

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0316.5
(318 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
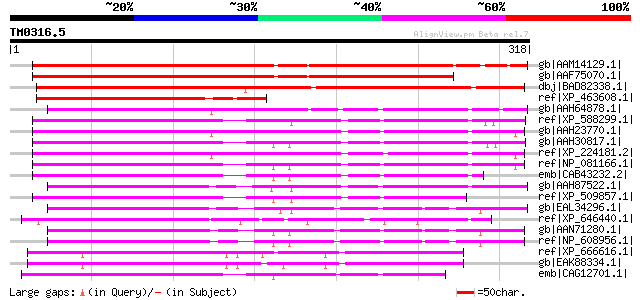
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM14129.1| unknown protein [Arabidopsis thaliana] gi|1581057... 295 1e-78
gb|AAF75070.1| F24B9.6 [Arabidopsis thaliana] gi|25372823|pir||H... 273 5e-72
dbj|BAD82338.1| leucine zipper factor-like [Oryza sativa (japoni... 260 3e-68
ref|XP_463608.1| P0456E05.27 [Oryza sativa (japonica cultivar-gr... 155 1e-36
gb|AAH64878.1| LOC395016 protein [Xenopus tropicalis] 132 1e-29
ref|XP_588299.1| PREDICTED: similar to Protein C14orf120 [Bos ta... 132 1e-29
gb|AAH23770.1| 1500001L15Rik protein [Mus musculus] 130 4e-29
gb|AAH30817.1| C14orf120 protein [Homo sapiens] gi|51471747|ref|... 130 7e-29
ref|XP_224181.2| PREDICTED: similar to Protein C14orf120 homolog... 129 9e-29
ref|NP_081166.1| hypothetical protein LOC68966 [Mus musculus] gi... 129 9e-29
emb|CAB43232.2| hypothetical protein [Homo sapiens] 129 9e-29
gb|AAH87522.1| LOC496169 protein [Xenopus laevis] 129 1e-28
ref|XP_509857.1| PREDICTED: similar to hypothetical protein DKFZ... 129 1e-28
gb|EAL34296.1| GA10717-PA [Drosophila pseudoobscura] 124 5e-27
ref|XP_646440.1| hypothetical protein DDB0190708 [Dictyostelium ... 119 2e-25
gb|AAN71280.1| RE04093p [Drosophila melanogaster] 114 5e-24
ref|NP_608956.1| CG11030-PA [Drosophila melanogaster] gi|2294569... 113 6e-24
ref|XP_666616.1| similar to CG11030-PA [Cryptosporidium hominis]... 112 1e-23
gb|EAK88334.1| conserved eukaryotic nuclear protein that shares ... 112 2e-23
emb|CAG12701.1| unnamed protein product [Tetraodon nigroviridis] 108 2e-22
>gb|AAM14129.1| unknown protein [Arabidopsis thaliana] gi|15810573|gb|AAL07174.1|
unknown protein [Arabidopsis thaliana]
gi|42571389|ref|NP_973785.1| leucine zipper
factor-related [Arabidopsis thaliana]
gi|18390818|ref|NP_563798.1| leucine zipper
factor-related [Arabidopsis thaliana]
Length = 312
Score = 295 bits (755), Expect = 1e-78
Identities = 162/304 (53%), Positives = 214/304 (70%), Gaps = 9/304 (2%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
+KEA LA++L+EMK LDVVR K+++LTA VK +PTA G SYLEAK+LLLL+YCQ L
Sbjct: 14 KKEAPQLASVLREMKNVLDVVRSKVEALTALVKANSFPTAGGISYLEAKHLLLLSYCQDL 73
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
VYY+LRKAKG SI+ HP+VRS+VEIR++LEKIRPIDKK QYQIQKL G
Sbjct: 74 VYYILRKAKGLSIDGHPLVRSLVEIRMFLEKIRPIDKKLQYQIQKLTTAGGPVTELAHSE 133
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNA 194
K + KSED+S Y+P PD+L +ED QE YRP KFAP SM +DKTSKQER+A
Sbjct: 134 GKGSCEAQKSEDLSNYKPKPDLLADKED--DQEDDGVYRPPKFAPMSM-EDKTSKQERDA 190
Query: 195 LRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEG-SSKEVERHIAKMEERAWQEEELF 253
R+EK +++ +++++ +++D+E++PEEIRDY G S E +R +A+ E + EEELF
Sbjct: 191 ARKEKHFFRQATENTYMKDVLDDLEDRPEEIRDYYGVESNEQKRFMAQYERQQKAEEELF 250
Query: 254 NRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNGRL 313
R P TK ++KREK LK S +GL LTE+F+D+IK F D+ GE+ + + R G
Sbjct: 251 TRAPRTKEDKKREKRLKSS-SGLHELTENFYDDIK---FLDKDGEKPRSFGR-NKRGGPF 305
Query: 314 KKRK 317
KKRK
Sbjct: 306 KKRK 309
>gb|AAF75070.1| F24B9.6 [Arabidopsis thaliana] gi|25372823|pir||H86213 protein
F24B9.6 [imported] - Arabidopsis thaliana
Length = 279
Score = 273 bits (698), Expect = 5e-72
Identities = 143/259 (55%), Positives = 187/259 (71%), Gaps = 4/259 (1%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
