

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0299.14
(88 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
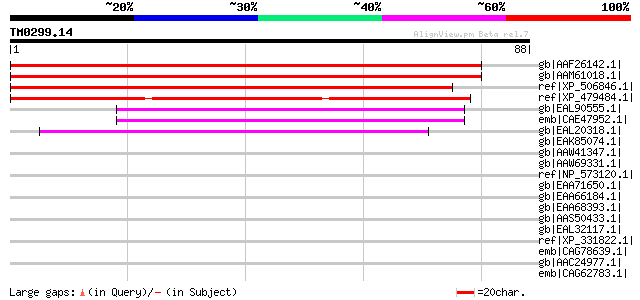
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF26142.1| unknown protein [Arabidopsis thaliana] gi|1523000... 146 1e-34
gb|AAM61018.1| unknown [Arabidopsis thaliana] gi|28972983|gb|AAO... 139 2e-32
ref|XP_506846.1| PREDICTED OSJNBa0016G10.13 gene product [Oryza ... 134 6e-31
ref|XP_479484.1| unknown protein [Oryza sativa (japonica cultiva... 118 3e-26
gb|EAL90555.1| mitochondrial hypoxia responsive domain protein [... 47 1e-04
emb|CAE47952.1| hypothetical protein, conserved [Aspergillus fum... 47 1e-04
gb|EAL20318.1| hypothetical protein CNBF1290 [Cryptococcus neofo... 45 5e-04
gb|EAK85074.1| hypothetical protein UM03929.1 [Ustilago maydis 5... 44 0.001
gb|AAW41347.1| mitochondrion protein, putative [Cryptococcus neo... 42 0.003
gb|AAW69331.1| hypothetical protein [Magnaporthe grisea] gi|3810... 41 0.007
ref|NP_573120.1| CG9921-PA [Drosophila melanogaster] gi|7293210|... 41 0.009
gb|EAA71650.1| hypothetical protein FG03448.1 [Gibberella zeae P... 40 0.019
gb|EAA66184.1| hypothetical protein AN1066.2 [Aspergillus nidula... 39 0.033
gb|EAA68393.1| hypothetical protein FG00663.1 [Gibberella zeae P... 39 0.043
gb|AAS50433.1| AAR068Cp [Ashbya gossypii ATCC 10895] gi|45184891... 38 0.074
gb|EAL32117.1| GA22125-PA [Drosophila pseudoobscura] 38 0.074
ref|XP_331822.1| hypothetical protein [Neurospora crassa] gi|289... 37 0.096
emb|CAG78639.1| unnamed protein product [Yarrowia lipolytica CLI... 37 0.13
gb|AAC24977.1| anaerobically inducible early gene 2 [Oryza sativa] 37 0.16
emb|CAG62783.1| unnamed protein product [Candida glabrata CBS138... 35 0.36
>gb|AAF26142.1| unknown protein [Arabidopsis thaliana]
gi|15230007|ref|NP_187206.1| hypoxia-responsive family
protein [Arabidopsis thaliana]
Length = 97
Score = 146 bits (368), Expect = 1e-34
Identities = 70/80 (87%), Positives = 74/80 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M E+KT+ E IRKWV DHKLRTVGCLWLSGITGSIAYNWS+P MKTSVKIIHARLHAQAL
Sbjct: 1 MVESKTKFEEIRKWVSDHKLRTVGCLWLSGITGSIAYNWSQPAMKTSVKIIHARLHAQAL 60
Query: 61 TLAALAGAAVVEYYDHRAEA 80
TLAALAGAAVVEYYDH+ EA
Sbjct: 61 TLAALAGAAVVEYYDHKTEA 80
>gb|AAM61018.1| unknown [Arabidopsis thaliana] gi|28972983|gb|AAO63816.1| unknown
protein [Arabidopsis thaliana]
gi|28393673|gb|AAO42249.1| unknown protein [Arabidopsis
thaliana] gi|15241035|ref|NP_198128.1|
hypoxia-responsive family protein [Arabidopsis
thaliana]
Length = 96
Score = 139 bits (349), Expect = 2e-32
Identities = 64/80 (80%), Positives = 74/80 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAE KT+V IR+W+++HKLRTVGCLWLSGI+GSIAYNWS+P MKTSV+IIHARLHAQAL
Sbjct: 1 MAEPKTKVAEIREWIIEHKLRTVGCLWLSGISGSIAYNWSKPAMKTSVRIIHARLHAQAL 60
Query: 61 TLAALAGAAVVEYYDHRAEA 80
TLAALAGAA VEYYDH++ A
Sbjct: 61 TLAALAGAAAVEYYDHKSGA 80
>ref|XP_506846.1| PREDICTED OSJNBa0016G10.13 gene product [Oryza sativa (japonica
cultivar-group)] gi|50910007|ref|XP_466492.1|
hypoxia-responsive family protein-like [Oryza sativa
(japonica cultivar-group)] gi|50726476|dbj|BAD34085.1|
