

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.8
(765 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
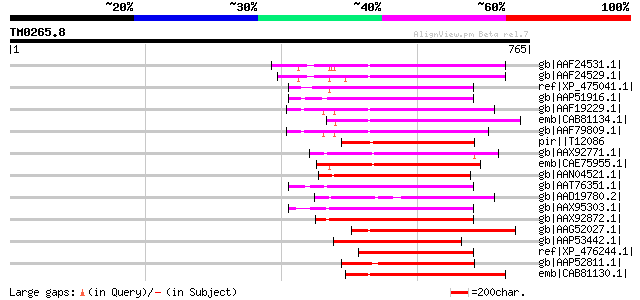
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF24531.1| F7F22.17 [Arabidopsis thaliana] 194 1e-47
gb|AAF24529.1| F7F22.15 [Arabidopsis thaliana] 187 9e-46
ref|XP_475041.1| OSJNBa0042F21.11 [Oryza sativa (japonica cultiv... 187 1e-45
gb|AAP51916.1| putative polyprotein [Oryza sativa (japonica cult... 182 5e-44
gb|AAF19229.1| Similar to Athila ORF 1 [Arabidopsis thaliana] gi... 181 1e-43
emb|CAB81134.1| putative athila transposon protein [Arabidopsis ... 178 7e-43
gb|AAF79809.1| T32E20.9 [Arabidopsis thaliana] 177 2e-42
pir||T12086 hypothetical protein - fava bean (fragment) gi|25222... 176 3e-42
gb|AAX92771.1| Transposable element protein, putative [Oryza sat... 175 4e-42
emb|CAE75955.1| B1159F04.18 [Oryza sativa (japonica cultivar-gro... 174 1e-41
gb|AAN04521.1| Hypothetical protein with similarity to putative ... 171 6e-41
gb|AAT76351.1| hypothetical protein [Oryza sativa (japonica cult... 166 3e-39
gb|AAD19780.2| hypothetical protein [Arabidopsis thaliana] 165 5e-39
gb|AAX95303.1| hypothetical protein [Oryza sativa (japonica cult... 164 8e-39
gb|AAX92872.1| Transposable element protein, putative [Oryza sat... 164 1e-38
gb|AAG52027.1| Athila ORF 1, putative; 43045-40843 [Arabidopsis ... 163 2e-38
gb|AAP53442.1| hypothetical protein similar to putative retroele... 163 2e-38
ref|XP_476244.1| hypothetical protein [Oryza sativa (japonica cu... 162 5e-38
gb|AAP52811.1| hypothetical protein similar to putative retroele... 157 1e-36
emb|CAB81130.1| AT4g07600 [Arabidopsis thaliana] gi|5724765|gb|A... 155 6e-36
>gb|AAF24531.1| F7F22.17 [Arabidopsis thaliana]
Length = 1799
Score = 194 bits (492), Expect = 1e-47
Identities = 132/397 (33%), Positives = 204/397 (51%), Gaps = 64/397 (16%)
Query: 387 GTDGQANVRKTPGMFPSDTVINPKENCSAITLRSGATL---SEPKQKIVENNNVNDEFVG 443
G ++ K P V NPKE AITLRSG L EPK + E++ D
Sbjct: 405 GHSTSSSATKQTSQLPGKAVQNPKEYAHAITLRSGKALPTREEPKT-VTEDSEDQD---- 459
Query: 444 DIEPLGENKKKQKEKEEEKRKVELEK---------------KFTK--------------- 473
GE+ +K++ ++ + LE+ FT+
Sbjct: 460 -----GEDLSLEKDQADKPLDLSLEQPLDLSLQQSLDPPLDSFTRPTTRPVIPAASPTAP 514
Query: 474 ------------VPFP-----PFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMP 516
VP P PFP K +K + F K++ + +P + L +P
Sbjct: 515 KPVAVKNKEKVFVPPPYKSQLPFPGRHKKALADKYRAMFAKNIKEVELRIPLVDALALIP 574
Query: 517 QYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKR-KDPGSFTLPVNFGASKQV 575
KF+K+++ + R+ E +V L+ ECS I+Q+K+ PK+ DPGSFTLP + G
Sbjct: 575 DSHKFLKDLIVE--RIQEVQGMVVLSHECSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFN 632
Query: 576 RALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFP 635
R LCDLG+SV+LMPLS+ +RL + K + L LADRS+ P G++E++ +R+G E P
Sbjct: 633 RCLCDLGASVSLMPLSVAKRLGFTQYKSCNISLILADRSVRIPHGLLENLPIRIGAVEIP 692
Query: 636 VDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKIGFNIFDLKPKPVE 694
DFV+++MDE+ K PLILGRPFLAT+ A I+V KG I L + D ++ F++ D KP
Sbjct: 693 TDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLNLGKDFRMTFDVKDAMKKPTI 752
Query: 695 KNDVFLVKMMEEWSDEKLKQFFLKEKAGASNKKKKED 731
+ +F ++ M++ +DE L++ ++ ++ K ED
Sbjct: 753 EGQLFWIEEMDQLADELLEELAEEDHLNSALTKSGED 789
>gb|AAF24529.1| F7F22.15 [Arabidopsis thaliana]
Length = 1862
