

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
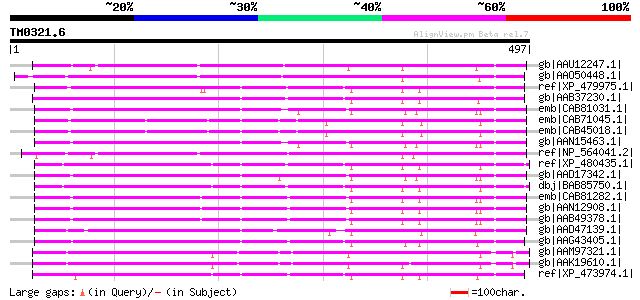
Query= TM0321.6
(497 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAU12247.1| homeodomain protein HOX3 [Gossypium hirsutum] 326 1e-87
gb|AAO50448.1| putative homeobox protein [Arabidopsis thaliana] ... 290 9e-77
ref|XP_479975.1| putative OCL5 protein [Oryza sativa (japonica c... 285 2e-75
gb|AAB37230.1| homeobox protein gi|2147484|pir||S71477 homeotic ... 281 3e-74
emb|CAB81031.1| putative homeotic protein [Arabidopsis thaliana]... 281 4e-74
emb|CAB71045.1| homeobox protein [Arabidopsis thaliana] gi|20465... 281 4e-74
emb|CAB45018.1| homeodomain GLABRA2 like 1 protein [Arabidopsis ... 281 4e-74
gb|AAN15463.1| Unknown protein [Arabidopsis thaliana] gi|1427606... 281 4e-74
ref|NP_564041.2| homeobox-leucine zipper family protein / lipid-... 280 1e-73
ref|XP_480435.1| roc1(homeobox protein) [Oryza sativa (japonica ... 278 4e-73
gb|AAD17342.1| contains similarity to homeobox domains (Pfam: PF... 278 4e-73
dbj|BAB85750.1| Roc1 [Oryza sativa] 278 4e-73
emb|CAB81282.1| L1 specific homeobox gene ATML1/ovule-specific h... 272 2e-71
gb|AAN12908.1| putative L1-specific homeobox gene ATML1/ovule-sp... 272 2e-71
gb|AAB49378.1| A20 271 3e-71
gb|AAD47139.1| Anthocyaninless2 [Arabidopsis thaliana] 271 3e-71
gb|AAG43405.1| homeobox 1 [Picea abies] 271 3e-71
gb|AAM97321.1| homeodomain protein GhHOX1 [Gossypium hirsutum] 270 6e-71
gb|AAK19610.1| BNLGHi8377 [Gossypium hirsutum] 270 6e-71
ref|XP_473974.1| OSJNBb0060E08.16 [Oryza sativa (japonica cultiv... 270 8e-71
>gb|AAU12247.1| homeodomain protein HOX3 [Gossypium hirsutum]
Length = 713
Score = 326 bits (835), Expect = 1e-87
Identities = 195/498 (39%), Positives = 297/498 (59%), Gaps = 31/498 (6%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITN----SE 78
++KS M +A AM+ELL LL T+EPLW+KS+ND L E Y + FP+ N S
Sbjct: 219 IDKSLMSDIAANAMEELLRLLQTNEPLWIKSTNDGKD-ALNLESYERIFPKPNNTHFKSP 277
Query: 79 NLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGA 138
N+ V E+S+DS +V+ +G+ LV +F+DS+KW ++ PTIVS AKTI+V G + H +
Sbjct: 278 NIRV--EASRDSGVVIMNGLALVDMFMDSNKWLELFPTIVSIAKTIEVISPGMLGTHRCS 335
Query: 139 LRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGC 198
L+LM+ E+ VLSPLVP+RE++ LRYCQQ+E G W IV+VS D AS+ + + PSGC
Sbjct: 336 LQLMYEELQVLSPLVPTREFYTLRYCQQIEQGLWAIVNVSYD-LPQFASQCRSHRLPSGC 394
Query: 199 RIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSE 258
I+++PNG VTW+E+VE+ + L RD+V + A+GA+RWL LQR E AC
Sbjct: 395 LIQDMPNGYSKVTWLERVEIEDKTPIHRLYRDLVHSGSAFGAERWLTTLQRMCEWFACLR 454
Query: 259 FSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLI 318
S T ST L G + +GR+S++ L+ RMV +F + T S +++
Sbjct: 455 VSST-STRDLGGVIPSPEGRRSMMKLAQRMVNNFCTSVGTSNSHRSTTLSGSNEVGVRVT 513
Query: 319 AHRNT---ILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS- 374
H+++ +GIV+ A ++ WLPV+P+N+F F KDE + +WD L+ G +++ AHI+
Sbjct: 514 VHKSSDPGQPNGIVLSAATTFWLPVSPQNVFNFFKDERTRPQWDVLSNGNAVQEVAHIAN 573
Query: 375 -INESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ N +S+L+ + +M+I QES ID G VVY P+D + ++G D S +
Sbjct: 574 GSHPGNCISVLRAFNTSHNNMLILQESCIDSSGSLVVYCPVDLPAINVAMSGEDPSYIPL 633
Query: 434 LPSGFTISRDGR----------------EGNEGSLLTMCFQVLVDEEGSSSQVRKKTLES 477
LPSGFTI+ DG + GSL+T+ FQ+LV S+++ ++
Sbjct: 634 LPSGFTITPDGHLEQGDGASTSSSTGHGRSSGGSLITVAFQILVSSL-PSAKLNLDSVTI 692
Query: 478 IDSSCMLMLQRIKDALHC 495
+++ +Q+IK AL+C
Sbjct: 693 VNNLIANTVQQIKAALNC 710
>gb|AAO50448.1| putative homeobox protein [Arabidopsis thaliana]
gi|28393178|gb|AAO42020.1| putative homeobox protein
[Arabidopsis thaliana] gi|15219456|ref|NP_177479.1|
homeobox-leucine zipper family protein / lipid-binding
START domain-containing protein [Arabidopsis thaliana]
gi|11120798|gb|AAG30978.1| homeobox protein, putative
[Arabidopsis thaliana] gi|25386239|pir||B96760 probable
homeobox protein T9L24.43 [imported] - Arabidopsis
thaliana
Length = 722
Score = 290 bits (741), Expect = 9e-77
Identities = 177/508 (34%), Positives = 287/508 (55%), Gaps = 28/508 (5%)
Query: 5 NDNVVNNPTRESQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVR 64
N+N+ + P + ++K M +A AM+ELL LL T+EPLW ++ D +L
Sbjct: 215 NNNLQSQPNLAISD----MDKPIMTGIALTAMEELLRLLQTNEPLWTRT--DGCRDILNL 268
Query: 65 EIYAKKFPRITN-SENLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKT 123
Y FPR +N +N E+S+ S +V + M LV +F+D KW ++ P+I++ +KT
Sbjct: 269 GSYENVFPRSSNRGKNQNFRVEASRSSGIVFMNAMALVDMFMDCVKWTELFPSIIAASKT 328
Query: 124 IQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFG 183
+ V G H GAL L++ E+ VLSPLV +RE+ LRYCQQ E G W++V+VS D
Sbjct: 329 LAVISSGMGGTHEGALHLLYEEMEVLSPLVATREFCELRYCQQTEQGSWIVVNVSYD-LP 387
Query: 184 NCASRSVAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRW 243
S S + +FPSGC I+++PNG VTWVE +E + + L R+I+ IA+GA RW
Sbjct: 388 QFVSHSQSYRFPSGCLIQDMPNGYSKVTWVEHIETEEKELVHELYREIIHRGIAFGADRW 447
Query: 244 LLELQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQN 303
+ LQR E+ A +S+ L G + +G++S++ L+ RM+ ++ ++
Sbjct: 448 VTTLQRMCERFASLSVP-ASSSRDLGGVILSPEGKRSMMRLAQRMISNYCLSVSRSNNTR 506
Query: 304 LLDPSNQLTTSIKLIAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAG 363
S I++ AH++ +G V+ A ++ WLP +P+N+F FLKDE + +WD L+
