVLFNEFRFPY-KELFGCASSPQLSSVGSVHSSIPLFSFP---------QL 232
T R+Y S+ V F E FP+ S Q S + + IPL P L
Sbjct: 715 ATNRIYTSRHVQFVESSFPFASPRTSETDSTQTMSQPTTTNVIPLLQRPPHIAPPTALPL 774
Query: 233 TTLSSPAPHSPNTTLPESS-----SPASQSSTPI------------LPEPTSPVPQSSPA 275
+ PHSP++ S + AS SS I + PTS P ++P
Sbjct: 775 CPIFHSPPHSPSSPASPPSEHVPLTAASSSSNAINDDNISSTGQVSVSGPTSQSPHTTPT 834
Query: 276 PISPALSSPAPQSSSTDSSSSVSPIVN------------------------PHNTHSMVT 311
+ + S +P ++T+ S + +P + P N H M T
Sbjct: 835 NQNTSPLSKSPNPTNTNQSQNSTPPTSPTTSVHQHSPTPSPLPQNPPLPPPPQNDHPMRT 894
Query: 312 RAKDGIVKPRLVPTLLLSQAE-----PTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVP 366
RAK+ I KP+ L S PT+ ALKDP W +AM +E NA N TW LV
Sbjct: 895 RAKNQITKPKTKFNLTTSLTSSKPTIPTTVAQALKDPNWRNAMSEEINAQMKNHTWDLVS 954
Query: 367 LPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRII 426
+ I CKW+F +K N DGS+ RYKARLVA+G++Q G DY ETFSPV+K T+R +
Sbjct: 955 PEEAKHVISCKWIFTLKYNVDGSIARYKARLVARGFNQQYGIDYSETFSPVIKSTTIRTV 1014
Query: 427 LSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLNKALYGLKQAP 485
L +AV + W IHQ+D+NNAFL G L EEVY+ QPPGF +DR S VCRLNKALYGLKQAP
Sbjct: 1015 LEVAVKRNWSIHQVDINNAFLQGTLNEEVYVSQPPGFIDRDRPSHVCRLNKALYGLKQAP 1074
Query: 486 RAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAK 545
RAW++ L+ LL+ GF S + SLF +YVLVYVDDIII G + ++Q A
Sbjct: 1075 RAWYQELRRFLLQAGFVNSLADASLFIYNRHNTFMYVLVYVDDIIIAGEN-ALVQAFNAS 1133
Query: 546 LNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQ 605
L + FSLK LG L YFLG++ + L L Q KYI DLL++ +M K +STPM
Sbjct: 1134 LASRFSLKDLGPLSYFLGIEATRT-SRGLHLMQRKYITDLLKKHNMLDTKPVSTPMSPTP 1192
Query: 606 KLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILR 665
KL+ + D + YR+++G+LQY+ TRP+I+++VN++ QF+ +P EHW A KRILR
Sbjct: 1193 KLSLLSGTALDDATEYRTVLGSLQYLAFTRPDIAFAVNRLSQFMHRPTNEHWQAAKRILR 1252
Query: 666 YLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQ 725
YL GT G+ L+ PL++ AF DADW D D + ST+ +V+ G + VSW SKKQ
Sbjct: 1253 YLAGTKSHGIFLRS---DTPLTIHAFSDADWGCDLDAYLSTNAYIVYFGGSPVSWSSKKQ 1309
Query: 726 SLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKT-PLILCDNMSTVALTHNPVLHT 784
VARSSTEAEYR +ANT +EL W+ SLL E+ I T P+I CDN+ L NPV H+
Sbjct: 1310 RSVARSSTEAEYRAVANTASELRWLCSLLLEMGISQTTVPVIYCDNIGATYLCANPVFHS 1369
Query: 785 RTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNVTD 840
R KH+ LD FVR + + +L + HV + DQ D LTK L RF +L +K+ V +
Sbjct: 1370 RMKHVALDYHFVRGYIQSGALRVSHVSTKDQLADALTKPLPRPRFTELNSKIGVQE 1425
>gb|AAC02664.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 707 bits (1826), Expect = 0.0
Identities = 412/899 (45%), Positives = 533/899 (58%), Gaps = 71/899 (7%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
YTRYTW++PL+KKS F+ F ++E + K+ + +D GGEF + + + GI H
Sbjct: 554 YTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGTLYSDNGGEFIAMRSFLASHGISH 613
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
T PHT NGI ERKHRHIVE GL LL+ A + EYW +AF T VYLINR+ T VL N
Sbjct: 614 MTTPPHTPELNGISERKHRHIVETGLTLLSTASMPKEYWSYAFATAVYLINRMLTPVLGN 673
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESP+ KLF + LRIFGC C+P+LRPY AHKL S CV LGYS + +Y CLD
Sbjct: 674 ESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHKLDNRSVPCVLLGYSLSQSAYLCLDR 733
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCA---SSPQLSSVGSVHSSIPLFSFP--------- 230
TGR+Y S+ V F E FP+ S P LS + S+PL + P
Sbjct: 734 ATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPLSQ-DTRPVSVPLLARPLTTAPPSSP 792
Query: 231 ----------QLTTLSSPAPHSPNTTLPESS---SPA---------------SQSSTPIL 262
Q LS PAP P+ +L +S SP+ + SS P+
Sbjct: 793 SCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSETGPTGSSPPLS 852
Query: 263 PEPTSPVPQS----SPAPISPALSSPAPQSS------STDSSSSVSPIVNPHN---THSM 309