Score = 187 bits (476), Expect = 9e-46
Identities = 128/353 (36%), Positives = 197/353 (55%), Gaps = 29/353 (8%)
Query: 396 KTPGMFPSDTVINPKENCSAITLRSGATL---SEPKQKIVENNNVNDEFVGDIEPLGENK 452
K P V NPKE AITL SG L EPK + E++ D GE+
Sbjct: 457 KQTSQLPGKAVQNPKEYAHAITLHSGKALPTREEPKT-VTEDSEDQD---------GEDL 506
Query: 453 KKQKEKEEEKRKVELEK----KFTKVPFPPFP--TNIAKRRLEKQFS----KFISMFKKL 502
+K++ ++ + LE+ + PP T R + S K +++ K
Sbjct: 507 SLKKDQADKPLDLSLEQPLDLSLQQSLDPPLDSFTRPTTRPVIPAASPTAPKPVAVKNKE 566
Query: 503 RVEL--PFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKR-KD 559
+VEL P + L +P KF+K+++ + R+ E +V L+ ECS I+Q+K+ PK+ D
Sbjct: 567 KVELRIPLVDALALIPDSHKFLKDLIVE--RIQEVQGMVVLSHECSAIIQKKIIPKKLSD 624
Query: 560 PGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPW 619
PGSFTLP + G R LCDLG+SV+LMPLS+ +RL + K + L LADRS+ P
Sbjct: 625 PGSFTLPCSLGPLAFNRCLCDLGASVSLMPLSVAKRLGFTQYKSCNISLILADRSVRIPH 684
Query: 620 GVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-D 678
G++E++ +R+G E P DFV+++MDE+ K PLILGRPFLAT+ A I+V KG I L + D
Sbjct: 685 GLLENLPIRIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLNLGKD 744
Query: 679 EKIGFNIFDLKPKPVEKNDVFLVKMMEEWSDEKLKQFFLKEKAGASNKKKKED 731
++ F++ D KP + +F ++ M++ +DE L++ ++ ++ K ED
Sbjct: 745 FRMTFDVKDAMKKPTIEGQLFWIEEMDQLADELLEELAEEDHLNSALTKSGED 797
>ref|XP_475041.1| OSJNBa0042F21.11 [Oryza sativa (japonica cultivar-group)]
gi|38347309|emb|CAE02304.2| OSJNBa0042F21.11 [Oryza
sativa (japonica cultivar-group)]
Length = 920
Score = 187 bits (475), Expect = 1e-45
Identities = 106/281 (37%), Positives = 164/281 (57%), Gaps = 25/281 (8%)
Query: 411 ENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGEN--KKKQKEKEEEKRKVELE 468
EN AIT R G + +P P G N K+ + + ++++ +
Sbjct: 619 ENVKAITTRGGKSTRDPPYP---------------NPAGTNGMSKETPSTDSDDKEIQPD 663
Query: 469 K----KFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKE 524
K ++ PFP K ++KQF+ F+ + +K+ + +P + ++ +P YA+++K+
Sbjct: 664 KTVPQEYCDTRLLPFPQQSRKPSVDKQFACFVEVIQKIHINVPLLDAMQ-VPTYARYLKD 722
Query: 525 ILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSS 584
IL+ KR L E+V+LTE CS ++ KLP K+KDPG T+ + GA + +ALCDLG+S
Sbjct: 723 ILNNKRPLPT-TEVVKLTEHCSNLILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGAS 781
Query: 585 VNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMD 644
V++MP +F++LN L PT M LQLAD S+ P G+ EDV ++ +F PVDFV++DMD
Sbjct: 782 VSVMPKDVFDKLNFTVLAPTPMRLQLADSSVHYPVGIAEDVPAKIRDFFIPVDFVVLDMD 841
Query: 645 EDSKIPLILGRPFLATSQAKINVGKGTISLRV--ADEKIGF 683
+ PLILGRPFL+T+ A I+VG G+I +EK F
Sbjct: 842 TGKETPLILGRPFLSTAGANIDVGTGSIRFHTNGKEEKFEF 882
>gb|AAP51916.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37530654|ref|NP_919629.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|20042899|gb|AAM08727.1| Putative polyprotein [Oryza
sativa] gi|18997221|gb|AAL83338.1| Putative polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 689
Score = 182 bits (461), Expect = 5e-44
Identities = 99/273 (36%), Positives = 164/273 (59%), Gaps = 11/273 (4%)
Query: 411 ENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKK 470
EN AIT+R G + +P N V + P ++ K+ + E + ++
Sbjct: 342 ENVKAITMRRGKSTRDPPYP----NPVGTNEISKGAPSNDSADKEVQPENT-----VPQE 392
Query: 471 FTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKR 530
+ FP + K +++QF++F+ + +K+ + +P + ++ +P YA+++K+IL+ KR