hypoxia-responsive family protein-like [Oryza sativa
(japonica cultivar-group)]
Length = 107
Score = 134 bits (337), Expect = 6e-31
Identities = 62/75 (82%), Positives = 70/75 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAEA ++ES+RKWV+DHKLR VGCLWL+GI+ SIAYNWSRPNMKTSVKIIHARLHAQAL
Sbjct: 9 MAEAPGKIESMRKWVIDHKLRAVGCLWLTGISSSIAYNWSRPNMKTSVKIIHARLHAQAL 68
Query: 61 TLAALAGAAVVEYYD 75
TLAAL G+A+VEYYD
Sbjct: 69 TLAALVGSAMVEYYD 83
>ref|XP_479484.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|33147046|dbj|BAC80135.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|22831150|dbj|BAC16011.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 99
Score = 118 bits (296), Expect = 3e-26
Identities = 59/78 (75%), Positives = 69/78 (87%), Gaps = 2/78 (2%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAE K+ ++S+R+WVVDHKLR V LWL+G+ SIAYNWSRP MKTSVKIIHA LHAQAL
Sbjct: 1 MAEEKSTMQSMREWVVDHKLRAV-TLWLTGVASSIAYNWSRPGMKTSVKIIHA-LHAQAL 58
Query: 61 TLAALAGAAVVEYYDHRA 78
TLAALAG+A+VEYYDHR+
Sbjct: 59 TLAALAGSALVEYYDHRS 76
>gb|EAL90555.1| mitochondrial hypoxia responsive domain protein [Aspergillus
fumigatus Af293]
Length = 268
Score = 47.0 bits (110), Expect = 1e-04
Identities = 24/59 (40%), Positives = 33/59 (55%)
Query: 19 KLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHR 77
K + VG W++ + GS A P + S KI+ AR++AQ LTLA L +A E D R
Sbjct: 116 KYKIVGATWVASMIGSFAMVSRNPYLTGSQKIVQARVYAQGLTLAVLVASAAFEISDQR 174
>emb|CAE47952.1| hypothetical protein, conserved [Aspergillus fumigatus]
gi|19309405|emb|CAD27304.1| hypothetical protein
[Aspergillus fumigatus]
Length = 225
Score = 47.0 bits (110), Expect = 1e-04
Identities = 24/59 (40%), Positives = 33/59 (55%)
Query: 19 KLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHR 77
K + VG W++ + GS A P + S KI+ AR++AQ LTLA L +A E D R
Sbjct: 116 KYKIVGATWVASMIGSFAMVSRNPYLTGSQKIVQARVYAQGLTLAVLVASAAFEISDQR 174
>gb|EAL20318.1| hypothetical protein CNBF1290 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57227662|gb|AAW44120.1|
mitochondrion protein, putative [Cryptococcus neoformans
var. neoformans JEC21] gi|58268542|ref|XP_571427.1|
mitochondrion protein, putative [Cryptococcus neoformans
var. neoformans JEC21]
Length = 226
Score = 45.1 bits (105), Expect = 5e-04
Identities = 20/66 (30%), Positives = 34/66 (51%)
Query: 6 TQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAAL 65
TQ+E + WV +HK + W+ + + P T+ K++ AR+ AQ LT+ L
Sbjct: 96 TQMEKAKNWVGNHKYSLISAAWVGSLGLAFGIVARNPYQTTAQKVVQARMWAQGLTVGLL 155
Query: 66 AGAAVV 71
G A++
Sbjct: 156 VGGALL 161
>gb|EAK85074.1| hypothetical protein UM03929.1 [Ustilago maydis 521]
gi|49074884|ref|XP_401544.1| hypothetical protein
UM03929.1 [Ustilago maydis 521]
Length = 211
Score = 43.9 bits (102), Expect = 0.001
Identities = 21/58 (36%), Positives = 33/58 (56%), Gaps = 1/58 (1%)
Query: 14 WVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVV 71
W ++K V W S + G AY ++P M + K++ AR+ AQ +TLA+L G A +
Sbjct: 118 WAKENKFSVVAATWASSMVGIFAYIHTQP-MSFAQKLVQARVWAQGITLASLVGMAAI 174
>gb|AAW41347.1| mitochondrion protein, putative [Cryptococcus neoformans var.