P+P P PQS SP SP +SP PQ S + SS S SP NP+ H+M
Sbjct: 853 PQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLTPTLTSSPSPSPPPNPNPPPIQHTM 912
Query: 310 VTRAKDGIVKPRLVPTLLLSQAEPTSAK--------LALKDPKWYSAMQDEFNALHSNQT 361
TR+K+ IVKP P +PT K AL DP W AM DE NA N T
Sbjct: 913 RTRSKNNIVKPN--PKFANLATKPTPLKPIIPKTVVEALLDPNWRQAMCDEINAQTRNGT 970
Query: 362 WTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPI 421
+ LVP ++ +GCKWVF +K ++G ++RYKARLVAKG+ Q G D+KETFSPV+K
Sbjct: 971 FDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVAKGFHQQYGHDFKETFSPVIKST 1030
Query: 422 TVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRS-LVCRLNKALYG 480
TVR +L +AVSKGW I Q+DVNNAFL G L +EVY+ QPPGF KD + VCRL KALYG
Sbjct: 1031 TVRSVLHIAVSKGWSIRQIDVNNAFLQGTLSDEVYVTQPPGFVDKDNAHHVCRLYKALYG 1090
Query: 481 LKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQ 540
LKQAPRAW++ L++ LL GF S + SLFTL +YVLVYVDD++ITG+ ++
Sbjct: 1091 LKQAPRAWYQELRSYLLTQGFVNSVADTSLFTLRHERTILYVLVYVDDMLITGSDTNIIT 1150
Query: 541 HLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTP 600
IA L A FSLK LG + YFLG++ K L L Q +Y+ DLLE+ +M A + TP
Sbjct: 1151 RFIANLAARFSLKDLGEMSYFLGIEATRT-SKGLHLMQKRYVLDLLEKTNMLAAHPVLTP 1209
Query: 601 MISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAV 660
M KL+ + PS YR+++G+LQY+ TRP+I+Y+VN++ Q++ P + HW A
Sbjct: 1210 MSPTPKLSLTSGKPLDKPSEYRAVLGSLQYLLFTRPDIAYAVNRLSQYMHCPTDLHWQAA 1269
Query: 661 KRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSW 720
KRILRYL GT G+ ++ PL+L A+ DADWA D D++ ST+ +++LG N +SW
Sbjct: 1270 KRILRYLAGTPSHGIFIR---ADTPLTLHAYSDADWAGDIDNYNSTNAYILYLGSNPISW 1326
Query: 721 CSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHN 779
SKKQ VARSSTEAEYR +AN +E+ WV SLL EL I P++ CDN+ L+ N
Sbjct: 1327 SSKKQKGVARSSTEAEYRAVANATSEIRWVCSLLTELGITLSSPPVVYCDNVGATYLSAN 1386
Query: 780 PVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
PV H+R KH+ LD FVRE V +L + HV + DQ D LTK L F L +K+ V
Sbjct: 1387 PVFHSRMKHIALDFHFVRESVQAGALRVTHVSTKDQLADALTKPLPRQPFTTLISKIGV 1445
>gb|AAC02666.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 704 bits (1817), Expect = 0.0
Identities = 411/899 (45%), Positives = 532/899 (58%), Gaps = 71/899 (7%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
YTRYTW++PL+KKS F+ F ++E + K+ + +D GGEF + + + GI H
Sbjct: 554 YTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGTLYSDNGGEFIAMRSFLASHGISH 613
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
T PHT NGI ERKHRHIVE GL LL+ A + EYW +AF T VYLINR+ T VL N
Sbjct: 614 MTTPPHTPELNGISERKHRHIVETGLTLLSTASMPKEYWSYAFATAVYLINRMLTPVLGN 673
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESP+ KLF + LRIFGC C+P+LRPY AHKL S CV LGYS + +Y CLD
Sbjct: 674 ESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHKLDNRSVPCVLLGYSLSQSAYLCLDR 733
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCA---SSPQLSSVGSVHSSIPLFSFP--------- 230
TGR+Y S+ V F E FP+ S P LS + S+PL + P
Sbjct: 734 ATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPLSQ-DTRPVSVPLLARPLTTAPPSSP 792
Query: 231 ----------QLTTLSSPAPHSPNTTLPESS---SPA---------------SQSSTPIL 262
Q LS PAP P+ +L +S SP+ + SS P+
Sbjct: 793 SCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSETGPTGSSPPLS 852
Query: 263 PEPTSPVPQS----SPAPISPALSSPAPQSS------STDSSSSVSPIVNPHN---THSM 309
P+P P PQS SP SP +SP PQ S + SS S SP NP+ H+M
Sbjct: 853 PQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLTPTLTSSPSPSPPPNPNPPPIQHTM 912
Query: 310 VTRAKDGIVKPRLVPTLLLSQAEPTSAK--------LALKDPKWYSAMQDEFNALHSNQT 361
TR+K+ IVKP P +PT K AL DP W AM DE NA N T
Sbjct: 913 RTRSKNNIVKPN--PKFANLATKPTPLKPIIPKTVVEALLDPNWRQAMCDEINAQTRNGT 970
Query: 362 WTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPI 421