Sbjct: 393 YCDTRLLSFPQRMRKPSVDEQFARFVEVIQKIHINVPLLDAMQ-VPTYARYLKDILNNKR 451
Query: 531 RLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPL 590
L E+V+LTE+CS ++ KLP K+KDPG T+ + GA + + C+LG+SV+++
Sbjct: 452 PLPT-TEVVKLTEQCSNLILHKLPEKKKDPGCPTITCSIGAQQFDQVSCNLGASVSVVLK 510
Query: 591 SMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIP 650
+F++LN L PT M LQLAD S+ P G+ EDV V++ +F PVDFV++DMD + P
Sbjct: 511 DVFDKLNFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFIPVDFVVLDMDTGKETP 570
Query: 651 LILGRPFLATSQAKINVGKGTISLRVADEKIGF 683
LILGRPFL+T+ A I+VG G+I + +K F
Sbjct: 571 LILGRPFLSTAGANIDVGTGSIRFHINGKKEKF 603
>gb|AAF19229.1| Similar to Athila ORF 1 [Arabidopsis thaliana]
gi|25518186|pir||B86490 F28L22.6 protein - Arabidopsis
thaliana
Length = 719
Score = 181 bits (458), Expect = 1e-43
Identities = 120/333 (36%), Positives = 179/333 (53%), Gaps = 31/333 (9%)
Query: 408 NPKENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGEN-KKKQKEKEE------ 460
N KE AITLRSG L + N N D D E +N +K EE
Sbjct: 356 NSKEYAHAITLRSGKELPTKESP---NQNTEDSLDQDGEDFCQNGNSAEKAIEEPILHQP 412
Query: 461 ------------EKRKVELEKKFTKVPFP-----PFPTNIAKRRLEKQFSKFISMFKKLR 503
EK K+ +P P PFP K ++K + K L
Sbjct: 413 TRPLAPAASPLVEKPAAAKTKENVFIPPPYKPPLPFPGRFKKVMIQKYKALLEKQLKNLE 472
Query: 504 VELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRK-DPGS 562
V +P + L +P K++K+++++ R+ E +V L+ ECS I+Q+K+ PK+ DPGS
Sbjct: 473 VTMPLVDCLALIPDSNKYVKDMITE--RIKEVQGMVVLSHECSAIIQQKIIPKKLGDPGS 530
Query: 563 FTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVV 622
FTLP G + LCDLG+SV+LMPL + ++L + KP + L LADRS+ G++
Sbjct: 531 FTLPCALGPLAFSKCLCDLGASVSLMPLPVAKKLGFNKYKPCNISLILADRSVRISHGLL 590
Query: 623 EDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKI 681
ED+ V +G E P DFV+++MDE+ K PLILGRPFLA ++A I+V KG I L + D K+
Sbjct: 591 EDLPVMIGVVEVPTDFVVLEMDEEPKDPLILGRPFLARARAIIDVKKGKIDLNLGRDLKM 650
Query: 682 GFNIFDLKPKPVEKNDVFLVKMMEEWSDEKLKQ 714
F+I + KP + ++F ++ M+ +D+ L++
Sbjct: 651 TFDITNTMKKPTIEGNIFWIEEMDMLADKMLEE 683
>emb|CAB81134.1| putative athila transposon protein [Arabidopsis thaliana]
gi|25354974|pir||B85075 probable athila transposon
protein [imported] - Arabidopsis thaliana
Length = 866
Score = 178 bits (451), Expect = 7e-43
Identities = 105/291 (36%), Positives = 169/291 (57%), Gaps = 7/291 (2%)
Query: 468 EKKFTKVPFPP---FPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKE 524
EK F P+ P FP K +K + F K++ + +P + L +P KF+K+
Sbjct: 444 EKVFVPPPYKPELPFPGRHKKALADKYRAMFAKNIKEVELRIPLVDALALIPDSHKFLKD 503
Query: 525 ILSKKRRLSEENEIVELTEECSVILQRKLPPKR-KDPGSFTLPVNFGASKQVRALCDLGS 583
++ + R+ E +V L+ CS I+Q+K+ PK+ DPGSFTLP + G R LCDLG+
Sbjct: 504 LIVE--RIQEVQGMVVLSHGCSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGA 561
Query: 584 SVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDM 643
SV+LM LS+ +RL + K + + L LADRS+ P G++++ + +G E P DFV+++M
Sbjct: 562 SVSLMALSVAKRLGFTQYKSSNISLILADRSVRIPHGLLKNFGITIGAVEIPTDFVVLEM 621
Query: 644 DEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKIGFNIFDLKPKPVEKNDVFLVK 702
DE+ K PLILGRPFLAT+ A I+V KG I L + D ++ F++ D KP + +F +K
Sbjct: 622 DEEPKDPLILGRPFLATAGAMIDVKKGKIDLSLGKDFRMTFDVKDAMKKPTIEGQLFWIK 681
Query: 703 MMEEWSDEKLKQFFLKEKAGASNKKKKEDT*INQTEQVYSMSMVVNSELKE 753
M++ +DE L++ ++ ++ K ED ++ Y + N ++E
Sbjct: 682 EMDQLADELLEELAEEDHLNSALTKSGEDGFLHLETFGYQKLLDSNKAMEE 732