Sbjct: 507 STVVSELNEVGIRVTAHKSPEPNGTVLCAATTFWLPNSPQNVFNFLKDERTRPQWDVLSN 566
Query: 364 GRPMRQKAHIS--INESNHVSILQTSSII-GEHMVIFQESSIDDLGGYVVYAPIDKLDAY 420
G +++ AHIS + N +S+L+ S+ +M+I QESS D G +VVY+P+D
Sbjct: 567 GNAVQEVAHISNGSHPGNCISVLRGSNATHSNNMLILQESSTDSSGAFVVYSPVDLAALN 626
Query: 421 MIVNGCDSSKKLILPSGFTISRDGREGN---------------EGSLLTMCFQVLVDEEG 465
+ ++G D S +L SGFTIS DG N GSL+T+ FQ++V
Sbjct: 627 IAMSGEDPSYIPLLSSGFTISPDGNGSNSEQGGASTSSGRASASGSLITVGFQIMVSNL- 685
Query: 466 SSSQVRKKTLESIDSSCMLMLQRIKDAL 493
++++ +++E++++ + +IK AL
Sbjct: 686 PTAKLNMESVETVNNLIGTTVHQIKTAL 713
>ref|XP_479975.1| putative OCL5 protein [Oryza sativa (japonica cultivar-group)]
gi|38636822|dbj|BAD03062.1| putative OCL5 protein [Oryza
sativa (japonica cultivar-group)]
gi|46390804|dbj|BAD16310.1| putative OCL5 protein [Oryza
sativa (japonica cultivar-group)]
Length = 828
Score = 285 bits (730), Expect = 2e-75
Identities = 173/486 (35%), Positives = 280/486 (57%), Gaps = 23/486 (4%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + EPLW + G L E YA+ FPR ++ +
Sbjct: 338 DKPLVIELAVAAMEELVRMAQLGEPLWAPALG---GEALGEEEYARTFPRGLGPKSPELR 394
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+++ +V+ + + LV++ +D +W + +IVS+A T++V G NH+GAL+LM
Sbjct: 395 SEASRETAVVIMNHVSLVEMLMDVGQWTALFSSIVSRAATLEVLSTGVAGNHNGALQLMS 454
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSF-----GNC--ASRSVAKKFPS 196
E + SPLVP+RE FLRYC+Q G W +VDVSLD G C A+ ++ PS
Sbjct: 455 AEFQMPSPLVPTRETQFLRYCKQHPDGTWAVVDVSLDGLRAGAGGGCQPAAARGHRRRPS 514
Query: 197 GCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKAC 256
GC I+E+PNG VTWVE VE + L + +V + +A+GA+RW+ L+R E+ A
Sbjct: 515 GCLIQEMPNGYSKVTWVEHVEADDQ-MVHNLYKPVVNSGMAFGARRWVATLERQCERLAS 573
Query: 257 SEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIK 316
+ S AS+ G G + +GR+S++ L+ RMV SF + S ++
Sbjct: 574 AMASNVASS-GDAGVITTSEGRRSMLKLAERMVASFCGGVTASTTHQWTTLSGSGAEDVR 632
Query: 317 LIAHRNTILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKA 371
++ R ++ D GI++ A +S WLPV P +F FL+D+ ++EWD L+ G +++ A
Sbjct: 633 VMT-RKSVDDPGRPPGIILNAATSFWLPVPPSRVFDFLRDDSTRSEWDILSNGGVVQEMA 691
Query: 372 HIS--INESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCD 427
HI+ + N VS+L+ ++ +M+I QE D G YV+YAP+D + +++NG D
Sbjct: 692 HIANGRDHGNAVSLLRVNNANSNQSNMLILQECCTDATGSYVIYAPVDVVAMNVVLNGGD 751
Query: 428 SSKKLILPSGFTISRDGREGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLMLQ 487
+LPSGF I DG +G GSLLT+ FQ+LVD ++++ ++ +++S ++
Sbjct: 752 PDYVALLPSGFAILPDGPDGGGGSLLTVAFQILVDSV-PTAKLSLGSVATVNSLIACTVE 810
Query: 488 RIKDAL 493
RIK A+
Sbjct: 811 RIKAAI 816
>gb|AAB37230.1| homeobox protein gi|2147484|pir||S71477 homeotic protein,
ovule-specific - Phalaenopsis sp
Length = 768
Score = 281 bits (720), Expect = 3e-74
Identities = 175/494 (35%), Positives = 278/494 (55%), Gaps = 28/494 (5%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSN-DVVGFVLVREIYAKKFPRITNSENLY 81
V+K ++ +A AAM+EL+ + EPLW S D +L E Y + FPR +
Sbjct: 274 VDKPMVIELAVAAMEELIRMAQLGEPLWTSSPGLDGGNEILNEEEYVQNFPRGIGPKPFG 333
Query: 82 VCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRL 141
+ E+S+++ +V+ S + LV++ +D+++W+ + IVS+ T++V G N++GAL++
Sbjct: 334 LKSEASRETAVVIMSHVNLVEILMDANQWSTMFSGIVSRGMTLEVLSTGVAGNYNGALQV 393
Query: 142 MWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIR 201
M E V SPLVP+RE +F+RYC+Q G W +VDVSLDS + ++ PSGC I+
Sbjct: 394 MTAEFQVPSPLVPTRESYFVRYCKQHPDGTWAVVDVSLDSLRPSSLMMRCRRRPSGCLIQ 453
Query: 202 ELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSY 261
E+PNG V WVE EV + + + +V + IA+GAKRW+ L R E+ A S
Sbjct: 454 EMPNGYSKVIWVEHFEVDDR-SVHSIYKPLVNSGIAFGAKRWVSTLDRQCERLASVMASS 512
Query: 262 TASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHR 321
S G G + +GRKS++ L+ RMV SF ++ S ++++ R
Sbjct: 513 IPS--GEIGVITTSEGRKSMLKLAERMVLSFCGGVSASTTHQWTTLSGSGAEDVRVMT-R 569
Query: 322 NTILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS-- 374
++ D GIV+ A +S WLPV+PK +F FL+DE ++EWD L+ G +++ AHI+
Sbjct: 570 KSVDDPGRPPGIVLNAATSFWLPVSPKRVFDFLRDESSRSEWDILSNGGVVQEMAHIANG 629
Query: 375 INESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKL 432
+ N VS+L+ +S +M+I QES D G YV+YAP+D + +++NG D
Sbjct: 630 RDHGNCVSLLRVNSTNSNQSNMLILQESCTDPTGSYVIYAPVDVVAMNVVLNGGDPDYVA 689
Query: 433 ILPSGFTISRDGREG-------------NEGSLLTMCFQVLVDEEGSSSQVRKKTLESID 479
+LPSGF I DG G GSLLT+ FQ+LVD ++++ ++ +++
Sbjct: 690 LLPSGFAILPDGSNGVHGGGSGIGEVGSGGGSLLTVAFQILVDSI-PTAKLSLGSVATVN 748
Query: 480 SSCMLMLQRIKDAL 493
S ++RIK A+
Sbjct: 749 SLIACTVERIKAAV 762
>emb|CAB81031.1| putative homeotic protein [Arabidopsis thaliana]
gi|25386232|pir||E85061 probable homeotic protein
[imported] - Arabidopsis thaliana
Length = 738
Score = 281 bits (718), Expect = 4e-74
Identities = 174/503 (34%), Positives = 287/503 (56%), Gaps = 43/503 (8%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLW+ + N V +L E Y + FPR + L +
Sbjct: 242 DKPIIVELAVAAMEELVRMAQTGDPLWLSTDNSVE--ILNEEEYFRTFPRGIGPKPLGLR 299
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ S +V+ + + LV++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 300 SEASRQSAVVIMNHINLVEILMDVNQWSCVFSGIVSRALTLEVLSTGVAGNYNGALQVMT 359
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS ++ PSGC I+EL