+KEA LA++L+EMK LDVVR K+++LTA VK +PTA G SYLEAK+LLLL+YCQ L
Sbjct: 14 KKEAPQLASVLREMKNVLDVVRSKVEALTALVKANSFPTAGGISYLEAKHLLLLSYCQDL 73
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
VYY+LRKAKG SI+ HP+VRS+VEIR++LEKIRPIDKK QYQIQKL G
Sbjct: 74 VYYILRKAKGLSIDGHPLVRSLVEIRMFLEKIRPIDKKLQYQIQKLTTAGGPVTELAHSE 133
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNA 194
K + KSED+S Y+P PD+L +ED QE YRP KFAP SM +DKTSKQER+A
Sbjct: 134 GKGSCEAQKSEDLSNYKPKPDLLADKED--DQEDDGVYRPPKFAPMSM-EDKTSKQERDA 190
Query: 195 LRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEG-SSKEVERHIAKMEERAWQEEELF 253
R+EK +++ +++++ +++D+E++PEEIRDY G S E +R +A+ E + EEELF
Sbjct: 191 ARKEKHFFRQATENTYMKDVLDDLEDRPEEIRDYYGVESNEQKRFMAQYERQQKAEEELF 250
Query: 254 NRVPLTKHERKREKYLKKS 272
R P TK ++KREK LK S
Sbjct: 251 TRAPRTKEDKKREKRLKSS 269
>dbj|BAD82338.1| leucine zipper factor-like [Oryza sativa (japonica cultivar-group)]
gi|56784387|dbj|BAD82426.1| leucine zipper factor-like
[Oryza sativa (japonica cultivar-group)]
Length = 335
Score = 260 bits (665), Expect = 3e-68
Identities = 139/303 (45%), Positives = 202/303 (65%), Gaps = 8/303 (2%)
Query: 17 EASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVY 76
+A L A LKEMKEGLD+V K+++LT KVK+ Q PTADG YLEAK+ LLL+YCQ +VY
Sbjct: 28 DAPKLLAALKEMKEGLDLVTGKVKALTRKVKKNQLPTADGIGYLEAKHHLLLSYCQDIVY 87
Query: 77 YLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKK 136
YLLRKAKG S+E HPVVRS+VEIRL+LEKIRPIDKK +YQIQKL ++ A +
Sbjct: 88 YLLRKAKGLSVEGHPVVRSLVEIRLFLEKIRPIDKKMEYQIQKLTNAADSGAAQEKVLNA 147
Query: 137 EPVASDK---SEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERN 193
E + D+ ED+ KYRPNPDM+ S+ D A Q+ YRP KF +MD + K+ +
Sbjct: 148 EAKSKDQPKDDEDLLKYRPNPDMMDSKIDPAGQDNDGIYRPPKFIAATMDDE--DKRHKQ 205
Query: 194 ALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEG-SSKEVERHIAKMEERAWQEEEL 252
A R++K + + + SS+ + +++D ++PEE+++ G S+E R++ + E + QEEEL
Sbjct: 206 ASRKDKTLARMATESSYFKEIIDDAADRPEELKETAGDESREFTRYMRQRELQEKQEEEL 265
Query: 253 FNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNGR 312
F R PLTK +++ EK+++K +GL GLT+ F ++ F D + +G ++ ++G
Sbjct: 266 FTRAPLTKRDKQTEKWMRKELHGLRGLTDGF--DLGINMFVDGDKDNDVGSTEPHFKSGG 323
Query: 313 LKK 315
+K
Sbjct: 324 RRK 326
>ref|XP_463608.1| P0456E05.27 [Oryza sativa (japonica cultivar-group)]
Length = 229
Score = 155 bits (393), Expect = 1e-36
Identities = 86/141 (60%), Positives = 101/141 (70%), Gaps = 6/141 (4%)
Query: 17 EASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVY 76
+A L A LKEMKEGLD+V K+++LT KVK+ Q PTADG YLEAK+ LLL+YCQ +VY
Sbjct: 28 DAPKLLAALKEMKEGLDLVTGKVKALTRKVKKNQLPTADGIGYLEAKHHLLLSYCQDIVY 87
Query: 77 YLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKK 136
YLLRKAKG S+E HPVVRS+VEIRL+LEKIRPIDKK +YQIQKL NAA S +K
Sbjct: 88 YLLRKAKGLSVEGHPVVRSLVEIRLFLEKIRPIDKKMEYQIQKL----TNAADSGAAQEK 143
Query: 137 EPVASDKSEDVSKYRPNPDML 157
E SE+V P L
Sbjct: 144 E--VDYCSEEVRSVTPTGGFL 162
>gb|AAH64878.1| LOC395016 protein [Xenopus tropicalis]
Length = 318
Score = 132 bits (333), Expect = 1e-29
Identities = 98/301 (32%), Positives = 152/301 (49%), Gaps = 17/301 (5%)
Query: 24 LLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVYYLLRKAK 83
L + +++ + V +Q+LT KV+ G Y T G S+LE K+ LLL Y Q L + +L K
Sbjct: 20 LFRTLQDQVTKVTAHVQALTQKVRSGIYNTDKGLSFLELKDQLLLFYLQDLTHLMLEKTN 79
Query: 84 GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMK------VGENAATSDLPSKKE 137