>gb|AAF79809.1| T32E20.9 [Arabidopsis thaliana]
Length = 1586
Score = 177 bits (448), Expect = 2e-42
Identities = 118/324 (36%), Positives = 173/324 (52%), Gaps = 31/324 (9%)
Query: 408 NPKENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGEN-KKKQKEKEE------ 460
N KE AITLRSG L + N N D D E +N +K EE
Sbjct: 459 NSKEYAHAITLRSGKELPTKESP---NQNTEDSLDQDGEDFCQNGNSAEKAIEEPILHQP 515
Query: 461 ------------EKRKVELEKKFTKVPFP-----PFPTNIAKRRLEKQFSKFISMFKKLR 503
EK K+ +P P PFP K ++K + K L
Sbjct: 516 TRPLAPAASPLVEKPAAAKTKENVFIPPPYKPPLPFPGRFKKVMIQKYKALLEKQLKNLE 575
Query: 504 VELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRK-DPGS 562
V +P + L +P K++K+++++ R+ E +V L+ ECS I+Q+K+ PK+ DPGS
Sbjct: 576 VTMPLVDCLALIPDSNKYVKDMITE--RIKEVQGMVVLSHECSAIIQQKIIPKKLGDPGS 633
Query: 563 FTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVV 622
FTLP G + LCDLG+SV+LMPL + ++L + KP + L LADRS+ G++
Sbjct: 634 FTLPCALGPLAFSKCLCDLGASVSLMPLPVAKKLGFNKYKPCNISLILADRSVRISHGLL 693
Query: 623 EDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKI 681
ED+ V +G E P DFV+++MDE+ K PLILGRPFLA ++A I+V KG I L + D K+
Sbjct: 694 EDLPVMIGVVEVPTDFVVLEMDEEPKDPLILGRPFLARARAIIDVKKGKIDLNLGRDLKM 753
Query: 682 GFNIFDLKPKPVEKNDVFLVKMME 705
F+I + KP + ++F ++ M+
Sbjct: 754 TFDITNTMKKPTIEGNIFWIEEMD 777
>pir||T12086 hypothetical protein - fava bean (fragment)
gi|2522230|dbj|BAA22788.1| retrotransposon-like gene~the
first amino acid was determined to be leucine [Vicia
faba]
Length = 238
Score = 176 bits (446), Expect = 3e-42
Identities = 89/198 (44%), Positives = 135/198 (67%), Gaps = 3/198 (1%)
Query: 489 EKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVI 548
EK F F+ MFKKL + + E LEKMP YAKF+K+I+SKKR + + + + TE CS I
Sbjct: 11 EKNFEIFLEMFKKLELNILVLEALEKMPTYAKFMKDIISKKRTI--DTDPIIHTETCSSI 68
Query: 549 LQ-RKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMM 607
LQ K+P K+KD G T P G + +LG+SV+L+P ++++L + ++ T M
Sbjct: 69 LQGMKIPVKKKDRGFVTTPCTIGDRSFKKYFINLGASVSLLPFYIYKKLGISNVQNTRMT 128
Query: 608 LQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINV 667
L+ AD S+ P+G+VEDVLV++ +F FPVDFV+++M ED +I LILGRPFL T + I++
Sbjct: 129 LRFADHSVKIPYGIVEDVLVKIDKFVFPVDFVVLEMPEDEEITLILGRPFLETRRCLIDI 188
Query: 668 GKGTISLRVADEKIGFNI 685
+GT++L+V DE++ ++
Sbjct: 189 EEGTVTLKVYDEELKIDV 206
>gb|AAX92771.1| Transposable element protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 689
Score = 175 bits (444), Expect = 4e-42
Identities = 104/289 (35%), Positives = 173/289 (58%), Gaps = 15/289 (5%)
Query: 442 VGDIEPLGENKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFISMFKK 501
+G + N KE + EK + +++ PFP + K +++QF++F+ + +K
Sbjct: 346 IGMSKEAPSNDSAGKEIQPEKT---VPQEYCNTRLLPFPQRMRKPSVDEQFARFVEVIQK 402
Query: 502 LRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPG 561
+ + +P + ++ +P YA ++K+IL+ KR L E+V+LTE+CS ++ KLP K+K PG
Sbjct: 403 IHINVPLLDAMQ-VPTYAHYLKDILNNKRPLPTM-EVVKLTEQCSNVILHKLPEKKKYPG 460
Query: 562 SFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGV 621
T+ + A + +ALCDLG+SV++MP +F++L L PT M LQLAD S+ P G+
Sbjct: 461 CPTITCSIRAQQFDQALCDLGASVSVMPKDVFDKLKFTVLAPTPMRLQLADSSVRYPAGI 520
Query: 622 VEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRV--ADE 679
EDV V++ +F PVDFV++DMD + PLILGRPFL+T A I++G G+I + +E
Sbjct: 521 AEDVPVKIRDFFTPVDFVVLDMDTGKETPLILGRPFLSTGGANIDMGTGSIHFHINRKEE 580