Sbjct: 360 AEFQVPSPLVPTRENYFVRYCKQHSDGSWAVVDVSLDSLRPSTPILRTRRRPSGCLIQEL 419
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
PNG VTW+E +EV + + + +V++ +A+GAKRW+ L+R E+ A S S
Sbjct: 420 PNGYSKVTWIEHMEVDDR-SVHNMYKPLVQSGLAFGAKRWVATLERQCERLASSMAS--- 475
Query: 264 STEGLEGEVKLL---DGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAH 320
+ G++ ++ +GRKS++ L+ RMV SF + S + ++++
Sbjct: 476 ---NIPGDLSVITSPEGRKSMLKLAERMVMSFCSGVGASTAHAWTTMSTTGSDDVRVMTR 532
Query: 321 RNTILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
++ +D GIV+ A +S W+PV PK +F FL+DE + EWD L+ G +++ AHI+
Sbjct: 533 KS--MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRKEWDILSNGGMVQEMAHIA 590
Query: 375 INE--SNHVSILQTSS--IIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSK 430
N VS+L+ +S +M+I QES D G YV+YAP+D + ++++G D
Sbjct: 591 NGHEPGNCVSLLRVNSGNSSQSNMLILQESCTDASGSYVIYAPVDIVAMNVVLSGGDPDY 650
Query: 431 KLILPSGFTISRDGR----EGNE--------------GSLLTMCFQVLVDEEGSSSQVRK 472
+LPSGF I DG +GN+ GSLLT+ FQ+LVD ++++
Sbjct: 651 VALLPSGFAILPDGSVGGGDGNQHQEMVSTTSSGSCGGSLLTVAFQILVDSV-PTAKLSL 709
Query: 473 KTLESIDSSCMLMLQRIKDALHC 495
++ +++S ++RIK A+ C
Sbjct: 710 GSVATVNSLIKCTVERIKAAVSC 732
>emb|CAB71045.1| homeobox protein [Arabidopsis thaliana] gi|20465805|gb|AAM20391.1|
putative homeobox protein [Arabidopsis thaliana]
gi|15292865|gb|AAK92803.1| putative homeobox protein
[Arabidopsis thaliana] gi|15233048|ref|NP_191674.1|
homeobox-leucine zipper family protein / homeodomain
GLABRA2 like protein 1 (HD-GL2-1) [Arabidopsis thaliana]
gi|11346374|pir||T47907 homeobox protein - Arabidopsis
thaliana
Length = 808
Score = 281 bits (718), Expect = 4e-74
Identities = 177/501 (35%), Positives = 286/501 (56%), Gaps = 37/501 (7%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGF-VLVREIYAKKFPRITNSENLYV 82
++S+ L +A AAM EL+ + T EPLWV+SS+ GF VL +E Y F R +
Sbjct: 313 QRSRYLDLALAAMDELVKMAQTREPLWVRSSDS--GFEVLNQEEYDTSFSRCVGPKQDGF 370
Query: 83 CEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLM 142
E+SK++ V+ + + LV+ +DS++WA++ P++VS+ T ++ G + +GAL LM
Sbjct: 371 VSEASKEAGTVIINSLALVETLMDSERWAEMFPSMVSRTSTTEIISSG-MGGRNGALHLM 429
Query: 143 WNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIRE 202
E+ +LSPLVP R+ FLR+C+Q G W +VDVS+DS +S S ++ PSGC +++
Sbjct: 430 HAELQLLSPLVPVRQVSFLRFCKQHAEGVWAVVDVSIDSIREGSSSS-CRRLPSGCLVQD 488
Query: 203 LPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYT 262
+ NG VTW+E E ++ + L R ++R +A+GA RW+ LQR E S T
Sbjct: 489 MANGYSKVTWIEHTEYDEN-HIHRLYRPLLRCGLAFGAHRWMAALQRQCECLTIL-MSST 546
Query: 263 ASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQ-----NLLDPSNQLTTSIKL 317
ST + +GRKS++ L+ RM +F + +Q N+ + + +
Sbjct: 547 VSTSTNPSPINC-NGRKSMLKLAKRMTDNFCGGVCASSLQKWSKLNVGNVDEDVRIMTRK 605
Query: 318 IAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--I 375
+ GI++ A +S+W+PV+P+ LF FL +E ++EWD L+ G PM++ AHI+
Sbjct: 606 SVNNPGEPPGIILNAATSVWMPVSPRRLFDFLGNERLRSEWDILSNGGPMKEMAHIAKGH 665
Query: 376 NESNHVSILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ SN VS+L+ S+I M+I QE+SID G VVYAP+D ++NG DS+ +
Sbjct: 666 DRSNSVSLLRASAINANQSSMLILQETSIDAAGAVVVYAPVDIPAMQAVMNGGDSAYVAL 725
Query: 434 LPSGFTISRDGREGNE-------------------GSLLTMCFQVLVDEEGSSSQVRKKT 474
LPSGF I +G+ G + GSLLT+ FQ+LV+ ++++ ++
Sbjct: 726 LPSGFAILPNGQAGTQRCAAEERNSIGNGGCMEEGGSLLTVAFQILVNSL-PTAKLTVES 784
Query: 475 LESIDSSCMLMLQRIKDALHC 495
+E++++ +Q+IK ALHC
Sbjct: 785 VETVNNLISCTVQKIKAALHC 805
>emb|CAB45018.1| homeodomain GLABRA2 like 1 protein [Arabidopsis thaliana]
Length = 808
Score = 281 bits (718), Expect = 4e-74
Identities = 177/501 (35%), Positives = 286/501 (56%), Gaps = 37/501 (7%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGF-VLVREIYAKKFPRITNSENLYV 82
++S+ L +A AAM EL+ + T EPLWV+SS+ GF VL +E Y F R +
Sbjct: 313 QRSRYLDLALAAMDELVKMAQTREPLWVRSSDS--GFEVLNQEEYDTSFSRCVGPKQDGF 370
Query: 83 CEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLM 142
E+SK++ V+ + + LV+ +DS++WA++ P++VS+ T ++ G + +GAL LM
Sbjct: 371 VSEASKEAGTVIINSLALVETLMDSERWAEMFPSMVSRTSTTEIISSG-MGGRNGALHLM 429
Query: 143 WNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIRE 202
E+ +LSPLVP R+ FLR+C+Q G W +VDVS+DS +S S ++ PSGC +++
Sbjct: 430 HAELQLLSPLVPVRQVSFLRFCKQHAEGVWAVVDVSIDSIREGSSSS-CRRLPSGCLVQD 488
Query: 203 LPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYT 262
+ NG VTW+E E ++ + L R ++R +A+GA RW+ LQR E S T
Sbjct: 489 MANGYSKVTWIEHTEYDEN-HIHRLYRPLLRCGLAFGAHRWMAALQRQCECLTIL-MSST 546
Query: 263 ASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQ-----NLLDPSNQLTTSIKL 317
ST + +GRKS++ L+ RM +F + +Q N+ + + +
Sbjct: 547 VSTSTNPSPINC-NGRKSMLKLAKRMTDNFCGGVCASSLQKWSKLNVGNVDKDVRIMTRK 605
Query: 318 IAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--I 375
+ GI++ A +S+W+PV+P+ LF FL +E ++EWD L+ G PM++ AHI+
Sbjct: 606 SVNNPGEPPGIILNAATSVWMPVSPRRLFDFLGNERLRSEWDILSNGGPMKEMAHIAKGH 665
Query: 376 NESNHVSILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ SN VS+L+ S+I M+I QE+SID G VVYAP+D ++NG DS+ +
Sbjct: 666 DRSNSVSLLRASAINANQSSMLILQETSIDAAGAVVVYAPVDIPAMQAVMNGGDSAYVAL 725
Query: 434 LPSGFTISRDGREGNE-------------------GSLLTMCFQVLVDEEGSSSQVRKKT 474
LPSGF I +G+ G + GSLLT+ FQ+LV+ ++++ ++