G SI+ +P + +VE+R LEK+RPID+K +YQI KL++ +GEN P+ +
Sbjct: 80 GKSIKGNPGILRLVELRTVLEKMRPIDQKLKYQIDKLVRASVTGSLGENDPLRFKPNPQN 139
Query: 138 PVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRR 197
++ D + D S PQ Y P + AP D D +++E + R
Sbjct: 140 LISKLSEADEGESDSGEDCAESGNAKKPQSKVKKYIPPRLAPVHYD-DTEAEREHRIIER 198
Query: 198 EKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVP 257
K++ + SS IR L + PEEIR EG + + RH + + R EE + R+
Sbjct: 199 AKKL---ALSSSTIRELKEQYSDAPEEIR--EGRAYHMMRHDKEEQHRINHEESMMVRLN 253
Query: 258 LTKHERKREKYLKKSRNGLDGLTESFFDEIKTLP-FEDETGEQVMGPSKGSNRNGRLKKR 316
+T+ E+ R+K + + L+ LT F +I L E T + V PS +R G K +
Sbjct: 254 MTRKEKARKKRVLAMTSQLNSLTH--FSDISALTGGEGRTDDLV--PSVKKSRKGPKKSK 309
Query: 317 K 317
K
Sbjct: 310 K 310
>ref|XP_588299.1| PREDICTED: similar to Protein C14orf120 [Bos taurus]
Length = 315
Score = 132 bits (332), Expect = 1e-29
Identities = 99/329 (30%), Positives = 153/329 (46%), Gaps = 48/329 (14%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + ALLK ++E + V ++Q+LT KV+ YPT G S LE K+ LLL Y L
Sbjct: 8 ESDLPNAVALLKNLQEQVMAVTAQVQTLTKKVQAKAYPTEKGLSLLEVKDQLLLMYLMDL 67
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
+ +L KA G S++ HP V +VEIR LEK+RP+D+K +YQI KL+K + S+
Sbjct: 68 SHLILDKASGGSLQGHPAVLRLVEIRTVLEKLRPLDQKLKYQIDKLVKTAVTGSLSE--- 124
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDA-------------------YRPV 175
D +++P+P ++S+ +E +A Y P
Sbjct: 125 ----------NDPLRFKPHPSNMMSKLSSEDEEEDEAEEGQSGASGKKSGKGTAKKYVPP 174
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEV 235
+ P D+ + ++++ R ++ L SS IR L + PEEIRD V
Sbjct: 175 RLVPVHYDETEAEREKKRLERAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHV 228
Query: 236 ERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTL----P 291
R + + R EE + R+ ++K E+ R K + L LT F +I L P
Sbjct: 229 TRQSQEDQHRINYEESMMVRLSVSKREKGRRKRANVMSSQLHSLTH--FSDISALTGGTP 286
Query: 292 FEDE----TGEQVMGPSKGSNRNGRLKKR 316
DE T ++ P KG + G ++R
Sbjct: 287 HLDEDQNPTKKRKKIPKKGRKKKGFRRRR 315
>gb|AAH23770.1| 1500001L15Rik protein [Mus musculus]
Length = 317
Score = 130 bits (328), Expect = 4e-29
Identities = 95/310 (30%), Positives = 144/310 (45%), Gaps = 17/310 (5%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + S LLK ++E + V +IQ+LT KV+ G Y T G S+LE K+ LLL Y L
Sbjct: 10 ESDVSSSITLLKNLQEQVMAVTAQIQALTTKVRAGTYSTEKGLSFLEVKDQLLLMYLMDL 69
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMK------VGENAA 128
+ +L KA G S++ HP V +VEIR LEK+RP+D+K +YQI KL+K + EN
Sbjct: 70 SHLILDKASGASLQGHPAVLRLVEIRTVLEKLRPLDQKLKYQIDKLVKTAVTGSLSENDP 129
Query: 129 TSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTS 188
P V+ SED + D + + + Y P + P D+ +
Sbjct: 130 LRFKPHPSNMVSKLSSEDEEESEAEEDQSEASGKKSAKGSAKKYVPPRLVPVHYDETEAE 189
Query: 189 KQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQ 248
++++ + ++ L SS IR L + PEEIRD V R + + R
Sbjct: 190 REQKRLEKAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHVTRQSQEDQHRVNY 243
Query: 249 EEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSN 308
EE + R+ ++K E+ + + L LT F +I L ++ P K
Sbjct: 244 EESMMVRLSVSKREKGLRRRASAMSSQLHSLTH--FSDISALTGGTAHLDEDQNPVKKRK 301
Query: 309 ---RNGRLKK 315
+ GR KK
Sbjct: 302 KLPKKGRKKK 311
>gb|AAH30817.1| C14orf120 protein [Homo sapiens] gi|51471747|ref|XP_033371.4|
PREDICTED: chromosome 14 open reading frame 120 [Homo
sapiens] gi|46395905|sp|Q8NEJ9|CN120_HUMAN Protein
C14orf120
Length = 315
