

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.4
(147 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
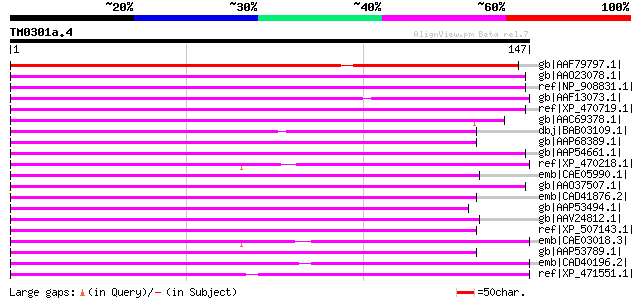
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF79797.1| T32E20.30 [Arabidopsis thaliana] 121 4e-27
gb|AAO23078.1| polyprotein [Glycine max] 107 5e-23
ref|NP_908831.1| putative polyprotein [Oryza sativa (japonica cu... 103 7e-22
gb|AAF13073.1| putative retroelement pol polyprotein [Arabidopsi... 102 2e-21
ref|XP_470719.1| putative polyprotein [Oryza sativa] gi|18921316... 101 4e-21
gb|AAC69378.1| putative retroelement pol polyprotein [Arabidopsi... 100 9e-21
dbj|BAB03109.1| retroelement pol polyprotein [Arabidopsis thalia... 99 2e-20
gb|AAP68389.1| putative polyprotein [Oryza sativa (japonica cult... 99 3e-20
gb|AAP54661.1| putative plant disease resistance polyprotein [Or... 99 3e-20
ref|XP_470218.1| Putative retroelement [Oryza sativa] gi|1545160... 99 3e-20
emb|CAE05990.1| OSJNBa0004L19.22 [Oryza sativa (japonica cultiva... 98 4e-20
gb|AAO37507.1| retrotransposon protein, putative, unclassified [... 97 6e-20
emb|CAD41876.2| OSJNBa0041A02.23 [Oryza sativa (japonica cultiva... 97 6e-20
gb|AAP53494.1| putative retroelement [Oryza sativa (japonica cul... 97 8e-20
gb|AAV24812.1| putative polyprotein [Oryza sativa (japonica cult... 96 1e-19
ref|XP_507143.1| PREDICTED OJ1449_C01.20 gene product [Oryza sat... 96 2e-19
emb|CAE03018.3| OSJNBa0091D06.3 [Oryza sativa (japonica cultivar... 96 2e-19
gb|AAP53789.1| hypothetical protein [Oryza sativa (japonica cult... 95 4e-19
emb|CAD40196.2| OSJNBb0043H09.2 [Oryza sativa (japonica cultivar... 94 5e-19
ref|XP_471551.1| OSJNBa0019J05.12 [Oryza sativa (japonica cultiv... 94 7e-19
>gb|AAF79797.1| T32E20.30 [Arabidopsis thaliana]
Length = 1397
Score = 121 bits (303), Expect = 4e-27
Identities = 60/144 (41%), Positives = 90/144 (61%), Gaps = 3/144 (2%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL R+NSV R C +L A Y+GP+ I++RIG VAYRL+LPEG ++H VFH S LK
Sbjct: 1249 VFLKLRPYRQNSVTKRVCQKLAAKYFGPFEIMERIGKVAYRLKLPEGSKIHLVFHVSQLK 1308
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+ +G++ + L + LT + V P +V+ +R+ E L + + WQG P E TW+
Sbjct: 1309 QVLGDHHQVIPLPEVLTADNEFVVVPEAVLETRY---NEDGLLEALVHWQGLPVHEDTWE 1365
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGG 144
+++ QFP L+DK+ + GG
Sbjct: 1366 IAKDLKKQFPGLALQDKLHVEGGG 1389
>gb|AAO23078.1| polyprotein [Glycine max]
Length = 1552
Score = 107 bits (268), Expect = 5e-23
Identities = 54/146 (36%), Positives = 80/146 (53%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V +KL R++S V R +L+ Y+GP+ ++ +IG VAY+L+LP R+HPVFH S LK
Sbjct: 1358 VLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLELPSAARIHPVFHVSQLK 1417
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
G L E + P ++ SR R + + Q+ +QW+ DE TW+
Sbjct: 1418 PFNGTAQDPYLPLPLTVTEMGPVMQPVKILASRIIIRGHNQIEQILVQWENGLQDEATWE 1477
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIV 146
D +I++ +P FNLEDKV G V
Sbjct: 1478 DIEDIKASYPTFNLEDKVVFKGEGNV 1503
>ref|NP_908831.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1522
Score = 103 bits (258), Expect = 7e-22
Identities = 52/146 (35%), Positives = 82/146 (55%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL + SVV R C +L Y+GP+PI++R+G+VAY+LQLP +VH VFH S LK