+ LVP ++ +GCKWVF +K ++G ++RYKARLVAKG+ Q G D+KETFSPV+K
Sbjct: 971 FDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVAKGFHQQYGHDFKETFSPVIKST 1030
Query: 422 TVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRS-LVCRLNKALYG 480
TVR +L +AVSKGW I Q+DVNNAFL G L +EVY+ QPPGF KD + VCRL KALYG
Sbjct: 1031 TVRSVLHIAVSKGWSIRQIDVNNAFLQGTLSDEVYVTQPPGFVDKDNAHHVCRLYKALYG 1090
Query: 481 LKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQ 540
LKQAPRAW++ L++ LL GF S + SLFTL +YVLVYVDD++ITG+ ++
Sbjct: 1091 LKQAPRAWYQELRSYLLTQGFVNSVADTSLFTLRHERTILYVLVYVDDMLITGSDTNIIT 1150
Query: 541 HLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTP 600
IA L A FSLK LG + YFLG++ K L L Q +Y+ DLLE+ +M A + TP
Sbjct: 1151 RFIANLAARFSLKDLGEMSYFLGIEATRT-SKGLHLMQKRYVLDLLEKTNMLAAHPVLTP 1209
Query: 601 MISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAV 660
M KL+ + PS YR+++G+LQY+ TRP+I+Y+VN++ Q++ P + HW A
Sbjct: 1210 MSPTPKLSLTSGKPLDKPSEYRAVLGSLQYLLFTRPDIAYAVNRLSQYMHCPTDLHWQAA 1269
Query: 661 KRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSW 720
KRILRYL GT G+ ++ PL+L A+ DADWA D D++ ST+ +++LG N +SW
Sbjct: 1270 KRILRYLAGTPSHGIFIR---ADTPLTLHAYSDADWAGDIDNYNSTNAYILYLGSNPISW 1326
Query: 721 CSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHN 779
SKKQ VARSSTEAEYR +AN +E+ WV SLL EL I P++ CDN+ L+ N
Sbjct: 1327 SSKKQKGVARSSTEAEYRAVANATSEIRWVCSLLTELGITLSSPPVVYCDNVGATYLSAN 1386
Query: 780 PVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
PV +R KH+ LD FVRE V +L + HV + DQ D LTK L F L +K+ V
Sbjct: 1387 PVFDSRMKHIALDFHFVRESVQAGALRVTHVSTKDQLADALTKPLPRQPFTTLISKIGV 1445
>gb|AAC02669.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 703 bits (1815), Expect = 0.0
Identities = 411/899 (45%), Positives = 532/899 (58%), Gaps = 71/899 (7%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
YTRYTW++PL+KKS F+ F ++E + K+ + +D GGEF + + + GI H
Sbjct: 554 YTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGTLYSDNGGEFIAMRSFLASHGISH 613
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
T PHT NGI ERKHRHIVE GL LL+ A + EYW +AF T VYLINR+ T VL N
Sbjct: 614 MTTPPHTPELNGISERKHRHIVETGLTLLSTASMPKEYWSYAFATAVYLINRMLTPVLGN 673
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESP+ KLF + LRIFGC C+P+LRPY AHKL S CV LGYS + +Y CLD
Sbjct: 674 ESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHKLDNRSVPCVLLGYSLSQSAYLCLDR 733
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCA---SSPQLSSVGSVHSSIPLFSFP--------- 230
TGR+Y S+ V F E FP+ S P LS + S+PL + P
Sbjct: 734 ATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPLSQ-DTRPVSVPLLARPLTTAPPSSP 792
Query: 231 ----------QLTTLSSPAPHSPNTTLPESS---SPA---------------SQSSTPIL 262
Q LS PAP P+ +L +S SP+ + SS P+
Sbjct: 793 SCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSETGPTGSSPPLS 852
Query: 263 PEPTSPVPQS----SPAPISPALSSPAPQSS------STDSSSSVSPIVNPHN---THSM 309
P+P P PQS SP SP +SP PQ S + SS S SP NP+ H+M
Sbjct: 853 PQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLTPTLTSSPSPSPPPNPNPPPIQHTM 912
Query: 310 VTRAKDGIVKPRLVPTLLLSQAEPTSAK--------LALKDPKWYSAMQDEFNALHSNQT 361
TR+K+ IVKP P +PT K AL DP W AM DE NA N T
Sbjct: 913 RTRSKNNIVKPN--PKFANLATKPTPLKPIIPKTVVEALLDPNWRQAMCDEINAQTRNGT 970
Query: 362 WTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPI 421
+ LVP ++ +GCKWVF +K ++G ++RYKARLVAKG+ Q G D+KETFSPV+K
Sbjct: 971 FDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVAKGFHQQYGHDFKETFSPVIKLT 1030
Query: 422 TVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRS-LVCRLNKALYG 480
TVR +L +AVSKGW I Q+DVNNAFL G L +EVY+ QPPGF KD + VCRL KALYG
Sbjct: 1031 TVRSVLHIAVSKGWSIRQIDVNNAFLQGTLSDEVYVTQPPGFVDKDNAHHVCRLYKALYG 1090
Query: 481 LKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQ 540