neoformans JEC21] gi|57223302|gb|AAW41346.1|
mitochondrion protein, putative [Cryptococcus neoformans
var. neoformans JEC21] gi|50260648|gb|EAL23301.1|
hypothetical protein CNBA4170 [Cryptococcus neoformans
var. neoformans B-3501A] gi|58259507|ref|XP_567166.1|
mitochondrion protein, putative [Cryptococcus neoformans
var. neoformans JEC21] gi|58259505|ref|XP_567165.1|
mitochondrion protein, putative [Cryptococcus neoformans
var. neoformans JEC21]
Length = 232
Score = 42.4 bits (98), Expect = 0.003
Identities = 21/68 (30%), Positives = 34/68 (49%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTL 62
E +++E W HK +G W + + + P TS K++ AR+ AQ LT+
Sbjct: 101 EKMSKLEKAGDWAARHKYGIIGGGWAASMALAFGIVARNPYQSTSQKVVQARMWAQGLTV 160
Query: 63 AALAGAAV 70
A L G+A+
Sbjct: 161 ALLVGSAM 168
>gb|AAW69331.1| hypothetical protein [Magnaporthe grisea]
gi|38109735|gb|EAA55559.1| hypothetical protein
MG01210.4 [Magnaporthe grisea 70-15]
gi|39950384|ref|XP_363284.1| hypothetical protein
MG01210.4 [Magnaporthe grisea 70-15]
Length = 253
Score = 41.2 bits (95), Expect = 0.007
Identities = 21/73 (28%), Positives = 41/73 (55%)
Query: 6 TQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAAL 65
+Q++ ++ W +++ V WL+ + ++A + T+ K++ AR++AQ LTLA L
Sbjct: 102 SQMQRLKDWGRENRYTIVFASWLASMGVALAAVGKDKYLTTAQKLVQARVYAQGLTLAVL 161
Query: 66 AGAAVVEYYDHRA 78
AV E D ++
Sbjct: 162 IATAVFETADAKS 174
>ref|NP_573120.1| CG9921-PA [Drosophila melanogaster] gi|7293210|gb|AAF48593.1|
CG9921-PA [Drosophila melanogaster]
gi|17945340|gb|AAL48726.1| RE16337p [Drosophila
melanogaster]
Length = 102
Score = 40.8 bits (94), Expect = 0.009
Identities = 22/77 (28%), Positives = 41/77 (52%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
+AE +T E +++ + ++ L +GCL + + YN+ N K S ++ +R+ AQ
Sbjct: 26 VAEVETTKEKLQRKIKENPLVPLGCLATTAALTAGLYNFRTGNRKMSQLMMRSRIAAQGF 85
Query: 61 TLAALAGAAVVEYYDHR 77
T+ AL V+ Y D +
Sbjct: 86 TVMALVVGVVMTYTDKK 102
>gb|EAA71650.1| hypothetical protein FG03448.1 [Gibberella zeae PH-1]
gi|46115212|ref|XP_383624.1| hypothetical protein
FG03448.1 [Gibberella zeae PH-1]
Length = 228
Score = 39.7 bits (91), Expect = 0.019
Identities = 25/75 (33%), Positives = 42/75 (55%), Gaps = 5/75 (6%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLS--GITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
E +T E + + +++ + V W++ GI +I SR M T+ K++ AR++AQ L
Sbjct: 99 ENETAYERFKDYAKENRYQIVFASWVASMGIAFAIV---SRQPMSTANKVVQARVYAQGL 155
Query: 61 TLAALAGAAVVEYYD 75
TLA L +A+ E D
Sbjct: 156 TLAVLIVSAIYEMRD 170
>gb|EAA66184.1| hypothetical protein AN1066.2 [Aspergillus nidulans FGSC A4]
gi|67517771|ref|XP_658670.1| hypothetical protein
AN1066_2 [Aspergillus nidulans FGSC A4]
gi|49086290|ref|XP_405203.1| hypothetical protein
AN1066.2 [Aspergillus nidulans FGSC A4]
Length = 236
Score = 38.9 bits (89), Expect = 0.033
Identities = 19/64 (29%), Positives = 31/64 (47%)
Query: 14 WVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEY 73
W + + V W++ + GS A P + K++ AR++AQ TLA L +A E
Sbjct: 111 WAREERYTIVFSTWVASMIGSFALVGRNPYLSGPQKLVQARVYAQGATLAVLIASAAFEI 170
Query: 74 YDHR 77
+ R
Sbjct: 171 SERR 174
>gb|EAA68393.1| hypothetical protein FG00663.1 [Gibberella zeae PH-1]
gi|46107560|ref|XP_380839.1| hypothetical protein
FG00663.1 [Gibberella zeae PH-1]
Length = 232
Score = 38.5 bits (88), Expect = 0.043
Identities = 24/73 (32%), Positives = 41/73 (55%), Gaps = 1/73 (1%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTL 62
E +T +E ++ +++ V WL+ + + A SR M T+ K++ AR++AQ LTL