Score = 130 bits (326), Expect = 7e-29
Identities = 98/329 (29%), Positives = 152/329 (45%), Gaps = 48/329 (14%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + LLK ++E + V +++SLT KV+ G YPT G S+LE K+ LLL Y L
Sbjct: 8 ESDLPSAVTLLKNLQEQVMAVTAQVKSLTQKVQAGAYPTEKGLSFLEVKDQLLLMYLMDL 67
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
+ +L KA G S++ H V +VEIR LEK+RP+D+K +YQI KL+K + S+
Sbjct: 68 THLILDKASGGSLQGHDAVLRLVEIRTVLEKLRPLDQKLKYQIDKLIKTAVTGSLSE--- 124
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSR---EDLAPQEGHD----------------AYRPV 175
D +++P+P ++S+ ED E D Y P
Sbjct: 125 ----------NDPLRFKPHPSNMMSKLSSEDEEEDEAEDDQSEASGKKSVKGVSKKYVPP 174
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEV 235
+ P D+ + ++++ R ++ L SS IR L + PEEIRD V
Sbjct: 175 RLVPVHYDETEAEREKKRLERAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHV 228
Query: 236 ERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLP---- 291
R + + R EE + R+ ++K E+ R K + L LT F +I L
Sbjct: 229 TRQSQEDQHRINYEESMMVRLSVSKREKGRRKRANVMSSQLHSLTH--FSDISALTGGTV 286
Query: 292 --FEDET--GEQVMGPSKGSNRNGRLKKR 316
ED+ ++ P KG + G ++R
Sbjct: 287 HLDEDQNPIKKRKKIPQKGRKKKGFRRRR 315
>ref|XP_224181.2| PREDICTED: similar to Protein C14orf120 homolog [Rattus norvegicus]
Length = 315
Score = 129 bits (325), Expect = 9e-29
Identities = 95/310 (30%), Positives = 144/310 (45%), Gaps = 17/310 (5%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E E S LLK ++E + V ++Q+LTAKV+ G Y T G S LE K+ LLL Y L
Sbjct: 8 ESEVSSSITLLKNLQEQVMAVTAQVQALTAKVRAGAYSTEKGLSLLEVKDQLLLMYLMDL 67
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMK------VGENAA 128
+ +L KA G S++ HP V +VEIR LEK+RP+D+K +YQI KL+K + EN
Sbjct: 68 SHLILDKASGGSLQGHPAVLRLVEIRTVLEKLRPLDQKLKYQIDKLVKTAVTGSLSENDP 127
Query: 129 TSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTS 188
P ++ SED + D + + + Y P + P D+ +
Sbjct: 128 LRFKPHPSNMMSKLSSEDEEENEAEEDQSEASGKKSVKGASKKYVPPRLVPVHYDETEAE 187
Query: 189 KQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQ 248
++++ + ++ L SS IR L + PEEIRD V R + + R
Sbjct: 188 REQKRLEKAKRRALS----SSVIRELKEQYSDAPEEIRDVR--HPHVTRQSQEDQHRINY 241
Query: 249 EEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSN 308
EE + R+ ++K E+ + + L LT F +I L ++ P K
Sbjct: 242 EESMMVRLSVSKREKGLRRRASAMSSQLHSLTH--FSDISALTGGTAHLDEDQNPVKKRK 299
Query: 309 ---RNGRLKK 315
+ GR KK
Sbjct: 300 KMPKKGRKKK 309
>ref|NP_081166.1| hypothetical protein LOC68966 [Mus musculus]
gi|46395974|sp|Q9DB96|CN120_MOUSE Protein C14orf120
homolog gi|31127183|gb|AAH52790.1| RIKEN cDNA 1500001L15
[Mus musculus] gi|12836794|dbj|BAB23816.1| unnamed
protein product [Mus musculus]
Length = 315
Score = 129 bits (325), Expect = 9e-29
Identities = 96/323 (29%), Positives = 149/323 (45%), Gaps = 43/323 (13%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + S LLK ++E + V +IQ+LT KV+ G Y T G S+LE K+ LLL Y L
Sbjct: 8 ESDVSSSITLLKNLQEQVMAVTAQIQALTTKVRAGTYSTEKGLSFLEVKDQLLLMYLMDL 67
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
+ +L KA G S++ HP V +VEIR LEK+RP+D+K +YQI KL+K + S+
Sbjct: 68 SHLILDKASGASLQGHPAVLRLVEIRTVLEKLRPLDQKLKYQIDKLVKTAVTGSLSE--- 124
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSR------EDLAPQEGHD-------------AYRPV 175
D +++P+P +VS+ E+ +EG Y P
Sbjct: 125 ----------NDPLRFKPHPSNMVSKLSSEDEEESEAEEGQSEASGKKSAKGSAKKYVPP 174
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEV 235
+ P D+ + ++++ + ++ L SS IR L + PEEIRD V