Sbjct: 1370 VFLKLQPYAQTSVVNRPCHKLAMKYFGPFPILERVGSVAYKLQLPPASQVHSVFHVSQLK 1429
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+V ++ L + +V + P ++ R + + + QV ++W P + TW+
Sbjct: 1430 PSVPSHVPVYTSLPDVVALDVEDLIPTEILDCRMVKKGNAAIVQVRVRWGSLPDNMATWE 1489
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIV 146
D IR++FP + + AGG V
Sbjct: 1490 DYDVIRTRFPAVSALGQAGSPAGGTV 1515
>gb|AAF13073.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1661
Score = 102 bits (254), Expect = 2e-21
Identities = 57/147 (38%), Positives = 82/147 (55%), Gaps = 2/147 (1%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL R++SV R +L+ Y+GP+ ++ RIG VAY+LQLPE +HPVFH S LK
Sbjct: 1483 VYLKLRPYRQSSVAHRKNEKLSQRYFGPFKVLHRIGQVAYKLQLPEHSTIHPVFHVSQLK 1542
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
AV + +L L+ + P ++ R + P+V +QW G E TW+
Sbjct: 1543 RAVPPSFTPQELPKILSPTLEWNTGPEKLLDIRQSNTNSG--PEVLVQWSGLSTLESTWE 1600
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIVR 147
L + Q+P F+LEDKV L G I R
Sbjct: 1601 PLLTLVQQYPDFDLEDKVSLLRGSIDR 1627
>ref|XP_470719.1| putative polyprotein [Oryza sativa] gi|18921316|gb|AAL82521.1|
putative polyprotein [Oryza sativa]
Length = 408
Score = 101 bits (251), Expect = 4e-21
Identities = 53/146 (36%), Positives = 81/146 (55%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL ++SVV R P+L ++GP+ I+++IG AYRLQLP+ +HPVFH S LK
Sbjct: 257 VFLKLQPYTQSSVVNRPYPKLAFKFFGPFEILEKIGQSAYRLQLPDSSLIHPVFHVSQLK 316
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
V +++ L H + A V P ++ R + +T QV I+W PA + TW+
Sbjct: 317 AHVPDHTPVFTSLPHPVVLDAADVWPEEILDRRLVKKGNATHVQVLIKWSSLPAHQATWE 376
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIV 146
D ++++FP + AGG V
Sbjct: 377 DYEVLKTRFPSAPAWGQAVSPAGGTV 402
>gb|AAC69378.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25411577|pir||F84519 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 945
Score = 100 bits (248), Expect = 9e-21
Identities = 49/142 (34%), Positives = 82/142 (57%), Gaps = 2/142 (1%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V +++ R+ ++ R +L+ +YGP+ + + G VAYRL LPEG R+HPVFH SLLK
Sbjct: 804 VLLRIQPYRQKTLFRRSSQKLSHRFYGPFQVASKHGEVAYRLTLPEGTRIHPVFHVSLLK 863
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
VG+ ++ L L + P +V+ R+ ++ + + + +QW+G ++ TW+
Sbjct: 864 PWVGDGEPDMGQLPPLRNNGELKLQPTAVLEVRWRSQDKKRVADLLVQWEGLHIEDATWE 923
Query: 121 DTLNIRSQFP--VFNLEDKVDL 140
+ + + FP V NLEDKV L
Sbjct: 924 EYDQLAASFPEFVLNLEDKVRL 945
>dbj|BAB03109.1| retroelement pol polyprotein [Arabidopsis thaliana]
gi|12322008|gb|AAG51046.1| gypsy/Ty-3 retroelement
polyprotein; 69905-74404 [Arabidopsis thaliana]
Length = 1499
Score = 99.4 bits (246), Expect = 2e-20
Identities = 51/132 (38%), Positives = 75/132 (56%), Gaps = 2/132 (1%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL R+ SVV R +L+ Y+GPY II R G VAY+L LP +VHPVFH S LK
Sbjct: 1368 VYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPSYSQVHPVFHVSQLK 1427
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
VGN S + L + ++V P V+ + RQ + +V ++W +P +E TW+
Sbjct: 1428 VLVGNVSTTVHLPSVM--QDVFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPLEEATWE 1485
Query: 121 DTLNIRSQFPVF 132
+++ FP F
Sbjct: 1486 FLFDLQKTFPEF 1497
>gb|AAP68389.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50917833|ref|XP_469313.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1246