Query: 680 KIGF-------NIFDLKPKPVEKN-DVFLVKMMEEWSDEKLKQFFLKEK 720
+ F ++ +K +P +N V V+ ++ S K Q FL+++
Sbjct: 581 RFEFQPRTEQCSMVKIKYRPNPQNIQVVEVEPPKKDSLVKFMQNFLEKE 629
>emb|CAE75955.1| B1159F04.18 [Oryza sativa (japonica cultivar-group)]
gi|38605826|emb|CAE05235.3| OSJNBa0011K22.17 [Oryza
sativa (japonica cultivar-group)]
gi|50923135|ref|XP_471928.1| OSJNBa0011K22.17 [Oryza
sativa (japonica cultivar-group)]
gi|50923097|ref|XP_471909.1| B1159F04.18 [Oryza sativa
(japonica cultivar-group)]
Length = 551
Score = 174 bits (441), Expect = 1e-41
Identities = 91/246 (36%), Positives = 154/246 (61%), Gaps = 6/246 (2%)
Query: 453 KKQKEKEEEKRKVELEK----KFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPF 508
K+ + ++V+LEK ++ F K +++QF++F + +K+ + +P
Sbjct: 246 KEVPSSDSADKEVQLEKIVPQEYCDTWLLSFHQRSRKPSVDEQFARFAEVIQKIHINVPL 305
Query: 509 SEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVN 568
+ ++ +P YA+++K++L+ KR L E+V+LTE+CS ++ KLP K+KDPG T+ +
Sbjct: 306 LDAMQ-VPTYARYLKDMLNNKRLLPTM-EVVKLTEQCSSVILHKLPEKKKDPGCPTITCS 363
Query: 569 FGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVR 628
GA + +A CDLG+S+++MP + ++LN L PT+M LQLAD S+ P G+ EDV V+
Sbjct: 364 IGAQQFDQAFCDLGASISVMPKDVPDKLNFTVLTPTLMCLQLADLSVHYPTGIAEDVPVK 423
Query: 629 VGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADEKIGFNIFDL 688
+ +F PVDFV++DMD + PLILGRPFL+T+ A I+VG G+I + ++ +
Sbjct: 424 IRDFFIPVDFVVLDMDMGKETPLILGRPFLSTAGANIDVGMGSIRFDINGKEKSLSFIRG 483
Query: 689 KPKPVE 694
+ +P E
Sbjct: 484 RAEPTE 489
>gb|AAN04521.1| Hypothetical protein with similarity to putative retroelements
[Oryza sativa (japonica cultivar-group)]
gi|19881695|gb|AAM01096.1| Hypothetical protein with
similarity to putative retroelement [Oryza sativa]
Length = 237
Score = 171 bits (434), Expect = 6e-41
Identities = 92/224 (41%), Positives = 146/224 (65%), Gaps = 4/224 (1%)
Query: 456 KEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKM 515
K K ++ V E +T++ PFP K +++QF+ F+ + +K+ +++P + ++ +
Sbjct: 14 KAKGLPEKTVPQEYCYTRLL--PFPQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAMQ-V 70
Query: 516 PQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQV 575
P YA ++K+IL+ KR L E+V+LT +CS ++ KLP K+KDPG T+ + GA +
Sbjct: 71 PTYACYLKDILNNKRPLPT-TEVVKLTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFD 129
Query: 576 RALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFP 635
+ALCDLG+SV++MP +F++LN L PT M LQLAD S+ P G+ EDV V++ +F
Sbjct: 130 QALCDLGASVSVMPKYVFDKLNFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFIL 189
Query: 636 VDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADE 679
VDFV++DMD ++ +ILGRPFL T+ A I+VG G+I + E
Sbjct: 190 VDFVVLDMDTGKEMSIILGRPFLGTAGANIHVGTGSIRFQYQRE 233
>gb|AAT76351.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 585
Score = 166 bits (420), Expect = 3e-39
Identities = 96/273 (35%), Positives = 157/273 (57%), Gaps = 11/273 (4%)
Query: 411 ENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKK 470
EN AIT R G + +P N G + N KE + EK + ++
Sbjct: 222 ENVKAITTRGGKSTRDPPYPNPAGTN------GMSKETPSNDSADKEIQPEKT---VPQE 272
Query: 471 FTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKR 530
+ PFP K +++ F++F+ + +++ + + + ++ +P YA+++K+IL+ KR
Sbjct: 273 YCDTRLLPFPQWSRKTSVDELFARFVEVIQRIHINVLLLDAMQ-VPIYARYLKDILNNKR 331
Query: 531 RLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPL 590
L E+V+LTE CS ++ KLP K+K T+ + GA + +ALCDLG+SV++MP
