

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0269b.2
(2064 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
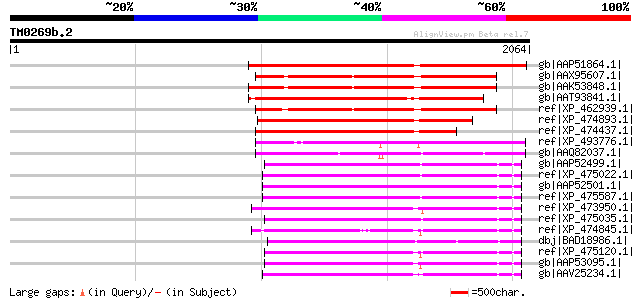
Sequences producing significant alignments: (bits) Value
gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sati... 1107 0.0
gb|AAX95607.1| RNase H, putative [Oryza sativa (japonica cultiva... 983 0.0
gb|AAK53848.1| Putative retroelement [Oryza sativa] 968 0.0
gb|AAT93841.1| putative polyprotein [Oryza sativa (japonica cult... 967 0.0
ref|XP_462939.1| putative gag-pol protein [Oryza sativa (japonic... 964 0.0
ref|XP_474893.1| OSJNBa0021F22.20 [Oryza sativa (japonica cultiv... 905 0.0
ref|XP_474437.1| OSJNBa0070M12.15 [Oryza sativa (japonica cultiv... 856 0.0
ref|XP_493776.1| unnamed protein product [Oryza sativa (japonica... 855 0.0
gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum] 794 0.0
gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonic... 708 0.0
ref|XP_475022.1| OSJNBb0093G06.3 [Oryza sativa (japonica cultiva... 708 0.0
gb|AAP52501.1| putative gag-pol precursor [Oryza sativa (japonic... 706 0.0
ref|XP_475587.1| putative polyprotein [Oryza sativa (japonica cu... 703 0.0
ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultiv... 702 0.0
ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultiva... 700 0.0
ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultiva... 697 0.0
dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera] 697 0.0
ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cu... 696 0.0
gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cul... 694 0.0
gb|AAV25234.1| putative polyprotein [Oryza sativa (japonica cult... 693 0.0
>gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37530550|ref|NP_919577.1| putative
retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|18581535|gb|AAK52540.2| Putative
retroelement pol polyprotein [Oryza sativa]
Length = 1580
Score = 1107 bits (2862), Expect = 0.0
Identities = 553/1117 (49%), Positives = 752/1117 (66%), Gaps = 31/1117 (2%)
Query: 950 RLDCIYDDGPLGFEKPVVDFSKKMEA-----QDPLEEVD--LGDGLEKRPTYISALIDPE 1002
+++ + D L E P F ME +D ++++D G GL I P
Sbjct: 479 KIEIVPADSQLKMENPSYCFEGVMEGSNVYTKDTVDDLDDKQGQGLMSADDLEEIDISPA 538
Query: 1003 L---KDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLV 1059
+ + +++LLKEF+DCF W+Y EMPG SR +VE +LP+K +P +QLPRR D+L
Sbjct: 539 IGQGRHLLIELLKEFRDCFTWEYYEMPGHSRSIVEHRLPLKPGVRPHQQLPRRCKADMLE 598
Query: 1060 KIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIA 1119
+K EI+RL FIR RY +W++++VPVIKKNGK+RVCIDFR LN ATPKDEY M +A
Sbjct: 599 PVKAEIKRLYDAGFIRPCRYAEWVSSIVPVIKKNGKVRVCIDFRYLNKATPKDEYPMLVA 658
Query: 1120 EMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGA 1179
+ +VD+A+GH+ LS +DG +GYNQIF+AEED+ KT FRCPGA+G +EWVV+ F LK+AGA
Sbjct: 659 DQLVDAASGHKILSFMDGNAGYNQIFMAEEDIHKTTFRCPGAIGLFEWVVITFVLKSAGA 718
Query: 1180 TYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAF 1239
TYQR MN I+HD I ++VYIDD+VVKS +DH+ LRK FER RKYGLKMNP KCAF
Sbjct: 719 TYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEIEDHIADLRKVFERTRKYGLKMNPTKCAF 778
Query: 1240 GVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKT 1299
GV AG FLGF+VH++GIE+ + AI PP K +LQ ++GKINF+RRFI+ LS K
Sbjct: 779 GVSAGQFLGFLVHERGIEVTQRSINAIKKIKPPEDKTELQEMIGKINFVRRFISILSGKL 838
Query: 1300 KSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGS 1359
+ F+PLLRLK + F W AE QKA D +K YLSSPP++ P +G P +LY+SA D +I S
Sbjct: 839 EPFTPLLRLKADQQFAWGAEQQKALDNIKEYLSSPPVLIPPQKGIPFRLYLSAGDKSISS 898
Query: 1360 MLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHY 1419
+L QE E KER +FYLSR L +AETRY+ +EKLCLCLYFSC +L++Y+ + V
Sbjct: 899 VLIQELE-RKERVVFYLSRRLLNAETRYSPVEKLCLCLYFSCTRLRHYLLSNECTVICKA 957
Query: 1420 DIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGI 1479
D++K+MLS PIL RIGKW +LTE+ L Y KA+KGQAIA+F+ D + I V +
Sbjct: 958 DVVKYMLSAPILKGRIGKWIFSLTEFDLRYESPKAIKGQAIANFIVDYR--DDSIGSVEV 1015
Query: 1480 QPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIAL 1539
PW LFFDGS +G GIG+ I+SPRG +F + IK +NN+AEYEA+ GL++L +
Sbjct: 1016 VPWTLFFDGSVCTHGCGIGLVIISPRGACFEFAYTIKPYATNNQAEYEAVLKGLQLLKEV 1075
Query: 1540 GAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEAN 1599
A + + GDS LVI QL EY+C ++ L Y K L+ +F L HVSR N EAN
Sbjct: 1076 EADTIEIMGDSLLVISQLAGEYECKNDTLIVYNEKCQELMREFRLVTLKHVSREQNIEAN 1135
Query: 1600 ELAQIASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQNPVGTTD 1659
+LAQ ASGY K +K + +++ VI +DWR + YLQ+P +
Sbjct: 1136 DLAQGASGY---KLMIKHV----------QIEVAVI---TADDWRYYVYQYLQDPSQSAS 1179
Query: 1660 RKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWIL 1719
RK +Y+A+ Y+++ +EL+ + +DG LLKCLS D A +AI VH+G+CG HQ+ KMKW+L
Sbjct: 1180 RKLRYKALKYILLDDELYYRTIDGVLLKCLSTDQAKVAIGEVHEGICGTHQSAHKMKWLL 1239
Query: 1720 FRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINP 1779
G +WPT+++DC Y K CQDCQ+ IQ PAS ++ IIKPWPFRGW +D+IG INP
Sbjct: 1240 RCAGYFWPTMLEDCFRYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMINP 1299
Query: 1780 CSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISS-IVYRFGLPESLTTDQGTVFV 1838
SS+ HK+I+VA DYFTKWVEAIPL+ V I+F+ I+YR G+P+++TTDQG++FV
Sbjct: 1300 PSSKGHKFILVATDYFTKWVEAIPLKKVDSGDAIQFVQEHIIYRVGIPQTITTDQGSIFV 1359
Query: 1839 GQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLW 1898
+ FA+S GIKLL S+PYYAQANGQ EA+NK+LI LIK+ + P+ WH L + LW
Sbjct: 1360 SDEFIQFADSMGIKLLNSSPYYAQANGQAEASNKSLIKLIKRKISDYPRQWHTRLAEALW 1419
Query: 1899 AYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDE 1958
+YR + S P++L Y +A+LP EV + S R + Q ++ +++Y N+M DE +L +
Sbjct: 1420 SYRMACHGSIQVPPYKLVYRHKAILPWEVRIGSRRTELQNDLTADEYSNLMADEREDLVQ 1479
Query: 1959 ERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPF 2018
RL L + + K+R+ + YNKKV K F G+ +WK++LPI +D ++ KW PNWEGPF
Sbjct: 1480 SRLRALAKVIKDKERVTRHYNKKVVPKDFSEGELIWKLILPIGTRDSKFGKWSPNWEGPF 1539
Query: 2019 TVEKVLVNNAYSIKELGGN-RQMTINGKYLKTYKPTL 2054
+ KV+ AY ++ L G +NGKYLK Y P++
Sbjct: 1540 QIHKVVSKGAYMLQGLDGEVYDRALNGKYLKKYYPSV 1576
>gb|AAX95607.1| RNase H, putative [Oryza sativa (japonica cultivar-group)]
Length = 2282
Score = 983 bits (2540), Expect = 0.0
Identities = 495/960 (51%), Positives = 654/960 (67%), Gaps = 30/960 (3%)
Query: 977 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELK 1036
D LEE+D+G G RPT+I + E + ++++LLKEF+DCFAW+Y EMPGLSR +VE +
Sbjct: 1334 DDLEEIDIGPGDRPRPTFICKNLSTEFRAKLIELLKEFRDCFAWEYYEMPGLSRSIVEHR 1393
Query: 1037 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKM 1096
LPIK +P +Q RR D+L +K EI+RL FIR RY +W++++VPVIKKN
Sbjct: 1394 LPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPCRYAEWVSSIVPVIKKN--- 1450
Query: 1097 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAF 1156
DLN ATPKDEY MP+A+ +VD+A+G++ LS +DG GYNQIF+AEED+ KTAF
Sbjct: 1451 -------DLNKATPKDEYPMPVADQLVDAASGNKILSYMDGNVGYNQIFMAEEDIHKTAF 1503
Query: 1157 RCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLL 1216
RCPGA+G +EWVVM FGLK+AGATYQR MN I+HD I ++VYIDD+VVKS +DH+
Sbjct: 1504 RCPGAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEIEDHIA 1563
Query: 1217 HLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKK 1276
LRK FER RKYGLKMNP KCAFGV G FLGF+VH++GIE+ + AI PP +K
Sbjct: 1564 DLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIEVTQRSVNAIKKIQPPENKT 1623
Query: 1277 QLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPI 1336
+LQ + GKINF+RRFI+ LS + + F+PLLRLK + F W AE QKA D++K YLSSPP+
Sbjct: 1624 ELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWGAEQQKALDDIKEYLSSPPV 1683
Query: 1337 MASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLC 1396
+ P +G +LY+SA + +IGS+L QE D KER +FYLSR L D ETRY+ +EKLCLC
Sbjct: 1684 LIPPQKGISFRLYLSAGEKSIGSVLIQE-LDGKERVVFYLSRRLLDVETRYSPMEKLCLC 1742
Query: 1397 LYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVK 1456
LYFSC +L +Y+ + V D+IK+MLS PIL R+GKW +LTE+ L Y KA+K
Sbjct: 1743 LYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGKWIFSLTEFDLRYESPKAIK 1802
Query: 1457 GQAIADFLADQTLPQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIK 1516
GQAIADF+ + + I V + PW FFDGS + GIG+ I+SPRG KF + IK
Sbjct: 1803 GQAIADFIVEHC--DDSIGSVEVVPWTSFFDGSVCTHDCGIGLVIISPRGACFKFAYTIK 1860
Query: 1517 SNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKAS 1576
+NN+AEYEA+ GL++L + A + + GDS LVI QL EY+C S+ L Y K
Sbjct: 1861 PYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDSLLVISQLAGEYECKSDTLMVYNEKCQ 1920
Query: 1577 NLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNPSDLDILVID 1636
L+ +F L HVSR N EAN+LAQ ASGY K +K +VK + I
Sbjct: 1921 ELMQEFRLVTLKHVSREQNIEANDLAQGASGY---KPMIK---DVKAE----------IA 1964
Query: 1637 NMAPNDWRKPIVDYLQNPVGTTDRKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFI 1696
+ DWR + YL NP + RK +Y+A+ Y ++ +EL+ + +DG LLKCLS D A +
Sbjct: 1965 AITAGDWRYDVHQYLHNPSQSASRKLRYKALKYTLLDDELYYRTIDGVLLKCLSADQAKV 2024
Query: 1697 AISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASE 1756
AI VH+G+CG HQ+ KMKW+L R +WPT+++DC +Y K CQDCQ+ IQ PAS
Sbjct: 2025 AIGEVHEGICGTHQSAHKMKWLLRRARYFWPTMLEDCFKYYKGCQDCQKFGAIQRAPASA 2084
Query: 1757 LHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFI 1816
++ IIKPWPFRGW +D+IG I+ SS+ HK+I+VA DYFTK VEAIPL+ V I+F+
Sbjct: 2085 MNPIIKPWPFRGWGIDMIGMISRPSSKGHKFILVATDYFTKLVEAIPLKKVDSGDAIQFV 2144
Query: 1817 SS-IVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLI 1875
I+YRFG+P+++TTDQG++FV + F +S GIKLL S+PYYAQANGQ EA+NK+LI
Sbjct: 2145 QEHIIYRFGIPQTITTDQGSIFVSDEFVQFVDSMGIKLLNSSPYYAQANGQAEASNKSLI 2204
Query: 1876 SLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQ 1935
LIK+ P+ WH L + LW+YR + S P++L YG EA+LP EV + S R +
Sbjct: 2205 KLIKRKNSDYPRQWHTRLDEALWSYRMACHGSIQVPPYKLVYGHEAILPWEVRIGSRRTE 2264
>gb|AAK53848.1| Putative retroelement [Oryza sativa]
Length = 2014
Score = 968 bits (2502), Expect = 0.0
Identities = 496/994 (49%), Positives = 661/994 (65%), Gaps = 37/994 (3%)
Query: 950 RLDCIYDDGPLGFEKPVVDFSKKMEA-----QDPLEEVD--LGDGLEKRPTYISALIDPE 1002
+++ + D L E P F +E +D ++++D G G I PE
Sbjct: 1032 KIEIVPADSQLKMENPSCYFEGVVEGSNVYIKDTVDDLDDKQGQGFISADDLEEIDIGPE 1091