Score = 98.6 bits (244), Expect = 3e-20
Identities = 46/132 (34%), Positives = 75/132 (55%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++K+ R+N+ R +L A +YGP+ ++++IG VAY+LQLPE +HPVFH S LK
Sbjct: 1113 VYLKMQPYRQNAFGLRGSLKLRAKFYGPFRVLEKIGKVAYKLQLPEEANIHPVFHVSQLK 1172
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+ VG ++ L L + + P +V+ R RQ + Q I W+ ++ TW+
Sbjct: 1173 KHVGRQAIPLPNLPLVGADGTVKTEPLAVLARRVIPRQNEPVVQWLIHWENLTPEDATWE 1232
Query: 121 DTLNIRSQFPVF 132
+ I++ FP F
Sbjct: 1233 NATFIQATFPHF 1244
>gb|AAP54661.1| putative plant disease resistance polyprotein [Oryza sativa
(japonica cultivar-group)] gi|37536144|ref|NP_922374.1|
putative plant disease resistance polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|10122041|gb|AAG13430.1| putative plant disease
resistance polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 921
Score = 98.6 bits (244), Expect = 3e-20
Identities = 52/146 (35%), Positives = 79/146 (53%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL ++SVV R P+L ++GP+ +++RIG AYRL LP VHPVFH S LK
Sbjct: 765 VFLKLQPYAQSSVVNRPYPKLAYKFFGPFTVLERIGVAAYRLDLPASSTVHPVFHVSQLK 824
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
V +++ L + A V P +++ R R + QV ++W PAD TW+
Sbjct: 825 AQVPDHTPVYTSLPSPVVLDAADVVPEAILERRLVKRGNAAHVQVLLKWSLLPADHATWE 884
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIV 146
D ++++FP + + AGG V
Sbjct: 885 DYHVVKTRFPDAPAWGQAETPAGGTV 910
>ref|XP_470218.1| Putative retroelement [Oryza sativa] gi|15451608|gb|AAK98732.1|
Putative retroelement [Oryza sativa]
Length = 1923
Score = 98.6 bits (244), Expect = 3e-20
Identities = 50/148 (33%), Positives = 83/148 (55%), Gaps = 5/148 (3%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL + SV +R C +L +YGP+ I+ R+G VAY+LQLP+ +HPVFH S LK
Sbjct: 843 VFLKLQPYVQKSVASRACHKLAFKFYGPFQILARVGTVAYKLQLPDDSTIHPVFHVSQLK 902
Query: 61 EAVG-NNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTW 119
A G + V+ +L L ++V+P ++ R + T+ QV + W A++ TW
Sbjct: 903 VAHGFKHQVQSRLPKFLK----STVYPLQILDQRLIRKGNRTVSQVLVYWSDSVAEDATW 958
Query: 120 KDTLNIRSQFPVFNLEDKVDLSAGGIVR 147
+D +++ +FP+ + G+V+
Sbjct: 959 EDREDLQQRFPMALAWGQATFQGEGVVK 986
>emb|CAE05990.1| OSJNBa0004L19.22 [Oryza sativa (japonica cultivar-group)]
Length = 1586
Score = 98.2 bits (243), Expect = 4e-20
Identities = 47/133 (35%), Positives = 74/133 (55%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL R+ + R +L + +YGP+ I++++G VAY+LQLPEG +HPVFH S LK
Sbjct: 1453 VYLKLQPYRQTAFGIRGSLKLRSKFYGPFKIMEKVGRVAYKLQLPEGSNIHPVFHVSQLK 1512
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+ +G+ +V + L + + P +V+ R R + Q + W E TW+
Sbjct: 1513 KHIGSRAVPMANLPSVGPDGQIKTEPVAVLKRRMIPRGGVAVTQWLVLWHNLSPSEATWE 1572
Query: 121 DTLNIRSQFPVFN 133
D I+S FP FN
Sbjct: 1573 DASMIQSMFPSFN 1585
>gb|AAO37507.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)] gi|50916505|ref|XP_468649.1|
putative polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1155
Score = 97.4 bits (241), Expect = 6e-20
Identities = 52/146 (35%), Positives = 78/146 (52%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL ++SVV R P+L ++GP+ +++RIG AYRL LP VHPVFH S LK
Sbjct: 999 VFLKLQPYAQSSVVNRPYPKLAYKFFGPFTVLERIGVAAYRLDLPASSTVHPVFHVSQLK 1058
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
V +++ L + A V P +++ R R + QV ++W PAD TW+
