

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.5
(404 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
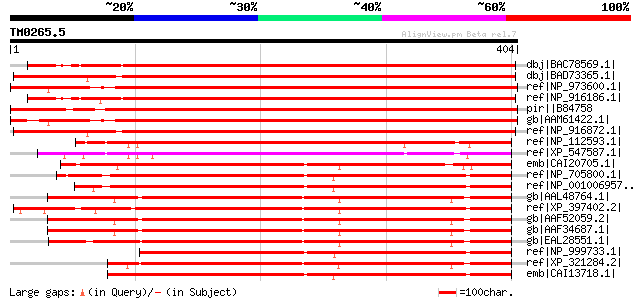
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAC78569.1| katanin [Oryza sativa (japonica cultivar-group)]... 581 e-164
dbj|BAD73365.1| vacuolar protein sorting factor 4B-like [Oryza s... 579 e-164
ref|NP_973600.1| katanin, putative [Arabidopsis thaliana] 576 e-163
ref|NP_916186.1| katanin p60 subunit A 1-like [Oryza sativa (jap... 572 e-162
pir||B84758 probable katanin [imported] - Arabidopsis thaliana 564 e-159
gb|AAM61422.1| putative katanin [Arabidopsis thaliana] gi|201970... 555 e-157
ref|NP_916872.1| putative katanin [Oryza sativa (japonica cultiv... 525 e-148
ref|NP_112593.1| hypothetical protein LOC83473 [Homo sapiens] gi... 360 6e-98
ref|XP_547587.1| PREDICTED: similar to RIKEN cDNA 3110023G01 [Ca... 301 2e-80
emb|CAI20705.1| novel protein similar to vertebrate katanin p60 ... 289 9e-77
ref|NP_705800.1| katanin p60 subunit A-like 1 [Mus musculus] gi|... 287 5e-76
ref|NP_001006957.1| katanin p60 subunit A-like 1 [Rattus norvegi... 287 5e-76
gb|AAL48764.1| RE17942p [Drosophila melanogaster] 285 2e-75
ref|XP_397402.2| PREDICTED: similar to ENSANGP00000018492 [Apis ... 284 3e-75
gb|AAF52059.2| CG10229-PA [Drosophila melanogaster] gi|24644145|... 284 4e-75
gb|AAF34687.1| putative microtubule severing protein katanin p60... 284 4e-75
gb|EAL28551.1| GA10173-PA [Drosophila pseudoobscura] 283 9e-75
ref|NP_999733.1| katanin p60 [Strongylocentrotus purpuratus] gi|... 282 1e-74
ref|XP_321284.2| ENSANGP00000018492 [Anopheles gambiae str. PEST... 281 2e-74
emb|CAI13718.1| katanin p60 subunit A-like 1 [Homo sapiens] gi|1... 281 2e-74
>dbj|BAC78569.1| katanin [Oryza sativa (japonica cultivar-group)]
gi|57899262|dbj|BAD87507.1| katanin [Oryza sativa
(japonica cultivar-group)]
Length = 386
Score = 581 bits (1498), Expect = e-164
Identities = 302/390 (77%), Positives = 334/390 (85%), Gaps = 11/390 (2%)
Query: 15 DFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQG-Q 73
DFK +++S+FG KK + +N NG V+NG+V KR+SD+A+YEQ Q Q
Sbjct: 2 DFKGFWESRFGGKKEQEPEQNGH---ANG-----VANGSVR-KRTSDLAVYEQFEQQARQ 52
Query: 74 NGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAK 133
+ + N D QK LLP FESAEMR LAE+L RDIIRGSPDVKWESIKGLENAK
Sbjct: 53 TEVRAAAIRDGNADAI-QKPLLPSFESAEMRNLAETLLRDIIRGSPDVKWESIKGLENAK 111
Query: 134 RLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSV 193
RLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASS+
Sbjct: 112 RLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSI 171
Query: 194 VSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQM 253
VSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQM
Sbjct: 172 VSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQM 231
Query: 254 DGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEES 313
DGLT+T++LVFVLAATNLPWELDAAMLRRLEKRILVPLPE EAR AMFEELLP +
Sbjct: 232 DGLTKTNDLVFVLAATNLPWELDAAMLRRLEKRILVPLPEAEARHAMFEELLPSTTSKLE 291
Query: 314 IPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDI 373