LKQAPRAW++ L++ LL GF S + SLFTL +YVLVYVDD++ITG+ ++
Sbjct: 1091 LKQAPRAWYQELRSYLLTQGFVNSVADTSLFTLRHERTILYVLVYVDDMLITGSDTNIIT 1150
Query: 541 HLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTP 600
IA L A FSLK LG + YFLG++ K L L Q +Y+ DLLE+ +M A + TP
Sbjct: 1151 RFIANLAARFSLKDLGEMSYFLGIEATRT-SKGLHLMQKRYVLDLLEKTNMLAAHPVLTP 1209
Query: 601 MISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAV 660
M KL+ + PS YR+++G+LQY+ TRP+I+Y+VN++ Q++ P + HW A
Sbjct: 1210 MSPTPKLSLTSGKPLDKPSEYRAVLGSLQYLLFTRPDIAYAVNRLSQYMHCPTDLHWQAA 1269
Query: 661 KRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSW 720
KRILRYL GT G+ ++ PL+L A+ DADWA D D++ ST+ +++LG N +SW
Sbjct: 1270 KRILRYLAGTPSHGIFIR---ADTPLTLHAYSDADWAGDIDNYNSTNAYILYLGSNPISW 1326
Query: 721 CSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHN 779
SKKQ VARSSTEAEYR +AN +E+ WV SLL EL I P++ CDN+ L+ N
Sbjct: 1327 SSKKQKGVARSSTEAEYRAVANATSEIRWVCSLLTELGITLSSPPVVYCDNVGATYLSAN 1386
Query: 780 PVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
PV +R KH+ LD FVRE V +L + HV + DQ D LTK L F L +K+ V
Sbjct: 1387 PVFDSRMKHIALDFHFVRESVQAGALRVTHVSTKDQLADALTKPLPRQPFTTLISKIGV 1445
>gb|AAF99727.1| F17L21.7 [Arabidopsis thaliana]
Length = 1534
Score = 701 bits (1809), Expect = 0.0
Identities = 399/889 (44%), Positives = 533/889 (59%), Gaps = 57/889 (6%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
YTRYTW++PLK+KS TFV F ++E + K++ + +D GGEF L + GI H
Sbjct: 645 YTRYTWLYPLKQKSQVRETFVAFKALVENRFQTKIRTLYSDNGGEFIALRQFLLTHGISH 704
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NGI ERKHRHI+E GL LL A I YW +AF T VYLINRLP+ VL+N
Sbjct: 705 LTSLPHTPEHNGIAERKHRHILETGLTLLTQASIPTSYWTYAFGTAVYLINRLPSSVLNN 764
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESP++KLF+ S + LR+FGC+C+P+LRPY HKL S+ CVFLGYS T +Y CLD
Sbjct: 765 ESPYSKLFKTSPNYLKLRVFGCSCFPWLRPYTNHKLERRSQPCVFLGYSLTQSAYLCLDR 824
Query: 183 PTGRLYISKDVLFNEFRFPYK--ELFGCASSPQLSSVGSVH----------SSIPLFSFP 230
+GR+Y S+ V F E +FP+ + ++S + S H SS PL P
Sbjct: 825 SSGRVYTSRHVQFVEDQFPFSISDTHSVSNSSPEEASPSCHQPPSRIPIQSSSPPLVQAP 884
Query: 231 Q-LTTLSSPAPHSPNTTLPESSS-----------------------PASQSSTPIL--PE 264
L LSS + PN SSS P S SS
Sbjct: 885 SSLPPLSSDSHRRPNAETSSSSSSTNNDVVVSKDNTQVDNRNNFIGPTSSSSAQSQNNSN 944
Query: 265 PTSPVP-QSSPAPI-SPALSSPAPQSSSTDSSSSVSPIVNPH-----NTHSMVTRAKDGI 317
P+S + Q+ P P SP + +P+SS + S+S+ S + NP N H M TRAK+ I
Sbjct: 945 PSSSIQTQNEPNPSPSPTPQNSSPESSPSSSTSATSTVPNPPPPPPTNNHPMRTRAKNHI 1004
Query: 318 VKPRLVPTLLLSQAE-----PTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRK 372
KP+ +LL + P + AL+D KW +AM +E NA N T+ LVP ++
Sbjct: 1005 TKPKTKLSLLAKTVQTRPQIPNTVNQALRDEKWRNAMGEEINAQIRNNTFELVPPKPNQN 1064
Query: 373 AIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVS 432
I KW+F +K +G+++RYKARLVA+G+ Q G Y ETFSPVVK +T+R++L LAVS
Sbjct: 1065 VISTKWIFTLKYLPNGTLDRYKARLVARGFRQQYGLHYSETFSPVVKSLTIRLVLQLAVS 1124
Query: 433 KGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLNKALYGLKQAPRAWFER 491
+ W I QLDVNNAFL G L +EVY+ QPPGF DR VCRL KALYGLKQAPRAW++
Sbjct: 1125 RSWTIKQLDVNNAFLQGTLTDEVYVTQPPGFIDPDRPHHVCRLKKALYGLKQAPRAWYQE 1184
Query: 492 LKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFS 551
L+ + GFT S + S+F + VY LVYVDDII+TG+S ++ I L+ FS
Sbjct: 1185 LRNFVCSLGFTNSLADTSVFVYINDIQIVYCLVYVDDIIVTGSSDALVMAFITALSRRFS 1244
Query: 552 LKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHG 611
LK L YFLG++ + L L Q KY+ DLL R M AK +STPM ++ KL+ +
Sbjct: 1245 LKDPTDLVYFLGIEATRT-SQGLHLMQHKYVYDLLSRMKMLDAKPVSTPMATHPKLSLYS 1303
Query: 612 ANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTV 671
+ +P YR+++G+LQY+ TRP+I+Y+VN++ QF+ +P + HW A KR+LRYL GT
