

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.2
(142 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
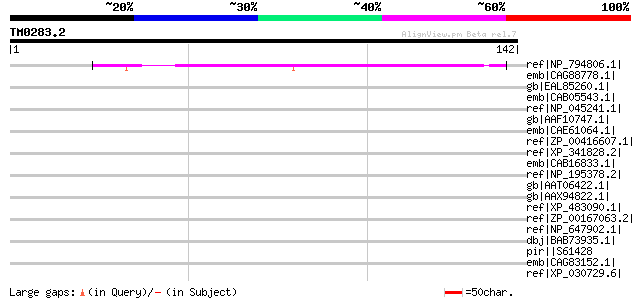
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_794806.1| amine oxidase, flavin-containing [Pseudomonas s... 44 6e-04
emb|CAG88778.1| unnamed protein product [Debaryomyces hansenii C... 39 0.020
gb|EAL85260.1| LEA domain protein [Aspergillus fumigatus Af293] 39 0.027
emb|CAB05543.1| Hypothetical protein K08H10.1 [Caenorhabditis el... 39 0.027
ref|NP_045241.1| 24 [Equid herpesvirus 4] gi|2605967|gb|AAC59539... 39 0.027
gb|AAF10747.1| hypothetical protein [Deinococcus radiodurans] gi... 39 0.035
emb|CAE61064.1| Hypothetical protein CBG04813 [Caenorhabditis br... 39 0.035
ref|ZP_00416607.1| Protein of unknown function DUF224 [Azotobact... 38 0.045
ref|XP_341828.2| PREDICTED: similar to Hypothetical protein MGC5... 37 0.077
emb|CAB16833.1| putative protein [Arabidopsis thaliana] gi|72706... 37 0.077
ref|NP_195378.2| late embryogenesis abundant domain-containing p... 37 0.077
gb|AAT06422.1| At4g36600 [Arabidopsis thaliana] 37 0.077
gb|AAX94822.1| transposon protein, putative, CACTA, En/Spm sub-c... 37 0.10
ref|XP_483090.1| hypothetical protein [Oryza sativa (japonica cu... 37 0.13
ref|ZP_00167063.2| COG2812: DNA polymerase III, gamma/tau subuni... 37 0.13
ref|NP_647902.1| CG15021-PA [Drosophila melanogaster] gi|2309300... 36 0.17
dbj|BAB73935.1| ferrichrome-iron receptor [Nostoc sp. PCC 7120] ... 36 0.17
pir||S61428 embryonic abundant protein group 3 precursor (clone ... 36 0.22
emb|CAG83152.1| unnamed protein product [Yarrowia lipolytica CLI... 36 0.22
ref|XP_030729.6| PREDICTED: hypothetical protein DKFZp434I1117 [... 36 0.22
>ref|NP_794806.1| amine oxidase, flavin-containing [Pseudomonas syringae pv. tomato
str. DC3000] gi|28855441|gb|AAO58501.1| amine oxidase,
flavin-containing [Pseudomonas syringae pv. tomato str.