Sbjct: 726 LPSGFAILPNGQAGTQRCAAEERNSIGNGGCMEEGGSLLTVAFQILVNSL-PTAKLTVES 784
Query: 475 LESIDSSCMLMLQRIKDALHC 495
+E++++ +Q+IK ALHC
Sbjct: 785 VETVNNLISCTVQKIKAALHC 805
>gb|AAN15463.1| Unknown protein [Arabidopsis thaliana] gi|14276060|dbj|BAB58961.1|
protodermal factor2 [Arabidopsis thaliana]
gi|17064998|gb|AAL32653.1| Unknown protein [Arabidopsis
thaliana] gi|15983372|gb|AAL11554.1| AT4g04890/T1J1_3
[Arabidopsis thaliana] gi|18412734|ref|NP_567274.1|
homeobox-leucine zipper protein protodermal factor 2
(PDF2) [Arabidopsis thaliana]
Length = 743
Score = 281 bits (718), Expect = 4e-74
Identities = 174/503 (34%), Positives = 287/503 (56%), Gaps = 43/503 (8%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLW+ + N V +L E Y + FPR + L +
Sbjct: 247 DKPIIVELAVAAMEELVRMAQTGDPLWLSTDNSVE--ILNEEEYFRTFPRGIGPKPLGLR 304
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ S +V+ + + LV++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 305 SEASRQSAVVIMNHINLVEILMDVNQWSCVFSGIVSRALTLEVLSTGVAGNYNGALQVMT 364
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS ++ PSGC I+EL
Sbjct: 365 AEFQVPSPLVPTRENYFVRYCKQHSDGSWAVVDVSLDSLRPSTPILRTRRRPSGCLIQEL 424
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
PNG VTW+E +EV + + + +V++ +A+GAKRW+ L+R E+ A S S
Sbjct: 425 PNGYSKVTWIEHMEVDDR-SVHNMYKPLVQSGLAFGAKRWVATLERQCERLASSMAS--- 480
Query: 264 STEGLEGEVKLL---DGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAH 320
+ G++ ++ +GRKS++ L+ RMV SF + S + ++++
Sbjct: 481 ---NIPGDLSVITSPEGRKSMLKLAERMVMSFCSGVGASTAHAWTTMSTTGSDDVRVMTR 537
Query: 321 RNTILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
++ +D GIV+ A +S W+PV PK +F FL+DE + EWD L+ G +++ AHI+
Sbjct: 538 KS--MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRKEWDILSNGGMVQEMAHIA 595
Query: 375 INE--SNHVSILQTSS--IIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSK 430
N VS+L+ +S +M+I QES D G YV+YAP+D + ++++G D
Sbjct: 596 NGHEPGNCVSLLRVNSGNSSQSNMLILQESCTDASGSYVIYAPVDIVAMNVVLSGGDPDY 655
Query: 431 KLILPSGFTISRDGR----EGNE--------------GSLLTMCFQVLVDEEGSSSQVRK 472
+LPSGF I DG +GN+ GSLLT+ FQ+LVD ++++
Sbjct: 656 VALLPSGFAILPDGSVGGGDGNQHQEMVSTTSSGSCGGSLLTVAFQILVDSV-PTAKLSL 714
Query: 473 KTLESIDSSCMLMLQRIKDALHC 495
++ +++S ++RIK A+ C
Sbjct: 715 GSVATVNSLIKCTVERIKAAVSC 737
>ref|NP_564041.2| homeobox-leucine zipper family protein / lipid-binding START
domain-containing protein [Arabidopsis thaliana]
gi|25386240|pir||D86314 hypothetical protein F2H15.14 -
Arabidopsis thaliana gi|9665069|gb|AAF97271.1| Strong
similarity to meristem L1 layer homeobox protein (ATML1)
from Arabidopsis thaliana gb|U37589 and contains
Transposase PF|01527, Homeobox PF|00046, and START
PF|01852 domains. EST gb|AI995645 comes from this gene
Length = 687
Score = 280 bits (715), Expect = 1e-73
Identities = 182/496 (36%), Positives = 287/496 (57%), Gaps = 19/496 (3%)
Query: 12 PTRESQESEVTVE--KSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAK 69
P+ SQ + V E KS M +A AM+ELL LL T+EPLW+K+ D VL E Y
Sbjct: 195 PSLPSQPNLVLSEMDKSLMTNIAVTAMEELLRLLQTNEPLWIKT--DGCRDVLNLENYEN 252
Query: 70 KFPRITNS----ENLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQ 125
F R + S NL + E+S+ S +V + + LV + ++S K ++ P+IV+ +KT+
Sbjct: 253 MFTRSSTSGGKKNNLGM--EASRSSGVVFTNAITLVDMLMNSVKLTELFPSIVASSKTLA 310
Query: 126 VFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNC 185
V G NH AL LM E+ VLSPLV +RE+ LRYCQQ+E G W IV+VS + F
Sbjct: 311 VISSGLRGNHGDALHLMIEELQVLSPLVTTREFCVLRYCQQIEHGTWAIVNVSYE-FPQF 369
Query: 186 ASRSVAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLL 245
S+S + +FPSGC I+++ NG VTWVE E + + + +DIV +A+GA+RW+
Sbjct: 370 ISQSRSYRFPSGCLIQDMSNGYSKVTWVEHGEFEEQEPIHEMFKDIVHKGLAFGAERWIA 429
Query: 246 ELQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLL 305
LQR E+ T+S + L G + +G++S++ L++RMV +F + T
Sbjct: 430 TLQRMCERFTNLLEPATSSLD-LGGVIPSPEGKRSIMRLAHRMVSNFCLSVGTSNNTRST 488
Query: 306 DPSNQLTTSIKLIAHRNT-ILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGG 364
S I++ +H++ +G+V+ A +S WLP++P+N+F FLKDE + +WD L+ G
Sbjct: 489 VVSGLDEFGIRVTSHKSRHEPNGMVLCAATSFWLPISPQNVFNFLKDERTRPQWDVLSNG 548
Query: 365 RPMRQKAHIS--INESNHVSILQ--TSSIIGEHMVIFQESSID-DLGGYVVYAPIDKLDA 419
+++ AHI+ N N +S+L+ +S +M+I QES ID V+Y P+D
Sbjct: 549 NSVQEVAHITNGSNPGNCISVLRGFNASSSQNNMLILQESCIDSSSAALVIYTPVDLPAL 608
Query: 420 YMIVNGCDSSKKLILPSGFTISRDGREGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESID 479
+ ++G D+S ILPSGF IS DG GSL+T+ FQ++V +++ +++E+++
Sbjct: 609 NIAMSGQDTSYIPILPSGFAISPDGSSKGGGSLITVGFQIMVSGL-QPAKLNMESMETVN 667
Query: 480 SSCMLMLQRIKDALHC 495
+ + +IK L+C
Sbjct: 668 NLINTTVHQIKTTLNC 683
>ref|XP_480435.1| roc1(homeobox protein) [Oryza sativa (japonica cultivar-group)]
gi|38637066|dbj|BAD03323.1| roc1(homeobox protein)
[Oryza sativa (japonica cultivar-group)]
gi|38636931|dbj|BAD03194.1| roc1(homeobox protein)
[Oryza sativa (japonica cultivar-group)]
Length = 784
Score = 278 bits (710), Expect = 4e-73
Identities = 178/496 (35%), Positives = 280/496 (55%), Gaps = 30/496 (6%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM EL+ + EPLW SS++ +L E YA+ FPR + +
Sbjct: 293 DKPMIVELAVAAMDELVQMAQLDEPLW-SSSSEPAAALLDEEEYARMFPRGLGPKQYGLK 351
