

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0278.5.1
(30 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
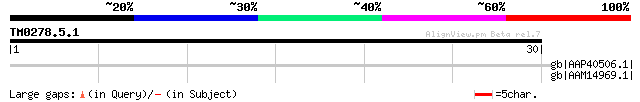
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP40506.1| putative small nuclear ribonucleoprotein Prp4p [A... 36 0.26
gb|AAM14969.1| putative small nuclear ribonucleoprotein Prp4p [A... 36 0.26
>gb|AAP40506.1| putative small nuclear ribonucleoprotein Prp4p [Arabidopsis
thaliana]
Length = 554
Score = 36.2 bits (82), Expect = 0.26
Identities = 17/30 (56%), Positives = 22/30 (72%), Gaps = 3/30 (10%)
Query: 4 IVTVSHDRTIKLWSSNMTNDNE---HTMDV 30
I TVSHDRTIKLW+S+ +D + TMD+
Sbjct: 523 IATVSHDRTIKLWTSSGNDDEDEEKETMDI 552
>gb|AAM14969.1| putative small nuclear ribonucleoprotein Prp4p [Arabidopsis
thaliana] gi|2618685|gb|AAB84332.1| putative small
nuclear ribonucleoprotein Prp4p [Arabidopsis thaliana]
gi|58652074|gb|AAW80862.1| At2g41500 [Arabidopsis
thaliana] gi|7488209|pir||T02445 probable U4/U6 small
nuclear ribonucleoprotein [imported] - Arabidopsis
thaliana gi|15227373|ref|NP_181681.1| WD-40 repeat
family protein / small nuclear ribonucleoprotein
Prp4p-related [Arabidopsis thaliana]
gi|3123130|sp|O22212|PRP4_ARATH Hypothetical Trp-Asp
repeats containing protein At2g41500
Length = 554
Score = 36.2 bits (82), Expect = 0.26
Identities = 17/30 (56%), Positives = 22/30 (72%), Gaps = 3/30 (10%)
Query: 4 IVTVSHDRTIKLWSSNMTNDNE---HTMDV 30
I TVSHDRTIKLW+S+ +D + TMD+
Sbjct: 523 IATVSHDRTIKLWTSSGNDDEDEEKETMDI 552
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.128 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,923,166
Number of Sequences: 2540612
Number of extensions: 804156
Number of successful extensions: 6479
Number of sequences better than 10.0: 2
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 6471
Number of HSP's gapped (non-prelim): 8
length of query: 30
length of database: 863,360,394
effective HSP length: 6
effective length of query: 24
effective length of database: 848,116,722
effective search space: 20354801328
effective search space used: 20354801328
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0278.5.1