Query: 1003 LKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIK 1062
+ ++++LLKEF+DCFAW+Y EMPGLSR +VE +LPIK +P +Q RR D+L +K
Sbjct: 1092 FRAKLIELLKEFRDCFAWEYYEMPGLSRSIVEHRLPIKPGVRPHQQPLRRCKADMLEPVK 1151
Query: 1063 EEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMM 1122
EI+RL FIR RY +W++++VPVIKKN DLN ATPKDEY MP+A+ +
Sbjct: 1152 VEIKRLYDACFIRPCRYAEWVSSIVPVIKKN----------DLNKATPKDEYPMPVADQL 1201
Query: 1123 VDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQ 1182
VD+A+G++ LS +DG GYNQIF+AEED+ KTAFRCPGA+G +EWVVM FGLK+AGATYQ
Sbjct: 1202 VDAASGNKILSYMDGNVGYNQIFMAEEDIHKTAFRCPGAIGLFEWVVMTFGLKSAGATYQ 1261
Query: 1183 RVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVI 1242
R MN I+HD I ++VYIDD+VVKS +DH+ LRK FER RKYGLKMNP KCAFGV
Sbjct: 1262 RAMNYIYHDLIGWLVEVYIDDVVVKSKEIEDHIADLRKVFERTRKYGLKMNPTKCAFGVS 1321
Query: 1243 AGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSF 1302
G FLGF+VH++GIE+ + AI PP +K +LQ + GKINF+RRFI+ LS + + F
Sbjct: 1322 VGQFLGFLVHERGIEVTQRSVNAIKKIQPPENKTELQEMNGKINFVRRFISYLSGRLEPF 1381
Query: 1303 SPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLA 1362
+PLLRLK + F W AE QKA D++K YLSSPP++ P +G +LY+SA + +IGS+L
Sbjct: 1382 TPLLRLKADQQFTWGAEQQKALDDIKEYLSSPPVLIPPQKGISFRLYLSAGEKSIGSVLI 1441
Query: 1363 QEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDII 1422
QE D KER +FYLSR L D ETRY+ +EKLCLCLYFSC +L +Y+ + V D+I
Sbjct: 1442 QE-LDGKERVVFYLSRRLLDVETRYSPMEKLCLCLYFSCTRLSHYLLSNECTVICKADVI 1500
Query: 1423 KHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPW 1482
K+MLS PIL R+GKW +LTE+ L Y KA+KGQAIADF+ + + I V + PW
Sbjct: 1501 KYMLSAPILKGRVGKWIFSLTEFDLRYESPKAIKGQAIADFIVEHC--DDSIGSVEVVPW 1558
Query: 1483 KLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAK 1542
FFDGS + GIG+ I+SPRG KF + IK +NN+AEYEA+ GL++L + A
Sbjct: 1559 TSFFDGSVCTHDCGIGLVIISPRGACFKFAYTIKPYATNNQAEYEAVLKGLQLLKEVEAD 1618
Query: 1543 NVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELA 1602
+ + GDS LVI QL EY+C S+ L Y K L+ +F L HVSR N EAN+LA
Sbjct: 1619 AIEIMGDSLLVISQLAGEYECKSDTLMVYNEKCQELMQEFRLVTLKHVSREQNIEANDLA 1678
Query: 1603 QIASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQNPVGTTDRKT 1662
Q ASGY K +K +VK + I + DWR + YL NP + RK
Sbjct: 1679 QGASGY---KPMIK---DVKAE----------IAAITAGDWRYDVHQYLHNPSQSASRKL 1722
Query: 1663 KYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQ 1722
+Y+A+ Y ++ +EL+ + +DG LLKCLS D A +AI VH+G+CG HQ+ KMKW+L R
Sbjct: 1723 RYKALKYTLLDDELYYRTIDGVLLKCLSADQAKVAIGEVHEGICGTHQSAHKMKWLLRRA 1782
Query: 1723 GMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSS 1782
+WPT+++DC +Y K CQDCQ+ IQ PAS ++ IIKPWPFRGW +D+IG I+ SS
Sbjct: 1783 RYFWPTMLEDCFKYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMISRPSS 1842
Query: 1783 RQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISS-IVYRFGLPESLTTDQGTVFVGQK 1841
+ HK+I+VA DYFTK VEAIPL+ V I+F+ I+YRFG+P+++TTDQG++FV +
Sbjct: 1843 KGHKFILVATDYFTKLVEAIPLKKVDSGDAIQFVQEHIIYRFGIPQTITTDQGSIFVSDE 1902
Query: 1842 VASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYR 1901
F +S GIKLL S+PYYAQANGQ EA+NK+LI LIK+ P+ WH L + LW+YR
Sbjct: 1903 FVQFVDSMGIKLLNSSPYYAQANGQAEASNKSLIKLIKRKNSDYPRQWHTRLDEALWSYR 1962
Query: 1902 NSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQ 1935
+ S P++L YG EA+LP EV + S R +
Sbjct: 1963 MACHGSIQVPPYKLVYGHEAILPWEVRIGSRRTE 1996
>gb|AAT93841.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1739
Score = 967 bits (2500), Expect = 0.0
Identities = 487/934 (52%), Positives = 643/934 (68%), Gaps = 42/934 (4%)
Query: 951 LDCIYDDGPLGFEKPVVDFSKKMEAQDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKL 1010
+D +YD GF M A D LEE+D+G G RPT++S + PE + ++++L
Sbjct: 844 VDDLYDKQGQGF----------MSADD-LEEIDIGPGDRPRPTFVSKNLSPEFRTKLIEL 892
Query: 1011 LKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLR 1070
LKEF+DCFAW+Y EMPGLSR +VE +LPIK +P +Q RR D+L +K EI+RL
Sbjct: 893 LKEFRDCFAWEYYEMPGLSRSIVEHRLPIKPGVRPRQQPLRRCKADMLEPVKAEIKRLYD 952
Query: 1071 CKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHE 1130
FIR RY W++++VPVIKKNGK+RVCIDFRDLN ATPKDEY MP+A+ +VD+A+G++
Sbjct: 953 AGFIRPWRYAKWVSSIVPVIKKNGKVRVCIDFRDLNKATPKDEYPMPVADQLVDAASGNK 1012
Query: 1131 YLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFH 1190
LS +DG +GYNQIF+AEED+ KTAFRCP A+G +EWVVM FGLK+AGATYQR MN I+H
Sbjct: 1013 ILSFMDGNAGYNQIFMAEEDIHKTAFRCPCAIGLFEWVVMTFGLKSAGATYQRAMNYIYH 1072
Query: 1191 DFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFV 1250
D I + VYIDD+VVKS +D + LRK FER RKYGLKMNP KCAFGV AG FLGF+
Sbjct: 1073 DLIGWLVGVYIDDVVVKSKEIEDQIADLRKVFERTRKYGLKMNPTKCAFGVSAGQFLGFL 1132
Query: 1251 VHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKK 1310
VH +GIE+ + AI PP +K +LQ ++GKI+F+RRFI+NLS + + F+PLLRLK
Sbjct: 1133 VHDRGIEVTQRSVNAIKKIQPPENKTELQEMIGKIHFVRRFISNLSGRLEPFTPLLRLKA 1192
Query: 1311 EDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKE 1370
+ F AE QKA D++K YLSSPP++ P +G P +LY+SA + +IGS+L QE E KE
Sbjct: 1193 DQQFTCGAEQQKALDDIKEYLSSPPVLIPPHKGIPFRLYLSAGEKSIGSVLIQELE-GKE 1251
Query: 1371 RAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPI 1430
R +FYLSR L +AETRY+ ++KLCLCLYFSC +L++Y+ + V D++K+MLS PI
Sbjct: 1252 RVVFYLSRRLLNAETRYSPVKKLCLCLYFSCTRLRHYLLSNECTVICKADVVKYMLSAPI 1311
Query: 1431 LHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGSS 1490
L R+GKW +LTE+ Y KA+KGQAIADF+ + + I V I PW LFFDGS
Sbjct: 1312 LKGRVGKWIFSLTEFDHRYESPKAIKGQAIADFIVEHR--DDSIGSVEIVPWTLFFDGSV 1369
Query: 1491 HKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGDS 1550
+G GIG+ I+SPRG +F + IK +NN+AEYEA+ GL++L + A + + GDS
Sbjct: 1370 CTHGCGIGLVIISPRGACFEFAYTIKPYATNNQAEYEAVLKGLQLLKKVEADTIEIMGDS 1429
Query: 1551 ELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYMV 1610
LVI QL EY+C ++ L Y K L+ +F R+ N EAN+LAQ ASGY
Sbjct: 1430 LLVISQLAGEYECKNDTLMVYNEKCQELMKEF---------RLQNIEANDLAQGASGY-- 1478
Query: 1611 DKCRLKELIEVKEKLNPSDLDILV-IDNMAPNDWRKPIVDYLQNPVGTTDRKTKYRAMSY 1669
P D+ V I M +DWR + YL NP + RK +Y+A+ Y
Sbjct: 1479 ---------------KPMIKDVKVEIAAMTADDWRYDVHRYLSNPSQSASRKLRYKALKY 1523
Query: 1670 VIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTI 1729
++ +EL+ + +DG LLKCLS D A +AI VH+G+CG HQ+ KMKW+L R G +W T+
Sbjct: 1524 TLLDDELYYRTIDGVLLKCLSADQAKVAIGEVHEGICGTHQSAHKMKWLLRRAGYFWSTM 1583
Query: 1730 MKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYII 1789
++DC Y K CQDCQ+ IQ PAS ++ IIKPWPFRGW +D+IG INP SS+ HK+I+
Sbjct: 1584 LEDCFRYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMINPPSSKGHKFIL 1643
Query: 1790 VAIDYFTKWVEAIPLQNVTQDTVIEFISS-IVYRFGLPESLTTDQGTVFVGQKVASFAES 1848
VA DYFTKWVEAIPL+ V I+F+ I+YRFG P+++TTDQG++FV + FA+S
Sbjct: 1644 VATDYFTKWVEAIPLKKVDSGDAIQFVQEYIIYRFGTPQTITTDQGSIFVSDEFVQFADS 1703
Query: 1849 WGIKLLTSTPYYAQANGQVEAANKTLISLIKKHV 1882
IKLL S+PYYAQANGQ EA+NK+LI LIK+ +
Sbjct: 1704 MSIKLLNSSPYYAQANGQAEASNKSLIKLIKRKI 1737
>ref|XP_462939.1| putative gag-pol protein [Oryza sativa (japonica cultivar-group)]
Length = 2273
Score = 964 bits (2492), Expect = 0.0
Identities = 488/960 (50%), Positives = 645/960 (66%), Gaps = 39/960 (4%)
Query: 977 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELK 1036
D LEE+D+G G RPT+I + E + ++++LLKEF+DCFAW+Y EMPGLSR +VE +
Sbjct: 1334 DDLEEIDIGPGDRPRPTFICKNLSTEFRAKLIELLKEFRDCFAWEYYEMPGLSRSIVEHR 1393
Query: 1037 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKM 1096
LPIK +P +Q RR D+L +K EI+RL FIR RY +W++N
Sbjct: 1394 LPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPCRYAEWVSN----------- 1442
Query: 1097 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAF 1156
LN ATPKDEY MP+A+ +VD+A+G++ LS +DG GYNQIF+AEED+ KTAF
Sbjct: 1443 --------LNKATPKDEYPMPVADQLVDAASGNKILSYMDGNVGYNQIFMAEEDIHKTAF 1494
Query: 1157 RCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLL 1216
RCPGA+G +EWVVM FGLK+AGATYQR MN I+HD I ++VYIDD+VVKS +DH+
Sbjct: 1495 RCPGAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEIEDHIA 1554
Query: 1217 HLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKK 1276
LRK FER RKYGLKMNP KCAFGV G FLGF+VH++GIE+ + AI PP +K
Sbjct: 1555 DLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIEVTQRSVNAIKKIQPPENKT 1614
Query: 1277 QLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPI 1336
+LQ + GKINF+RRFI+ LS + + F+PLLRLK + F W AE QKA D++K YLSSPP+
Sbjct: 1615 ELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWGAEQQKALDDIKEYLSSPPV 1674
Query: 1337 MASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLC 1396
+ P +G +LY+SA + +IGS+L QE D KER +FYLSR L D ETRY+ +EKLCLC
Sbjct: 1675 LIPPQKGISFRLYLSAGEKSIGSVLIQE-LDGKERVVFYLSRRLLDVETRYSPMEKLCLC 1733
Query: 1397 LYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVK 1456
LYFSC +L +Y+ + V D+IK+MLS PIL R+GKW +LTE+ L Y KA+K
Sbjct: 1734 LYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGKWIFSLTEFDLRYESPKAIK 1793
Query: 1457 GQAIADFLADQTLPQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIK 1516
GQAIADF+ + + I V + PW FFDGS + GIG+ I+SPRG KF + IK
Sbjct: 1794 GQAIADFIVEHC--DDSIGSVEVVPWTSFFDGSVCTHDCGIGLVIISPRGACFKFAYTIK 1851
Query: 1517 SNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKAS 1576
+NN+AEYEA+ GL++L + A + + GDS LVI QL EY+C S+ L Y K
Sbjct: 1852 PYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDSLLVISQLAGEYECKSDTLMVYNEKCQ 1911
Query: 1577 NLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNPSDLDILVID 1636
L+ +F L HVSR N EAN+LAQ ASGY K +K +VK + I
Sbjct: 1912 ELMQEFRLVTLKHVSREQNIEANDLAQGASGY---KPMIK---DVKAE----------IA 1955
Query: 1637 NMAPNDWRKPIVDYLQNPVGTTDRKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFI 1696
+ DWR + YL NP + RK +Y+A+ Y ++ +EL+ + +DG LLKCLS D A +
Sbjct: 1956 AITAGDWRYDVHQYLHNPSQSASRKLRYKALKYTLLDDELYYRTIDGVLLKCLSADQAKV 2015
Query: 1697 AISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASE 1756
AI VH+G+CG HQ+ KMKW+L R +WPT+++DC +Y K CQDCQ+ IQ PAS
Sbjct: 2016 AIGEVHEGICGTHQSAHKMKWLLRRARYFWPTMLEDCFKYYKGCQDCQKFGAIQRAPASA 2075
Query: 1757 LHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFI 1816
++ IIKPWPFRGW +D+IG I+ SS+ HK+I+VA DYFTK VEAIPL+ V I+F+