+PYD LV +TEGYSGSDIRL+CKE AMQPLRRLMS LE R++LVPEEELP+VGP++PEDI
Sbjct: 292 VPYDTLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLEARDELVPEEELPEVGPLKPEDI 351
Query: 374 QAALKNTRPSAHLHAHKYDKFNADYGSQIL 403
+ AL+NTRPSAHLHAH+Y+KFN DYGSQIL
Sbjct: 352 EVALRNTRPSAHLHAHRYEKFNQDYGSQIL 381
>dbj|BAD73365.1| vacuolar protein sorting factor 4B-like [Oryza sativa (japonica
cultivar-group)] gi|56201862|dbj|BAD73312.1| vacuolar
protein sorting factor 4B-like [Oryza sativa (japonica
cultivar-group)]
Length = 410
Score = 579 bits (1492), Expect = e-164
Identities = 297/409 (72%), Positives = 337/409 (81%), Gaps = 13/409 (3%)
Query: 4 EEPMPTRWSFQDFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSS--- 60
+EP TRW+F+DF++YY+ + G ++ E+ D G G + + +G+ S R S
Sbjct: 5 DEPSITRWTFEDFEVYYEVRLGIRREPGGDEDGDGDGGGGRGYAPLGSGSAGSTRPSAAH 64
Query: 61 -----DMAIYEQL-RSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDI 114
D+A++EQ R + + + + PQKSLLP FESAEMR LAE+L RDI
Sbjct: 65 ANGGADLAVFEQFERLERKVELRNGAIEAGP----PQKSLLPSFESAEMRNLAETLLRDI 120
Query: 115 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLA 174
IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYF GLLSPWKGILLFGPPGTGKTMLA
Sbjct: 121 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFKGLLSPWKGILLFGPPGTGKTMLA 180
Query: 175 KAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGE 234
KAVATECKTTFFNISASS+VSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQRGE
Sbjct: 181 KAVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRGE 240
Query: 235 ARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEP 294
ARSEHEASRRLKTELLIQMDGLT+TD+LVFVLAATNLPWELDAAMLRRLEKRILVPLPE
Sbjct: 241 ARSEHEASRRLKTELLIQMDGLTKTDDLVFVLAATNLPWELDAAMLRRLEKRILVPLPEQ 300
Query: 295 EARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRE 354
EAR AMFEELLP P +IPYD+LV +TEGYSGSDIRL+CKE AMQPLRRLMS LE R+
Sbjct: 301 EARHAMFEELLPSVPGTMNIPYDVLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLEGRQ 360
Query: 355 DLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQIL 403
+ VPE+ELP+VGP+ EDI+ AL+NTRPSAHLH H+Y+KFN DYGS +L
Sbjct: 361 EEVPEDELPEVGPVTTEDIELALRNTRPSAHLHVHRYEKFNQDYGSHVL 409
>ref|NP_973600.1| katanin, putative [Arabidopsis thaliana]
Length = 393
Score = 576 bits (1485), Expect = e-163
Identities = 294/409 (71%), Positives = 337/409 (81%), Gaps = 21/409 (5%)
Query: 1 MADEEPMPTRWSFQDFKLYYDSKFGRKKV----VQNGENADKAVGNGSSMSVVSNGNVHS 56
MA +EP TRWSF +FK +YD+KFGRKK+ V N + + NG++ V +N + +
Sbjct: 1 MATDEPSQTRWSFLEFKTFYDAKFGRKKLPEEDVSNKDQPEDGSSNGNNGDVNNNSSPVT 60
Query: 57 KRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIR 116
+ + A+ NG N + E+P+KS+ PPFESAE RTLAESLSRDIIR
Sbjct: 61 NQDGNTAL--------ANG--------NVIREKPKKSMFPPFESAETRTLAESLSRDIIR 104
Query: 117 GSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKA 176
G+P++KWESIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKA
Sbjct: 105 GNPNIKWESIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKA 164
Query: 177 VATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG-EA 235
VATEC TTFFNISASSVVSKWRGDSEKL++VLF LARHHAPSTIFLDEIDAIISQRG E