Sbjct: 1059 AQVPDHTPVYTSLPSPVVLDAADVVPEAILERRLVKRGNAAHVQVLLKWSLLPADHATWE 1118
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIV 146
D ++++FP + AGG V
Sbjct: 1119 DYHVVKTRFPDAPAWGQAKTPAGGTV 1144
>emb|CAD41876.2| OSJNBa0041A02.23 [Oryza sativa (japonica cultivar-group)]
gi|50928515|ref|XP_473785.1| OSJNBa0041A02.23 [Oryza
sativa (japonica cultivar-group)]
Length = 1373
Score = 97.4 bits (241), Expect = 6e-20
Identities = 49/132 (37%), Positives = 75/132 (56%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++K+ R+N+ R +L A +YGP+ I++RIG VAY LQLPE +HPVFH S LK
Sbjct: 1240 VYLKMQPYRQNAFGLRGSLKLRAKFYGPFRILKRIGKVAYHLQLPEEAGIHPVFHVSQLK 1299
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+ VGN V L L + + P +V+ R R+ + Q + W+ ++ TW+
Sbjct: 1300 KHVGNLVVPLPNLPLVDEDGNIKTEPVAVLARRVVPRRNEPVTQWLVHWENLSPEDATWE 1359
Query: 121 DTLNIRSQFPVF 132
D+ I++ FP F
Sbjct: 1360 DSAFIQATFPHF 1371
>gb|AAP53494.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37533810|ref|NP_921207.1| putative retroelement
[Oryza sativa (japonica cultivar-group)]
gi|21672107|gb|AAM74469.1| Putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|18652523|gb|AAL77156.1| Putative polyprotein [Oryza
sativa]
Length = 999
Score = 97.1 bits (240), Expect = 8e-20
Identities = 45/130 (34%), Positives = 76/130 (57%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL + S+ R C +++ Y+GPY +++R+G VAYRL LP G ++HPV H SLLK
Sbjct: 860 VYLKLQPFAQLSIAQRSCQKISFRYFGPYTVLERVGQVAYRLDLPAGSQIHPVVHVSLLK 919
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
AV +++ L ++ + P V+ R ++ +P IQW G PA+ TW+
Sbjct: 920 AAVKPDTIVSPELPLQITDDSVLLQPEVVLRRSLIKRGKAAVPYGLIQWTGVPAELATWE 979
Query: 121 DTLNIRSQFP 130
+ +++ +FP
Sbjct: 980 NLRHLKLRFP 989
>gb|AAV24812.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1475
Score = 96.3 bits (238), Expect = 1e-19
Identities = 45/133 (33%), Positives = 77/133 (57%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL R+++ R +L + +YGP+ ++Q+IG +AY+LQLP+ ++HPVFH S LK
Sbjct: 1342 VYLKLKPYRQSAFGIRGSLKLRSKFYGPFKVLQKIGQLAYKLQLPDDAQIHPVFHVSQLK 1401
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+ +G +++ + L + + P +V+ R R+ + Q I WQ E TW+
Sbjct: 1402 KHLGKHAIPMSNLPSVGPDGQIKTEPLAVLQRRMVPRKGVAVTQWLILWQNLSPAEATWE 1461
Query: 121 DTLNIRSQFPVFN 133
D I++ FP FN
Sbjct: 1462 DASVIQAMFPSFN 1474
>ref|XP_507143.1| PREDICTED OJ1449_C01.20 gene product [Oryza sativa (japonica
cultivar-group)]
Length = 299
Score = 95.9 bits (237), Expect = 2e-19
Identities = 46/132 (34%), Positives = 76/132 (56%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL R+N++ R +L + +YGP+ + Q+IG VAY+LQLPEG +HPVFH S LK
Sbjct: 166 VYLKLQPYRQNALGVRGSLKLRSKFYGPFRVEQKIGQVAYKLQLPEGANIHPVFHVSQLK 225
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+ +G ++V + L + + P +++ R R+ + Q + W+ + TW+
Sbjct: 226 KHLGRHAVPMPNLPLVGPDGQIKAEPVAILQRRMIPRRGEPVVQWLVLWRNLSPADATWE 285
Query: 121 DTLNIRSQFPVF 132
D IR+ FP F
Sbjct: 286 DATFIRATFPNF 297
>emb|CAE03018.3| OSJNBa0091D06.3 [Oryza sativa (japonica cultivar-group)]
gi|50926772|ref|XP_473325.1| OSJNBa0091D06.3 [Oryza
sativa (japonica cultivar-group)]
Length = 753
Score = 95.9 bits (237), Expect = 2e-19
Identities = 54/148 (36%), Positives = 78/148 (52%), Gaps = 5/148 (3%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL + SV R +L Y+GPY + +IG VAY+LQLP +HPVFH S LK