Sbjct: 2076 MNPIIKPWPFRGWGIDMIGMISRPSSKGHKFILVATDYFTKLVEAIPLKKVDSGDAIQFV 2135
Query: 1817 SS-IVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLI 1875
I+YRFG+P+++TTDQG++FV + F +S GIKLL S+PYYAQANGQ EA+NK+LI
Sbjct: 2136 QEHIIYRFGIPQTITTDQGSIFVSDEFVQFVDSMGIKLLNSSPYYAQANGQAEASNKSLI 2195
Query: 1876 SLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQ 1935
LIK+ P+ WH L + LW+YR + S P++L YG EA+LP EV + S R +
Sbjct: 2196 KLIKRKNSDYPRQWHTRLDEALWSYRMACHGSIQVPPYKLVYGHEAILPWEVRIGSRRTE 2255
>ref|XP_474893.1| OSJNBa0021F22.20 [Oryza sativa (japonica cultivar-group)]
gi|21743195|emb|CAD40050.1| OSJNBa0085C10.2 [Oryza sativa
(japonica cultivar-group)] gi|38346487|emb|CAE03726.2|
OSJNBa0021F22.20 [Oryza sativa (japonica cultivar-group)]
Length = 861
Score = 905 bits (2339), Expect = 0.0
Identities = 448/854 (52%), Positives = 595/854 (69%), Gaps = 22/854 (2%)
Query: 987 GLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPV 1046
G + RPT+IS + E + ++++LLKEF+DCFAW+Y EMPGLSR +VE +LPIK +P
Sbjct: 26 GDQPRPTFISKNLSSEFRTKLIELLKEFRDCFAWEYYEMPGLSRSIVEHRLPIKPGVRPR 85
Query: 1047 KQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLN 1106
+Q PRR D+L +K EI+RL FI RY +W++++VPVIKKNGK+RVCIDFRDLN
Sbjct: 86 QQPPRRCKADMLEPVKAEIKRLYDAGFIHPYRYAEWVSSIVPVIKKNGKVRVCIDFRDLN 145
Query: 1107 AATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYE 1166
ATPKDEY MP+A+ + D+A+G++ LS +D +GYNQIF+AEED+ KTAFRCP A+G +E
Sbjct: 146 KATPKDEYPMPVADQLFDAASGNKILSFMDRNTGYNQIFMAEEDIHKTAFRCPSAIGLFE 205
Query: 1167 WVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMR 1226
WVVM F LK+AGATYQR MN I+HD I + ++V IDD+VVKS DDH+ LRK FER R
Sbjct: 206 WVVMTFDLKSAGATYQRAMNYIYHDLIGSLVEVDIDDVVVKSKEVDDHIADLRKVFERTR 265
Query: 1227 KYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKIN 1286
KYGLKMNP KCAFGV A FLGF+VH++GIE+ + AI PP +K +LQ ++GKIN
Sbjct: 266 KYGLKMNPTKCAFGVSARQFLGFLVHERGIEVTQRSVNAIQKIKPPENKTELQEMIGKIN 325
Query: 1287 FLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPM 1346
F+RRFI+NLS + + F+PLLRLK + F W AE QKA D +K YLSSPP++ P +G P
Sbjct: 326 FVRRFISNLSGRLEPFTPLLRLKADQQFTWGAEQQKALDNIKEYLSSPPVLIPPQKGIPF 385
Query: 1347 KLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKY 1406
+LY+SA + + GS+L QE E KER +FY+SR L DAETRY+ +EKLCLCLYF C +L++
Sbjct: 386 RLYLSAGEKSNGSVLIQELE-GKERVVFYISRRLLDAETRYSPMEKLCLCLYFLCTRLRH 444
Query: 1407 YIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLAD 1466
Y+ + V D++++MLS PIL +IG W +LTE+ L Y KA+KGQAIADF+ +
Sbjct: 445 YLLSNECTVICKADVVRYMLSAPILKGQIGNWIFSLTEFDLRYESPKAIKGQAIADFIVE 504
Query: 1467 QTLPQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEY 1526
+ I V I PW LFFDGS +G GIG+ I+SPRG +F + IKS +NN+AEY
Sbjct: 505 HR--DDSIGSVEIVPWTLFFDGSVCTHGCGIGLVIISPRGACFEFAYTIKSYATNNQAEY 562
Query: 1527 EALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEAR 1586
EA+ GL++L + A + + GDS LVI QL EY+C S+ Y K L+ +F
Sbjct: 563 EAVLKGLQLLKEVEANIIEIMGDSLLVISQLAGEYECKSDTFIVYNEKCLELMKEFRLVT 622
Query: 1587 LGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNPSDLDILV-IDNMAPNDWRK 1645
L HVSR N EAN+LAQ ASGY P D+ V I M+ +DWR
Sbjct: 623 LKHVSREQNLEANDLAQGASGY-----------------KPMIKDVKVEIAAMSADDWRY 665
Query: 1646 PIVDYLQNPVGTTDRKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGL 1705
+ YL NP + RK +Y+A+ Y ++G+EL+ + +DG LLKCLS D A +AI VH+G+
Sbjct: 666 DVHQYLSNPSQSASRKLRYKALKYTLLGDELYYRVIDGVLLKCLSADQAKVAIGEVHEGI 725
Query: 1706 CGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWP 1765
CG HQ+ KMKW+L R G +WPT+++ Y K CQDCQ+ IQ PAS ++ IIKPWP
Sbjct: 726 CGTHQSAHKMKWLLRRTGYFWPTMLEVYFRYYKGCQDCQKFGAIQRAPASAMNPIIKPWP 785
Query: 1766 FRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISS-IVYRFG 1824
FRGW +D+IG INP SS+ HK+I+VA DYFTKWVEAIPL+ V I+F+ I+Y+FG
Sbjct: 786 FRGWEIDMIGMINPPSSKGHKFILVATDYFTKWVEAIPLKKVDSGDAIQFVQEHIIYQFG 845
Query: 1825 LPESLTTDQGTVFV 1838
LP+++T DQG++F+
Sbjct: 846 LPQTITMDQGSIFM 859
>ref|XP_474437.1| OSJNBa0070M12.15 [Oryza sativa (japonica cultivar-group)]
gi|38345832|emb|CAE01862.2| OSJNBa0070M12.15 [Oryza
sativa (japonica cultivar-group)]
Length = 1685
Score = 856 bits (2212), Expect = 0.0
Identities = 420/800 (52%), Positives = 555/800 (68%), Gaps = 21/800 (2%)
Query: 977 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELK 1036
D L+EVD+G G RPT+IS + E + ++++LLKEF+DCFAW+Y EMPGLSR +VE +
Sbjct: 904 DDLDEVDIGPGDRPRPTFISKNLSSEFRTKLMELLKEFRDCFAWEYYEMPGLSRSIVEHR 963
Query: 1037 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKM 1096
LP+K +P +Q PRR D+L +K EI+RL FI RY +W++++VPVIKKNGK+
Sbjct: 964 LPLKPGIRPHQQPPRRCKADMLEPVKAEIKRLYDAGFIHPCRYAEWVSSIVPVIKKNGKV 1023
Query: 1097 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAF 1156
RVCIDFRDLN ATPKDEY MP+A+ +VD+A+GH+ LS +DG +GYNQIF+ EED+ KT F
Sbjct: 1024 RVCIDFRDLNKATPKDEYPMPVADQLVDAASGHKILSFMDGNAGYNQIFMTEEDIHKTVF 1083
Query: 1157 RCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLL 1216
RCPGA+G +EWVVM FGLK+AGATYQR MN I+HD I ++VYIDD+VVKS +DH+
Sbjct: 1084 RCPGAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGWLVEVYIDDVVVKSREIEDHIA 1143
Query: 1217 HLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKK 1276
LRK FER RKYGLKMNP KCAFGV AG FLGF+VH++GIEI + AI PP +K
Sbjct: 1144 DLRKVFERTRKYGLKMNPTKCAFGVSAGQFLGFLVHERGIEIPQRSINAIKKIKPPGNKT 1203
Query: 1277 QLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPI 1336
+LQ ++GKINF+RRFI+NLS K + F+PLLRL+ + F W AE QKA D +K YLSSPP+
Sbjct: 1204 ELQEMIGKINFVRRFISNLSGKLEPFTPLLRLRADQQFTWGAEQQKALDNIKEYLSSPPV 1263
Query: 1337 MASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLC 1396
+ P +G P LY+SA D +IGS+L QE E KER +FYLSR L DAETRY+ +EKLCLC
Sbjct: 1264 LIPPQKGIPFWLYLSAGDKSIGSVLIQELE-GKERVVFYLSRRLLDAETRYSPVEKLCLC 1322
Query: 1397 LYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVK 1456
LYFSC KL++Y+ + V D+IK+MLS PIL R+GKW +LTE+ L + KA+K
Sbjct: 1323 LYFSCTKLRHYLLSNECTVVCKADVIKYMLSAPILKVRVGKWIFSLTEFDLRHESPKAIK 1382
Query: 1457 GQAIADFLADQTLPQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIK 1516
GQA+ADF+ I V I PW LFFDGS +G GIG+ I++P G +F + I
Sbjct: 1383 GQAVADFIVGHR--DGSIGLVDIMPWVLFFDGSVCSHGYGIGLVIIAPWGASFEFAYAIT 1440
Query: 1517 SNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKAS 1576
+ +NN+AEYEA+ GL++L + A + + GDS LVI QL EY+C + L Y K
Sbjct: 1441 HHVTNNQAEYEAVLKGLQLLQEVAADAIEIMGDSLLVISQLAGEYECKDDTLMVYYEKCR 1500
Query: 1577 NLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNPSDLDI-LVI 1635
L+++F L H+SR N EAN+L+Q ASGY P DI + I
Sbjct: 1501 TLISEFRLVTLRHISREQNIEANDLSQGASGY-----------------KPMTKDIEIEI 1543
Query: 1636 DNMAPNDWRKPIVDYLQNPVGTTDRKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAF 1695
+ +DWR + YLQNP + RK +Y+A+ Y ++ ++L+ + +DG LLKCLS D A
Sbjct: 1544 ATITADDWRYDVFQYLQNPSQSALRKLRYKALKYTLLDDDLYYRTIDGVLLKCLSADQAK 1603
Query: 1696 IAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPAS 1755
+AI +H+G+CG HQ+ KMKW+L R G +WPT+++DC Y K CQDCQ+ IQ PAS
Sbjct: 1604 VAIGEIHEGICGTHQSAHKMKWLLRRAGYFWPTMLEDCFRYYKGCQDCQKFGAIQRAPAS 1663
Query: 1756 ELHSIIKPWPFRGWALDLIG 1775
++ II+PWPFRGW +D+IG
Sbjct: 1664 AMNPIIQPWPFRGWGIDMIG 1683
>ref|XP_493776.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
gi|9927274|dbj|BAB08213.2| Similar to Arabidopsis
thaliana chromosome II BAC F26H6; putative retroelement
pol polyprotein (AC006920) [Oryza sativa (japonica
cultivar-group)]
Length = 2876
Score = 855 bits (2208), Expect = 0.0
Identities = 458/1110 (41%), Positives = 667/1110 (59%), Gaps = 47/1110 (4%)
Query: 977 DPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELK 1036
D L E++LG + RP ++S ++ E ++ L EF+DCFAW Y EMPGL V K
Sbjct: 1776 DELVEMNLGTEDDPRPIFVSGMLTEEEREDYRSFLMEFRDCFAWTYKEMPGLDSRVATHK 1835
Query: 1037 LPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKM 1096
L I +PVKQ PRR P+ ++ E++RL+ FI+ +Y WLAN+VPV KKNG++
Sbjct: 1836 LAIDPQFRPVKQPPRRLRPEFQDQVIAEVDRLINVGFIKEIQYPRWLANIVPVEKKNGQV 1895
Query: 1097 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAF 1156
RVC+DFRDLN A PKD++ +PI EM+VDS G+ LS GYNQI + D TAF
Sbjct: 1896 RVCVDFRDLNRACPKDDFPLPITEMVVDSTTGYGALS------GYNQIKMDLLDAFDTAF 1949
Query: 1157 RCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLL 1216
R P G + + VMPFGLKNAGATYQR M + D I ++ Y+DD+VVK+ + H
Sbjct: 1950 RTPK--GNFYYTVMPFGLKNAGATYQRAMQFVLDDLIHHSVECYVDDMVVKTKDHEHHQE 2007
Query: 1217 HLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKK 1276
LR FER+R++ LKMNPLKCAF V +G FLGFV+ +GIEI K KAIL+ PP K
Sbjct: 2008 DLRIVFERLRRHQLKMNPLKCAFAVQSGVFLGFVIRHRGIEIEPKKIKAILNMPPPQELK 2067
Query: 1277 QLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPI 1336
L+ L GK+ ++RRFI+NLS + + FS L+ KK F W+ E Q FD +K YL +PP+
Sbjct: 2068 DLRKLQGKLAYIRRFISNLSGRIQPFSKLM--KKGTPFVWDEECQNGFDSIKRYLLNPPV 2125
Query: 1337 MASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLC 1396
+A+P++G+P+ LYI+ +IG++LAQ +++ KE A +YLSR + AE Y+ IEKLCL
Sbjct: 2126 LAAPVKGRPLILYIATQPASIGALLAQHNDEGKEVACYYLSRTMVGAEQNYSPIEKLCLA 2185
Query: 1397 LYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVK 1456
L F+ KL++Y+ + + + D I+++LS+P+L R+GKWAL + EY +T+ P KA+K
Sbjct: 2186 LIFALKKLRHYMLAHQIQLIARADPIRYVLSQPVLTGRLGKWALLMMEYDITFVPQKAIK 2245
Query: 1457 GQAIADFLADQTLP-----------QEIITYVGIQPWKLFFDGSSHKN----GT-----G 1496
GQA+A+FLA +P +EI T + W+L+FDG+S K+ GT G
Sbjct: 2246 GQALAEFLATHPMPDDSPLIANLPDEEIFTAELQEQWELYFDGASRKDINPDGTPRRRAG 2305
Query: 1497 IGMFIVSPRGIPTKFKFRI-KSNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIK 1555
G+ +P+G F + K CSNNEAEYEAL GL + +++ +++ GDS L+I+
Sbjct: 2306 AGLVFKTPQGGVIYHSFSLLKEECSNNEAEYEALIFGLLLALSMEVRSLRAHGDSRLIIR 2365
Query: 1556 QLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRL 1615
Q+ Y+ L Y T A L+ KF+ + HV R N A+ LA++A+ +
Sbjct: 2366 QINNIYEVRKPELVPYYTVARRLMDKFEHIEVIHVPRSKNAPADALAKLAAALVFQGDNP 2425
Query: 1616 KELIEVKEK--------LNPSDLDILVIDNMAPNDWRKPIVDYLQN---PVGTTDRKTKY 1664
+++ V+E+ L P +++I++ ++ DWR+P +DY ++ P +R+
Sbjct: 2426 AQIV-VEERWLLPAVLELIPEEVNIIITNSAEEEDWRQPFLDYFKHGSLPEDPVERRQLQ 2484
Query: 1665 RAM-SYVIMGNELFKKNV-DGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQ 1722
R + SY+ L+K++ LL+C+ +A + VH G+CG HQ+G KM +