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ +V+ + LV++ +D +++A + +IVS+A T +V G N++GAL++M
Sbjct: 352 SEASRHGAVVIMTHSNLVEILMDVNQFATVFSSIVSRASTHEVLSTGVAGNYNGALQVMS 411
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+ G W +VDVSLDS + ++ PSGC I+E+
Sbjct: 412 MEFQVPSPLVPTRESYFVRYCKNNSDGTWAVVDVSLDSLRPSPVQKCRRR-PSGCLIQEM 470
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
PNG VTWVE VEV S + + + +V + +A+GAKRW+ L R E+ A + S
Sbjct: 471 PNGYSKVTWVEHVEVDDS-SVHNIYKPLVNSGLAFGAKRWVGTLDRQCERLASAMASNIP 529
Query: 264 STEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNT 323
+ G G + ++GRKS++ L+ RMV SF + S ++++ R +
Sbjct: 530 N--GDLGVITSVEGRKSMLKLAERMVASFCGGVTASVAHQWTTLSGSGAEDVRVMT-RKS 586
Query: 324 ILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--IN 376
+ D GIV+ A +S WLPV P +F FL+DE ++EWD L+ G +++ AHI+ +
Sbjct: 587 VDDPGRPPGIVLNAATSFWLPVPPAAVFDFLRDETSRSEWDILSNGGAVQEMAHIANGRD 646
Query: 377 ESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLIL 434
N VS+L+ +S +M+I QES D G YVVYAP+D + +++NG D +L
Sbjct: 647 HGNSVSLLRVNSANSNQSNMLILQESCTDASGSYVVYAPVDIVAMNVVLNGGDPDYVALL 706
Query: 435 PSGFTISRDGREGNE-------------GSLLTMCFQVLVDEEGSSSQVRKKTLESIDSS 481
PSGF I DG GN GSLLT+ FQ+LVD ++++ ++ +++S
Sbjct: 707 PSGFAILPDGPSGNAQAAVGENGSGSGGGSLLTVAFQILVDSV-PTAKLSLGSVATVNSL 765
Query: 482 CMLMLQRIKDALHCSD 497
++RIK A+ C D
Sbjct: 766 IACTVERIKAAV-CRD 780
>gb|AAD17342.1| contains similarity to homeobox domains (Pfam: PF00046, Score,36.5,
E=6.9e-08, N=1) [Arabidopsis thaliana]
Length = 772
Score = 278 bits (710), Expect = 4e-73
Identities = 174/516 (33%), Positives = 285/516 (54%), Gaps = 50/516 (9%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLW+ + N V +L E Y + FPR + L +
Sbjct: 257 DKPIIVELAVAAMEELVRMAQTGDPLWLSTDNSVE--ILNEEEYFRTFPRGIGPKPLGLR 314
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ S +V+ + + LV++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 315 SEASRQSAVVIMNHINLVEILMDVNQWSCVFSGIVSRALTLEVLSTGVAGNYNGALQVMT 374
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS ++ PSGC I+EL
Sbjct: 375 AEFQVPSPLVPTRENYFVRYCKQHSDGSWAVVDVSLDSLRPSTPILRTRRRPSGCLIQEL 434
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACS------ 257
PNG VTW+E +EV + + + +V++ +A+GAKRW+ L+R E+ A S
Sbjct: 435 PNGYSKVTWIEHMEVDDR-SVHNMYKPLVQSGLAFGAKRWVATLERQCERLASSMASNIP 493
Query: 258 ----------EFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDP 307
+F Y + + +GRKS++ L+ RMV SF +
Sbjct: 494 GDLSGRGYSDQFKYVGAKLNENVMITSPEGRKSMLKLAERMVMSFCSGVGASTAHAWTTM 553
Query: 308 SNQLTTSIKLIAHRNTILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFL 361
S + ++++ ++ +D GIV+ A +S W+PV PK +F FL+DE + EWD L
Sbjct: 554 STTGSDDVRVMTRKS--MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRKEWDIL 611
Query: 362 AGGRPMRQKAHISINE--SNHVSILQTSS--IIGEHMVIFQESSIDDLGGYVVYAPIDKL 417
+ G +++ AHI+ N VS+L+ +S +M+I QES D G YV+YAP+D +
Sbjct: 612 SNGGMVQEMAHIANGHEPGNCVSLLRVNSGNSSQSNMLILQESCTDASGSYVIYAPVDIV 671
Query: 418 DAYMIVNGCDSSKKLILPSGFTISRDGR----EGNE--------------GSLLTMCFQV 459
++++G D +LPSGF I DG +GN+ GSLLT+ FQ+
Sbjct: 672 AMNVVLSGGDPDYVALLPSGFAILPDGSVGGGDGNQHQEMVSTTSSGSCGGSLLTVAFQI 731
Query: 460 LVDEEGSSSQVRKKTLESIDSSCMLMLQRIKDALHC 495
LVD ++++ ++ +++S ++RIK A+ C
Sbjct: 732 LVDSV-PTAKLSLGSVATVNSLIKCTVERIKAAVSC 766
>dbj|BAB85750.1| Roc1 [Oryza sativa]
Length = 784
Score = 278 bits (710), Expect = 4e-73
Identities = 178/496 (35%), Positives = 280/496 (55%), Gaps = 30/496 (6%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM EL+ + EPLW SS++ +L E YA+ FPR + +
Sbjct: 293 DKPMIVELAVAAMDELVQMAQLDEPLW-SSSSEPAAALLDEEEYARMFPRGLGPKQYGLK 351
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ +V+ + LV++ +D +++A + +IVS+A T +V G N++GAL++M
Sbjct: 352 SEASRHGAVVIMTHSNLVEILMDVNQFATVFSSIVSRASTHEVLSTGVAGNYNGALQVMS 411
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+ G W +VDVSLDS + ++ PSGC I+E+
Sbjct: 412 MEFQVPSPLVPTRESYFVRYCKNNSDGTWAVVDVSLDSLRPSPVQKCRRR-PSGCLIQEM 470
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
PNG VTWVE VEV S + + + +V + +A+GAKRW+ L R E+ A + S
Sbjct: 471 PNGYSKVTWVEHVEVDDS-SVHNIYKPLVNSGLAFGAKRWVGTLDRQCERLASAMASNIP 529
Query: 264 STEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNT 323
+ G G + ++GRKS++ L+ RMV SF + S ++++ R +
Sbjct: 530 N--GDLGVITSVEGRKSMLKLAERMVASFCGGVTASVAHQWTTLSGSGAEDVRVMT-RKS 586
Query: 324 ILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--IN 376
+ D GIV+ A +S WLPV P +F FL+DE ++EWD L+ G +++ AHI+ +
Sbjct: 587 VDDPGRPPGIVLNAATSFWLPVPPAAVFDFLRDETSRSEWDILSNGGAVQEMAHIANGRD 646
Query: 377 ESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLIL 434
N VS+L+ +S +M+I QES D G YVVYAP+D + +++NG D +L
Sbjct: 647 HGNSVSLLRVNSANSNQSNMLILQESCTDASGSYVVYAPVDIVAMNVVLNGGDPDYVALL 706
Query: 435 PSGFTISRDGREGNE-------------GSLLTMCFQVLVDEEGSSSQVRKKTLESIDSS 481
PSGF I DG GN GSLLT+ FQ+LVD ++++ ++ +++S
Sbjct: 707 PSGFAILPDGPSGNAQAAVGENGSGSGGGSLLTVAFQILVDSV-PTAKLSLGSVATVNSL 765
Query: 482 CMLMLQRIKDALHCSD 497
++RIK A+ C D
Sbjct: 766 IACTVERIKAAV-CRD 780
>emb|CAB81282.1| L1 specific homeobox gene ATML1/ovule-specific homeobox protein A20
[Arabidopsis thaliana] gi|4455283|emb|CAB36819.1| L1
specific homeobox gene ATML1/ovule-specific homeobox
protein A20 [Arabidopsis thaliana]