DC3000]
Length = 625
Score = 44.3 bits (103), Expect = 6e-04
Identities = 40/119 (33%), Positives = 54/119 (44%), Gaps = 13/119 (10%)
Query: 24 SPATSPSP-SQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGL--GA 80
+P +P+P SQKS+ + FS++FG D+P P T + + GF + L GA
Sbjct: 467 APPKAPAPASQKSEG---------NFFSNLFGGGSADKPAEPTTPQASKPGFFSRLFGGA 517
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
APAP P AP+P+ A A K V E AK A A +AKK
Sbjct: 518 QEPAPAPAPLEKAPEPAQAPAPVAAPVPVVAPVVQPAKPPVKAESAKKPAAKA-DAAKK 575
>emb|CAG88778.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50423773|ref|XP_460471.1| unnamed protein product
[Debaryomyces hansenii]
Length = 708
Score = 39.3 bits (90), Expect = 0.020
Identities = 32/126 (25%), Positives = 53/126 (41%), Gaps = 18/126 (14%)
Query: 19 ICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGL 78
I +P+ PA SP P PP A P + P +P+ + +Q+ T
Sbjct: 505 ISQPNQPAVSPIPPVHQ--PPPLAQ---------SNPQTSYLPSIPQPSSQIQSSTHTPN 553
Query: 79 GAPVEAPAPGPFASAPQPSSEAAQQAFSY-------GKDKAYDIKDKAEVAYEDAKDKAE 131
A V+ P+P P +S P P + Q++ S G + + K +V + +K A+
Sbjct: 554 QASVKPPSPPPQSSLPPPQNSPPQKSTSIPSIQQNNGTKRDLEDKKDTKVTKKQSKKSAK 613
Query: 132 SAYQSA 137
S S+
Sbjct: 614 SQQNSS 619
>gb|EAL85260.1| LEA domain protein [Aspergillus fumigatus Af293]
Length = 1236
Score = 38.9 bits (89), Expect = 0.027
Identities = 19/33 (57%), Positives = 24/33 (72%)
Query: 110 DKAYDIKDKAEVAYEDAKDKAESAYQSAKKTVT 142
DKA +I ++A+ A E AKDKAE + AKKTVT
Sbjct: 372 DKAREIPEQADDAVEQAKDKAEETTEQAKKTVT 404
Score = 37.0 bits (84), Expect = 0.10
Identities = 19/49 (38%), Positives = 24/49 (48%)
Query: 94 PQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKKTVT 142
P+ + A QA + A KDK E + A DKAE + AK TVT
Sbjct: 255 PEQAENVASQAKGTADETANQAKDKVEETVDQADDKAEETTEQAKNTVT 303
>emb|CAB05543.1| Hypothetical protein K08H10.1 [Caenorhabditis elegans]
gi|17562388|ref|NP_505575.1| plant Late Embryo Abundant
related, contains Thrombospondin type 3 repeat (77.0 kD)
(lea-1) [Caenorhabditis elegans]
gi|2353333|gb|AAB69446.1| Ce-LEA [Caenorhabditis
elegans] gi|7505592|pir||T23507 hypothetical protein
K08H10.1 - Caenorhabditis elegans
Length = 733
Score = 38.9 bits (89), Expect = 0.027
Identities = 21/48 (43%), Positives = 27/48 (55%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A + + A A+ KDKA D KDKA A++ KDKA A+ S K
Sbjct: 486 ADAYNSAKDKASDAWDKTKDKAGDAKDKAADAWDTTKDKAGDAWDSTK 533
Score = 37.0 bits (84), Expect = 0.10
Identities = 19/48 (39%), Positives = 29/48 (59%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
+ A + + A A+ KDKA + KDKA A+++ KDKA +A+ S K
Sbjct: 599 SDAYNSAKDKASDAWDKTKDKAGEAKDKAGDAWDNTKDKAGNAWDSTK 646
Score = 34.7 bits (78), Expect = 0.50
Identities = 20/52 (38%), Positives = 28/52 (53%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKKTVT 142
A A + + A A+ KD A D KDKA A DAKDK++S + A ++
Sbjct: 515 ADAWDTTKDKAGDAWDSTKDHAADAKDKASDAAGDAKDKSKSLTEKAGDAIS 566
Score = 33.9 bits (76), Expect = 0.85
Identities = 16/51 (31%), Positives = 27/51 (52%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
G + S + +S+ A ++ D ++++KA AY AKDKA A+ K
Sbjct: 265 GAYDSVKEKASDVADSFKAHSTDSKDNVENKAADAYNTAKDKASDAWDKTK 315
Score = 33.9 bits (76), Expect = 0.85
