

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0288.8
(846 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
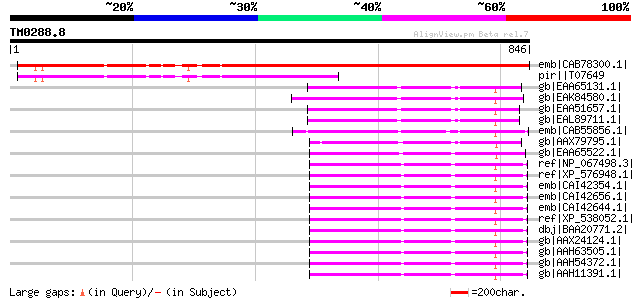
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB78300.1| polyubiquitin-like protein [Arabidopsis thaliana... 767 0.0
pir||T07649 hypothetical protein T1P17.160 - Arabidopsis thalian... 362 2e-98
gb|EAA65131.1| hypothetical protein AN1966.2 [Aspergillus nidula... 204 1e-50
gb|EAK84580.1| hypothetical protein UM03442.1 [Ustilago maydis 5... 204 1e-50
gb|EAA51657.1| hypothetical protein MG03252.4 [Magnaporthe grise... 202 5e-50
gb|EAL89711.1| ubiquitin-protein ligase (Tom1), putative [Asperg... 199 2e-49
emb|CAB55856.1| SPAC1805.15c [Schizosaccharomyces pombe] gi|1911... 199 3e-49
gb|AAX79795.1| ubiquitin-protein ligase, putative [Trypanosoma b... 199 4e-49
gb|EAA65522.1| hypothetical protein AN1339.2 [Aspergillus nidula... 196 4e-48
ref|NP_067498.3| HECT, UBA and WWE domain containing 1 [Mus musc... 195 6e-48
ref|XP_576948.1| PREDICTED: similar to mKIAA0312 protein [Rattus... 195 6e-48
emb|CAI42354.1| upstream regulatory element binding protein 1 (U... 195 6e-48
emb|CAI42656.1| upstream regulatory element binding protein 1 (U... 195 6e-48
emb|CAI42644.1| upstream regulatory element binding protein 1 (U... 195 6e-48
ref|XP_538052.1| PREDICTED: similar to E3 ubiquitin protein liga... 195 6e-48
dbj|BAA20771.2| KIAA0312 [Homo sapiens] 195 6e-48
gb|AAX24124.1| LASU1 [Mus musculus] 195 6e-48
gb|AAH63505.1| HUWE1 protein [Homo sapiens] 195 6e-48
gb|AAH54372.1| Huwe1 protein [Mus musculus] 195 6e-48
gb|AAH11391.1| Huwe1 protein [Mus musculus] 195 6e-48
>emb|CAB78300.1| polyubiquitin-like protein [Arabidopsis thaliana]
gi|5823568|emb|CAB41727.2| polyubiquitin-like protein
[Arabidopsis thaliana] gi|25407523|pir||H85134
polyubiquitin-like protein [imported] - Arabidopsis
thaliana gi|15235410|ref|NP_192994.1| ubiquitin-protein
ligase, putative [Arabidopsis thaliana]
Length = 873
Score = 767 bits (1981), Expect = 0.0
Identities = 428/870 (49%), Positives = 561/870 (64%), Gaps = 61/870 (7%)
Query: 14 KRKHDEIDDDEGGVSD-IDPIRMKKDDA----AKAVYSAVHHRS---------------- 52
KRK DE D GV++ ++ ++ ++ DA A A + + RS
Sbjct: 28 KRKRDEDSSDYVGVAESLEMLKKQEIDADHMAASAQQTLISWRSGENSRSLSSSGECSSS 87
Query: 53 -----IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQS 107
LQ FVRMM G TIV+ A DTV+ +H+RI+ IP EQR+IY GKQLQ
Sbjct: 88 NRPESTRLQIFVRMMSGGKTIVIHAEKYDTVEKLHQRIEWKTKIPALEQRVIYKGKQLQR 147
Query: 108 EQTLAECSIGKDASLQLVGRMRSTGHPHAWQVINDMVSLVIRLCHNEPVHDANKTIKGLI 167
E +L SI +DASLQLV RM+ST HP AWQ I+D++ + R+ E + I I
Sbjct: 148 ENSLTYYSIEQDASLQLVARMQSTEHPVAWQTIDDIMYTISRMYKGE---NLQSNINEKI 204
Query: 168 TSYLNLSPKNDNDSASGYFQIFMSAT--AVLVSLYLSPYTGNKDCADSSIRHFLTTCKTT 225
++ + P ++S + Y IF +++ A LV LY S NK CA SS++ FL+ C
Sbjct: 205 VTFFAMIPVESDESIAKYLNIFSNSSVPAALVMLYASSLERNKSCAKSSVKLFLSNC--- 261
Query: 226 VSPPLHTQ--CARVVLDFCTLLKKRVSSEDPLYGFCRSTFGSLLETARVYCADSEGDNVK 283
V+ P + + C +VL+FC LL+K V + LY CR+T GS+LET DN
Sbjct: 262 VALPKNQKNYCLPIVLEFCKLLRK-VCPDQKLYVTCRNTLGSMLETF---------DNPH 311
Query: 284 GSILVQ------EIFPFVSELAKSLLVDLDLSIDSPSNAGTFLINVADFTAFLVPLRARI 337
G Q EIFPF +EL LL +L N+G + ++F LR I
Sbjct: 312 GVYNDQYETFGVEIFPFFTELTGLLLNEL------AQNSGPSFCDFQKVSSFWQQLRKVI 365
Query: 338 MEQQALRGSMPTQKRHKEALLVEEIEGLHLLYNDLLNKTDQCLQKLEQTWAGQGMMQGED 397
+ A +P + L EI LH L+ LL D C+ ++E + A + + E
Sbjct: 366 ELKVAF--PIPIVLPMQSTALEAEIRHLHRLFGSLLTTMDLCMCRVESSLADKEVGNSET 423
Query: 398 IYPAWSHYLSILKELYLISKLYDGAEEKLWKVLTRQRSVVCLLIVRYAKRTDEHQWILEH 457