gi|7446299|pir||T05850 homeobox protein ATML1,
L1-specific - Arabidopsis thaliana
Length = 718
Score = 272 bits (696), Expect = 2e-71
Identities = 175/513 (34%), Positives = 285/513 (55%), Gaps = 50/513 (9%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLWV S N V +L E Y + FPR + + +
Sbjct: 212 DKPMIVELAVAAMEELVRMAQTGDPLWVSSDNSVE--ILNEEEYFRTFPRGIGPKPIGLR 269
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S++S +V+ + + L+++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 270 SEASRESTVVIMNHINLIEILMDVNQWSSVFCGIVSRALTLEVLSTGVAGNYNGALQVMT 329
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS + + +++ PSGC I+EL
Sbjct: 330 AEFQVPSPLVPTRENYFVRYCKQHSDGIWAVVDVSLDSL-RPSPITRSRRRPSGCLIQEL 388
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
NG VTWVE +EV + + + +V +A+GAKRW+ L R E+ A S S
Sbjct: 389 QNGYSKVTWVEHIEVDDR-SVHNMYKPLVNTGLAFGAKRWVATLDRQCERLASSMASNIP 447
Query: 264 STEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNT 323
+ + + +GRKS++ L+ RMV SF + S + ++++ ++
Sbjct: 448 ACD--LSVITSPEGRKSMLKLAERMVMSFCTGVGASTAHAWTTLSTTGSDDVRVMTRKS- 504
Query: 324 ILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--I 375
+D GIV+ A +S W+PV PK +F FL+DE ++EWD L+ G +++ AHI+
Sbjct: 505 -MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRSEWDILSNGGLVQEMAHIANGR 563
Query: 376 NESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ N VS+L+ +S +M+I QES D G YV+YAP+D + ++++G D +
Sbjct: 564 DPGNSVSLLRVNSGNSGQSNMLILQESCTDASGSYVIYAPVDIIAMNVVLSGGDPDYVAL 623
Query: 434 LPSGFTISRDGR--------------------EGNE-----------GSLLTMCFQVLVD 462
LPSGF I DG EGN GSLLT+ FQ+LVD
Sbjct: 624 LPSGFAILPDGSARGGGGSANASAGAGVEGGGEGNNLEVVTTTGSCGGSLLTVAFQILVD 683
Query: 463 EEGSSSQVRKKTLESIDSSCMLMLQRIKDALHC 495
++++ ++ +++S ++RIK AL C
Sbjct: 684 SV-PTAKLSLGSVATVNSLIKCTVERIKAALAC 715
>gb|AAN12908.1| putative L1-specific homeobox gene ATML1/ovule-specific homeobox
protein A20 [Arabidopsis thaliana]
gi|20268701|gb|AAM14054.1| putative L1-specific homeobox
gene ATML1/ovule-specific homeobox protein A20
[Arabidopsis thaliana] gi|22328861|ref|NP_193906.2| L1
specific homeobox gene (ML1) / ovule-specific homeobox
protein A20 [Arabidopsis thaliana]
Length = 762
Score = 272 bits (696), Expect = 2e-71
Identities = 175/513 (34%), Positives = 285/513 (55%), Gaps = 50/513 (9%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLWV S N V +L E Y + FPR + + +
Sbjct: 256 DKPMIVELAVAAMEELVRMAQTGDPLWVSSDNSVE--ILNEEEYFRTFPRGIGPKPIGLR 313
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S++S +V+ + + L+++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 314 SEASRESTVVIMNHINLIEILMDVNQWSSVFCGIVSRALTLEVLSTGVAGNYNGALQVMT 373
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS + + +++ PSGC I+EL
Sbjct: 374 AEFQVPSPLVPTRENYFVRYCKQHSDGIWAVVDVSLDSL-RPSPITRSRRRPSGCLIQEL 432
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
NG VTWVE +EV + + + +V +A+GAKRW+ L R E+ A S S
Sbjct: 433 QNGYSKVTWVEHIEVDDR-SVHNMYKPLVNTGLAFGAKRWVATLDRQCERLASSMASNIP 491
Query: 264 STEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNT 323
+ + + +GRKS++ L+ RMV SF + S + ++++ ++
Sbjct: 492 ACD--LSVITSPEGRKSMLKLAERMVMSFCTGVGASTAHAWTTLSTTGSDDVRVMTRKS- 548
Query: 324 ILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--I 375
+D GIV+ A +S W+PV PK +F FL+DE ++EWD L+ G +++ AHI+
Sbjct: 549 -MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRSEWDILSNGGLVQEMAHIANGR 607
Query: 376 NESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ N VS+L+ +S +M+I QES D G YV+YAP+D + ++++G D +
Sbjct: 608 DPGNSVSLLRVNSGNSGQSNMLILQESCTDASGSYVIYAPVDIIAMNVVLSGGDPDYVAL 667
Query: 434 LPSGFTISRDGR--------------------EGNE-----------GSLLTMCFQVLVD 462
LPSGF I DG EGN GSLLT+ FQ+LVD
Sbjct: 668 LPSGFAILPDGSARGGGGSANASAGAGVEGGGEGNNLEVVTTTGSCGGSLLTVAFQILVD 727
Query: 463 EEGSSSQVRKKTLESIDSSCMLMLQRIKDALHC 495
++++ ++ +++S ++RIK AL C
Sbjct: 728 SV-PTAKLSLGSVATVNSLIKCTVERIKAALAC 759
>gb|AAB49378.1| A20
Length = 718
Score = 271 bits (694), Expect = 3e-71
Identities = 175/513 (34%), Positives = 285/513 (55%), Gaps = 50/513 (9%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLWV S N V +L E Y + FPR + + +
Sbjct: 212 DKPMIVELAVAAMEELVRMAQTGDPLWVSSDNSVE--ILNEEEYFRTFPRGIGPKPIGLR 269
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S++S +V+ + + L+++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 270 SEASRESTVVIMNHINLIEILMDVNQWSSVFCGIVSRALTLEVLSTGVRGNYNGALQVMT 329
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS + + +++ PSGC I+EL
Sbjct: 330 AEFQVPSPLVPTRENYFVRYCKQHSDGIWAVVDVSLDSL-RPSPITRSRRRPSGCLIQEL 388
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
NG VTWVE +EV + + + +V +A+GAKRW+ L R E+ A S S
Sbjct: 389 QNGYSKVTWVEHIEVDDR-SVHNMYKPLVNTGLAFGAKRWVATLDRQCERLASSMASNIP 447
Query: 264 STEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNT 323
+ + + +GRKS++ L+ RMV SF + S + ++++ ++
Sbjct: 448 ACD--LSVITSPEGRKSMLKLAERMVMSFCTGVGASTADAWTTLSTTGSDDVRVMTRKS- 504
Query: 324 ILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--I 375
+D GIV+ A +S W+PV PK +F FL+DE ++EWD L+ G +++ AHI+
Sbjct: 505 -MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRSEWDILSNGGLVQEMAHIANGR 563
Query: 376 NESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ N VS+L+ +S +M+I QES D G YV+YAP+D + ++++G D +