Sbjct: 175 RLVPVHYDETEAEREQKRLEKAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHV 228
Query: 236 ERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDE 295
R + + R EE + R+ ++K E+ + + L LT F +I L
Sbjct: 229 TRQSQEDQHRVNYEESMMVRLSVSKREKGLRRRASAMSSQLHSLTH--FSDISALTGGTA 286
Query: 296 TGEQVMGPSKGSN---RNGRLKK 315
++ P K + GR KK
Sbjct: 287 HLDEDQNPVKKRKKLPKKGRKKK 309
>emb|CAB43232.2| hypothetical protein [Homo sapiens]
Length = 311
Score = 129 bits (325), Expect = 9e-29
Identities = 91/295 (30%), Positives = 139/295 (46%), Gaps = 40/295 (13%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + LLK ++E + V +++SLT KV+ G YPT G S+LE K+ LLL Y L
Sbjct: 8 ESDLPSAVTLLKNLQEQVMAVTAQVKSLTQKVQAGAYPTEKGLSFLEVKDQLLLMYLMDL 67
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
+ +L KA G S++ H V +VEIR LEK+RP+D+K +YQI KL+K + S+
Sbjct: 68 THLILDKASGGSLQGHDAVLRLVEIRTVLEKLRPLDQKLKYQIDKLIKTAVTGSLSE--- 124
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSR---EDLAPQEGHD----------------AYRPV 175
D +++P+P ++S+ ED E D Y P
Sbjct: 125 ----------NDPLRFKPHPSNMMSKLSSEDEEEDEAEDDQSEASGKKSVKGVSKKYVPP 174
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEV 235
+ P D+ + ++++ R ++ L SS IR L + PEEIRD V
Sbjct: 175 RLVPVHYDETEAEREKKRLERAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHV 228
Query: 236 ERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTL 290
R + + R EE + R+ ++K E+ R K + L LT F +I L
Sbjct: 229 TRQSQEDQHRINYEESMMVRLSVSKREKGRRKRANVMSSQLHSLTH--FSDISAL 281
>gb|AAH87522.1| LOC496169 protein [Xenopus laevis]
Length = 316
Score = 129 bits (324), Expect = 1e-28
Identities = 97/313 (30%), Positives = 153/313 (47%), Gaps = 40/313 (12%)
Query: 24 LLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVYYLLRKAK 83
L +++ + V +Q LT KV+ Y T G S+LE K+ LLL Y Q L + +L K
Sbjct: 17 LFNTLQDQITKVTAHVQDLTQKVRSSIYNTDKGLSFLELKDQLLLFYLQDLTHLMLEKTN 76
Query: 84 GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDK 143
G SI+ +P + +VE+R LEK+RPID+K +YQI KL+K AA + + +P+
Sbjct: 77 GKSIKGNPGILRLVELRTVLEKMRPIDQKLKYQIDKLVK----AAVTGSLGENDPL---- 128
Query: 144 SEDVSKYRPNPDMLVS-------REDLAPQEGHDA------------YRPVKFAPTSMDQ 184
+++PNP L+S RE + +EG + Y P + AP D
Sbjct: 129 -----RFKPNPQNLMSKLSEPDERESDSGEEGAEGGVAKKPQSKVKRYIPPRLAPVHYDD 183
Query: 185 DKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEE 244
+ ++ R R +K L SS IR L + PEEIR EG + + RH + +
Sbjct: 184 TEAEREHRIVERAKKLALS----SSTIRELKEQYSDAPEEIR--EGRAYHMMRHDKEEQH 237
Query: 245 RAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPS 304
R EE + R+ +T+ E+ R+K + + L+ LT F +I L + E ++
Sbjct: 238 RINHEESMMVRLNMTRKEKARKKRVLSMTSQLNSLTH--FSDISALTGGEGRAEDMVPFM 295
Query: 305 KGSNRNGRLKKRK 317
K S + + K+K
Sbjct: 296 KKSKKGPKKSKKK 308
>ref|XP_509857.1| PREDICTED: similar to hypothetical protein DKFZp564O092.1 - human
(fragment) [Pan troglodytes]
Length = 452
Score = 129 bits (323), Expect = 1e-28
Identities = 88/285 (30%), Positives = 135/285 (46%), Gaps = 38/285 (13%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + LLK ++E + V +++SLT KV+ G YPT G S+LE K+ LLL Y L
Sbjct: 33 ESDLPSAVTLLKNLQEQVMAVTAQVKSLTQKVQAGAYPTEKGLSFLEVKDQLLLMYLMDL 92
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
+ +L KA G S++ H V +VEIR LEK+RP+D+K +YQI KL+K + S+
Sbjct: 93 THLILDKASGGSLQGHDAVLRLVEIRTVLEKLRPLDQKLKYQIDKLIKTAVTGSLSE--- 149
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSR---EDLAPQEGHDA----------------YRPV 175