+ +WS YLSILK + +S +Y GA+ +L +L + + L+V++AKR D+HQWI E+
Sbjct: 424 MSSSWSQYLSILKIINSMSNIYQGAKGQLAVMLNKNKVSFSALVVKFAKRGDDHQWIFEY 483
Query: 458 KCVTNFESRRHLAMMMFPEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAGLFME 517
K TNFE+RRHLAM++FP+VKED+EE+HEMLIDRS LLSESFEYI A +LH GLFME
Sbjct: 484 KEATNFEARRHLAMLLFPDVKEDFEEMHEMLIDRSNLLSESFEYIVGASPEALHGGLFME 543
Query: 518 FKNEEATGPGVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFS 577
FKNEEATGPGVLREWF LVCQ +FNP+N LF+ +D RRF PN ASKV PLH ++F F+
Sbjct: 544 FKNEEATGPGVLREWFYLVCQEIFNPKNTLFLRSADDFRRFSPNPASKVDPLHPDFFEFT 603
Query: 578 GRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDS 637
GRVIALALMHKVQVG++ DRVFFLQLAG I+LED+KD D +Y+SCKQILEMD +F DS
Sbjct: 604 GRVIALALMHKVQVGVLFDRVFFLQLAGLKISLEDIKDTDRIMYNSCKQILEMDPEFFDS 663
Query: 638 DA-LGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHF 696
+A LGLTFV E EELG +ELCP GK VNSKNR++Y+DLLI+ RF T I EQV F
Sbjct: 664 NAGLGLTFVLETEELGKRDTIELCPDGKLKAVNSKNRKQYVDLLIERRFATPILEQVKQF 723
Query: 697 SKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITW 756
S+GFTD+LS S + FF+ L LEDLD ML G E+ IS++DWKAHTEYNG+KETD QI W
Sbjct: 724 SRGFTDMLSHSVPPRSFFKRLYLEDLDGMLRGGENPISIDDWKAHTEYNGFKETDRQIDW 783
Query: 757 FWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLASRLYIYKSPEPEDRLPSSHTCFYRM 816
FW+I+ +MT E+++ +LFFWTS K++PVEGF GL+S+LYIY+ E DRLP SHTCFYR+
Sbjct: 784 FWKILKKMTEEEQRSILFFWTSNKFVPVEGFRGLSSKLYIYRLYEANDRLPLSHTCFYRL 843
Query: 817 CFPAYSSMTVMQERLEVITQEHIGCSFGTW 846
C P Y ++T+M++RL +I Q+H+ SFG W
Sbjct: 844 CIPRYPTITLMEQRLRLIAQDHVSSSFGKW 873
>pir||T07649 hypothetical protein T1P17.160 - Arabidopsis thaliana (fragment)
Length = 561
Score = 362 bits (930), Expect = 2e-98
Identities = 232/558 (41%), Positives = 316/558 (56%), Gaps = 60/558 (10%)
Query: 14 KRKHDEIDDDEGGVSD-IDPIRMKKDDA----AKAVYSAVHHRS---------------- 52
KRK DE D GV++ ++ ++ ++ DA A A + + RS
Sbjct: 28 KRKRDEDSSDYVGVAESLEMLKKQEIDADHMAASAQQTLISWRSGENSRSLSSSGECSSS 87
Query: 53 -----IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQS 107
LQ FVRMM G TIV+ A DTV+ +H+RI+ IP EQR+IY GKQLQ
Sbjct: 88 NRPESTRLQIFVRMMSGGKTIVIHAEKYDTVEKLHQRIEWKTKIPALEQRVIYKGKQLQR 147
Query: 108 EQTLAECSIGKDASLQLVGRMRSTGHPHAWQVINDMVSLVIRLCHNEPVHDANKTIKGLI 167
E +L SI +DASLQLV RM+ST HP AWQ I+D++ + R+ E + I I
Sbjct: 148 ENSLTYYSIEQDASLQLVARMQSTEHPVAWQTIDDIMYTISRMYKGE---NLQSNINEKI 204
Query: 168 TSYLNLSPKNDNDSASGYFQIFMSAT--AVLVSLYLSPYTGNKDCADSSIRHFLTTCKTT 225
++ + P ++S + Y IF +++ A LV LY S NK CA SS++ FL+ C
Sbjct: 205 VTFFAMIPVESDESIAKYLNIFSNSSVPAALVMLYASSLERNKSCAKSSVKLFLSNC--- 261
Query: 226 VSPPLHTQ--CARVVLDFCTLLKKRVSSEDPLYGFCRSTFGSLLETARVYCADSEGDNVK 283
V+ P + + C +VL+FC LL+K V + LY CR+T GS+LET DN
Sbjct: 262 VALPKNQKNYCLPIVLEFCKLLRK-VCPDQKLYVTCRNTLGSMLETF---------DNPH 311
Query: 284 GSILVQ------EIFPFVSELAKSLLVDLDLSIDSPSNAGTFLINVADFTAFLVPLRARI 337
G Q EIFPF +EL LL +L N+G + ++F LR I
Sbjct: 312 GVYNDQYETFGVEIFPFFTELTGLLLNEL------AQNSGPSFCDFQKVSSFWQQLRKVI 365
Query: 338 MEQQALRGSMPTQKRHKEALLVEEIEGLHLLYNDLLNKTDQCLQKLEQTWAGQGMMQGED 397
+ A +P + L EI LH L+ LL D C+ ++E + A + + E
Sbjct: 366 ELKVAF--PIPIVLPMQSTALEAEIRHLHRLFGSLLTTMDLCMCRVESSLADKEVGNSET 423
Query: 398 IYPAWSHYLSILKELYLISKLYDGAEEKLWKVLTRQRSVVCLLIVRYAKRTDEHQWILEH 457
+ +WS YLSILK + +S +Y GA+ +L +L + + L+V++AKR D+HQWI E+
Sbjct: 424 MSSSWSQYLSILKIINSMSNIYQGAKGQLAVMLNKNKVSFSALVVKFAKRGDDHQWIFEY 483
Query: 458 KCVTNFESRRHLAMMMFPEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAGLFME 517
K TNFE+RRHLAM++FP+VKED+EE+HEMLIDRS LLSESFEYI A +LH GLFME