Sbjct: 1304 GIALDEPGEYRTVIGSLQYLAFTRPDIAYAVNRLSQFMHRPTDIHWQAAKRVLRYLAGTA 1363
Query: 672 DFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARS 731
G+ L+ + PLSL AF DADWA D DD ST+ +V+LG ++W SKKQ VARS
Sbjct: 1364 THGILLRS---NSPLSLHAFSDADWAGDNDDFVSTNAYIVYLGSTPIAWSSKKQKGVARS 1420
Query: 732 STEAEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHNPVLHTRTKHME 790
STEAEYR +ANT +E+ WV SLL EL I K P+I CDN+ L+ NPV H+R KH+
Sbjct: 1421 STEAEYRAVANTTSEIRWVCSLLTELGITLPKMPVIYCDNVGATYLSANPVFHSRMKHLA 1480
Query: 791 LDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNVT 839
LD F+R+ V +L + H+ + DQ D LTK L F +K+ V+
Sbjct: 1481 LDYHFIRDNVSAGALRVSHISTHDQLADALTKPLPRQHFLQFSSKIGVS 1529
>gb|AAK51235.1| polyprotein [Arabidopsis thaliana]
Length = 1453
Score = 687 bits (1773), Expect = 0.0
Identities = 372/863 (43%), Positives = 526/863 (60%), Gaps = 33/863 (3%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKP--LTPYFTNLGI 61
Y+RY+W +PLK KSD + FV F ++E Q N K+KV Q+DGGGEF + + T+ GI
Sbjct: 544 YSRYSWFYPLKAKSDFFAVFVAFQNLVENQFNTKIKVFQSDGGGEFTSNLMKKHLTDCGI 603
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
HR +CP+T QNGI ERKHRH VE+GL+++ H+H L++W AF T +L N LP+ L
Sbjct: 604 QHRISCPYTPQQNGIAERKHRHFVELGLSMMFHSHTPLQFWVEAFFTASFLSNMLPSPSL 663
Query: 122 DNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCL 181
N SP L + + LR+FG ACYP LRP HK S +CVFLGY++ +K Y+CL
Sbjct: 664 GNVSPLEALLKQKPNYAMLRVFGTACYPCLRPLGEHKFEPRSLQCVFLGYNSQYKGYRCL 723
Query: 182 -DPTGRLYISKDVLFNEFRFPYKELF---------GCASSPQLSSVGSVHSSIPLFSFPQ 231
PTGR+YIS+ V+F+E FP+K+ + S+ Q S + S IP +
Sbjct: 724 YPPTGRVYISRHVIFDEETFPFKQKYQFLVPQYESSLLSAWQSSIPQADQSLIPQAEEGK 783
Query: 232 LTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPA-PQSSS 290
+ +L+ P NT ++ PA + + E S + +L+ Q+
Sbjct: 784 IESLAKPPSIQKNTIQDTTTQPAILTEGVLNEEEEE---DSFEETETESLNEETHTQNDE 840
Query: 291 TDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQ---AEPTSAKLALKDPKWYS 347
+ + P NTH M TR+K GI K LL S+ EP S AL P W +
Sbjct: 841 AEVTVEEEVQQEPENTHPMTTRSKAGIHKSNTRYALLTSKFSVEEPKSIDEALNHPGWNN 900
Query: 348 AMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEG 407
A+ DE +H TW+LV +GC+WVF+ K DGSV++ KARLVAKG+ Q EG
Sbjct: 901 AVNDEMRTIHMLHTWSLVQPTEDMNILGCRWVFKTKLKPDGSVDKLKARLVAKGFHQEEG 960
Query: 408 FDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKD 467
DY ETFSPVV+ T+R++L +A +KGW+I QLDV+NAFL+G+L+E VYM+QPPGF ++
Sbjct: 961 LDYLETFSPVVRTATIRLVLDVATAKGWNIKQLDVSNAFLHGELKEPVYMLQPPGFVDQE 1020
Query: 468 R-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYV 526
+ S VCRL KALYGLKQAPRAWF+ + LL FGF+ S +PSLFT + +G ++ +L+YV
Sbjct: 1021 KPSYVCRLTKALYGLKQAPRAWFDTISNYLLDFGFSCSKSDPSLFTYHKNGKTLVLLLYV 1080
Query: 527 DDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLL 586
DDI++TG+ +LQ L+ LN FS+K LG YFLGV+++ + L L Q+ Y KD+L
Sbjct: 1081 DDILLTGSDHNLLQELLMSLNKRFSMKDLGAPSYFLGVEIES-SPEGLFLHQTAYAKDIL 1139
Query: 587 ERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVC 646
+A+M+ + TP+ Q + ++ +P+ +RS+ G LQY+TITRP+I ++VN +C
Sbjct: 1140 HQAAMSNCNSMPTPL--PQHIENLNSDLFPEPTYFRSLAGKLQYLTITRPDIQFAVNFIC 1197
Query: 647 QFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRST 706
Q + P + +KRILRY+KGT+ GLH+K +Q LSL+A+ D+DWA + RST
Sbjct: 1198 QRMHSPTTADFGLLKRILRYVKGTIHLGLHIKK---NQNLSLVAYSDSDWAGCKETRRST 1254
Query: 707 SGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTP-L 765
+G LG NL+SW +K+Q V++SSTEAEYR L EL W+ LL+++ + P L
Sbjct: 1255 TGFCTLLGCNLISWSAKRQETVSKSSTEAEYRALTAVAQELTWLSFLLRDIGVTQTHPTL 1314
Query: 766 ILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALS 825
+ CDN+S V L+ NP LH R+KH + D ++RE+V + +H+ + Q DI TK L
Sbjct: 1315 VKCDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGLVETKHISATLQLADIFTKPLP 1374