Identities = 18/48 (37%), Positives = 26/48 (53%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A + + A A+ KDKA + KDK A++ KDKA A+ + K
Sbjct: 297 ADAYNTAKDKASDAWDKTKDKAGEAKDKMGDAWDTTKDKAGDAWDTTK 344
Score = 33.1 bits (74), Expect = 1.5
Identities = 17/38 (44%), Positives = 22/38 (57%)
Query: 101 AQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A++ KDKA D DK + DAKDKA A+ + K
Sbjct: 485 AADAYNSAKDKASDAWDKTKDKAGDAKDKAADAWDTTK 522
Score = 32.7 bits (73), Expect = 1.9
Identities = 15/51 (29%), Positives = 27/51 (52%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
G + + + +S+ A ++ D ++++KA AY AKDKA A+ K
Sbjct: 454 GAYDTVKEKASDIADSFKAHSTDSKDNVENKAADAYNSAKDKASDAWDKTK 504
Score = 32.3 bits (72), Expect = 2.5
Identities = 15/51 (29%), Positives = 27/51 (52%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
G + S + +S+ A ++ + ++++KA AY AKDKA A+ K
Sbjct: 567 GAYDSVKEKASDIADSFKAHSTNSKDNVENKASDAYNSAKDKASDAWDKTK 617
Score = 32.0 bits (71), Expect = 3.2
Identities = 17/42 (40%), Positives = 24/42 (56%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
+ + A A+ KDKA D K KA A++ KDKA A+ + K
Sbjct: 332 TKDKAGDAWDTTKDKAGDGKGKAGDAWDTTKDKASDAWDTTK 373
Score = 31.6 bits (70), Expect = 4.2
Identities = 16/38 (42%), Positives = 22/38 (57%)
Query: 101 AQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A++ KDKA D DK + +AKDKA A+ + K
Sbjct: 598 ASDAYNSAKDKASDAWDKTKDKAGEAKDKAGDAWDNTK 635
Score = 31.2 bits (69), Expect = 5.5
Identities = 18/50 (36%), Positives = 24/50 (48%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
G A + + A A+ KDKA + KDK A++ KDKA A A
Sbjct: 352 GKAGDAWDTTKDKASDAWDTTKDKAGEAKDKMGEAWDHTKDKAGEAKDKA 401
Score = 31.2 bits (69), Expect = 5.5
Identities = 23/64 (35%), Positives = 31/64 (47%), Gaps = 7/64 (10%)
Query: 82 VEAPAPGPFASAPQPSSEA---AQQAFSYGKDKAYDI----KDKAEVAYEDAKDKAESAY 134
VE A + SA +S+A + KDKA D KDKA A++ KDKA A+
Sbjct: 594 VENKASDAYNSAKDKASDAWDKTKDKAGEAKDKAGDAWDNTKDKAGNAWDSTKDKASDAW 653
Query: 135 QSAK 138
+ K
Sbjct: 654 DTTK 657
Score = 30.4 bits (67), Expect = 9.4
Identities = 14/30 (46%), Positives = 22/30 (72%)
Query: 103 QAFSYGKDKAYDIKDKAEVAYEDAKDKAES 132
+A+ + KDKA + KDKA A +DA+ K++S
Sbjct: 385 EAWDHTKDKAGEAKDKASDAADDAQGKSKS 414
>ref|NP_045241.1| 24 [Equid herpesvirus 4] gi|2605967|gb|AAC59539.1| 24 [Equine
herpesvirus 4] gi|11278231|pir||T42567 tegument protein
24 - equine herpesvirus 4 (strain NS80567)
Length = 3534
Score = 38.9 bits (89), Expect = 0.027
Identities = 39/130 (30%), Positives = 56/130 (43%), Gaps = 15/130 (11%)
Query: 22 PDSPATSPSPSQKSDSP-PSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGA 80
P PA +P+PS+ + +P PS + A P + P ++
Sbjct: 2827 PSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSK---PAAAPAPSK 2883
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFS---------YGKDKAYD-IKDKA-EVAYEDAKDK 129
P APAP A+AP PS AA A S KD+A D KD+A + A + AKD+
Sbjct: 2884 PAAAPAPSKPAAAPAPSKPAAAPAPSKPQNTLVAIVAKDQAKDQAKDQAKDQAKDQAKDQ 2943
Query: 130 AESAYQSAKK 139
A+ + K
Sbjct: 2944 AKDQAKDQAK 2953
Score = 33.1 bits (74), Expect = 1.5
Identities = 33/113 (29%), Positives = 44/113 (38%), Gaps = 6/113 (5%)
Query: 22 PDSPATSPSPSQKSDSP-PSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGA 80