Sbjct: 165 VATECNTTFFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEG 224
Query: 236 RSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPE 295
RSEHEASRRLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKRILVPLP+PE
Sbjct: 225 RSEHEASRRLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPE 284
Query: 296 ARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRED 355
AR MFE L+P QP +E +P+D+LV ++EGYSGSDIR+LCKE AMQPLRR ++ LE RED
Sbjct: 285 ARRGMFEMLIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLAILEDRED 344
Query: 356 LVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQILQ 404
+VPE+ELPK+GPI PEDI AL NTRPSAHLHAH YDKFN DYGSQIL+
Sbjct: 345 VVPEDELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQILK 393
>ref|NP_916186.1| katanin p60 subunit A 1-like [Oryza sativa (japonica
cultivar-group)]
Length = 428
Score = 572 bits (1474), Expect = e-162
Identities = 302/412 (73%), Positives = 334/412 (80%), Gaps = 33/412 (8%)
Query: 15 DFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQ--- 71
DFK +++S+FG KK + +N NG V+NG+V KR+SD+A+YEQ Q
Sbjct: 22 DFKGFWESRFGGKKEQEPEQNGH---ANG-----VANGSVR-KRTSDLAVYEQFEQQVFL 72
Query: 72 --------------------GQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLS 111
Q + + N D QK LLP FESAEMR LAE+L
Sbjct: 73 DRRLVSCVVLDPRSVLLFQARQTEVRAAAIRDGNADAI-QKPLLPSFESAEMRNLAETLL 131
Query: 112 RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT 171
RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT
Sbjct: 132 RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT 191
Query: 172 MLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQ 231
MLAKAVATECKTTFFNISASS+VSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQ
Sbjct: 192 MLAKAVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQ 251
Query: 232 RGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPL 291
RGEARSEHEASRRLKTELLIQMDGLT+T++LVFVLAATNLPWELDAAMLRRLEKRILVPL
Sbjct: 252 RGEARSEHEASRRLKTELLIQMDGLTKTNDLVFVLAATNLPWELDAAMLRRLEKRILVPL 311
Query: 292 PEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLE 351
PE EAR AMFEELLP + +PYD LV +TEGYSGSDIRL+CKE AMQPLRRLMS LE
Sbjct: 312 PEAEARHAMFEELLPSTTSKLEVPYDTLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLE 371
Query: 352 QREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQIL 403
R++LVPEEELP+VGP++PEDI+ AL+NTRPSAHLHAH+Y+KFN DYGSQIL
Sbjct: 372 ARDELVPEEELPEVGPLKPEDIEVALRNTRPSAHLHAHRYEKFNQDYGSQIL 423
>pir||B84758 probable katanin [imported] - Arabidopsis thaliana
Length = 393
Score = 564 bits (1453), Expect = e-159
Identities = 289/405 (71%), Positives = 329/405 (80%), Gaps = 13/405 (3%)
Query: 1 MADEEPMPTRWSFQDFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSS 60
MA +EP TRWSF +FK +YD+KFGRKK+ + + +GSS NGN ++
Sbjct: 1 MATDEPSQTRWSFLEFKTFYDAKFGRKKLPEEDVSNKDQPEDGSS-----NGNNGDVNNN 55
Query: 61 DMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIRGSPD 120
+ Q + + + +KS+ PPFESAE RTLAESLSRDIIRG+P+
Sbjct: 56 SSPVTNQR-------CYVYSLGSDECFFCRKKSMFPPFESAETRTLAESLSRDIIRGNPN 108
Query: 121 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATE 180
+KWESIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKAVATE
Sbjct: 109 IKWESIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKAVATE 168
Query: 181 CKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG-EARSEH 239
C TTFFNISASSVVSKWRGDSEKL++VLF LARHHAPSTIFLDEIDAIISQRG E RSEH
Sbjct: 169 CNTTFFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEGRSEH 228
Query: 240 EASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVA 299
EASRRLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKRILVPLP+PEAR
Sbjct: 229 EASRRLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPEARRG 288
Query: 300 MFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPE 359
MFE L+P QP +E +P+D+LV ++EGYSGSDIR+LCKE AMQPLRR ++ LE RED+VPE
Sbjct: 289 MFEMLIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLAILEDREDVVPE 348
Query: 360 EELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQILQ 404
+ELPK+GPI PEDI AL NTRPSAHLHAH YDKFN DYGSQIL+
Sbjct: 349 DELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQILK 393
>gb|AAM61422.1| putative katanin [Arabidopsis thaliana] gi|20197082|gb|AAC26698.2|
putative katanin [Arabidopsis thaliana]
gi|18403587|ref|NP_565791.1| katanin, putative
[Arabidopsis thaliana]
Length = 384
Score = 555 bits (1431), Expect = e-157
Identities = 289/409 (70%), Positives = 329/409 (79%), Gaps = 30/409 (7%)
Query: 1 MADEEPMPTRWSFQDFKLYYDSKFGRKKV----VQNGENADKAVGNGSSMSVVSNGNVHS 56
MA +EP TRWSF FGRKK+ V N + + NG++ V +N + +
Sbjct: 1 MATDEPSQTRWSFL---------FGRKKLPEEDVSNKDQPEDGSSNGNNGDVNNNSSPVT 51
Query: 57 KRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIR 116
+ + A+ NG N + E+P+KS+ PPFESAE RTLAESLSRDIIR
Sbjct: 52 NQDGNTAL--------ANG--------NVIREKPKKSMFPPFESAETRTLAESLSRDIIR 95
Query: 117 GSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKA 176
G+P++KWESIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKA
Sbjct: 96 GNPNIKWESIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKA 155
Query: 177 VATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG-EA 235
VATEC TTFFNISASSVVSKWRGDSEKL++VLF LARHHAPSTIFLDEIDAIISQRG E
Sbjct: 156 VATECNTTFFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEG 215
Query: 236 RSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPE 295
RSEHEASRRLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKRILVPLP+PE
Sbjct: 216 RSEHEASRRLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPE 275
Query: 296 ARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRED 355
AR MFE L+P QP +E +P+D+LV ++EGYSGSDIR+LCKE AMQPLRR ++ LE RED
Sbjct: 276 ARRGMFEMLIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLAILEDRED 335
Query: 356 LVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQILQ 404
+VPE+ELPK+GPI PEDI AL NTRPSAHLHAH YDKFN DYGSQIL+
Sbjct: 336 VVPEDELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQILK 384
>ref|NP_916872.1| putative katanin [Oryza sativa (japonica cultivar-group)]
Length = 411
Score = 525 bits (1353), Expect = e-148
Identities = 277/410 (67%), Positives = 322/410 (77%), Gaps = 14/410 (3%)
Query: 4 EEPMPTRWSFQDFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSS--- 60