Query: 826 PLRFEDLRTKLNVTDKFTLAQPP 848
F DLR KL V A+PP
Sbjct: 1375 RRAFIDLRIKLGV------AEPP 1391
>emb|CAB79576.1| putative protein [Arabidopsis thaliana] gi|3269282|emb|CAA19715.1|
putative protein [Arabidopsis thaliana]
gi|7444417|pir||T05745 hypothetical protein M4I22.20 -
Arabidopsis thaliana
Length = 1318
Score = 681 bits (1758), Expect = 0.0
Identities = 386/874 (44%), Positives = 522/874 (59%), Gaps = 49/874 (5%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
Y+R++W++PLK KSD Y F+ FHK++E QL++K+ V Q DGGGEF + + GI
Sbjct: 356 YSRFSWIYPLKLKSDFYNIFLAFHKLVENQLSQKISVFQCDGGGEFVSHKFLQHLQSHGI 415
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
+ +CPHT QNG+ ERKHRH+VE+GL++L +H+ ++W AF T +LIN LPT L
Sbjct: 416 QQQLSCPHTPQQNGLAERKHRHLVELGLSMLFQSHVPHKFWVEAFFTANFLINLLPTSAL 475
Query: 122 -DNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKC 180
++ SP+ KL+ D LR FG AC+P LR Y +K + S +CVFLGY+ +K Y+C
Sbjct: 476 KESISPYEKLYDKKPDYTSLRSFGSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYRC 535
Query: 181 L-DPTGRLYISKDVLFNEFRFP----YKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTL 235
L PTGRLYIS+ V+F+E +P YK L +P L++ S P +T
Sbjct: 536 LYPPTGRLYISRHVIFDESVYPFSHTYKHLHPQPRTPLLAAWLRSSDS------PAPSTS 589
Query: 236 SSPAPHSPNTTLPESSSPASQSSTPILPE--PTSPVPQSSPAPI--SPAL---------- 281
+SP+ SP T + P Q TP+LP P S V +S SP
Sbjct: 590 TSPSSRSPLFTSADFP-PLPQRKTPLLPTLVPISSVSHASNITTQQSPDFDSERTTDFDS 648
Query: 282 -----SSPAPQSSSTDSSSSVSPIVNPHNTHS------MVTRAKDGIVKPRLVPTLL--- 327
SS + Q+ S + VN H TH+ MVTRAK GI KP L
Sbjct: 649 ASIGDSSHSSQAGSDSEETIQQASVNVHQTHASTNVHPMVTRAKVGISKPNPRYVFLSHK 708
Query: 328 LSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENAD 387
+S EP + ALK P W AM +E QTW+LVP S +G KWVFR K +AD
Sbjct: 709 VSYPEPKTVTAALKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVFRTKLHAD 768
Query: 388 GSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFL 447
G++N+ KAR+VAKG+ Q EG DY ET+SPVV+ TVR++L LA + W I Q+DV NAFL
Sbjct: 769 GTLNKLKARIVAKGFLQEEGIDYLETYSPVVRTPTVRLVLHLATALNWDIKQMDVKNAFL 828
Query: 448 NGKLEEEVYMVQPPGF-EAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCC 506
+G L+E VYM QP GF + VC L+K++YGLKQ+PRAWF++ LL+FGF S
Sbjct: 829 HGDLKETVYMTQPAGFVDPSKPDHVCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCSKS 888
Query: 507 NPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQV 566
+PSLF + + +L+YVDD++ITGNS L L+A LN EF + +G L YFLG+QV
Sbjct: 889 DPSLFIYAHNNNLILLLLYVDDMVITGNSSQTLTSLLAALNKEFRMTDMGQLHYFLGIQV 948
Query: 567 QHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVG 626
Q L ++Q KY +DLL ASM + + TP+ H +DP+ +RSI G
Sbjct: 949 QR-QQNGLFMSQQKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEELFSDPTYFRSIAG 1007
Query: 627 ALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPL 686
LQY+T+TRP+I ++VN VCQ + QP + +KRILRY+KGT+ G+ S P
Sbjct: 1008 KLQYLTLTRPDIQFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITMGISY---SRDSPT 1064
Query: 687 SLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAE 746
L A+ D+DW RS G F+G NLVSW SKK V+RSSTEAEY++L++ +E
Sbjct: 1065 LLQAYSDSDWGNCKQTRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLSDAASE 1124
Query: 747 LLWVQSLLQELHIP-HKTPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSL 805
+LW+ +LL+EL IP TP + CDN+S V LT NP H RTKH ++D FVRE+V K+L
Sbjct: 1125 ILWLSTLLRELRIPLPDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHFVRERVALKAL 1184
Query: 806 IIQHVPSLDQRDDILTKALSPLRFEDLRTKLNVT 839
+++H+P +Q DI TK+L F LR KL VT
Sbjct: 1185 VVKHIPGSEQIADIFTKSLPYEAFIHLRGKLGVT 1218
>gb|AAF02855.1| Similar to retrotransposon proteins [Arabidopsis thaliana]
gi|25301689|pir||C96578 hypothetical protein T18A20.5
[imported] - Arabidopsis thaliana
Length = 1522