P PA +P+PS+ + +P PS + A P + P ++
Sbjct: 2755 PSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSK---PAAAPAPSK 2811
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESA 133
P APAP A+AP PS AA A S K A K A +K A A
Sbjct: 2812 PAAAPAPSKPAAAPAPSKPAAAPAPS--KPAAAPAPSKPAAAPAPSKPAAAPA 2862
Score = 32.7 bits (73), Expect = 1.9
Identities = 33/113 (29%), Positives = 44/113 (38%), Gaps = 15/113 (13%)
Query: 22 PDSPATSPSPSQKSDSP-PSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGA 80
P PA +P+PS+ + +P PS + A P + P ++
Sbjct: 2719 PSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSK------------ 2766
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESA 133
P APAP A+AP PS AA A S K A K A +K A A
Sbjct: 2767 PAAAPAPSKPAAAPAPSKPAAAPAPS--KPAAAPAPSKPAAAPAPSKPAAAPA 2817
>gb|AAF10747.1| hypothetical protein [Deinococcus radiodurans]
gi|7472204|pir||B75429 hypothetical protein -
Deinococcus radiodurans (strain R1)
gi|32363429|sp|Q9RV58|UB72_DEIRA Protein DR1172
gi|15806191|ref|NP_294896.1| hypothetical protein DR1172
[Deinococcus radiodurans R1]
Length = 298
Score = 38.5 bits (88), Expect = 0.035
Identities = 20/46 (43%), Positives = 27/46 (58%)
Query: 95 QPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKKT 140
Q + AQQA S KDK D+K A A + AKDKA+ Q+ K++
Sbjct: 197 QNVKQGAQQAASDAKDKVQDVKADASRAADQAKDKAQDVAQNVKQS 242
Score = 35.4 bits (80), Expect = 0.29
Identities = 18/39 (46%), Positives = 24/39 (61%)
Query: 101 AQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
AQQA + KDK D+K A A + AKDKA+ Q+ K+
Sbjct: 163 AQQAAANVKDKVQDVKADASKAADQAKDKAQDVAQNVKQ 201
Score = 31.6 bits (70), Expect = 4.2
Identities = 16/43 (37%), Positives = 24/43 (55%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
+S+AA QA +D A ++K A+ A DAKDK + A +
Sbjct: 181 ASKAADQAKDKAQDVAQNVKQGAQQAASDAKDKVQDVKADASR 223
>emb|CAE61064.1| Hypothetical protein CBG04813 [Caenorhabditis briggsae]
Length = 732
Score = 38.5 bits (88), Expect = 0.035
Identities = 22/58 (37%), Positives = 34/58 (57%), Gaps = 10/58 (17%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAE----------VAYEDAKDKAESAYQSAK 138
A A + +AA+ A+ + K+KA ++KDKAE +E AKDKAE+A+ + K
Sbjct: 599 ADAYNKAKDAAEDAWDHTKNKAEEVKDKAEDKKDDYKERSGPWETAKDKAETAWDNTK 656
Score = 32.0 bits (71), Expect = 3.2
Identities = 17/42 (40%), Positives = 26/42 (61%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAES 132
A A + AA A+ KDKA ++K+KA+ +DAKD+ +S
Sbjct: 297 ADAYNSAKGAAGDAWDATKDKAGEMKEKAQDKADDAKDEGKS 338
Score = 30.4 bits (67), Expect = 9.4
Identities = 16/40 (40%), Positives = 23/40 (57%)
Query: 99 EAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
EA ++ D K KA AY+DAKDKA +A+++ K
Sbjct: 352 EATKERAQGAADTYNTAKYKAADAYDDAKDKAGNAWEATK 391
>ref|ZP_00416607.1| Protein of unknown function DUF224 [Azotobacter vinelandii AvOP]
gi|67088995|gb|EAM08461.1| Protein of unknown function
DUF224 [Azotobacter vinelandii AvOP]
Length = 451
Score = 38.1 bits (87), Expect = 0.045
Identities = 19/45 (42%), Positives = 25/45 (55%), Gaps = 3/45 (6%)
Query: 57 RNDEPVLPRTTENLQTGFGTGLGAPVEAPAPGPFASAPQPSSEAA 101
RN P+L R +E L FG G P+ PAP PF + P ++ AA
Sbjct: 156 RNASPLLARLSERL---FGLAAGRPLPEPAPAPFVATPAATAPAA 197