Sbjct: 332 PLPT-TEVVKLTEHCSNVILHKLPEKKKYSECPTITCSIGAQQFDQALCDLGASVSVMPK 390
Query: 591 SMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIP 650
+F+RLN L PT M LQ+AD S+ G+ EDV +++ +F PVDFV++DMD + P
Sbjct: 391 DVFDRLNFTVLAPTPMRLQMADSSVRYQAGIAEDVPIKIWDFFIPVDFVVLDMDTRKETP 450
Query: 651 LILGRPFLATSQAKINVGKGTISLRVADEKIGF 683
LILGRPFL+T+ A I+V +I + +++ F
Sbjct: 451 LILGRPFLSTAGANIDVETSSICFHINEKEEKF 483
>gb|AAD19780.2| hypothetical protein [Arabidopsis thaliana]
Length = 1048
Score = 165 bits (418), Expect = 5e-39
Identities = 106/265 (40%), Positives = 150/265 (56%), Gaps = 14/265 (5%)
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFS 509
+N K+ + EEK K + + VP PFP K + EK++S F + ++L+V+LPF
Sbjct: 73 DNTGKESDTLEEKAKKTVLPPY--VPKLPFPGRQRKIQREKEYSLFDEIMRQLQVKLPFL 130
Query: 510 EVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNF 569
++++ + Y K +K+IL+ KR L E + ++ + ECS ILQ R G +T
Sbjct: 131 DLVQNVSIYRKHLKDILTNKRTLEEGHVLI--SHECSAILQNVALFSRMI-GEYTFD--- 184
Query: 570 GASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRV 629
R LCDLG+ V+LMP S+ +RL PT M L L DRSI P GV EDV VRV
Sbjct: 185 ------RCLCDLGAGVSLMPFSVAKRLGDTNFTPTKMSLVLGDRSISFPVGVAEDVQVRV 238
Query: 630 GEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADEKIGFNIFDLK 689
G F P DFVII++DE+ + LILGRPFL A I+V K I+LR+ D FN+ +
Sbjct: 239 GNFYIPTDFVIIELDEEPRHRLILGRPFLNIVAALIDVRKSKINLRIGDIVQEFNMERIM 298
Query: 690 PKPVEKNDVFLVKMMEEWSDEKLKQ 714
KP + F V +M+E +E L +
Sbjct: 299 SKPTTECQTFWVDIMDELVNELLAE 323
>gb|AAX95303.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 553
Score = 164 bits (416), Expect = 8e-39
Identities = 97/275 (35%), Positives = 153/275 (55%), Gaps = 26/275 (9%)
Query: 411 ENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKK 470
EN AIT R G G + N KE + EK ++
Sbjct: 279 ENVKAITTRGGTN-------------------GIAKEAPSNDSADKEVQPEKTVPQI--- 316
Query: 471 FTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKR 530
+ PFP K ++Q ++F+ + +K+ + +P + ++ +P YA+++K+IL+ KR
Sbjct: 317 YCDTRLLPFPQRSRKPLADEQLARFVEVIQKIHINVPLLDAMQ-VPTYARYLKDILNNKR 375
Query: 531 RLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPL 590
L E+V+LTE+C+ ++ KLP K+KDP T+ + A + RALCDLG+SV++MP
Sbjct: 376 LLPT-TEVVKLTEQCNDVILHKLPEKKKDPRCPTITCSIRAHQFDRALCDLGASVSVMPK 434
Query: 591 SMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIP 650
+F++LN L T M LQLA+ S+ P G+ E+V ++ +F P DFV++DMD + P
Sbjct: 435 DVFDKLNFTVLASTPMRLQLANSSVRYPAGIAEEVPFKIRDFFIPFDFVVLDMDTGKETP 494
Query: 651 LILGRPFLATSQAKINVGKGTISLRV--ADEKIGF 683
LILGR FL+T+ A I+VG G+I + +EK F
Sbjct: 495 LILGRSFLSTTGANIDVGTGSILFHINGKEEKFEF 529
>gb|AAX92872.1| Transposable element protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 548
Score = 164 bits (414), Expect = 1e-38
Identities = 89/235 (37%), Positives = 144/235 (60%), Gaps = 7/235 (2%)
Query: 451 NKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSE 510
N KE + EK ++ + PFP K ++Q ++F+ + +K+ + +P +
Sbjct: 295 NDSADKEVQPEKTVPQI---YCDTRLLPFPQRSRKPLADEQLARFVEVIQKIHINVPLLD 351
Query: 511 VLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFG 570
++ +P YA+++K+IL+ KR L E+V+LTE+C+ ++ KLP K+KDP T+ +
Sbjct: 352 AMQ-VPTYARYLKDILNNKRLLPT-TEVVKLTEQCNDVILHKLPEKKKDPRCPTITCSIR 409
Query: 571 ASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVG 630
A + RALCDLG+SV++MP +F++LN L T M LQLA+ S+ P G+ E+V ++