Score = 679 bits (1753), Expect = 0.0
Identities = 384/885 (43%), Positives = 532/885 (59%), Gaps = 51/885 (5%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
Y+R+TW +PLK KSD ++TFV F K++E QL K+K+ Q DGGGEF + + GI
Sbjct: 545 YSRFTWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFLKHLQDHGI 604
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
+CP+T QNG+ ERKHRHIVE+GL+++ + + L+YW +F T ++IN LPT L
Sbjct: 605 QQNMSCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVINLLPTSSL 664
Query: 122 DN-ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKC 180
DN ESP+ KL+ + + LR+FGCACYP LR Y + K S +CVFLGY+ +K Y+C
Sbjct: 665 DNNESPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRSLKCVFLGYNEKYKGYRC 724
Query: 181 L-DPTGRLYISKDVLFNEFRFPYKELFGCA----SSPQLSS-VGSVH---------SSIP 225
L PTGR+YIS+ V+F+E P++ ++ +P L + S H S P
Sbjct: 725 LYPPTGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVTPTQPDQSRYP 784
Query: 226 LFSFPQLTTL---SSPAPHSPNTTLPESSSPASQSSTPIL-----PEPTSPV-------- 269
+ S PQ T ++PA + T P +S SQ + I PE T+ +
Sbjct: 785 VSSIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISVVSGSPERTTGLDSASIGDS 844
Query: 270 ---PQSSPAPISPALSSPA--PQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVP 324
P + + SPA SSPA PQ S + + N H+MVTR K+GI KP
Sbjct: 845 YHSPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTNEHAMVTRGKEGISKPNKRY 904
Query: 325 TLL---LSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFR 381
LL +S EP + ALK P W +AMQ+E +TWTLVP + +G WVFR
Sbjct: 905 VLLTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVLGSMWVFR 964
Query: 382 IKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLD 441
K +ADGS+++ KARLVAKG+ Q EG DY ET+SPVV+ TVR+IL +A W + Q+D
Sbjct: 965 TKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVATVLKWELKQMD 1024
Query: 442 VNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFG 500
V NAFL+G L E VYM QP GF K + VC L+K+LYGLKQ+PRAWF+R LL+FG
Sbjct: 1025 VKNAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYGLKQSPRAWFDRFSNFLLEFG 1084
Query: 501 FTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDY 560
F S +PSLF S+ + +L+YVDD++ITGN+ L HL+A LN EF +K +G + Y
Sbjct: 1085 FICSLFDPSLFVYSSNNDVILLLLYVDDMVITGNNSQSLTHLLAALNKEFRMKDMGQVHY 1144
Query: 561 FLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSL 620
FLG+Q+Q +D L ++Q KY +DLL ASMA + TP+ + +DP+
Sbjct: 1145 FLGIQIQ-TYDGGLFMSQQKYAEDLLITASMANCSPMPTPLPLQLDRVSNQDEVFSDPTY 1203
Query: 621 YRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPC 680
+RS+ G LQY+T+TRP+I ++VN VCQ + QP + +KRILRY+KGTV G+
Sbjct: 1204 FRSLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLKRILRYIKGTVSMGIQYNSN 1263
Query: 681 S------LHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTE 734
S L A+ D+D+A + RS G F+G N++SW SKKQ V+RSSTE
Sbjct: 1264 SSSVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFMGQNIISWSSKKQPTVSRSSTE 1323
Query: 735 AEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHNPVLHTRTKHMELDI 793
AEYR+L+ T +E+ W+ S+L+E+ + TP + CDN+S V LT NP H RTKH ++D
Sbjct: 1324 AEYRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLSAVYLTANPAFHARTKHFDVDH 1383
Query: 794 FFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
++RE+V K+L+++H+P Q DI TK+L F LR KL V
Sbjct: 1384 HYIRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRLRFKLGV 1428
>gb|AAD21687.1| Strong similarity to gi|3600044 T12H20.12 protease homolog from
Arabidopsis thaliana BAC gb|AF080119 and is a member of
the reverse transcriptase family PF|00078
gi|25301706|pir||C86438 hypothetical protein F28K20.17 -
Arabidopsis thaliana
Length = 1415
Score = 675 bits (1741), Expect = 0.0
Identities = 358/852 (42%), Positives = 520/852 (61%), Gaps = 46/852 (5%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