Sbjct: 2485 RRLPSYIYKAGVLYKRSYGQEVLLRCVDRSEANRVLQEVHHGVCGGHQSGPKMYHSIRLV 2544
Query: 1723 GMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSS 1782
G YWP IM DC++ AK C CQ H +H P + LH + WPF W +D+IG INP SS
Sbjct: 2545 GYYWPGIMADCLKTAKTCHGCQIHDNFKHQPPAPLHPTVPSWPFDAWGIDVIGLINPPSS 2604
Query: 1783 RQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISS-IVYRFGLPESLTTDQGTVFVGQK 1841
R H++I+ A DYF+KW EA+PL+ V VI F+ I+YRFG+P +T+D F QK
Sbjct: 2605 RGHRFILTATDYFSKWAEAVPLREVKSSDVINFLERHIIYRFGVPHRITSDNAKAFKSQK 2664
Query: 1842 VASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYR 1901
+ F E + IK ST YY QANG EA NKTL ++KK V + ++WH L + LWAYR
Sbjct: 2665 IYRFMEKYKIKWNYSTGYYPQANGMAEAFNKTLGKILKKTVDKHRRDWHDRLYEALWAYR 2724
Query: 1902 NSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERL 1961
+ + T ATP+ L YG EAVLP+E+ L S R+ +E+ ++ + EL ++EERL
Sbjct: 2725 VTVRTPTQATPYSLVYGNEAVLPLEIQLPSLRVAIHDELTKDEQIRLRFQELDAVEEERL 2784
Query: 1962 LVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVE 2021
L L + + +AY+K V+ + F G+ V + PI K K+ P WEGP+ +E
Sbjct: 2785 GALQNLELYRQNMVRAYDKLVKQRVFRKGELVLVLRRPIVVTHKMKGKFEPKWEGPYVIE 2844
Query: 2022 KVLVNNAYSIKELGGNRQM-TINGKYLKTY 2050
+ AY + + G++ M ING++LK Y
Sbjct: 2845 QAYDGGAYQLIDHQGSQPMPPINGRFLKKY 2874
>gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 794 bits (2051), Expect = 0.0
Identities = 425/1113 (38%), Positives = 660/1113 (59%), Gaps = 48/1113 (4%)
Query: 976 QDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVEL 1035
Q+ LE V+LG R I A ++ +K R++++L+E+ + FAW Y +MPGL ++V
Sbjct: 1158 QEELEVVNLGTEDATREIKIGAALEDSVKRRLIEMLREYVEIFAWSYQDMPGLDTDIVVH 1217
Query: 1036 KLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGK 1095
+LP++E VKQ RR PD+ KIKEE+++ F+ Y W+AN+VPV KK+GK
Sbjct: 1218 RLPLREGCPSVKQKLRRTSPDMATKIKEEVQKQWDAGFLAVTSYPPWMANIVPVPKKDGK 1277
Query: 1096 MRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTA 1155
+R+C+D+RDLN A+PKD++ +P +++VD+ A S +DG+SGYNQI +A ED+ KT
Sbjct: 1278 VRMCVDYRDLNRASPKDDFPLPHIDVLVDNTAQSSVFSFMDGFSGYNQIKMAPEDMEKTT 1337
Query: 1156 FRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHL 1215
F P GT+ + VMPFGLKNAGATYQR M T+FHD + ++VY+DD++ KS + ++HL
Sbjct: 1338 FITP--WGTFCYKVMPFGLKNAGATYQRAMTTLFHDMMHKEIEVYVDDMIAKSQTEEEHL 1395
Query: 1216 LHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSK 1275
++L+K F+R+RK+ L++NP KC FGV +G LGF+V +KGIE++ K KAI + P ++
Sbjct: 1396 VNLQKLFDRLRKFKLRLNPNKCTFGVRSGKLLGFIVSEKGIEVDPAKVKAIQEMPEPKTE 1455
Query: 1276 KQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPP 1335
KQ++ LG++N++ RFI++L+ + LLR K A +W + QKAFD++K YL PP
Sbjct: 1456 KQVRGFLGRLNYIARFISHLTATCEPIFKLLR--KNQAIKWNDDCQKAFDKIKEYLQKPP 1513
Query: 1336 IMASPIRGKPMKLYISATDGTIGSMLAQEDEDS-KERAIFYLSRVLNDAETRYTMIEKLC 1394
I+ P+ G+P+ +Y+S T+ ++G +L + DE KE AI+YLS+ D ETRY+++EK C
Sbjct: 1514 ILIPPVPGRPLIMYLSVTENSMGCVLGRHDESGRKEHAIYYLSKKFTDCETRYSLLEKTC 1573
Query: 1395 LCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKA 1454
L ++ +L+ Y+ ++ S D +K++ KP L R+ +W + LTEY + Y KA
Sbjct: 1574 CALAWAARRLRQYMLNHTTLLISKMDPVKYIFEKPALTGRVARWQMILTEYDIQYTSQKA 1633
Query: 1455 VKGQAIADFLADQTL----------PQEIITYVGIQ---------------PWKLFFDGS 1489
+KG ++D+LA+Q + P E I Y+ ++ W L FDG+
Sbjct: 1634 IKGSILSDYLAEQPIEDYQPMMFEFPDEDIMYLKMKDCKEPLVEEGPDPDDKWTLMFDGA 1693
Query: 1490 SHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGD 1549
+ NG G+G +++P+G F R+ + +NNEAEYEA G+E I L K + + GD
Sbjct: 1694 VNMNGNGVGAVLINPKGAHMPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKTLDIFGD 1753
Query: 1550 SELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYM 1609
S LV+ Q+ ++ +L Y +L F + +L HV R +NQ A+ LA ++S
Sbjct: 1754 SALVVNQVNGDWNTNQPHLIPYRDYTRRILTFFKKVKLYHVPRDENQMADALATLSSMIK 1813
Query: 1610 VDKCRLKELIEVKEKLNPSDL---DILVIDNMAPNDWRKPIVDYLQN---PVGTT--DRK 1661
V+ + V P+ + + +VID W I ++L+ P G + D+K
Sbjct: 1814 VNWWNHVPHVAVNRLERPAYVFAAESVVIDE---KPWYYDIKNFLKTQEYPEGASKNDKK 1870
Query: 1662 TKYRAMS--YVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWIL 1719
T R Y+ + L+K+N D LL+C+ +A + + VH+G G H G M L
Sbjct: 1871 TLRRLAGSFYLNQDDVLYKRNFDMVLLRCMDRPEADMLMQEVHEGSFGTHAGGHAMAKKL 1930
Query: 1720 FRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINP 1779
R G YW T+ DC +YA++C CQ +A HVP S L+ + PWPF W +D+IG+I P
Sbjct: 1931 LRAGYYWMTMESDCFKYARKCHKCQIYADRVHVPPSPLNVMNSPWPFAMWGIDMIGKIEP 1990
Query: 1780 CSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFI-SSIVYRFGLPESLTTDQGTVFV 1838
+S H++I+VAIDYFTKWVEA N+T+ V FI I+ R+G+PE + TD G+
Sbjct: 1991 TASNGHRFILVAIDYFTKWVEAASYANITKQVVTRFIKKEIICRYGVPERIITDNGSNLN 2050
Query: 1839 GQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLW 1898
+ + + + I+ S+PY + NG VEAANK + +++K V K+WH+ L L
Sbjct: 2051 NKMMKELCKDFKIEHHNSSPYRPKMNGAVEAANKNIKKIVRKMVVTY-KDWHEMLPFALH 2109
Query: 1899 AYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDE 1958
YR S + STGATP+ L YG EAVLPVEV + S R+ ++ ++ +EL ++E
Sbjct: 2110 GYRTSVRTSTGATPYSLVYGMEAVLPVEVEIPSLRVLLDVKLDEAEWIRTRFNELSLIEE 2169
Query: 1959 ERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPF 2018
RL V+ + R+ +A+++KVR +S+ +GD V K +LP ++ KW PN+EGP+
Sbjct: 2170 RRLAVVCHGQLYQRRMKRAFDQKVRPRSYQIGDLVLKRILPPGTDNR--GKWTPNYEGPY 2227
Query: 2019 TVEKVLVNNAYSIKELGG-NRQMTINGKYLKTY 2050
V+KV A + + G + +N +K Y
Sbjct: 2228 VVKKVFSGGALMLTTMDGEDFPSPVNSDVVKKY 2260
>gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37531820|ref|NP_920212.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|22128689|gb|AAM92802.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 2017
Score = 708 bits (1828), Expect = 0.0
Identities = 397/1041 (38%), Positives = 594/1041 (56%), Gaps = 35/1041 (3%)
Query: 1015 KDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFI 1074
KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+ +LL FI
Sbjct: 976 KDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFI 1035
Query: 1075 RTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSL 1134
+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS AG E LS
Sbjct: 1036 KEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSF 1095
Query: 1135 LDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIE 1194
LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++ F I
Sbjct: 1096 LDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIG 1153
Query: 1195 TFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKK 1254
++ Y+DD+VVK+ +DD +L L ++F +R + +K+NP KC FGV +G LGF+V +
Sbjct: 1154 RNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHR 1213
Query: 1255 GIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAF 1314
GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L LKK D F
Sbjct: 1214 GIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LKKTDDF 1271
Query: 1315 RWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDED-----SK 1369
+W E QKAF++ K L+ PP++ASP +P+ LY+SAT + ++L E E+
Sbjct: 1272 QWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKV 1331
Query: 1370 ERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKP 1429
+R I+++S VL D++TRY ++KL + + KL +Y + V V + + + +L
Sbjct: 1332 QRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP-LGDILHNR 1390
Query: 1430 ILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGS 1489
+ RI KWAL L L++ P ++K QA+ADF+A+ T QE T ++ W + FDGS
Sbjct: 1391 EANGRIAKWALELMSLDLSFKPRISIKSQALADFVAEWTECQEDTTVKKMEHWTMHFDGS 1450
Query: 1490 SHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGD 1549
+GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG K ++V+GD
Sbjct: 1451 KRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGD 1510
Query: 1550 SELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELA------Q 1603
S+LV+ Q+ KE+ C+ +N+ Y + L KFD L HV R DN+ A+ LA +
Sbjct: 1511 SQLVVNQVMKEWSCLDDNMKAYRQEVRKLEDKFDGLELSHVLRHDNEAADRLANFGSKRE 1570
Query: 1604 IASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQNPVGTTDR--- 1660
+A + + + K+ + + + M DWR+P++ +L + D+
Sbjct: 1571 VAPSDVFVEHLYTPTVPHKDTTQAAGIHDVA---MVETDWREPLIRFLTSQELPQDKDEA 1627
Query: 1661 -KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWIL 1719
+ R+ YV+ EL+KK+ G L +C+S ++ + +H G+CG H A +
Sbjct: 1628 ERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKA 1687
Query: 1720 FRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINP 1779
+RQG +WPT + D + + C+ CQ A H+PA EL +I WPF W LD++G
Sbjct: 1688 YRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKK 1747
Query: 1780 CSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTTDQGTVFVG 1839
+ ++ VAID F+KW+EA P+ +T D +F +IV+RFG+P + TD GT F G
Sbjct: 1748 AVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPNRIITDNGTQFTG 1806
Query: 1840 QKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK----NWHQTLGQ 1895
F E +GIK+ ++ + +NGQVE AN ++ IK V + K W Q L
Sbjct: 1807 GVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPS 1866
Query: 1896 VLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVN 1955
VLW+ R +P +TG +PF L YG EA+LP EV +S R + E E Y +D+L
Sbjct: 1867 VLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEEDRVDDLHR 1923
Query: 1956 LDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWE 2015
L+E R L R + + +N+ VR+++F++GD V + + + + K P WE
Sbjct: 1924 LEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QTTRDRHKLSPLWE 1979
Query: 2016 GPFTVEKVLVNNAYSIKELGG 2036
GPF + +V +Y +K G
Sbjct: 1980 GPFIISEVTRPGSYRLKREDG 2000
>ref|XP_475022.1| OSJNBb0093G06.3 [Oryza sativa (japonica cultivar-group)]
gi|38347562|emb|CAE04995.2| OSJNBb0093G06.3 [Oryza sativa
(japonica cultivar-group)]
Length = 1986
Score = 708 bits (1827), Expect = 0.0
Identities = 398/1049 (37%), Positives = 596/1049 (55%), Gaps = 35/1049 (3%)
Query: 1007 MVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1066
++ L+ KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+
Sbjct: 937 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 996
Query: 1067 RLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1126
+LL FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS
Sbjct: 997 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1056