Sbjct: 564 DPGNSVSLLRVNSGNSGQSNMLILQESCTDASGSYVIYAPVDIIAMNVVLSGGDPDYVAL 623
Query: 434 LPSGFTISRDGR--------------------EGNE-----------GSLLTMCFQVLVD 462
LPSGF I DG EGN GSLLT+ FQ+LVD
Sbjct: 624 LPSGFAILPDGSARGGGGSANASAGAGVEGGGEGNNLEVVTTTGSCGGSLLTVAFQILVD 683
Query: 463 EEGSSSQVRKKTLESIDSSCMLMLQRIKDALHC 495
++++ ++ +++S ++RIK AL C
Sbjct: 684 SV-PTAKLSLGSVATVNSLIKCTVERIKAALAC 715
>gb|AAD47139.1| Anthocyaninless2 [Arabidopsis thaliana]
Length = 801
Score = 271 bits (693), Expect = 3e-71
Identities = 171/499 (34%), Positives = 281/499 (56%), Gaps = 43/499 (8%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+KS +L +A AM EL+ L + EPLWVKS D L ++ Y + F ++++ +
Sbjct: 317 QKSVLLELALTAMDELVKLAQSEEPLWVKSL-DGERDELNQDEYMRTF---SSTKPTGLA 372
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ S +V+ + + LV+ +DS++W ++ P V++A T V G +GAL+LM
Sbjct: 373 TEASRTSGMVIINSLALVETLMDSNRWTEMFPCNVARATTTDVISGGMAGTINGALQLMN 432
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSF-GNCASRSVAKKFPSGCRIRE 202
E+ VLSPLVP R FLR+C+Q G W +VDVS+D N V ++ PSGC +++
Sbjct: 433 AELQVLSPLVPVRNVNFLRFCKQHAEGVWPVVDVSIDPVRENSGGAPVIRRLPSGCVVQD 492
Query: 203 LPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYT 262
+ NG VTWVE E ++ Q+ L R ++R+ + +G++RWL LQR E C +
Sbjct: 493 VSNGYSKVTWVEHAEYDEN-QIHQLYRPLLRSGLGFGSQRWLATLQRQCE---CLAILMS 548
Query: 263 ASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNL-------LDPSNQLTTSI 315
+S + GRKS++ L+ RM +F ++ + N +DP ++ T
Sbjct: 549 SSVTSHDNTSITPGGRKSMLKLAQRMTFNFCSGISAPSVHNWSKLTVGNVDPDVRVMT-- 606
Query: 316 KLIAHRNTILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQK 370
R ++ D GIV+ A +S+WLP P+ L+ FL++E + EWD L+ G PM++
Sbjct: 607 -----RKSVDDPGEPPGIVLSAATSVWLPAAPQRLYDFLRNERMRCEWDILSNGGPMQEM 661
Query: 371 AHISINESNHVSILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDS 428
AHI+ + VS+L+++++ M+I QE+ ID G VVYAP+D ++++NG DS
Sbjct: 662 AHITKGQDQGVSLLRSNAMNANQSSMLILQETCIDASGALVVYAPVDIPAMHVVMNGGDS 721
Query: 429 SKKLILPSGFTISRDG------------REGNEGSLLTMCFQVLVDEEGSSSQVRKKTLE 476
S +LPSGF +S DG R GSLLT+ FQ+LV+ ++++ +++E
Sbjct: 722 SYVALLPSGFAVSSDGGIDGGGSGDGDQRPVGGGSLLTVAFQILVNNL-PTAKLTVESVE 780
Query: 477 SIDSSCMLMLQRIKDALHC 495
++++ +Q+I+ AL C
Sbjct: 781 TVNNLISCTVQKIRAALQC 799
>gb|AAG43405.1| homeobox 1 [Picea abies]
Length = 763
Score = 271 bits (693), Expect = 3e-71
Identities = 170/492 (34%), Positives = 275/492 (55%), Gaps = 28/492 (5%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
EK ++ +A AAM+EL+ + EPLW D +L + Y + FPR +
Sbjct: 273 EKPMVVELAVAAMEELVRMAQLGEPLWTSHPEDSTD-ILNEDEYIRTFPRGIGPRPYGLK 331
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+++ +V+ + + LV+ +D ++W+ + P IVS+ T+ VF G N++GAL++M
Sbjct: 332 AEASRETAVVIMNAINLVETLMDVNQWSSMFPGIVSRPFTVDVFSTGVAGNYNGALQVMH 391
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSF-GNCASRSVAKKFPSGCRIRE 202
E V SPLVP+RE +F+RYC+Q W +VDVSLDS GN +S ++ PSGC I+E
Sbjct: 392 AEFQVPSPLVPTREIYFVRYCKQHSDSIWAVVDVSLDSLRGNSSSVIRCRRRPSGCLIQE 451
Query: 203 LPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYT 262
+PN VTWVE VE + + R +V + +A+GAKRW+ LQR E+ A S
Sbjct: 452 MPNSYSKVTWVEHVEADDR-AVHHIYRQLVNSGMAFGAKRWIATLQRQCERLASVLASNI 510
Query: 263 ASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRN 322
+ + G + +GRKS++ L+ RMV SF ++ S ++++ R
Sbjct: 511 PARD--LGVIPSPEGRKSILKLAERMVTSFCAGVSASTAHTWTTLSGSGAEDVRVMT-RK 567
Query: 323 TILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISINE 377
+I D GI++ A +S+WLPV PK +F FL+DE + EWD L+ G +++ HI+ +
Sbjct: 568 SIDDPGRPPGIILSAATSLWLPVPPKKVFDFLRDENSRNEWDILSNGGLVQEVDHIANGQ 627
Query: 378 --SNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
N VS+L+ +++ +M+I QES D G +V+YAP+D + ++++G D +
Sbjct: 628 DPGNCVSLLRVNTVNSNQSNMLILQESCTDASGSFVIYAPVDIVAMNVVLSGGDPDYVAL 687
Query: 434 LPSGFTISRDGRE------------GNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSS 481
LPSGF I D + G GSLLT+ FQ+LVD ++++ ++ +++S
Sbjct: 688 LPSGFAILPDSPKCMAVTNSGINDLGTGGSLLTVAFQILVDSV-PTAKLSLGSVATVNSL 746
Query: 482 CMLMLQRIKDAL 493
+ RIK A+
Sbjct: 747 ISCTVDRIKAAV 758
>gb|AAM97321.1| homeodomain protein GhHOX1 [Gossypium hirsutum]
Length = 753
Score = 270 bits (691), Expect = 6e-71
Identities = 175/503 (34%), Positives = 273/503 (53%), Gaps = 38/503 (7%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYV 82
+EKS+++ + N AM+EL + EPLWV+S + +L + Y K+ ++S
Sbjct: 260 LEKSRIMEIVNQAMEELQKMATAGEPLWVRSV-ETGREILNYDEYVKELSVESSSNGRPK 318
Query: 83 CE-ESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRL 141
E+S+++ +V LV+ F+D+++W ++ P I+SKA T+ V G N +GA++L
Sbjct: 319 RSIEASRETGVVFLDLPRLVQSFMDANQWKEMFPCIISKAATVDVICHGEAPNKNGAVQL 378
Query: 142 MWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAK--KFPSGCR 199
M+ E+ +L+PLVP+RE +F+RYC+Q+ A +W IVDVS+D S+ K K PSGC
Sbjct: 379 MFAELQMLTPLVPTREVYFVRYCKQLSAEQWAIVDVSIDKVEENIDASLVKCRKRPSGCI 438
Query: 200 IRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEF 259
I++ NG C V WVE +E Q + L R IVR+ +A+GA+ W+ LQ E+
Sbjct: 439 IQDTTNGHCKVIWVEHLEC-QKNTVHTLYRTIVRSGLAFGARHWMATLQHQCERLVFF-M 496
Query: 260 SYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIA 319
+ T+ G V L GRKS++ L+ RM SF + S + I++ +