Y+RY+W +PL KS+ + F++F K++E QLN K+KV Q+DGGGEF L + + GI
Sbjct: 541 YSRYSWFYPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGI 600
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
HR +CP+T QNG+ ERKHRH+VE+GL++L H+H ++W +F T Y+INRLP+ VL
Sbjct: 601 HHRISCPYTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVL 660
Query: 122 DNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCL 181
N SP+ LF D LR+FG ACYP LRP +K S +CVFLGY++ +K Y+C
Sbjct: 661 KNLSPYEALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCF 720
Query: 182 -DPTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAP 240
PTG++YIS++V+FNE P+KE + S +P +S P L
Sbjct: 721 YPPTGKVYISRNVIFNESELPFKEKY--------------QSLVPQYSTPLLQAWQ---- 762
Query: 241 HSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPI 300
H+ + + ++P S PI + + + ++ L+ P P S++ S V+P+
Sbjct: 763 HNKISEISVPAAPVQLFSKPI------DLNTYAGSQVTEQLTDPEPTSNNEGSDEEVNPV 816
Query: 301 VNPH--------NTHSMVTRAKDGIVKPRLVPTLLLSQ---AEPTSAKLALKDPKWYSAM 349
N+H+M TR+K GI KP L+ S+ AEP + A+K P W A+
Sbjct: 817 AEEIAANQEQVINSHAMTTRSKAGIQKPNTRYALITSRMNTAEPKTLASAMKHPGWNEAV 876
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFD 409
+E N +H TW+LVP + KWVF+ K + DGS+++ KARLVAKG+ Q EG D
Sbjct: 877 HEEINRVHMLHTWSLVPPTDDMNILSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVD 936
Query: 410 YKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGF-EAKDR 468
Y ETFSPVV+ T+R++L ++ SKGW I QLDV+NAFL+G+L+E V+M QP GF + +
Sbjct: 937 YLETFSPVVRTATIRLVLDVSTSKGWPIKQLDVSNAFLHGELQEPVFMYQPSGFIDPQKP 996
Query: 469 SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDD 528
+ VCRL KA+YGLKQAPRAWF+ LL +GF S +PSLF + G +Y+L+YVDD
Sbjct: 997 THVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGFVCSKSDPSLFVCHQDGKILYLLLYVDD 1056
Query: 529 IIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLER 588
I++TG+ +L+ L+ L FS+K LG YFLG+Q++ + L L Q+ Y D+L++
Sbjct: 1057 ILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYFLGIQIED-YANGLFLHQTAYATDILQQ 1115
Query: 589 ASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQF 648
A M+ + TP+ Q+L + A+P+ +RS+ G LQY+TITRP+I ++VN +CQ
Sbjct: 1116 AGMSDCNPMPTPL--PQQLDNLNSELFAEPTYFRSLAGKLQYLTITRPDIQFAVNFICQR 1173
Query: 649 LSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSG 708
+ P + +KRILRY+KGT+ GL P + L+L A+ D+D A + RST+G
Sbjct: 1174 MHSPTTSDFGLLKRILRYIKGTIGMGL---PIKRNSTLTLSAYSDSDHAGCKNTRRSTTG 1230
Query: 709 SLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPL-IL 767
+ LG NL+SW +K+Q V+ SSTEAEYR L E+ W+ LL++L IP P +
Sbjct: 1231 FCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYAAREITWISFLLRDLGIPQYLPTQVY 1290
Query: 768 CDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPL 827
CDN+S V L+ NP LH R+KH + D ++RE+V + QH+ + Q D+ TK+L
Sbjct: 1291 CDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGLIETQHISATFQLADVFTKSLPRR 1350
Query: 828 RFEDLRTKLNVT 839
F DLR+KL V+
Sbjct: 1351 AFVDLRSKLGVS 1362
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,488,402,541
Number of Sequences: 2540612
Number of extensions: 65901795
Number of successful extensions: 480212
Number of sequences better than 10.0: 6145
Number of HSP's better than 10.0 without gapping: 2337
Number of HSP's successfully gapped in prelim test: 4248
Number of HSP's that attempted gapping in prelim test: 401555
Number of HSP's gapped (non-prelim): 36037
length of query: 856
length of database: 863,360,394
effective HSP length: 137
effective length of query: 719
effective length of database: 515,296,550
effective search space: 370498219450
effective search space used: 370498219450
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0322.2