>ref|XP_341828.2| PREDICTED: similar to Hypothetical protein MGC59495 [Rattus
norvegicus]
Length = 525
Score = 37.4 bits (85), Expect = 0.077
Identities = 32/87 (36%), Positives = 41/87 (46%), Gaps = 17/87 (19%)
Query: 24 SPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRND-----EPVL--PRTTENLQTGFGT 76
S ++SPSP++ S S +S S F P N EPVL P T+ + T
Sbjct: 187 SSSSSPSPNEASSSSFPISSDGLSPCSVTFNPDSNKSSSPKEPVLGVPPTSTSAAT---- 242
Query: 77 GLGAPVEAPA---PGPFASAPQPSSEA 100
AP+ AP PGP ASAP P + A
Sbjct: 243 ---APISAPQVSIPGPPASAPPPCTSA 266
>emb|CAB16833.1| putative protein [Arabidopsis thaliana] gi|7270608|emb|CAB80326.1|
putative protein [Arabidopsis thaliana]
gi|25407743|pir||B85432 hypothetical protein AT4g36600
[imported] - Arabidopsis thaliana
Length = 347
Score = 37.4 bits (85), Expect = 0.077
Identities = 31/111 (27%), Positives = 48/111 (42%), Gaps = 17/111 (15%)
Query: 30 SPSQKSDSPPSFASWAYDKFSHVF-GPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAPG 88
S S+K+ FA YDK +H Y E V+ T+ + A G
Sbjct: 197 SASEKAGQAKDFA---YDKAAHAKDAAYNKAEDVIKMATDT---------SGEAKDSAYG 244
Query: 89 PFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
+ + E ++ A DKA+D+++ A A + AKDKA AY+S +
Sbjct: 245 TY----ERFKEGSKNAKDIASDKAHDVRETAGRAVDYAKDKANDAYESGSE 291
Score = 35.4 bits (80), Expect = 0.29
Identities = 20/41 (48%), Positives = 24/41 (57%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
+ EAA+ A +Y DKA D A A + A DKA SAY SA
Sbjct: 75 AEEAAESAKNYAYDKAGSAYDNAGYAKDFASDKAGSAYDSA 115
>ref|NP_195378.2| late embryogenesis abundant domain-containing protein / LEA
domain-containing protein [Arabidopsis thaliana]
Length = 335
Score = 37.4 bits (85), Expect = 0.077
Identities = 31/111 (27%), Positives = 48/111 (42%), Gaps = 17/111 (15%)
Query: 30 SPSQKSDSPPSFASWAYDKFSHVF-GPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAPG 88
S S+K+ FA YDK +H Y E V+ T+ + A G
Sbjct: 185 SASEKAGQAKDFA---YDKAAHAKDAAYNKAEDVIKMATDT---------SGEAKDSAYG 232
Query: 89 PFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
+ + E ++ A DKA+D+++ A A + AKDKA AY+S +
Sbjct: 233 TY----ERFKEGSKNAKDIASDKAHDVRETAGRAVDYAKDKANDAYESGSE 279
Score = 35.4 bits (80), Expect = 0.29
Identities = 20/41 (48%), Positives = 24/41 (57%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
+ EAA+ A +Y DKA D A A + A DKA SAY SA
Sbjct: 63 AEEAAESAKNYAYDKAGSAYDNAGYAKDFASDKAGSAYDSA 103
>gb|AAT06422.1| At4g36600 [Arabidopsis thaliana]
Length = 350
Score = 37.4 bits (85), Expect = 0.077
Identities = 31/111 (27%), Positives = 48/111 (42%), Gaps = 17/111 (15%)
Query: 30 SPSQKSDSPPSFASWAYDKFSHVF-GPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAPG 88
S S+K+ FA YDK +H Y E V+ T+ + A G
Sbjct: 200 SASEKAGQAKDFA---YDKAAHAKDAAYNKAEDVIKMATDT---------SGEAKDSAYG 247
Query: 89 PFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
+ + E ++ A DKA+D+++ A A + AKDKA AY+S +
Sbjct: 248 TY----ERFKEGSKNAKDIASDKAHDVRETAGRAVDYAKDKANDAYESGSE 294
Score = 35.4 bits (80), Expect = 0.29
Identities = 20/41 (48%), Positives = 24/41 (57%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
+ EAA+ A +Y DKA D A A + A DKA SAY SA
Sbjct: 78 AEEAAESAKNYAYDKAGSAYDNAGYAKDFASDKAGSAYDSA 118
Score = 30.4 bits (67), Expect = 9.4