D +++P+P ++S+ ED E D Y P
Sbjct: 150 ----------NDPLRFKPHPSNMMSKLSSEDEEEDEAEDGQSEASGQKSVKGVSKKYVPP 199
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEV 235
+ P D+ + ++++ R ++ L SS IR L + PEEIRD V
Sbjct: 200 RLVPVHYDETEAEREKKRLERAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHV 253
Query: 236 ERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLT 280
R + + R EE + R+ ++K E+ R K + L LT
Sbjct: 254 TRQSQEDQHRINYEESMMVRLSVSKREKGRRKRANVMSSQLHSLT 298
>gb|EAL34296.1| GA10717-PA [Drosophila pseudoobscura]
Length = 305
Score = 124 bits (310), Expect = 5e-27
Identities = 101/305 (33%), Positives = 156/305 (51%), Gaps = 34/305 (11%)
Query: 24 LLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVYYLLRKAK 83
LL EM + V ++ + +VK G+ T G S+LE K +LL+Y +L Y +LRK
Sbjct: 22 LLGEMNSNVKQVTDLVEGMLQRVKRGELTTEYGLSFLEVKYHMLLDYLINLTYVVLRKCS 81
Query: 84 GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDK 143
G +IE P + ++EIR LEKIRPID K +YQI KL+K AT+ + S +P+
Sbjct: 82 GETIEGDPSIERLIEIRTVLEKIRPIDHKLRYQIDKLVK----TATTGVSSSTDPIL--- 134
Query: 144 SEDVSKYRPNPDMLVSREDLA--PQEGHDA-----YRPVKFAPTSMDQD-KTSKQERNAL 195
Y+PNPD ++SR A P++ A Y P + P D D K + +E+ AL
Sbjct: 135 ------YKPNPDEMMSRAGAAKKPRKAATAGKSGVYVPPRIKPVYYDGDEKDADKEKKAL 188
Query: 196 RREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNR 255
R K K + SS ++ L + + P EI GS + A+ E++ ++E L R
Sbjct: 189 DRAK---KRAITSSMLQDLKEEYLDAPTEIS--SGSRAQQILSNAQKEKQEYEETYLM-R 242
Query: 256 VPLTKHERKREKYLKKSRNGLDGLTESFFDEI--KTLPFEDETGEQVMGPSKG-SNRNGR 312
+P+TK E+ R++ L L L + EI ++ D + ++ + S+G + G
Sbjct: 243 LPVTKAEKHRQRKL----TTLGTLGDEILGEISRESALRGDGSKKRKLAKSRGKGKKKGS 298
Query: 313 LKKRK 317
KKRK
Sbjct: 299 GKKRK 303
>ref|XP_646440.1| hypothetical protein DDB0190708 [Dictyostelium discoideum]
gi|60474399|gb|EAL72336.1| hypothetical protein
DDB0190708 [Dictyostelium discoideum]
Length = 687
Score = 119 bits (297), Expect = 2e-25
Identities = 101/316 (31%), Positives = 157/316 (48%), Gaps = 36/316 (11%)
Query: 8 LNNHDSKEK------EASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLE 61
++N KEK E+ L LL++ K ++ V+ I KVK Q PT+ G S+LE
Sbjct: 208 ISNFTKKEKLQYLITESPLLLDLLEDFKVKMNEVKTSILPALEKVKSNQLPTSKGISFLE 267
Query: 62 AKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLM 121
K LLL+YC ++ Y+L+ K+ G SI++HPV+ +++ R +EKI+P+DKK +YQI KL+
Sbjct: 268 TKYQLLLSYCLNITYFLMLKSSGVSIKDHPVIDQLIKCRTMIEKIQPLDKKLKYQIDKLL 327
Query: 122 KVGENAATSDLPSKKEPVA-----------SDKSEDVSKYRPNPDMLVSREDLAPQEGHD 170
K N T SK +P+ D+ ED + N D +A + G
Sbjct: 328 K-SANTGTLVGSSKDDPLQHRPNLSSMGGDDDEDEDDEQGLDN-DYDDENSRIAQRAG-- 383
Query: 171 AYRPVKFAPTS---MDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRD 227
Y+ K +DQ+ ++++ R+K+ + K + I + D KPEE D
Sbjct: 384 VYQAPKLYGAKGGVIDQEDAIAKKKSKEIRDKQKSIKGKMAKEIEEIYGD---KPEEEDD 440
Query: 228 Y-----EGSSKEVERHIAKMEERAWQ--EEELFNRVPLTKHERKREK-YLKKSRNGLDGL 279
Y GSS + +ERA + EE + RV L+K E+K K KK +GL+ L
Sbjct: 441 YLDGSGGGSSSRYHGDDEEDDERALKNYEESNYTRVMLSKKEQKAMKSKQKKLSDGLNDL 500
Query: 280 TESFFDEIKTLPFEDE 295
+ F D + ED+
Sbjct: 501 AD-FSDLTSLMENEDD 515
>gb|AAN71280.1| RE04093p [Drosophila melanogaster]
Length = 332
Score = 114 bits (284), Expect = 5e-24
Identities = 98/332 (29%), Positives = 151/332 (44%), Gaps = 63/332 (18%)
Query: 24 LLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVYYLLRKAK 83
LL EM + V ++ + +VK G+ T G S+LE K +LL+Y +L Y +LRK
Sbjct: 22 LLGEMNSNVKQVTDLVEGMLQRVKRGELTTEYGLSFLEVKYHMLLDYLINLTYVVLRKCS 81