Sbjct: 313 VYLKLQPYVQTSVAVRANHKLAFKYFGPYQVTAKIGTVAYQLQLPSSTSIHPVFHVSQLK 372
Query: 61 EAVG-NNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTW 119
A+G V+ QL +V P ++ R T R S + QV IQW ++ TW
Sbjct: 373 SAIGFTGPVQHQLPASTAPLQV----PLRILDRRLTKRGNSAVAQVLIQWSASVPEDATW 428
Query: 120 KDTLNIRSQFPVFNLEDKVDLSAGGIVR 147
+D ++RS+FP + + G+VR
Sbjct: 429 EDLDDLRSRFPAALAWGQANSQGEGLVR 456
>gb|AAP53789.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37534400|ref|NP_921502.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1611
Score = 94.7 bits (234), Expect = 4e-19
Identities = 46/132 (34%), Positives = 75/132 (55%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL R+ ++ R +L + YYGP+ +I+++GAVAY+LQLP+G +HPVFH S LK
Sbjct: 1478 VYLKLQPYRQVAMGIRGSLKLRSKYYGPFKVIEKMGAVAYKLQLPDGAGIHPVFHVSQLK 1537
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
+ +G ++ + L + + P +V+ R R + Q I W+ E TW+
Sbjct: 1538 KHLGARAIPMPNLPAIGPDGQIKTEPAAVLQRRMIPRHNEPVTQWLILWENLTPAEATWE 1597
Query: 121 DTLNIRSQFPVF 132
D I++ FP F
Sbjct: 1598 DASYIQAAFPNF 1609
>emb|CAD40196.2| OSJNBb0043H09.2 [Oryza sativa (japonica cultivar-group)]
gi|50921815|ref|XP_471268.1| OSJNBb0043H09.2 [Oryza
sativa (japonica cultivar-group)]
Length = 1409
Score = 94.4 bits (233), Expect = 5e-19
Identities = 52/147 (35%), Positives = 75/147 (50%), Gaps = 3/147 (2%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL + SV R +L Y+GP+ ++ R+GAVAYRLQLP +HPVFH S L+
Sbjct: 1213 VYLKLQPYIQTSVTKRANHKLEFRYFGPFQVLDRVGAVAYRLQLPSYSAIHPVFHVSQLR 1272
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
A G + + L GE P + R T + T+ Q+ W PA+E TW+
Sbjct: 1273 AAAGFSKPVMSQLPSSLGELQI---PLQFLDKRLTKKGNKTVVQLLTHWSDSPAEEATWE 1329
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIVR 147
D +R++FP + GIVR
Sbjct: 1330 DMEELRARFPGALAWGQASHQEWGIVR 1356
>ref|XP_471551.1| OSJNBa0019J05.12 [Oryza sativa (japonica cultivar-group)]
gi|38346427|emb|CAD40214.2| OSJNBa0019J05.12 [Oryza
sativa (japonica cultivar-group)]
Length = 1817
Score = 94.0 bits (232), Expect = 7e-19
Identities = 50/147 (34%), Positives = 75/147 (51%), Gaps = 3/147 (2%)
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
V++KL + SV R +L Y+GP+ I +IG VAY+LQLP +HPVFH S LK
Sbjct: 1477 VYLKLQPYVQTSVAVRANHKLAFKYFGPFQITAQIGTVAYKLQLPPTSSIHPVFHVSQLK 1536
Query: 61 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 120
A G + HL A P+ ++ R + ST+ QV +QW P ++ TW+
Sbjct: 1537 SATGFTG---HVQSHLPSSISALQVPWRILDRRLAKKGNSTISQVLVQWSESPPEDATWE 1593
Query: 121 DTLNIRSQFPVFNLEDKVDLSAGGIVR 147
D + ++FP + + G+VR
Sbjct: 1594 DYEELHARFPAALAWGQANSQGVGLVR 1620
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 261,008,597
Number of Sequences: 2540612
Number of extensions: 10333139
Number of successful extensions: 24512
Number of sequences better than 10.0: 668
Number of HSP's better than 10.0 without gapping: 500
Number of HSP's successfully gapped in prelim test: 168
Number of HSP's that attempted gapping in prelim test: 23457
Number of HSP's gapped (non-prelim): 682
length of query: 147
length of database: 863,360,394
effective HSP length: 123
effective length of query: 24
effective length of database: 550,865,118
effective search space: 13220762832
effective search space used: 13220762832
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0301a.4