Identities = 34/126 (26%), Positives = 45/126 (34%), Gaps = 18/126 (14%)
Query: 28 SPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAP 87
S + D S+A W DK S G + + + +N + A A
Sbjct: 46 SDAAHDTKDKTASWAGWVSDKISTGLGGKKAEAEEAAESAKNY--AYDKAGSAYDNAGYA 103
Query: 88 GPFASAPQPSS-EAAQQAFSYGKDKAYDIKD---------------KAEVAYEDAKDKAE 131
FAS S+ ++A A Y DKA D KD KA AYE A +
Sbjct: 104 KDFASDKAGSAYDSAHNAKHYAYDKAGDAKDMAYDKTGQAKYMAYDKAGSAYEKAGQAKD 163
Query: 132 SAYQSA 137
AY A
Sbjct: 164 MAYDKA 169
>gb|AAX94822.1| transposon protein, putative, CACTA, En/Spm sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 873
Score = 37.0 bits (84), Expect = 0.10
Identities = 32/121 (26%), Positives = 50/121 (40%), Gaps = 2/121 (1%)
Query: 21 RPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGA 80
R SP PSP Q P A + + + P + P P+ T L A
Sbjct: 353 RAPSPPAPPSPPQAPPPSPPHAPAPFPRHAPAPSPPQAPAPTPPQAPA--LTPPQAPLPA 410
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKKT 140
P ++ AP AP +++ A+ S KD YD + VAY ++ K + +S +K
Sbjct: 411 PSKSRAPQAPPPAPTRATKKAKFDASKNKDPRYDCTQEELVAYVASEVKRQFKPRSPEKK 470
Query: 141 V 141
+
Sbjct: 471 I 471
>ref|XP_483090.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|42408489|dbj|BAD09669.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 301
Score = 36.6 bits (83), Expect = 0.13
Identities = 33/143 (23%), Positives = 57/143 (39%), Gaps = 8/143 (5%)
Query: 1 MGNAKIVLVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDE 60
M A + LV + LA C + + A +P+ + KS S + +S ++ S P ++ E
Sbjct: 103 MARADLALVAVLLAACAAVALA-AEAQAPAAAPKSSSSSNSSSGSHTSPSKAPSPSKSPE 161
Query: 61 P------VLPRTTENLQTGFGTGLGAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYD 114
P + G AP EAP A +P+ E+ + S KD +
Sbjct: 162 KSGKAPAAAPPKAAAAKAPSGKS-EAPSEAPDAESGAESPEAGEESGKSPASAPKDSSSS 220
Query: 115 IKDKAEVAYEDAKDKAESAYQSA 137
++ E + D+ D + A
Sbjct: 221 SSEEEEASSPDSGDMEDETAAEA 243
>ref|ZP_00167063.2| COG2812: DNA polymerase III, gamma/tau subunits [Ralstonia eutropha
JMP134]
Length = 730
Score = 36.6 bits (83), Expect = 0.13
Identities = 32/96 (33%), Positives = 39/96 (40%), Gaps = 14/96 (14%)
Query: 21 RPDSPATSPSPSQK-----SDSPPSFASWAYD--------KFSHVFGPYRNDEPVLPRTT 67
RP PA +P+P++K +PP AS D VFGPY N+ LP
Sbjct: 506 RPVPPAAAPAPARKPAPQAESAPPWEASGGSDIPPWEDLPPDLPVFGPYDNEPVSLPTKA 565
Query: 68 ENLQTGFGTGLGAPVEAPAPGPFASAPQPSSEAAQQ 103
AP EAP P P A A P AA +
Sbjct: 566 AAPARASKPEAPAP-EAPRPRPVAKAVPPRPPAAAE 600
>ref|NP_647902.1| CG15021-PA [Drosophila melanogaster] gi|23093008|gb|AAF47902.2|
CG15021-PA [Drosophila melanogaster]
gi|17945382|gb|AAL48746.1| RE17165p [Drosophila
melanogaster]
Length = 420
Score = 36.2 bits (82), Expect = 0.17
Identities = 24/75 (32%), Positives = 30/75 (40%), Gaps = 11/75 (14%)
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAP 81
PD P P+PS+ PP + + +GP P P+ T G G P
Sbjct: 194 PDQPKPRPTPSRPQPPPPPPPR---PQPTPGYGPPPPPPPPKPQPTP--------GYGPP 242
Query: 82 VEAPAPGPFASAPQP 96
P PGP APQP
Sbjct: 243 TPPPGPGPAQPAPQP 257
>dbj|BAB73935.1| ferrichrome-iron receptor [Nostoc sp. PCC 7120]
gi|17229728|ref|NP_486276.1| ferrichrome-iron receptor
[Nostoc sp. PCC 7120] gi|25318835|pir||AE2085
ferrichrome-iron receptor [imported] - Nostoc sp.