Query: 84 GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDK 143
G +IE P + ++EIR LEKIRPID K +YQI KL+K AT+ + S +P+
Sbjct: 82 GETIEGDPSIERLIEIRTVLEKIRPIDHKLRYQIDKLVKT----ATTGVSSSTDPIL--- 134
Query: 144 SEDVSKYRPNPDMLVSR-----------EDLAPQEGHD---------------------- 170
Y+PNPD ++S ED + +E D
Sbjct: 135 ------YKPNPDEMMSSAGGAGRDEDDGEDDSDEEDEDDDEEDEDEAGAAKMPRKAATAG 188
Query: 171 ---AYRPVKFAPTSMDQD-KTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIR 226
Y P + P D D + + +E+ AL R K K + SS ++ L + + P EI
Sbjct: 189 KSGVYVPPRIKPVYYDGDERDADKEKKALDRAK---KRAITSSMLQDLKEEYLDAPTEIS 245
Query: 227 DYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDE 286
GS + A+ E++ ++E L R+P+TK E+ R++ L L L + E
Sbjct: 246 S--GSRAQQMLSQAQKEKQEYEETYLM-RLPVTKAEKHRQRKL----TTLGTLGDEILGE 298
Query: 287 I---KTLPFEDETGEQVMGPSKGSNRNGRLKK 315
I L + ++ KG R G+ +K
Sbjct: 299 ISRESALRGDGSKKRKLAKKGKGKKRGGKKRK 330
>ref|NP_608956.1| CG11030-PA [Drosophila melanogaster] gi|22945693|gb|AAF52286.2|
CG11030-PA [Drosophila melanogaster]
Length = 332
Score = 113 bits (283), Expect = 6e-24
Identities = 98/332 (29%), Positives = 151/332 (44%), Gaps = 63/332 (18%)
Query: 24 LLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVYYLLRKAK 83
LL EM + V ++ + +VK G+ T G S+LE K +LL+Y +L Y +LRK
Sbjct: 22 LLGEMNSNVKQVTDLVEGMLQRVKRGELTTEYGLSFLEVKYHMLLDYLINLTYVVLRKCS 81
Query: 84 GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDK 143
G +IE P + ++EIR LEKIRPID K +YQI KL+K AT+ + S +P+
Sbjct: 82 GETIEGDPSIERLIEIRTVLEKIRPIDHKLRYQIDKLVKT----ATTGVSSSTDPIL--- 134
Query: 144 SEDVSKYRPNPDMLVSR-----------EDLAPQEGHD---------------------- 170
Y+PNPD ++S ED + +E D
Sbjct: 135 ------YKPNPDDMMSSAGGAGRDEDDGEDDSDEEDEDDDEEDEDEAGAAKMPRKAATAG 188
Query: 171 ---AYRPVKFAPTSMDQD-KTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIR 226
Y P + P D D + + +E+ AL R K K + SS ++ L + + P EI
Sbjct: 189 KSGVYVPPRIKPVYYDGDERDADKEKKALDRAK---KRAITSSMLQDLKEEYLDAPTEIS 245
Query: 227 DYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDE 286
GS + A+ E++ ++E L R+P+TK E+ R++ L L L + E
Sbjct: 246 S--GSRAQQMLSQAQKEKQEYEETYLM-RLPVTKAEKHRQRKL----TTLGTLGDEILGE 298
Query: 287 I---KTLPFEDETGEQVMGPSKGSNRNGRLKK 315
I L + ++ KG R G+ +K
Sbjct: 299 ISRESALRGDGSKKRKLAKKGKGKKRGGKKRK 330
>ref|XP_666616.1| similar to CG11030-PA [Cryptosporidium hominis]
gi|54657645|gb|EAL36386.1| similar to CG11030-PA
[Cryptosporidium hominis]
Length = 692
Score = 112 bits (280), Expect = 1e-23
Identities = 92/297 (30%), Positives = 145/297 (47%), Gaps = 35/297 (11%)
Query: 12 DSKEKEASPLAALLKEMKEGLDVVRHKIQSLT--AKVKEGQ-YPTADGFSYLEAKNLLLL 68
D E S LA +L +KE + V+ ++Q L K KEG T G YL++KN LLL
Sbjct: 184 DGIEGLDSELAPILTSIKEKVTEVKERMQVLLNLVKTKEGSGLVTEKGMEYLDSKNTLLL 243
Query: 69 NYCQSLVYYLLRKAK-GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENA 127
Y L YY++ K +I+EHP++ +V +R +EK++PID K Q QI +++++ E +
Sbjct: 244 MYIGYLCYYMMLKTSPDINIKEHPILLRLVTLRTMMEKLKPIDVKLQPQIDRILELAEKS 303
Query: 128 ATSD--LPSKKEP---VASDKSEDVSKYRPN------------------PDMLVSREDLA 164
+ D L S P V D+ D S N D+ V+ ED
Sbjct: 304 SQVDNFLSSAPRPDRFVFEDEDNDNSDIEENDIFKHDLRDEFDESTNIDSDVEVTEED-- 361
Query: 165 PQEGHDAYRPVKFAPTSMDQDKTSKQER--NALRREKEILKESKHSSFIRTLMNDIEEKP 222
G Y+ K P + K SK E+ L RE++ L + IR + + I E P
Sbjct: 362 SDGGKGVYKAPKNLPVEFNDKKLSKTEKMMKELERERQRL---LRTDIIRQMRSSIHEGP 418
Query: 223 EEI-RDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDG 278