Sbjct: 410 AHQFDRALCDLGASVSVMPKDVFDKLNFTVLASTPMRLQLANSSVRYPAGIAEEVPFKIR 469
Query: 631 EFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRV--ADEKIGF 683
+F P DFV++DMD + PLILGR FL+T+ A I+VG G+I + +EK F
Sbjct: 470 DFFIPFDFVVLDMDTGKETPLILGRSFLSTTGANIDVGTGSILFHINGKEEKFEF 524
>gb|AAG52027.1| Athila ORF 1, putative; 43045-40843 [Arabidopsis thaliana]
gi|25405058|pir||F96491 hypothetical protein T4I21.13
[imported] - Arabidopsis thaliana
Length = 530
Score = 163 bits (413), Expect = 2e-38
Identities = 93/243 (38%), Positives = 150/243 (61%), Gaps = 4/243 (1%)
Query: 505 ELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKL-PPKRKDPGSF 563
++P + L +P K++K+++++ R+ E +V L+ ECS I+Q+K+ K +DPGSF
Sbjct: 120 DMPLVDCLALIPNEHKYVKDLITE--RIKEVQGMVVLSHECSAIIQQKIVQEKLEDPGSF 177
Query: 564 TLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVE 623
TLP + +LCDLG+SV++MPLSM +L + KP + L LADR+ P+G++E
Sbjct: 178 TLPCSIRQLTFSNSLCDLGASVSIMPLSMARKLGFVQYKPCDLTLILADRTSRRPFGLLE 237
Query: 624 DVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADE-KIG 682
DV V + E P+DFV+++MDE+SK PLILGRPFLA++ A I+V +G I+L + ++ K+
Sbjct: 238 DVPVMINGVEVPIDFVVLEMDEESKDPLILGRPFLASAGAVIDVKQGKINLNLGEDFKMK 297
Query: 683 FNIFDLKPKPVEKNDVFLVKMMEEWSDEKLKQFFLKEKAGASNKKKKEDT*INQTEQVYS 742
F I + KP + FLV+ M + ++E L++ K+ + K E + Y
Sbjct: 298 FEIRNTMKKPTIEGQTFLVEEMGQLANELLEELIDKDHLQTALTKNSEAGFFSSETLSYE 357
Query: 743 MSM 745
MS+
Sbjct: 358 MSL 360
>gb|AAP53442.1| hypothetical protein similar to putative retroelements [Oryza
sativa (japonica cultivar-group)]
gi|37533706|ref|NP_921155.1| hypothetical protein
similar to putative retroelements [Oryza sativa
(japonica cultivar-group)]
Length = 725
Score = 163 bits (412), Expect = 2e-38
Identities = 82/189 (43%), Positives = 129/189 (67%), Gaps = 2/189 (1%)
Query: 478 PFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENE 537
PFP K +++QF+ F+ + +K+ +++P + ++ +P YA ++K+IL+ KR L E
Sbjct: 65 PFPQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAMQ-VPTYACYLKDILNNKRPLPT-TE 122
Query: 538 IVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLN 597
+V+LT +CS ++ KLP K+KDPG T+ + GA + +ALCDLG+SV++MP +F++LN
Sbjct: 123 VVKLTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVFDKLN 182
Query: 598 VGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPF 657
L PT M LQLAD S+ P G+ EDV V++ +F VDFV++DMD ++ +ILGRPF
Sbjct: 183 FTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIILGRPF 242
Query: 658 LATSQAKIN 666
L T+ A I+
Sbjct: 243 LGTAGANIH 251
>ref|XP_476244.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|54287542|gb|AAV31286.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|46575996|gb|AAT01357.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 256
Score = 162 bits (409), Expect = 5e-38
Identities = 83/170 (48%), Positives = 119/170 (69%), Gaps = 1/170 (0%)
Query: 514 KMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASK 573
++P YA+++K IL+ KR L E+V+LTE CS ++ KL K+KDPG T+ + GA +
Sbjct: 2 QVPTYARYLKVILNNKRPLPT-TEVVKLTEHCSNLILHKLQEKKKDPGCPTITCSIGAQQ 60
Query: 574 QVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFE 633
V ALCDLG+SV++MP ++F++LN L PT M LQLAD S+ P G+VEDV V++ F
Sbjct: 61 FVHALCDLGASVSVMPKNVFDKLNFMVLAPTPMCLQLADSSVRYPAGIVEDVPVKIRNFF 120
Query: 634 FPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADEKIGF 683