Query: 1127 AGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMN 1186
AG E LS LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++
Sbjct: 1057 AGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQ 1114
Query: 1187 TIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1246
F I ++ Y+DD+VVK+ +DD + L ++F +R + +K+NP KC FGV +G
Sbjct: 1115 RCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKL 1174
Query: 1247 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLL 1306
LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L
Sbjct: 1175 LGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL- 1233
Query: 1307 RLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDE 1366
LKK D F+W E QKAF++ K L+ PP++ASP +P+ LY+SAT + ++L E E
Sbjct: 1234 -LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQVVSTVLVVERE 1292
Query: 1367 D-----SKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDI 1421
+ +R I+++S VL D++TRY ++KL + + KL +Y + V V + +
Sbjct: 1293 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP- 1351
Query: 1422 IKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQP 1481
+ +L + RI KWAL L +++ P ++K QA+ADF+A+ T QE I+
Sbjct: 1352 LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQEDTPVEKIEH 1411
Query: 1482 WKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGA 1541
W + FDGS +GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG
Sbjct: 1412 WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGI 1471
Query: 1542 KNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANEL 1601
K ++V+GDS+LV+ Q+ KE+ CI +N+ Y + L KFD L HV R +N+ A+ L
Sbjct: 1472 KRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRL 1531
Query: 1602 A------QIASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQNPV 1655
A + A + + + K+ +D +V M DWR+P + +L +
Sbjct: 1532 ANFGSKREAAPSDVFVEHLYTPTVPHKDTTQDADTHDVV---MVEADWREPFIRFLSSQE 1588
Query: 1656 GTTDR----KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQA 1711
D+ + R+ YV+ +EL+KK+ G L +C+S ++ + +H G+CG H A
Sbjct: 1589 LPQDKDEAERISRRSKLYVMHESELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAA 1648
Query: 1712 GIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWAL 1771
+ +RQG +WPT + D + + C+ CQ A H+PA EL +I WPF W L
Sbjct: 1649 ARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGL 1708
Query: 1772 DLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTT 1831
D++G + ++ VAID F+KW+EA P+ +T D +F +IV+RFG+P + T
Sbjct: 1709 DMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPNRIIT 1767
Query: 1832 DQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK---- 1887
D GT F G F E +GIK+ ++ + +NGQVE AN ++ IK V + K
Sbjct: 1768 DNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAG 1827
Query: 1888 NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWN 1947
W Q L VLW+ R +P +TG +PF L YG EA+LP EV +S R + E E Y
Sbjct: 1828 KWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEE 1884
Query: 1948 MMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRY 2007
+D+L L+E R L R + + +N+ VR+++F++GD V + + + +
Sbjct: 1885 DRVDDLHRLEEAREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QTTRDR 1940
Query: 2008 RKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
K P WEGPF + +V +Y +K G
Sbjct: 1941 HKLSPLWEGPFIISEVTRPGSYRLKREDG 1969
>gb|AAP52501.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37531824|ref|NP_920214.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|22128685|gb|AAM92798.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 1986
Score = 706 bits (1822), Expect = 0.0
Identities = 400/1049 (38%), Positives = 593/1049 (56%), Gaps = 35/1049 (3%)
Query: 1007 MVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1066
++ L+ KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+
Sbjct: 937 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 996
Query: 1067 RLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1126
+LL FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS
Sbjct: 997 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1056
Query: 1127 AGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMN 1186
AG E LS LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++
Sbjct: 1057 AGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQ 1114
Query: 1187 TIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1246
F I ++ Y+DD+VVK+ +DD + L ++F +R + +K+NP KC FGV +G
Sbjct: 1115 RCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKL 1174
Query: 1247 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLL 1306
LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L
Sbjct: 1175 LGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL- 1233
Query: 1307 RLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDE 1366
LKK D F+W E QKAF++ K L+ PP++ASP +P+ LY+SAT + ++L E E
Sbjct: 1234 -LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQVVSTVLVVERE 1292
Query: 1367 D-----SKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDI 1421
+ +R I+++S VL D++ RY ++KL + + KL +Y + V V + +
Sbjct: 1293 EEGHVQKVQRPIYFVSEVLADSKARYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP- 1351
Query: 1422 IKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQP 1481
+ +L + RI KWAL L +++ P ++K QA+ADF+A+ T QE I+
Sbjct: 1352 LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQEDTPVEKIEH 1411
Query: 1482 WKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGA 1541
W + FDGS +GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG
Sbjct: 1412 WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGI 1471
Query: 1542 KNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANEL 1601
K ++V+GDS+LV+ Q+ KE+ CI +N+ Y + L KFD L HV R +N+ A+ L
Sbjct: 1472 KRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRL 1531
Query: 1602 AQIASGYMVDKCRLKELIE------VKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQNPV 1655
A G + +E V K D D I M DWR+P + +L +
Sbjct: 1532 ANF--GSKRETAPSDVFVEHLYTPTVPHKDTTQDADTRNI-AMVEADWREPFIRFLTSQE 1588
Query: 1656 GTTDR----KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQA 1711
D+ + R+ YV+ +EL+KK+ G L +C+S ++ + +H G+CG H A
Sbjct: 1589 LPQDKDEAERISRRSKLYVLHESELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAA 1648
Query: 1712 GIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWAL 1771
+ +RQG +WPT + D + + C+ CQ A H+PA EL +I WPF W L
Sbjct: 1649 ARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGL 1708
Query: 1772 DLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTT 1831
D++G + ++ VAID F+KW+EA P+ +T D +F +IV+RFG+P + T
Sbjct: 1709 DMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPNRIIT 1767
Query: 1832 DQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK---- 1887
D GT F G F E +GIK+ ++ + +NGQVE AN ++ IK V + K
Sbjct: 1768 DNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAG 1827
Query: 1888 NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWN 1947
W Q L VLW+ R +P +TG +PF L YG EA+LP EV +S R + E E Y
Sbjct: 1828 KWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEE 1884
Query: 1948 MMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRY 2007
+D+L L+E R L R + + +N+ VR+++F++GD V + + + +
Sbjct: 1885 DRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QTTRDR 1940
Query: 2008 RKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
K P WEGPF + +V +Y +K G
Sbjct: 1941 HKLSPLWEGPFIISEVTRPGSYRLKREDG 1969
>ref|XP_475587.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46391119|gb|AAS90646.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1991
Score = 703 bits (1814), Expect = 0.0
Identities = 395/1049 (37%), Positives = 593/1049 (55%), Gaps = 35/1049 (3%)
Query: 1007 MVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1066
++ L+ KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+
Sbjct: 942 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 1001
Query: 1067 RLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1126
+LL FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS
Sbjct: 1002 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1061
Query: 1127 AGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMN 1186
AG E LS LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++
Sbjct: 1062 AGCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQ 1119
Query: 1187 TIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1246
F I ++ Y+DD+VVK+ +DD +L L ++F +R + +K+NP KC FGV +G
Sbjct: 1120 RCFSTQIGRNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFRMKLNPEKCTFGVPSGKL 1179
Query: 1247 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLL 1306
LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L
Sbjct: 1180 LGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL- 1238
Query: 1307 RLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDE 1366
LKK D F+W E QKAF++ K L+ PP++ASP +P+ LY+SAT + ++L E E
Sbjct: 1239 -LKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQVVSTVLVVERE 1297
Query: 1367 D-----SKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDI 1421
+ +R I+++S VL D++TRY ++KL + + KL +Y + V V + +
Sbjct: 1298 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP- 1356
Query: 1422 IKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQP 1481
+ +L + RI KWAL L +++ P ++K QA+ADF+A+ T QE I+
Sbjct: 1357 LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQEDTPVEKIEH 1416
Query: 1482 WKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGA 1541
W + FDGS +GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG
Sbjct: 1417 WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGI 1476
Query: 1542 KNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANEL 1601
K ++V+GDS+LV+ Q+ KE+ CI +N+ Y + L KFD L HV R +N+ A+ L
Sbjct: 1477 KRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRL 1536
Query: 1602 A------QIASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQN-- 1653
A + A + + + K+ +D + M DWR+P++ +L +
Sbjct: 1537 ANFGSKREAAPSDVFVEHLYSPTVPHKDATQVADTHDIA---MVEADWREPLIRFLTSQE 1593
Query: 1654 --PVGTTDRKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQA 1711
+ R+ YV+ EL+KK+ G L +C+S ++ + +H G+CG H A
Sbjct: 1594 LPQYKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAA 1653