(strain PCC 7120)
Length = 858
Score = 36.2 bits (82), Expect = 0.17
Identities = 16/41 (39%), Positives = 21/41 (51%)
Query: 33 QKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTG 73
Q +DS +ASW +FG RN+EP P T E + G
Sbjct: 633 QPTDSTSIYASWTNSFNPQIFGKTRNNEPFKPETAEQFEVG 673
>pir||S61428 embryonic abundant protein group 3 precursor (clone PM10) - soybean
gi|414977|gb|AAA91965.1| 51 kDa seed maturation protein
Length = 473
Score = 35.8 bits (81), Expect = 0.22
Identities = 27/103 (26%), Positives = 46/103 (44%), Gaps = 2/103 (1%)
Query: 36 DSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAPGPFASAPQ 95
++ S+ WA +K S G +++D+ TT N + + T + A +
Sbjct: 61 EAAESWTEWAKEKLSEGLG-FKHDQESKESTT-NKVSDYATDTAQKSKDYATDTAQKSKD 118
Query: 96 PSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
+ +AAQ++ Y D A KD A + +KD A A Q +K
Sbjct: 119 YAGDAAQKSKDYAGDAAQKTKDYASDTAQTSKDYAGDAAQKSK 161
>emb|CAG83152.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50546863|ref|XP_500901.1| hypothetical protein
[Yarrowia lipolytica]
Length = 914
Score = 35.8 bits (81), Expect = 0.22
Identities = 29/115 (25%), Positives = 51/115 (44%), Gaps = 2/115 (1%)
Query: 26 ATSPSPSQKSDSPP-SFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEA 84
A + PS SP + A A + +HV P + T+ T T L +P A
Sbjct: 442 ARNYKPSGARRSPQITRAEMASPRSTHVTPASPRGSPAVADTSPAAPTAPATALASPASA 501
Query: 85 PAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
A P A+ QP+ +AA + ++ ++KA++ E+ + E ++A+K
Sbjct: 502 SA-SPVAAPSQPAMDAAAALLAQQEETMRQAREKAKLRKEEEQRVEEERRKAARK 555
>ref|XP_030729.6| PREDICTED: hypothetical protein DKFZp434I1117 [Homo sapiens]
Length = 828
Score = 35.8 bits (81), Expect = 0.22
Identities = 27/74 (36%), Positives = 28/74 (37%), Gaps = 15/74 (20%)
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVL-PRTTENLQTGFGTGLGA 80
PDSP TS + SP SF S AY PVL P T N T
Sbjct: 55 PDSPVTSVDLETPAHSPLSFVSTAY--------------PVLGPGVTVNPGTSLSVFTAL 100
Query: 81 PVEAPAPGPFASAP 94
P PAPGP P
Sbjct: 101 PFATPAPGPAHRPP 114
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.311 0.128 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 274,697,673
Number of Sequences: 2540612
Number of extensions: 13224641
Number of successful extensions: 57636
Number of sequences better than 10.0: 468
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 354
Number of HSP's that attempted gapping in prelim test: 56374
Number of HSP's gapped (non-prelim): 1267
length of query: 142
length of database: 863,360,394
effective HSP length: 118
effective length of query: 24
effective length of database: 563,568,178
effective search space: 13525636272
effective search space used: 13525636272
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0283.2