PVDFV++DMD + PLILGRPFL+T+ A +VG G+IS + +++ F
Sbjct: 121 IPVDFVVLDMDMGKETPLILGRPFLSTAGANSDVGTGSISFHINGKELKF 170
>gb|AAP52811.1| hypothetical protein similar to putative retroelements [Oryza
sativa (japonica cultivar-group)]
gi|37532444|ref|NP_920524.1| hypothetical protein
similar to putative retroelements [Oryza sativa
(japonica cultivar-group)] gi|21672049|gb|AAM74411.1|
Hypothetical protein similar to putative retroelements
[Oryza sativa (japonica cultivar-group)]
Length = 322
Score = 157 bits (397), Expect = 1e-36
Identities = 82/198 (41%), Positives = 128/198 (64%), Gaps = 10/198 (5%)
Query: 490 KQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVIL 549
+QF++ + + +K+ + +P + ++ +P YA+++K+IL+ KR L TE CS ++
Sbjct: 123 QQFARSVEVIQKIHINVPLLDAMQ-VPTYARYLKDILNNKRPLPT-------TEHCSNVI 174
Query: 550 QRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQ 609
KLP K+KDP T+ + GA + +ALCDLG+SV++MP +F++LN L PT M LQ
Sbjct: 175 LHKLPEKKKDPRCPTITCSIGAQQFDQALCDLGASVSVMPKDVFDKLNFTVLAPTPMRLQ 234
Query: 610 LADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGK 669
LAD S+ P G+ EDV V++ +F PVDFV++DMD + LILG PFL+T+ A I+VG
Sbjct: 235 LADSSVHCPVGIEEDVPVKIWDFFIPVDFVVLDMDTGKETSLILGHPFLSTAGANIDVGM 294
Query: 670 GTISLRV--ADEKIGFNI 685
G+I + +EK F +
Sbjct: 295 GSIRFHINGKEEKFEFQL 312
>emb|CAB81130.1| AT4g07600 [Arabidopsis thaliana] gi|5724765|gb|AAD48069.1| contains
similarity to Pfam family PF00078 -943 Reverse
transcriptase (RNA-dependent DNA polymerase); score
65.8, E=9.4e-16, N=1; may be a pseudogene [Arabidopsis
thaliana] gi|25407387|pir||F85074 hypothetical protein
AT4g07600 [imported] - Arabidopsis thaliana
Length = 630
Score = 155 bits (391), Expect = 6e-36
Identities = 90/239 (37%), Positives = 145/239 (60%), Gaps = 4/239 (1%)
Query: 495 FISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLP 554
F K++ + +P + L + KF+K+++ + R+ E +V L+ ECS I+Q+K+
Sbjct: 2 FAKNIKEVELRIPLVDALALILDTHKFLKDLIVE--RIQEVQGMVVLSHECSAIIQKKIV 59
Query: 555 PKR-KDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADR 613
PK+ DPGSFTLP G R LCDLG+ V+ MPLS+ +RL + K + L LADR
Sbjct: 60 PKKLSDPGSFTLPCFLGTVAFNRCLCDLGALVSPMPLSIAKRLGFTQYKSCNISLILADR 119
Query: 614 SIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTIS 673
S+ G+++++ +R+G E DFVI++MDE+ K PLIL RPFLAT+ A I+V KG I
Sbjct: 120 SVRISHGLLKNLPIRIGAAEISTDFVILEMDEEPKDPLILRRPFLATAGAMIDVKKGKID 179
Query: 674 LRVA-DEKIGFNIFDLKPKPVEKNDVFLVKMMEEWSDEKLKQFFLKEKAGASNKKKKED 731
L + D ++ F+I D KP + +F ++ M++ +DE L++ ++ ++ K +D
Sbjct: 180 LNLGKDFRMTFDIKDAMKKPNIEGQLFWIEEMDQLADELLEELAQEDHLNSALTKTGQD 238
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.337 0.147 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,200,891,936
Number of Sequences: 2540612
Number of extensions: 49707643
Number of successful extensions: 289150
Number of sequences better than 10.0: 401
Number of HSP's better than 10.0 without gapping: 120
Number of HSP's successfully gapped in prelim test: 294
Number of HSP's that attempted gapping in prelim test: 285696
Number of HSP's gapped (non-prelim): 1878
length of query: 765
length of database: 863,360,394
effective HSP length: 136
effective length of query: 629
effective length of database: 517,837,162
effective search space: 325719574898
effective search space used: 325719574898
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0265.8