Sbjct: 484 KEATNFEARRHLAMLLFPDVKEDFEEMHEMLIDRSNLLSESFEYIVGASPEALHGGLFME 543
Query: 518 FKNEEATGPGVLREWFLL 535
FKNEEATGPGVLREWF L
Sbjct: 544 FKNEEATGPGVLREWFYL 561
>gb|EAA65131.1| hypothetical protein AN1966.2 [Aspergillus nidulans FGSC A4]
gi|67523019|ref|XP_659570.1| hypothetical protein
AN1966_2 [Aspergillus nidulans FGSC A4]
gi|49088592|ref|XP_406103.1| hypothetical protein
AN1966.2 [Aspergillus nidulans FGSC A4]
Length = 4022
Score = 204 bits (519), Expect = 1e-50
Identities = 120/355 (33%), Positives = 188/355 (52%), Gaps = 17/355 (4%)
Query: 486 EMLIDRSQLLSESFEYIARADTASL-HAGLFMEFKNEEATGPG-VLREWFLLVCQALFNP 543
++ + R Q+ +SF + L H L + F EE G V REWF ++ + +FNP
Sbjct: 3666 QLAVRRDQVFLDSFRALYFKSAEELKHGKLNVRFHGEEGVDAGGVTREWFQVLARGMFNP 3725
Query: 544 QNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQL 603
ALF+ DR F+PN S V+P HL +F F GR+I AL + R + +
Sbjct: 3726 DYALFIPVAADRTTFHPNRLSGVNPEHLMFFKFIGRIIGKALYEGRVLDCHFSRAVYKCI 3785
Query: 604 AGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGG 663
G++++++D++ D Y S +LE D +D + TF E ++ G+ + ++L G
Sbjct: 3786 LGRNVSIKDMETLDLDYYKSLLWMLENDI----TDIITETFAVETDDFGEKQTIDLIENG 3841
Query: 664 KNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLD 723
+N+ V +N+E+Y+ ++ R V S+ EQ+ +F KGF +I+ F Q LEL
Sbjct: 3842 RNIPVTQENKEEYVQKVVDYRLVASVREQLDNFLKGFHEIIPPELISIFNEQELEL---- 3897
Query: 724 WMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLP 783
++ G + I V+DWKA+TEY+ Y + QI WFW V E+R LL F T +P
Sbjct: 3898 -LISGLPE-IDVDDWKANTEYHNYSASSPQIQWFWRAVRSFDKEERAKLLQFVTGTSKVP 3955
Query: 784 VEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEV 833
+ GF L SR I++ +DRLPSSHTCF ++ P Y S +++RL +
Sbjct: 3956 LNGFKELEGMNGVSRFNIHRDYGNKDRLPSSHTCFNQLDLPEYDSYETLRQRLYI 4010
>gb|EAK84580.1| hypothetical protein UM03442.1 [Ustilago maydis 521]
gi|49073738|ref|XP_401057.1| hypothetical protein
UM03442.1 [Ustilago maydis 521]
Length = 571
Score = 204 bits (519), Expect = 1e-50
Identities = 125/385 (32%), Positives = 196/385 (50%), Gaps = 18/385 (4%)
Query: 460 VTNFESRRHLAMMMFPEVKEDYEELHEMLIDRSQLLSESFEYIARADTASL-HAGLFMEF 518
V +F+++++ + + D+ + + R+ + +SF Y +R + H L + F
Sbjct: 190 VLDFDNKKNYFTQQLHKGRRDHYTPLSLSVRRNSVFEDSFRYFSRKTGPEVKHGKLNVRF 249
Query: 519 KNEEATGPG-VLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFS 577
NEE G V REWF ++ +A+FNP ALF C DR + PN S V+P HL +F F
Sbjct: 250 TNEEGIDAGGVTREWFQVLARAMFNPDYALFQPCAADRTTYQPNRMSYVNPDHLSFFKFV 309
Query: 578 GRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDS 637
GR+I A+ + R F+ + GK + D++ DP + S + +L D +
Sbjct: 310 GRIIGKAIYDGRLLDAYFTRSFYKHILGKPVDYRDLESIDPEYFKSLEWMLSNDI----T 365
Query: 638 DALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFS 697
D L LTF + EE G+TKVV+L P G ++ V N+++Y+ L+ + R SI Q+ F
Sbjct: 366 DILDLTFSVDDEEFGETKVVDLKPNGTSISVTEANKQEYVRLVTEQRLTKSIKSQIDAFL 425
Query: 698 KGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWF 757
GF +I+ S + F Q LEL ++ G D I V+ WK +TE +GY D + W+
Sbjct: 426 GGFNEIIPSDLIRIFSEQELEL-----LISGLPD-IDVDAWKNNTELHGYSSGDAVVQWW 479
Query: 758 WEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTC 812
W V ++ LL F T +P+EGF L R I+K+ DRLP++HTC
Sbjct: 480 WRAVRSFDQTEKAKLLQFITGTSKVPLEGFAHLQGVQGTQRFNIHKA-YGADRLPAAHTC 538
Query: 813 FYRMCFPAYSSMTVMQERLEVITQE 837
F ++ P Y S ++ L + E
Sbjct: 539 FNQLDLPQYESYEKLRSSLLLAMNE 563
>gb|EAA51657.1| hypothetical protein MG03252.4 [Magnaporthe grisea 70-15]
gi|39942344|ref|XP_360709.1| hypothetical protein
MG03252.4 [Magnaporthe grisea 70-15]
Length = 4048
Score = 202 bits (513), Expect = 5e-50