Sbjct: 497 ATNVPTKDSTG-VATLAGRKSILKLAQRMTWSFCHSIGASSYHTWNKVSTKTGEDIRVSS 555
Query: 320 HRNT----ILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISI 375
+N G++V AVSS+WLPV+P LF FL+DE R++EWD ++ G P++ A+++
Sbjct: 556 RKNLNDPGEPHGVIVCAVSSVWLPVSPTLLFDFLRDESRRSEWDIMSNGGPVQSIANLAK 615
Query: 376 NESNHVSI-LQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLIL 434
+ ++ +Q M + Q+S + VV+A +D ++ GCDSS IL
Sbjct: 616 GKDRGNAVTIQAMKSKENSMWVLQDSCTNAFESMVVFAHVDVTGIQSVITGCDSSNMAIL 675
Query: 435 PSGFTISRDGREGNE---------------GSLLTMCFQVLVDEEGSSSQVRKKTLESID 479
PSGF+I DG E GSLLT+ FQ+L +SS K T+ES++
Sbjct: 676 PSGFSILPDGLESRPLVISSRHEKSNDTEGGSLLTVAFQILT----NSSPTAKLTMESVE 731
Query: 480 S-----SCMLMLQRIKDALHCSD 497
S SC L+ IK +L C D
Sbjct: 732 SVNTIVSC--TLRNIKTSLQCED 752
>gb|AAK19610.1| BNLGHi8377 [Gossypium hirsutum]
Length = 758
Score = 270 bits (691), Expect = 6e-71
Identities = 178/504 (35%), Positives = 275/504 (54%), Gaps = 40/504 (7%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYV 82
+EKS+++ + N AM+EL + EPLWV+S + +L + Y K+ ++S
Sbjct: 265 LEKSRIMEIVNQAMEELQKMATAGEPLWVRSV-ETGREILNYDEYVKELSVESSSNGRPK 323
Query: 83 CE-ESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRL 141
E+S+++ +V LV+ F+D+++W ++ P I+SKA T+ V G N +GA++L
Sbjct: 324 RSIEASRETGVVFLDLPRLVQSFMDANQWKEMFPCIISKAATVDVICHGEAPNKNGAVQL 383
Query: 142 MWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAK--KFPSGCR 199
M+ E+ +L+PLVP+RE +F+RYC+Q+ A +W IVDVS+D S+ K K PSGC
Sbjct: 384 MFAELQMLTPLVPTREVYFVRYCKQLSAEQWAIVDVSIDKVEENIDASLVKCRKRPSGCI 443
Query: 200 IRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEF 259
I++ NG C V WVE +E Q + L R IVR+ +A+GA+ W+ LQ E+
Sbjct: 444 IQDKTNGHCKVIWVEHLEC-QKNTVHTLYRTIVRSGLAFGARHWMATLQHQCERLVFF-M 501
Query: 260 SYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIA 319
+ T+ G V L GRKS++ L+ RM SF + S + I++ +
Sbjct: 502 ATNVPTKDSTG-VATLAGRKSILKLAQRMTWSFCHSIGASSYHTWNKVSTKTGEDIRVSS 560
Query: 320 HRNT----ILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS- 374
+N G++V AVSS+WLPV+P LF FL+DE R++EWD ++ G P++ A+++
Sbjct: 561 RKNLNDPGEPHGVIVCAVSSVWLPVSPTLLFDFLRDESRRSEWDIMSNGGPVQSIANLAK 620
Query: 375 -INESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
++ N V+I Q M + Q+S + VV+A +D ++ GCDSS I
Sbjct: 621 GKDQGNAVTI-QAMKSKENSMWVLQDSCTNAFESMVVFAHVDVTGIQSVITGCDSSNMAI 679
Query: 434 LPSGFTISRDGREGNE---------------GSLLTMCFQVLVDEEGSSSQVRKKTLESI 478
LPSGF+I DG E GSLLT+ FQ+L +SS K T+ES+
Sbjct: 680 LPSGFSILPDGLESRPLVISSRHEKSNDTEGGSLLTVAFQILT----NSSPTAKLTMESV 735
Query: 479 DS-----SCMLMLQRIKDALHCSD 497
+S SC L+ IK +L C D
Sbjct: 736 ESVNTIVSC--TLRNIKTSLQCED 757
>ref|XP_473974.1| OSJNBb0060E08.16 [Oryza sativa (japonica cultivar-group)]
gi|39545845|emb|CAE04753.3| OSJNBb0060E08.16 [Oryza
sativa (japonica cultivar-group)]
Length = 781
Score = 270 bits (690), Expect = 8e-71
Identities = 176/496 (35%), Positives = 280/496 (55%), Gaps = 31/496 (6%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLW-VKSSNDVVGFV---LVREIYAKKFPRITNSE 78
V+K ++ +A AAM+EL+ + EPLW V D L E YA+ FPR +
Sbjct: 285 VDKPMIVELAVAAMEELVRMAQLDEPLWSVAPPLDATAAAMETLSEEEYARMFPRGLGPK 344
Query: 79 NLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGA 138
+ E+S+DS +V+ + LV++ +D++++A + IVS+A T++V G N++GA
Sbjct: 345 QYGLRSEASRDSAVVIMTHANLVEILMDANQYAAVFSNIVSRAITLEVLSTGVAGNYNGA 404
Query: 139 LRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGC 198
L++M E V SPLVP+RE +F+RYC+Q G W +VDVSLDS ++ PSGC
Sbjct: 405 LQVMSVEFQVPSPLVPTRESYFVRYCKQNADGTWAVVDVSLDSLRPSPVLKCRRR-PSGC 463
Query: 199 RIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSE 258
I+E+PNG VTWVE VEV + + + +V + +A+GA+RW+ L R E+ A
Sbjct: 464 LIQEMPNGYSKVTWVEHVEVDDR-SVHNIYKLLVNSGLAFGARRWVGTLDRQCERLASVM 522
Query: 259 FSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLI 318
S +++ G + +GRKS++ L+ RMV SF + S ++++
Sbjct: 523 ASNIPTSD--IGVITSSEGRKSMLKLAERMVVSFCGGVTASVAHQWTTLSGSGAEDVRVM 580
Query: 319 AHRNTILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHI 373
R ++ D GIV+ A +S WLPV PK +F FL+DE ++EWD L+ G +++ AHI
Sbjct: 581 T-RKSVDDPGRPPGIVLNAATSFWLPVPPKRVFDFLRDESSRSEWDILSNGGIVQEMAHI 639
Query: 374 S--INESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSS 429
+ ++ N VS+L+ +S +M+I QES D G YV+YAP+D + +++NG D
Sbjct: 640 ANGRDQGNCVSLLRVNSSNSNQSNMLILQESCTDASGSYVIYAPVDVVAMNVVLNGGDPD 699
Query: 430 KKLILPSGFTISRDGRE------------GNEGSLLTMCFQVLVDEEGSSSQVRKKTLES 477
+LPSGF I DG G+ GSLLT+ FQ+LVD ++++ ++ +
Sbjct: 700 YVALLPSGFAILPDGPAHDGGDGDGGVGVGSGGSLLTVAFQILVDSV-PTAKLSLGSVAT 758
Query: 478 IDSSCMLMLQRIKDAL 493
++S ++RIK A+
Sbjct: 759 VNSLIACTVERIKAAV 774
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 779,504,022
Number of Sequences: 2540612
Number of extensions: 30732933
Number of successful extensions: 64757
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 96
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 64187
Number of HSP's gapped (non-prelim): 119
length of query: 497
length of database: 863,360,394
effective HSP length: 132
effective length of query: 365
effective length of database: 527,999,610
effective search space: 192719857650
effective search space used: 192719857650
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0321.6