EE+ R+ ++ER +++ER EE+ R+P TK +++ EK K+ + ++G
Sbjct: 419 EEVGREETEQLPQLERLQRQIKERINFEEDNMMRLPKTKKDKREEKLYKRLMSQVEG 475
>gb|EAK88334.1| conserved eukaryotic nuclear protein that shares a domain with
yeast Lcp5p, a component of the U3 small nucleolar
ribonucleoprotein [Cryptosporidium parvum]
gi|66362330|ref|XP_628129.1| conserved eukaryotic
nuclear protein that shares a domain with yeast Lcp5p
[Cryptosporidium parvum]
Length = 692
Score = 112 bits (279), Expect = 2e-23
Identities = 91/298 (30%), Positives = 145/298 (48%), Gaps = 37/298 (12%)
Query: 12 DSKEKEASPLAALLKEMKEGLDVVRHKIQSLT--AKVKEGQ-YPTADGFSYLEAKNLLLL 68
D E S LA +L +KE + V+ ++Q L K KEG T G YL++KN LLL
Sbjct: 184 DGIEGLDSELAPILTSIKEKVTEVKERMQVLLNLVKTKEGSGLVTEKGMEYLDSKNTLLL 243
Query: 69 NYCQSLVYYLLRKAK-GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENA 127
Y L YY++ K +I+EHP++ +V +R +EK++PID K Q QI +++++ E +
Sbjct: 244 MYIGYLCYYMMLKTSPDINIKEHPILLRLVTLRTMMEKLKPIDVKLQPQIDRILELAEKS 303
Query: 128 ATSD--LPSKKEP---VASDKSEDVSKYRPNPDMLVSREDLAPQ---------------- 166
+ D L S P V D+ D S N +S+ DL +
Sbjct: 304 SQVDNFLSSAPRPDRFVFEDEDNDNSDIEENE---ISKHDLGDEFDESTNIDSDVELTEE 360
Query: 167 ---EGHDAYRPVKFAPTSMDQDKTSKQER--NALRREKEILKESKHSSFIRTLMNDIEEK 221
G Y+ K P + K SK E+ L RE++ L + IR + + I E
Sbjct: 361 DSDGGKGVYKAPKNLPVEFNDKKLSKTEKMMKELERERQRL---LRTDIIRQMRSSIHEG 417
Query: 222 PEEI-RDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDG 278
PEE+ R+ ++ER +++ER EE+ R+P TK +++ EK K+ + ++G
Sbjct: 418 PEEVGREEAEQLPQLERLQRQIKERINFEEDNMMRLPKTKKDKREEKLYKRLMSQVEG 475
>emb|CAG12701.1| unnamed protein product [Tetraodon nigroviridis]
Length = 311
Score = 108 bits (270), Expect = 2e-22
Identities = 80/273 (29%), Positives = 124/273 (45%), Gaps = 30/273 (10%)
Query: 8 LNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLL 67
L ++D EK+ LL + E + V ++ L +VK+G + T+ G S+L+ + LL
Sbjct: 4 LVDNDLIEKDLPKCVQLLNALTEQVAAVTAHVRELLRQVKDGAFQTSKGLSFLDLRYHLL 63
Query: 68 LNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENA 127
L Y Q L + + KA+G I++ + IV +R LEK+RPID K +YQI KL++
Sbjct: 64 LFYLQDLTHLISIKAEGQKIKDSDALNRIVTVRTVLEKMRPIDHKLKYQIDKLVRTAVTG 123
Query: 128 ATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSR-------------EDLAPQEGHDAYRP 174
A ++ +D + RPNP+ L+S+ E A G Y P
Sbjct: 124 ALAE-------------DDPLQLRPNPENLISKLSESEESEAEEQAEKKAAPSGGRKYIP 170
Query: 175 VKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKE 234
K AP D D T + A + + + SS I+ L + PEEIR E +
Sbjct: 171 PKIAPVHYDGDMTDADRKKAQAERQR--RAALRSSVIQELRQQYSDAPEEIR--EKRDFQ 226
Query: 235 VERHIAKMEERAWQEEELFNRVPLTKHERKREK 267
ER + R EE + R+ + K + K
Sbjct: 227 TERDSREELHRKNYEESMMVRLSVPKRAKNARK 259
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.311 0.130 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 536,480,631
Number of Sequences: 2540612
Number of extensions: 23140052
Number of successful extensions: 94509
Number of sequences better than 10.0: 1332
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 1290
Number of HSP's that attempted gapping in prelim test: 90464
Number of HSP's gapped (non-prelim): 3497
length of query: 318
length of database: 863,360,394
effective HSP length: 128
effective length of query: 190
effective length of database: 538,162,058
effective search space: 102250791020
effective search space used: 102250791020
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0316.5