Identities = 118/353 (33%), Positives = 188/353 (52%), Gaps = 17/353 (4%)
Query: 486 EMLIDRSQLLSESFEYIARADTASLHAG-LFMEFKNEEATGPG-VLREWFLLVCQALFNP 543
++ + R + +SF+ + + G L + F EE G V REWF ++ + +F+P
Sbjct: 3692 QLSVRRDHVFHDSFKSLYFKKGEEMKYGKLNIRFHGEEGVDAGGVTREWFQVLSRQMFDP 3751
Query: 544 QNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQL 603
LF +DR F+PN S ++ HL +F F GR+I AL + R + ++
Sbjct: 3752 NYVLFTPVSSDRTTFHPNKLSSINDEHLMFFKFIGRIIGKALYEGRVLDCYFSRAVYKRI 3811
Query: 604 AGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGG 663
GK ++++D++ DP Y S +LE D +D + TF E + G TK V+LC G
Sbjct: 3812 LGKPVSVKDMESFDPEYYKSLVWMLENDI----TDIITETFAVEDDAFGVTKTVDLCENG 3867
Query: 664 KNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLD 723
+N+ V N+ +Y+ L+++ + + S+ +Q++ F +GF DI+ + F Q LEL
Sbjct: 3868 RNIPVTEDNKHEYVRLVVEHKLLASVKDQMAEFLQGFHDIIPAELIAIFNEQELEL---- 3923
Query: 724 WMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLP 783
++ G D I V+DWKAHTEY+ Y+ + QI WFW V E+R LL F T +P
Sbjct: 3924 -LISGLPD-IDVDDWKAHTEYHNYQPSSQQIQWFWRAVRSFDKEERAKLLQFVTGTSKVP 3981
Query: 784 VEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERL 831
+ GF L SR I++ DRLPSSHTCF ++ P Y S V+++++
Sbjct: 3982 LNGFKELEGMNGVSRFNIHRDYGRPDRLPSSHTCFNQLDLPEYESYDVLRKQI 4034
>gb|EAL89711.1| ubiquitin-protein ligase (Tom1), putative [Aspergillus fumigatus
Af293]
Length = 4037
Score = 199 bits (507), Expect = 2e-49
Identities = 120/354 (33%), Positives = 188/354 (52%), Gaps = 19/354 (5%)
Query: 486 EMLIDRSQLLSESFE--YIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFN 542
++ + R Q+ +SF+ Y AD + L + F EE G V REWF ++ + +FN
Sbjct: 3681 QLSVRRDQVFLDSFKSLYFKTADELK-YGKLNVRFHGEEGVDAGGVTREWFQVLARGMFN 3739
Query: 543 PQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQ 602
P ALF+ DR F+PN S V+ HL +F F GR+I AL + R +
Sbjct: 3740 PNYALFIPVAADRTTFHPNRLSGVNSEHLMFFKFIGRIIGKALYEGRVLDCHFSRAVYKC 3799
Query: 603 LAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPG 662
+ G+ ++++D++ D Y S +LE D +D + TF E ++ G+ +V++L
Sbjct: 3800 ILGRSVSIKDMETLDLDYYKSLLWMLENDI----TDIITETFAVETDDFGEKQVIDLIEN 3855
Query: 663 GKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDL 722
G+N+ V +N+E+Y+ ++ R V S+ +Q+ +F KGF DI+ F Q LEL
Sbjct: 3856 GRNIPVTQENKEEYVQRVVDYRLVKSVKDQLDNFLKGFHDIIPPDLISIFNEQELEL--- 3912
Query: 723 DWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYL 782
++ G + I V+DWKA+TEY+ Y + QI WFW V E+R LL F T +
Sbjct: 3913 --LISGLPE-IDVDDWKANTEYHNYSASSPQIQWFWRAVRSFDKEERAKLLQFVTGTSKV 3969
Query: 783 PVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERL 831
P+ GF L S+ I++ +DRLPSSHTCF ++ P Y S +++RL
Sbjct: 3970 PLNGFKELEGMNGVSKFNIHRDYGNKDRLPSSHTCFNQLDLPEYDSYETLRQRL 4023
>emb|CAB55856.1| SPAC1805.15c [Schizosaccharomyces pombe]
gi|19114838|ref|NP_593926.1| hypothetical protein
SPAC1805.15c [Schizosaccharomyces pombe 972h-]
gi|46397059|sp|Q9UTG2|PUB2_SCHPO E3 ubiquitin--protein
ligase pub2
Length = 671
Score = 199 bits (506), Expect = 3e-49
Identities = 124/390 (31%), Positives = 202/390 (51%), Gaps = 19/390 (4%)
Query: 462 NFESRRHLAMMMF-PEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAGLFMEFKN 520
N E +R +A M PE+ + +L ++ + R+ ++++ I++ + + L + F+N
Sbjct: 294 NDEYQRKIAYMYDRPEMAVNDAQL-QLKVSRATTFEDAYDIISKLSVSDMKKKLLIRFRN 352
Query: 521 EEATG-PGVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGR 579
E+ GV RE+F ++ A+FNP +LF +D + S V+P YF F GR
Sbjct: 353 EDGLDYGGVSREFFYILSHAIFNPGYSLFEYATDDNYGLQISPLSSVNPDFRSYFRFVGR 412
Query: 580 VIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDA 639