Query: 1712 GIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWAL 1771
+ +RQG +WPT + D + + C+ CQ A H+PA EL +I WPF W L
Sbjct: 1654 ARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGL 1713
Query: 1772 DLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTT 1831
D++G + ++ +AID F+KW+EA P+ +T D +F +IV+RFG+P + T
Sbjct: 1714 DMVGPFKKAVG-GYTHLFMAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPNRIIT 1772
Query: 1832 DQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK---- 1887
D GT F G F E +GIK+ ++ + +NGQVE AN ++ IK V + K
Sbjct: 1773 DNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQRIKARVFDRLKPYAG 1832
Query: 1888 NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWN 1947
W L VLW+ R +P +TG +PF L YG EA+LP EV +S R + E E Y
Sbjct: 1833 KWVSQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEE 1889
Query: 1948 MMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRY 2007
+D+L L+E R L R + + +N+ VR+++F++GD V + + + +
Sbjct: 1890 DRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QTTRDR 1945
Query: 2008 RKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
K P WEGPF + +V +Y +K G
Sbjct: 1946 HKLSPLWEGPFIISEVTRPGSYRLKREDG 1974
>ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultivar-group)]
gi|38344177|emb|CAE03508.2| OSJNBa0053K19.16 [Oryza
sativa (japonica cultivar-group)]
Length = 2010
Score = 702 bits (1813), Expect = 0.0
Identities = 407/1116 (36%), Positives = 612/1116 (54%), Gaps = 70/1116 (6%)
Query: 963 EKPVVDFSKKMEAQDPLEEVDLGDGL----EKRPTYISALIDPELKDRMVKL-------L 1011
+K D ++ E E++ L E+ T IS + E K + + L
Sbjct: 906 DKESCDMAQTREMASAREDIRLAAATASEGEEPATKISKSGESEAKTKKIPLDPSDPAKT 965
Query: 1012 KEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRC 1071
KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+ +LL
Sbjct: 966 ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAA 1025
Query: 1072 KFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEY 1131
FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS AG E
Sbjct: 1026 GFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGREL 1085
Query: 1132 LSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHD 1191
LS LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++ F
Sbjct: 1086 LSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQRCFST 1143
Query: 1192 FIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVV 1251
I ++ Y+DD+VVK+ +DD + L ++F +R + +K+NP KC FGV +G +GF+V
Sbjct: 1144 QIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLVGFMV 1203
Query: 1252 HKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKE 1311
+GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L LKK
Sbjct: 1204 SHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LKKT 1261
Query: 1312 DAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDED---- 1367
D F+W E QKAF++ K L+ PP++ASP +P+ LY+SAT + ++L E E+
Sbjct: 1262 DNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHV 1321
Query: 1368 -SKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHML 1426
+R I+++S VL D++TRY ++KL + + KL +Y + V V + + + +L
Sbjct: 1322 QKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP-LGDIL 1380
Query: 1427 SKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFF 1486
+ RI KWAL L +++ P ++K QA+ADF+A+ T QE ++ W + F
Sbjct: 1381 HNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAENMEHWTMHF 1440
Query: 1487 DGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVV 1546
DGS +GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG K ++V
Sbjct: 1441 DGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIV 1500
Query: 1547 KGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIAS 1606
+GDS+LV+ Q+ KE+ C+ +N+ Y + L KFD L HV R +N+ A+ LA S
Sbjct: 1501 RGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGS 1560
Query: 1607 GYMVDKCRLKELIEVKEKLNPSDLDILVIDN------------------MAPNDWRKPIV 1648
K ++ PSD+ + + M DWR+P++
Sbjct: 1561 ---------------KREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVETDWREPLI 1605
Query: 1649 DYLQNPVGTTDR----KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDG 1704
+L + D+ + R+ YV+ EL+KK+ G L +C+S ++ + +H G
Sbjct: 1606 RFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLQDIHSG 1665
Query: 1705 LCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPW 1764
+CG H A + +RQG +WPT + D + + C+ CQ A H+PA EL +I W
Sbjct: 1666 ICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSW 1725
Query: 1765 PFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFG 1824
PF W LD++G + ++ VAID F+KW+EA P+ +T D +F +IV+RFG
Sbjct: 1726 PFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFG 1784
Query: 1825 LPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGR 1884
+P + TD GT F G F E +GIK+ ++ + +NGQVE AN ++ IK V
Sbjct: 1785 VPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFD 1844
Query: 1885 KPK----NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEI 1940
+ K W Q L VLW+ R +P +TG +PF L YG EA+LP EV +S R + E
Sbjct: 1845 RLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE- 1903
Query: 1941 PSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPI 2000
E Y +D+L L+E R L R + + +N+ VR+++F++GD V + +
Sbjct: 1904 --ERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI--- 1958
Query: 2001 NKKDKRYRKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
+ + K P WEGPF + +V +Y +K G
Sbjct: 1959 -QTTRDRHKLSPLWEGPFIISEVTRPGSYRLKREDG 1993
>ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultivar-group)]
gi|38347303|emb|CAE02298.2| OSJNBa0042F21.5 [Oryza sativa
(japonica cultivar-group)]
Length = 1950
Score = 700 bits (1806), Expect = 0.0
Identities = 394/1042 (37%), Positives = 593/1042 (56%), Gaps = 37/1042 (3%)
Query: 1015 KDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFI 1074
KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+ +LL FI
Sbjct: 909 KDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFI 968
Query: 1075 RTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSL 1134
+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS AG E LS
Sbjct: 969 KEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSF 1028
Query: 1135 LDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIE 1194
LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++ F I
Sbjct: 1029 LDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIG 1086
Query: 1195 TFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKK 1254
++ Y+DD+VVK+ +DD + L ++F +R + +K+NP KC FGV +G LGF+V +
Sbjct: 1087 RNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHR 1146
Query: 1255 GIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAF 1314
GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L LKK D F
Sbjct: 1147 GIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LKKTDNF 1204
Query: 1315 RWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDED-----SK 1369
+W E QKAF++ K L+ PPI+ASP +P+ LY+SAT + ++L E E+
Sbjct: 1205 QWGPEAQKAFEDFKKLLTEPPILASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKV 1264
Query: 1370 ERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKP 1429
+R I+++S VL D++TRY ++KL + + KL +Y + V V + + + +L
Sbjct: 1265 QRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP-LGDILHNR 1323
Query: 1430 ILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGS 1489
+ RI KWAL L +++ P ++K QA+ADF+A+ T QE ++ W + FDGS
Sbjct: 1324 EANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAENMEHWTMHFDGS 1383
Query: 1490 SHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGD 1549
+GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG K ++V+GD
Sbjct: 1384 KRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGD 1443
Query: 1550 SELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELA------Q 1603
S+LV+ Q+ KE+ C+ +N+ Y + L KFD L HV R +N+ A+ LA +
Sbjct: 1444 SQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGSKRE 1503
Query: 1604 IASGYMVDKCRLKELIEVKEKLNPSDL-DILVIDNMAPNDWRKPIVDYLQNPVGTTDR-- 1660
+A + + + K+ +D D+ +++ DWR+P + +L + D+
Sbjct: 1504 MAPSDVFVEHLYTPTVPHKDTTQDADTHDVALVE----ADWREPFIRFLTSQELPQDKDE 1559
Query: 1661 --KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWI 1718
+ R+ Y + EL+KK+ G L +C+S ++ + +H G+CG H A +
Sbjct: 1560 AERISRRSKLYAMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGK 1619
Query: 1719 LFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEIN 1778
+RQG +WPT + D + + C+ CQ A H+PA EL +I WPF W LD++G
Sbjct: 1620 AYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK 1679
Query: 1779 PCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTTDQGTVFV 1838
+ ++ VAID F+KW+EA P+ +T D +F +IV+RFG+P + TD GT F
Sbjct: 1680 KAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPNRIITDNGTQFT 1738
Query: 1839 GQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK----NWHQTLG 1894
G F E +GIK+ ++ + +NGQVE AN ++ IK V + K W Q L
Sbjct: 1739 GGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLP 1798
Query: 1895 QVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELV 1954
VLW+ R +P +TG +PF L YG EA+LP EV +S R + E E Y +D+L
Sbjct: 1799 SVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEEDRVDDLH 1855
Query: 1955 NLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNW 2014
L+E R L R + + +N+ VR+++F++GD V + + + + K P W
Sbjct: 1856 RLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QTTRDRHKLSPLW 1911
Query: 2015 EGPFTVEKVLVNNAYSIKELGG 2036
EGPF + +V +Y +K G
Sbjct: 1912 EGPFIISEVTRPGSYRLKREDG 1933
>ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultivar-group)]
gi|21741410|emb|CAD40114.1| OSJNBa0035O13.3 [Oryza sativa
(japonica cultivar-group)]
Length = 2008
Score = 697 bits (1800), Expect = 0.0
Identities = 413/1109 (37%), Positives = 612/1109 (54%), Gaps = 91/1109 (8%)
Query: 963 EKPVVDFSKKMEAQDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDY 1022
E PV SK E++ +++ L D + T SALI L+ KD FAW
Sbjct: 939 EIPVTKTSKSGESEAKTKKIPL-DPSDPTKTAESALIT---------FLQNNKDIFAWKP 988
Query: 1023 DEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDW 1082
+MPG+ REV+E L +KED KP+KQ RRF D IKEE+ +LL FI+ + DW