Sbjct: 99 ENQTAMERFMEYGKENRYSIVFVSWLASMGVAFAMV-SRSPMNTANKVVQARVYAQGLTL 157
Query: 63 AALAGAAVVEYYD 75
A L +A+ E D
Sbjct: 158 AVLIISAIFEVND 170
>gb|AAS50433.1| AAR068Cp [Ashbya gossypii ATCC 10895] gi|45184891|ref|NP_982609.1|
AAR068Cp [Eremothecium gossypii]
Length = 221
Score = 37.7 bits (86), Expect = 0.074
Identities = 26/93 (27%), Positives = 46/93 (48%), Gaps = 8/93 (8%)
Query: 1 MAEAKTQVE-----SIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARL 55
+AEAK E + + +++K + +G LW + + GS P M K + AR+
Sbjct: 92 LAEAKRWAELPLGDKLLESGLNNKYKIIGSLWAASMYGSWRLVDRDPIMTKVQKAVQARM 151
Query: 56 HAQALTLAALAGAAVVEYYDHRAEAKRAKNSEQ 88
+AQALT+ L + ++ Y + + +N EQ
Sbjct: 152 YAQALTVVMLLASVMLSMYGAK---RNPENKEQ 181
>gb|EAL32117.1| GA22125-PA [Drosophila pseudoobscura]
Length = 102
Score = 37.7 bits (86), Expect = 0.074
Identities = 21/71 (29%), Positives = 36/71 (50%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTL 62
E +T E +++ + ++ L +GCL + YN+ N K S ++ R+ AQ T+
Sbjct: 28 EVETTKEKLQRKIKENPLVPIGCLATTAALTMGLYNFRTGNRKMSQLMMRTRIAAQGFTV 87
Query: 63 AALAGAAVVEY 73
AAL V+ Y
Sbjct: 88 AALIIGVVMTY 98
>ref|XP_331822.1| hypothetical protein [Neurospora crassa] gi|28926826|gb|EAA35790.1|
hypothetical protein [Neurospora crassa]
Length = 241
Score = 37.4 bits (85), Expect = 0.096
Identities = 19/73 (26%), Positives = 36/73 (49%)
Query: 6 TQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAAL 65
T + +W +++ V W++ + ++A + S K++ AR++AQ LTLA L
Sbjct: 103 TSTQRFMEWGRENRYSIVFASWIAAMGAAMAIVNRNKYLSASQKLVQARVYAQGLTLAVL 162
Query: 66 AGAAVVEYYDHRA 78
A E D ++
Sbjct: 163 IATAAFETADAKS 175
>emb|CAG78639.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50556840|ref|XP_505828.1| hypothetical protein
[Yarrowia lipolytica]
Length = 279
Score = 37.0 bits (84), Expect = 0.13
Identities = 20/65 (30%), Positives = 34/65 (51%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M K Q ++ +HK + V W+ + GS+ Y + M ++ KI+ AR++AQA
Sbjct: 89 MEHRKEQQAGFMHYLKEHKYKCVFGGWVVSMAGSMWYVFRDKYMTSTQKIVQARMYAQAS 148
Query: 61 TLAAL 65
T+ L
Sbjct: 149 TIILL 153
>gb|AAC24977.1| anaerobically inducible early gene 2 [Oryza sativa]
Length = 127
Score = 36.6 bits (83), Expect = 0.16
Identities = 15/39 (38%), Positives = 25/39 (63%), Gaps = 1/39 (2%)
Query: 10 SIRKWVVDHKLRTVGCLWLSGITGSIAYNWSR-PNMKTS 47
S++ WV +HKL ++G LW + + S+AY + P M+ S
Sbjct: 7 SVQSWVEEHKLASIGGLWATAVGASVAYGRRKTPQMRDS 45
>emb|CAG62783.1| unnamed protein product [Candida glabrata CBS138]
gi|50294788|ref|XP_449805.1| unnamed protein product
[Candida glabrata]
Length = 225
Score = 35.4 bits (80), Expect = 0.36
Identities = 18/72 (25%), Positives = 36/72 (50%)
Query: 17 DHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDH 76
++K + + W + + GS P M T+ K + AR++AQ +T+A L + + Y+
Sbjct: 114 NNKYKIITGAWAASLYGSWVLVDRDPIMTTAQKAVQARMYAQFITVALLLASVGLSMYEA 173
Query: 77 RAEAKRAKNSEQ 88
+ + K E+
Sbjct: 174 KLHPDKKKQGEK 185
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.127 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 131,656,224
Number of Sequences: 2540612
Number of extensions: 3798907
Number of successful extensions: 11314
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 11290
Number of HSP's gapped (non-prelim): 41
length of query: 88
length of database: 863,360,394
effective HSP length: 64
effective length of query: 24
effective length of database: 700,761,226
effective search space: 16818269424
effective search space used: 16818269424
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0299.14