V+ LA+ H+ + + F+ ++ K + LEDVKD D Y S K I D D ++
Sbjct: 413 VMGLAIYHRRYLDVQFVLPFYKRILQKPLCLEDVKDVDEVYYESLKWIKNNDVD----ES 468
Query: 640 LGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKG 699
L L F E G++ V+L P G+N+ VN++N+ Y+ L + + VTS EQ + G
Sbjct: 469 LCLNFSVEENRFGESVTVDLIPNGRNIAVNNQNKMNYLKALTEHKLVTSTEEQFNALKGG 528
Query: 700 FTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWE 759
+++ S Q F +LD +L+G D I V+DWK T+Y Y ETD + WFWE
Sbjct: 529 LNELIPDSVLQIF-----NENELDTLLNGKRD-IDVQDWKRFTDYRSYTETDDIVIWFWE 582
Query: 760 IVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL----ASRLYIYKSPEPEDRLPSSHTCFYR 815
++ + E++ LL F T LP+ GF + R + + +LP +HTCF R
Sbjct: 583 LLSEWSPEKKAKLLQFATGTSRLPLSGFKDMHGSDGPRKFTIEKVGHISQLPKAHTCFNR 642
Query: 816 MCFPAYSSMTVMQERLEVITQEHIGCSFGT 845
+ P Y+S ++++L + QE G FGT
Sbjct: 643 LDIPPYNSKEELEQKLTIAIQETAG--FGT 670
>gb|AAX79795.1| ubiquitin-protein ligase, putative [Trypanosoma brucei]
Length = 4304
Score = 199 bits (505), Expect = 4e-49
Identities = 122/353 (34%), Positives = 185/353 (51%), Gaps = 23/353 (6%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEE-ATGPGVLREWFLLVCQALFNPQNAL 547
I+R Q L +SF +++ + + H + + F EE A G+ REWF L+ + + N AL
Sbjct: 3955 INRGQCLRDSFAQLSKLN--NFHGSISVRFSGEEGADAGGLTREWFQLIAEEMVNDNYAL 4012
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F+ + + PN S V+PLHL+YF F+G V+ +A+ H+V + + R + + G
Sbjct: 4013 FIHS-REGMTYQPNPLSGVNPLHLQYFRFAGIVVGMAVAHRVAIDVHFTRAVYRHMIGIQ 4071
Query: 608 ITLEDVKDADPFLYSSCKQILEMD-SDFIDSDALGLTFVREVEELGDTKVVELCPGGKNL 666
T D+K DP LY + K +L D SD LGL F E+ G T+ VEL P G +
Sbjct: 4072 PTFGDLKSVDPELYENLKWLLVNDVSD------LGLFFTVSCEKFGVTEEVELIPNGSQV 4125
Query: 667 IVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWML 726
V + N+ +Y+ L + I EQ+ F KGF ++ + + F Q LEL ++
Sbjct: 4126 AVTNANKSQYVRLRCEFCMTRQIEEQLQEFLKGFYAVIPRKEIRNFSAQELEL-----VI 4180
Query: 727 HGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEG 786
G D I VED + +T Y+GY T +QI WFWE+V M+ E R LL F T +P G
Sbjct: 4181 CGMPD-IDVEDLRLNTTYDGYTSTSLQIRWFWEVVAAMSKEDRANLLQFATGASRVPHGG 4239
Query: 787 FCGLAS------RLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEV 833
F L S R + + + + LP +HTCF ++ P YSS ++++L V
Sbjct: 4240 FSNLESSNGSPQRFTVSRWADSAELLPQAHTCFNKIALPEYSSCEELRKKLMV 4292
>gb|EAA65522.1| hypothetical protein AN1339.2 [Aspergillus nidulans FGSC A4]
gi|67521764|ref|XP_658943.1| hypothetical protein
AN1339_2 [Aspergillus nidulans FGSC A4]
gi|49086950|ref|XP_405476.1| hypothetical protein
AN1339.2 [Aspergillus nidulans FGSC A4]
Length = 821
Score = 196 bits (497), Expect = 4e-48
Identities = 120/358 (33%), Positives = 187/358 (51%), Gaps = 17/358 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R+ + +S+ I R + L L ++F E+ G L RE+F L+ +FNP L
Sbjct: 471 VRRNNIFEDSYAEIMRQSASDLKKRLMIKFDGEDGLDYGGLSREFFFLLSHEMFNPFYCL 530
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F +D N S V+P HL YF F GRV+ LA+ H+ + F+ + K
Sbjct: 531 FEYSAHDNYTLQINPHSGVNPEHLNYFKFIGRVVGLAIFHRRFLDSFFIGAFYKMMLRKK 590
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
++L+D++ D L+ + LE D + I + LTF + E+ G+ + +EL PGG+++
Sbjct: 591 VSLQDMEGVDEDLHRNLTWTLENDIEGI----IDLTFTVDDEKFGERRTIELKPGGEDIP 646
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V ++N+ +Y++L+ + + V + EQ + F GF +++ + F + LEL L
Sbjct: 647 VTNENKHEYVELVTEWKIVKRVEEQFNAFMSGFNELIPADLVNVFDERELEL------LI 700
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
G I V+DWK HT+Y GY+E D I FW+IV AEQ+ LL F T +PV GF
Sbjct: 701 GGIADIDVDDWKKHTDYRGYQEQDEVIQNFWKIVRTWDAEQKSRLLQFTTGTSRIPVNGF 760