Sbjct: 989 SDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDW 1048
Query: 1083 LANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYN 1142
LAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS AG E LS LD YSGY+
Sbjct: 1049 LANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYH 1108
Query: 1143 QIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYID 1202
QI + E D KT+F P G Y +V MPFGLKNAGATYQR++ F I ++ Y+D
Sbjct: 1109 QIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVD 1166
Query: 1203 DIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNK 1262
D+VVK+ +DD + L ++F +R + +K+NP KC FGV +G LGF+V +GI+ N K
Sbjct: 1167 DVVVKTKQKDDLISDLEETFVSIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEK 1226
Query: 1263 AKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQK 1322
AIL+ PP+++K +Q L G + L RF++ L E+ F L LKK D F+W E QK
Sbjct: 1227 VTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LKKTDNFQWGPEAQK 1284
Query: 1323 AFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDED-----SKERAIFYLS 1377
AF++ K L+ PP++ASP +P+ LY+SAT + ++LA E E+ +R I+++S
Sbjct: 1285 AFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLAVEREEEGHVQKVQRPIYFVS 1344
Query: 1378 RVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGK 1437
VL D++TRY ++KL LY V + V F DI+ + + + RI K
Sbjct: 1345 EVLADSKTRYPQVQKL---LYGHSVTV--------VTSFPLGDILHNREA----NGRIAK 1389
Query: 1438 WALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGSSHKNGTGI 1497
WAL L +++ P ++K QA+ADF+A+ T QE ++ W + FDGS +GTG
Sbjct: 1390 WALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAENMEHWTMHFDGSKRLSGTGA 1449
Query: 1498 GMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQL 1557
G+ ++SP G + I + S+N AEYEAL GL I I+LG K ++V+GDS+LV+ Q+
Sbjct: 1450 GVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQV 1509
Query: 1558 TKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKE 1617
KE+ C+ +N+ Y + L KFD L HV R +N+ A+ LA S
Sbjct: 1510 MKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGS----------- 1558
Query: 1618 LIEVKEKLNPSDL----------------------DILVIDNMAPNDWRKPIVDYLQNPV 1655
K ++ PSD+ D+ V++ DWR+P++ +L +
Sbjct: 1559 ----KREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAVVE----TDWREPLIRFLTSQE 1610
Query: 1656 GTTDR----KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQA 1711
D+ + R+ YV+ EL+KK+ G L +C+S ++ + +H G+CG H A
Sbjct: 1611 LPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAA 1670
Query: 1712 GIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWAL 1771
+ +RQG +WPT + D + + C+ CQ A H+PA EL +I WPF W L
Sbjct: 1671 ARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGL 1730
Query: 1772 DLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTT 1831
D++G + ++ VAID F+KW+EA P+ +T D +F +IV+RFG+P + T
Sbjct: 1731 DMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPNRIIT 1789
Query: 1832 DQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK---- 1887
D GT F G F E +GIK+ ++ + +NGQVE AN ++ IK V + K
Sbjct: 1790 DNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAG 1849
Query: 1888 NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWN 1947
W Q L VLW+ R +P +TG +PF L YG EA+LP EV +S R + E E Y
Sbjct: 1850 KWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEE 1906
Query: 1948 MMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRY 2007
+D+L L+E R L R + + +N+ VR+++F++GD V + + + +
Sbjct: 1907 DRVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QTTRDR 1962
Query: 2008 RKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
K P WEGPF + +V +Y +K G
Sbjct: 1963 HKLSPLWEGPFIISEVTRPGSYRLKREDG 1991
>dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera]
Length = 1027
Score = 697 bits (1798), Expect = 0.0
Identities = 386/1024 (37%), Positives = 594/1024 (57%), Gaps = 26/1024 (2%)
Query: 1025 MPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLA 1084
M G+ + +L + +PV+Q RRFHPD I+ EI++LL FIR Y DWLA
Sbjct: 1 MKGIHPSITSHRLNVVSTARPVRQRIRRFHPDRQRVIRNEIDKLLEAGFIREVSYPDWLA 60
Query: 1085 NVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQI 1144
NVV V KK GK RVC+D+ +LN A PKD + +P + +VDS +G LS LD +SGY+QI
Sbjct: 61 NVVVVPKKEGKWRVCVDYTNLNNACPKDSFPLPRIDQIVDSTSGQGMLSFLDAFSGYHQI 120
Query: 1145 FIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDI 1204
++ +D K AF P L Y+ VMPFGLKNAGATYQR+M IF I ++VYIDDI
Sbjct: 121 PMSPDDEEKIAFITPHDLYCYK--VMPFGLKNAGATYQRLMTKIFKPLIGHSVEVYIDDI 178
Query: 1205 VVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAK 1264
VVKS +R+ H+LHL++ F +R+YG+K+NP KCAFGV A FLGF+V ++GIE++ ++ K
Sbjct: 179 VVKSKTREQHILHLQEVFYLLRRYGMKLNPSKCAFGVSARKFLGFMVSQRGIEVSPDQVK 238
Query: 1265 AILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAF 1324
A+++T PP +KK+LQ L GK+ L RFIA ++ + F L ++K W Q A
Sbjct: 239 AVMETPPPRNKKELQRLTGKLVALGRFIARFIDELRPF--FLAIRKAGTHGWTDNCQNAL 296
Query: 1325 DELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAE 1384
+ +K YL PPI++SPI + + +Y++ ++ I ++L + +++ I+Y+SR L D E
Sbjct: 297 ERIKHYLMQPPILSSPIPKEKLYMYLAVSEWAISAVLFRCPSPKEQKPIYYVSRALADVE 356
Query: 1385 TRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTE 1444
TRY+ +E + L L + KL+ Y + V+V + ++++L KP L R+ +WA+ L+E
Sbjct: 357 TRYSKMELISLALRSAAQKLRPYFQAHPVIVLTDQP-LRNILHKPDLTGRMLQWAIELSE 415
Query: 1445 YSLTYAPLKAVKGQAIADFLADQT-LPQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVS 1503
+ + + P ++KGQ +ADF+ + + P + + W L DG+S +G+G+G+ + S
Sbjct: 416 FGIEFQPRLSMKGQVMADFVLEYSRKPGQHEGSRKKEWWTLRVDGASRSSGSGVGLLLQS 475
Query: 1504 PRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKC 1563
P G + R+ + SNNEAEYEA+ SGL++ +AL + + DS+LV+K + +EY+
Sbjct: 476 PTGEHLEQAIRLGFSASNNEAEYEAILSGLDLALALSVSKLRIFSDSQLVVKHVQEEYEA 535
Query: 1564 ISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKE 1623
+ARY K N L +F E + + R DN+ A+ LA IA+ + + L+ +
Sbjct: 536 KDARMARYLAKVRNTLQQFTEWTIEKIKRADNRRADALAGIAASLSIKEA---ILLPIHV 592
Query: 1624 KLNPSDLDILVIDNM-AP----NDWRKPIVDYLQNPVGTTD----RKTKYRAMSYVIMGN 1674
+ NPS +I + AP +W I +Y++ D K + +A + ++G
Sbjct: 593 QTNPSVSEISICSTTEAPQADDQEWMNDITEYIRTGTLPGDPKQAHKVRVQAARFTLIGG 652
Query: 1675 ELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCM 1734
L+K++ G L+CL +A ++ +H+G+ G H G + QG YWPT+ K+
Sbjct: 653 HLYKRSFTGPYLRCLGHSEAQYVLAELHEGIYGNHSGGRSLAHRAHSQGYYWPTMKKEAA 712
Query: 1735 EYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDY 1794
Y KRC CQR+A I H+P++ L SI PWPF W +D++ + P + Q K+++VA DY
Sbjct: 713 AYVKRCDKCQRYAPIPHMPSTTLKSISGPWPFAQWGMDIVRPL-PTAPAQKKFLLVATDY 771
Query: 1795 FTKWVEAIPLQNVTQDTVIEFI-SSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKL 1853
F+KWVEA + V +F+ +I+ RFG+P+++ D G F +F I+
Sbjct: 772 FSKWVEAEAYASTKDKDVTKFVWKNIICRFGIPQTIIADNGPQFDSIAFRNFCSELNIRN 831
Query: 1854 LTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPF 1913
STP Y Q+NGQ EA NKTLI+ +KK + + W + L VLWAYR +P TG TPF
Sbjct: 832 SYSTPRYPQSNGQAEATNKTLITALKKRLEQAKGKWVEELPGVLWAYRTTPGRPTGNTPF 891
Query: 1914 RLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVN-LDEERLLVLDTLTRQKD 1972
LAYG +AV+P+E+ L + ++ + NM L ++ DE R + +
Sbjct: 892 ALAYGMDAVIPIEIGLPTIWTNAAKQSDA----NMQLGRNLDWTDEVRESASIRMADYQQ 947
Query: 1973 RIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVEKVLVNNAYSIK 2032
R + YN+KVR +S G V + N + K+ NWEGP+ V K N AY ++
Sbjct: 948 RASAHYNRKVRPRSLKNGTLVLRKFFE-NTTEVGAGKFQANWEGPYIVSKASDNGAYHLQ 1006
Query: 2033 ELGG 2036
+L G
Sbjct: 1007 KLDG 1010
>ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46063437|gb|AAS79740.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1756
Score = 696 bits (1795), Expect = 0.0
Identities = 393/1057 (37%), Positives = 592/1057 (55%), Gaps = 67/1057 (6%)
Query: 1015 KDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFI 1074
KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+ +LL FI
Sbjct: 715 KDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFVQDRKDAIKEELTKLLAAGFI 774
Query: 1075 RTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSL 1134
+ + DWLAN V V KK G+ R+C+D+ DLN PKD + +P + +VDS AG E LS
Sbjct: 775 KEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCELLSF 834
Query: 1135 LDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIE 1194
LD YSGY+QI + E D KT+F P +G Y +V MPFGLKNAGATYQR++ F I
Sbjct: 835 LDCYSGYHQIRLKESDCLKTSFITP--IGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIG 892
Query: 1195 TFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKK 1254
++ Y+DD+V+K+ +DD + L ++F +R + +K+NP KC FGV +G LGF+V +
Sbjct: 893 RNVEAYVDDVVIKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHR 952
Query: 1255 GIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAF 1314
GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L LKK D F
Sbjct: 953 GIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LKKTDDF 1010
Query: 1315 RWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDED-----SK 1369
+W E QKAF++ K +L+ PP++ASP +P+ LY+SAT + ++L E E+
Sbjct: 1011 QWGPEAQKAFEDFKKFLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKV 1070
Query: 1370 ERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKP 1429
+R I+++S VL D++TRY ++KL + + KL +Y + V V + + + +L
Sbjct: 1071 QRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP-LGDILHNR 1129
Query: 1430 ILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGS 1489
+ RI KWAL L +++ P ++K QA+ADF+A+ T QE ++ W + FDGS
Sbjct: 1130 EANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAEKMEHWTMHFDGS 1189
Query: 1490 SHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGD 1549
+GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG K ++V+GD
Sbjct: 1190 KRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGD 1249
Query: 1550 SELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYM 1609
S+LV+ Q+ KE+ C+ +N+ Y + L KFD L HV R +N+ A+ LA S
Sbjct: 1250 SQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGS--- 1306
Query: 1610 VDKCRLKELIEVKEKLNPSDL----------------------DILVIDNMAPNDWRKPI 1647
K ++ PSD+ D+ +++ DWR+P
Sbjct: 1307 ------------KREVAPSDVFVEHLYTPTVPHKDTTQIAGTHDVALVE----ADWREPF 1350
Query: 1648 VDYLQNPVGTTDR----KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHD 1703