Query: 788 CGLAS-----RLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
L R I KS +P LP SHTCF R+ P Y S V++ +L + +E +G
Sbjct: 761 KDLQGSDGPRRFTIEKSGDP-IALPKSHTCFNRLDLPPYKSHEVLEHKLSIAVEETLG 817
>ref|NP_067498.3| HECT, UBA and WWE domain containing 1 [Mus musculus]
Length = 4378
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 4025 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 4084
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 4085 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 4144
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 4145 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 4201
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 4202 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 4255
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 4256 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 4315
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 4316 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 4372
Query: 843 FG 844
G
Sbjct: 4373 EG 4374
>ref|XP_576948.1| PREDICTED: similar to mKIAA0312 protein [Rattus norvegicus]
Length = 4147
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 3794 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 3853
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 3854 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 3913
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 3914 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 3970
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 3971 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 4024
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 4025 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 4084
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 4085 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 4141
Query: 843 FG 844
G
Sbjct: 4142 EG 4143
>emb|CAI42354.1| upstream regulatory element binding protein 1 (UREB1) [Homo sapiens]
gi|57210033|emb|CAI42654.1| upstream regulatory element
binding protein 1 (UREB1) [Homo sapiens]
gi|57160758|emb|CAI39580.1| upstream regulatory element
binding protein 1 (UREB1) [Homo sapiens]
gi|60549639|gb|AAX24125.1| LASU1 [Homo sapiens]
gi|61676188|ref|NP_113584.3| HECT, UBA and WWE domain
containing 1 [Homo sapiens]
Length = 4374
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 4021 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 4080
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 4081 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 4140
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 4141 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 4197
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 4198 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 4251
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 4252 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 4311
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 4312 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 4368
Query: 843 FG 844
G
Sbjct: 4369 EG 4370
>emb|CAI42656.1| upstream regulatory element binding protein 1 (UREB1) [Homo sapiens]
gi|57160759|emb|CAI39581.1| upstream regulatory element
binding protein 1 (UREB1) [Homo sapiens]
Length = 3407
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 3054 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 3113
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 3114 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 3173