+ +L + D+ + R+ YV+ EL+KK+ G L +C+S ++ + +H
Sbjct: 1351 IRFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHS 1410
Query: 1704 GLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKP 1763
G+CG H A + +RQG +WPT + D + + C+ CQ A H+PA EL +I
Sbjct: 1411 GICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLS 1470
Query: 1764 WPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRF 1823
WPF W LD++G + ++ VAID F+KW+EA P+ +T D +F +IV+RF
Sbjct: 1471 WPFAVWGLDMVGPFKKAVG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRF 1529
Query: 1824 GLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVG 1883
G+P + TD G F G F E +GIK+ ++ + +NGQVE AN ++ IK V
Sbjct: 1530 GVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVF 1589
Query: 1884 RKPK----NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEE 1939
+ K W + L VLW+ R +P +TG +PF L YG EA+LP EV +S R + E
Sbjct: 1590 DRLKPYAGKWVKQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE 1649
Query: 1940 IPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLP 1999
E Y +D+L L+E R L R + + +N+ VR+++F++GD V + +
Sbjct: 1650 ---ERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI-- 1704
Query: 2000 INKKDKRYRKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
+ + K P WEGPF + +V +Y +K G
Sbjct: 1705 --QTTRDRHKLSPLWEGPFIISEVTRPGSYRLKREDG 1739
>gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37533012|ref|NP_920808.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|19881590|gb|AAM00991.1| Putative retroelement [Oryza
sativa]
Length = 2017
Score = 694 bits (1791), Expect = 0.0
Identities = 395/1053 (37%), Positives = 588/1053 (55%), Gaps = 59/1053 (5%)
Query: 1015 KDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFI 1074
KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+ +LL FI
Sbjct: 976 KDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFI 1035
Query: 1075 RTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSL 1134
+ + DWLAN V V KK G+ R+C+D+ DLN PKD + +P + +VDS AG E LS
Sbjct: 1036 KEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCELLSF 1095
Query: 1135 LDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIE 1194
LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++ F I
Sbjct: 1096 LDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIG 1153
Query: 1195 TFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKK 1254
++ Y+DD+VVK+ +DD + L ++F +R + +K+NP KC FGV +G LGF+V +
Sbjct: 1154 RNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHR 1213
Query: 1255 GIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAF 1314
GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L LKK D F
Sbjct: 1214 GIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL--LKKTDDF 1271
Query: 1315 RWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDED-----SK 1369
W E QKAF++ K L+ PP++ASP +P+ LY+SAT + ++L E E+
Sbjct: 1272 HWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEDGHVQKV 1331
Query: 1370 ERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKP 1429
+R I+++S VL D++TRY ++KL + + KL +Y + V V + + + +L
Sbjct: 1332 QRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHLVTVVTSFP-LGDILHNR 1390
Query: 1430 ILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGS 1489
+ RI KWAL L +++ P ++K QA+ADF+A+ T QE ++ W + FDGS
Sbjct: 1391 EANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAEKVEHWTMHFDGS 1450
Query: 1490 SHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGD 1549
+GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG K ++V+GD
Sbjct: 1451 KRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGD 1510
Query: 1550 SELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYM 1609
S+LV+ Q+ KE+ C+ +N+ Y + L KFD L HV R +N+ A+ LA S
Sbjct: 1511 SQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNKAADRLANFGS--- 1567
Query: 1610 VDKCRLKELIEVKEKLNPSDL-------------DILVIDN-----MAPNDWRKPIVDYL 1651
K ++ PSD+ D + M DWR+P++ +L
Sbjct: 1568 ------------KREVAPSDVFVEHLYAPTVPHKDTTQVAGTHDVAMVEADWREPLIRFL 1615
Query: 1652 QNPVGTTDR----KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCG 1707
+ D+ + R+ YV+ EL+KK+ G L +C+S ++ + +H G+CG
Sbjct: 1616 TSQELPQDKDEAERISRRSRLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICG 1675
Query: 1708 AHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFR 1767
H A + +RQG +WPT + D + + C+ CQ A H+PA EL +I WPF
Sbjct: 1676 NHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFA 1735
Query: 1768 GWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPE 1827
W LD++G + ++ VAID F+KW+EA P+ +T D +F +IV+RFG+P
Sbjct: 1736 VWGLDMVGPFKKAFG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPN 1794
Query: 1828 SLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK 1887
+ TD G F G F E +GIK+ ++ + +NGQVE AN ++ IK V + K
Sbjct: 1795 RIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLK 1854
Query: 1888 ----NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSE 1943
W + L VLW+ R +P +TG +PF L YG EA+LP EV +S R + E E
Sbjct: 1855 PYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---E 1911
Query: 1944 DYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKK 2003
Y +D+L L+E R L R + + +N+ VR+++F++GD V + + +
Sbjct: 1912 RYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QT 1967
Query: 2004 DKRYRKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
+ K P WEGPF + +V +Y +K G
Sbjct: 1968 TRDRHKLSPLWEGPFIISEVTRPGSYRLKREDG 2000
>gb|AAV25234.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1953
Score = 693 bits (1789), Expect = 0.0
Identities = 394/1040 (37%), Positives = 587/1040 (55%), Gaps = 50/1040 (4%)
Query: 1007 MVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1066
++ L+ KD FAW +MPG+ REV+E L +KED KP+KQ RRF D IKEE+
Sbjct: 937 LITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELT 996
Query: 1067 RLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1126
+LL FI+ + DWLAN V V KK G+ R+C+D+ DLN + PKD + +P + +VDS
Sbjct: 997 KLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDST 1056
Query: 1127 AGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMN 1186
A E LS LD YSGY+QI + E D KT+F P G Y +V MPFGLKNAGATYQR++
Sbjct: 1057 ASCELLSFLDCYSGYHQIRLKESDCLKTSFITP--FGAYCYVTMPFGLKNAGATYQRMIQ 1114
Query: 1187 TIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1246
F I ++ Y+DD+VVK+ +DD + L ++F +R + +K+NP KC FGV +G
Sbjct: 1115 RCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKL 1174
Query: 1247 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLL 1306
LGF+V +GI+ N K AIL+ PP+++K +Q L G + L RF++ L E+ F L
Sbjct: 1175 LGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKL- 1233
Query: 1307 RLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDE 1366
LKK D F+W E Q+AF++ K L+ PP++ASP +P+ LY+SAT + ++L E E
Sbjct: 1234 -LKKTDNFQWGPEAQRAFEDFKQLLTKPPVLASPHPQEPLLLYVSATSQVVSTVLVVERE 1292
Query: 1367 D-----SKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDI 1421
+ +R I+++S VL D++TRY ++KL + + KL +Y + V V + +
Sbjct: 1293 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFP- 1351
Query: 1422 IKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQP 1481
+ +L + RI KWAL L +++ P ++K QA+ADF+A+ T QE I+
Sbjct: 1352 LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVAEWTECQEDTPVEKIEH 1411
Query: 1482 WKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGA 1541
W + FDGS +GTG G+ ++SP G + I + S+N AEYEAL GL I I+LG
Sbjct: 1412 WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGI 1471
Query: 1542 KNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANEL 1601
K ++V+GDS+LV+ Q+ KE+ CI +N+ Y + L KFD L HV R +N+ A+ L
Sbjct: 1472 KRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRL 1531
Query: 1602 AQIASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIV-DYLQNPVGTTDR 1660
A S K + PSD+ R P+ D L +R
Sbjct: 1532 ANFGS---------------KREAAPSDV-----------FRRTPLFPDKLPQDKDEAER 1565
Query: 1661 KTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILF 1720
++ R+ YV+ +EL+K++ G L +C+S ++ + +H G+CG H A + +
Sbjct: 1566 ISR-RSKLYVMHESELYKRSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAY 1624
Query: 1721 RQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPC 1780
RQG +WPT + D + + C+ CQ A H+PA EL +I WPF W LD++G
Sbjct: 1625 RQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKA 1684
Query: 1781 SSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTTDQGTVFVGQ 1840
+ ++ VAID F+KW+EA P+ +T D +F +IV+RFG+P + TD GT F G
Sbjct: 1685 VG-GYTHLFVAIDKFSKWIEAKPVVTITADNARDFFINIVHRFGVPNRIITDNGTQFTGG 1743
Query: 1841 KVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK----NWHQTLGQV 1896
F E +GIK+ ++ + +NGQVE AN ++ IK V + K W Q L V
Sbjct: 1744 VFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSV 1803
Query: 1897 LWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNL 1956
LW+ R +P +TG +PF L YG EA+LP EV +S R + E E Y +D+L L
Sbjct: 1804 LWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFRE---ERYEEDRVDDLHRL 1860
Query: 1957 DEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEG 2016
+E R L R + + +N+ VR+++F++GD V + + + + K P WEG
Sbjct: 1861 EEVREAALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKI----QTTRDRHKLSPLWEG 1916
Query: 2017 PFTVEKVLVNNAYSIKELGG 2036
PF + +V +Y +K G
Sbjct: 1917 PFIISEVTRPGSYRLKREDG 1936
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.334 0.144 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,189,631,835
Number of Sequences: 2540612
Number of extensions: 129921435
Number of successful extensions: 554950
Number of sequences better than 10.0: 34747
Number of HSP's better than 10.0 without gapping: 2915
Number of HSP's successfully gapped in prelim test: 31832
Number of HSP's that attempted gapping in prelim test: 482198
Number of HSP's gapped (non-prelim): 63718
length of query: 2064
length of database: 863,360,394
effective HSP length: 144
effective length of query: 1920
effective length of database: 497,512,266
effective search space: 955223550720
effective search space used: 955223550720
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 83 (36.6 bits)
Lotus: description of TM0269b.2