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 3174 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 3230
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 3231 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 3284
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 3285 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 3344
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 3345 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 3401
Query: 843 FG 844
G
Sbjct: 3402 EG 3403
>emb|CAI42644.1| upstream regulatory element binding protein 1 (UREB1) [Homo sapiens]
gi|57160756|emb|CAI39578.1| upstream regulatory element
binding protein 1 (UREB1) [Homo sapiens]
Length = 1196
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 843 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 902
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 903 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 962
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 963 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 1019
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 1020 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 1073
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 1074 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 1133
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 1134 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 1190
Query: 843 FG 844
G
Sbjct: 1191 EG 1192
>ref|XP_538052.1| PREDICTED: similar to E3 ubiquitin protein ligase URE-B1 (HSPC272)
[Canis familiaris]
Length = 4405
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 4052 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 4111
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 4112 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 4171
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 4172 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 4228
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 4229 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 4282
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 4283 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 4342
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 4343 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 4399
Query: 843 FG 844
G
Sbjct: 4400 EG 4401
>dbj|BAA20771.2| KIAA0312 [Homo sapiens]
Length = 3192
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 2839 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 2898
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 2899 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 2958
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 2959 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 3015
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 3016 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 3069
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 3070 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 3129
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 3130 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 3186
Query: 843 FG 844
G
Sbjct: 3187 EG 3188
>gb|AAX24124.1| LASU1 [Mus musculus]
Length = 4377
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 4024 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 4083
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 4084 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 4143
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 4144 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 4200
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 4201 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 4254
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 4255 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 4314
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 4315 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 4371
Query: 843 FG 844
G
Sbjct: 4372 EG 4373
>gb|AAH63505.1| HUWE1 protein [Homo sapiens]
Length = 388
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 35 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 94
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 95 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 154
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 155 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 211
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 212 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 265
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 266 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 325
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 326 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 382
Query: 843 FG 844
G
Sbjct: 383 EG 384
>gb|AAH54372.1| Huwe1 protein [Mus musculus]
Length = 444
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 91 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 150
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 151 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 210
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 211 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 267
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 268 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 321
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 322 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 381
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 382 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 438
Query: 843 FG 844
G
Sbjct: 439 EG 440
>gb|AAH11391.1| Huwe1 protein [Mus musculus]
Length = 1080
Score = 195 bits (495), Expect = 6e-48
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 727 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 786
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 787 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 846
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 847 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 903
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 904 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 957
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 958 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 1017
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 1018 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 1074
Query: 843 FG 844
G
Sbjct: 1075 EG 1076
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,419,820,630
Number of Sequences: 2540612
Number of extensions: 59328446
Number of successful extensions: 138811
Number of sequences better than 10.0: 1719
Number of HSP's better than 10.0 without gapping: 1488
Number of HSP's successfully gapped in prelim test: 232
Number of HSP's that attempted gapping in prelim test: 133795
Number of HSP's gapped (non-prelim): 3128
length of query: 846
length of database: 863,360,394
effective HSP length: 137
effective length of query: 709
effective length of database: 515,296,550
effective search space: 365345253950
effective search space used: 365345253950
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0288.8