+EP TRW+F+DF++YY+ + G ++ E+ D G G + + +G+ S R S
Sbjct: 5 DEPSITRWTFEDFEVYYEVRLGIRREPGGDEDGDGDGGGGRGYAPLGSGSAGSTRPSAAH 64
Query: 61 -----DMAIYEQL-RSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDI 114
D+A++EQ R + + + + PQKSLLP FESAEMR LAE+L RDI
Sbjct: 65 ANGGADLAVFEQFERLERKVELRNGAIEAGP----PQKSLLPSFESAEMRNLAETLLRDI 120
Query: 115 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFT-GLLSPWKGILLFGPPGTGKTML 173
IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPK ++ + I + +TML
Sbjct: 121 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKVLQRSAITLERYIAFWSTRNRKETML 180
Query: 174 AKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG 233
AKAVATECKTTFFNISASS+VSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQRG
Sbjct: 181 AKAVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRG 240
Query: 234 EARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPE 293
EARSEHEASRRLKTELLIQMDGLT+TD+LVFVLAATNLPWELDAAMLRRLEKRILVPLPE
Sbjct: 241 EARSEHEASRRLKTELLIQMDGLTKTDDLVFVLAATNLPWELDAAMLRRLEKRILVPLPE 300
Query: 294 PEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQR 353
EAR AMFEELLP P +IPYD+LV +TEGYSGSDIRL+CKE AMQPLRRLMS LE R
Sbjct: 301 QEARHAMFEELLPSVPGTMNIPYDVLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLEGR 360
Query: 354 EDLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQIL 403
++ VPE+ELP+VGP+ EDI+ AL+NTRPSAHLH H+Y+KFN DYGS +L
Sbjct: 361 QEEVPEDELPEVGPVTTEDIELALRNTRPSAHLHVHRYEKFNQDYGSHVL 410
>ref|NP_112593.1| hypothetical protein LOC83473 [Homo sapiens]
gi|34190544|gb|AAH34999.2| Similar to mouse
4933439B08Rik protein [Homo sapiens]
Length = 466
Score = 360 bits (923), Expect = 6e-98
Identities = 194/364 (53%), Positives = 256/364 (70%), Gaps = 20/364 (5%)
Query: 53 NVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKS--LLPPFES-----AEMRT 105
N H +R + ++ L + G T++++ N D P S LL P + +EMR
Sbjct: 106 NAHPRRGQ-IIDFQGLLTDAIKGA-TSELALNTFDHNPDPSERLLKPLSAFIGMNSEMRE 163
Query: 106 LAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGP 165
LA +SRDI +P++KW I GL+ AK+L+KEAVV PI+YP+ FTG+LSPWKG+LL+GP
Sbjct: 164 LAAVVSRDIYLHNPNIKWNDIIGLDAAKQLVKEAVVYPIRYPQLFTGILSPWKGLLLYGP 223
Query: 166 PGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEI 225
PGTGKT+LAKAVATECKTTFFNISAS++VSKWRGDSEKLV+VLF+LAR+HAPSTIFLDE+
Sbjct: 224 PGTGKTLLAKAVATECKTTFFNISASTIVSKWRGDSEKLVRVLFELARYHAPSTIFLDEL 283
Query: 226 DAIISQRGEAR-SEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLE 284
++++SQRG A EHE S R+KTELL+QMDGL R+++LVFVLAA+NLPWELD AMLRRLE
Sbjct: 284 ESVMSQRGTASGGEHEGSLRMKTELLVQMDGLARSEDLVFVLAASNLPWELDCAMLRRLE 343
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEES------IPYDLLVNQTEGYSGSDIRLLCKEV 338
KRILV LP EAR AM LPP + + Y +L +TEGYSGSDI+L+C+E
Sbjct: 344 KRILVDLPSREARQAMIYHWLPPVSKSRALELHTELEYSVLSQETEGYSGSDIKLVCREA 403
Query: 339 AMQPLRRLMSQLEQREDLVPEEELPKV--GPIRPEDIQAALKNTRPSAHLHAHKYDKFNA 396
AM+P+R++ LE + +LP++ + D L +T+PSA A +Y +
Sbjct: 404 AMRPVRKIFDALENHQS--ESSDLPRIQLDIVTTADFLDVLTHTKPSAKNLAQRYSDWQR 461
Query: 397 DYGS 400
++ S
Sbjct: 462 EFES 465
>ref|XP_547587.1| PREDICTED: similar to RIKEN cDNA 3110023G01 [Canis familiaris]
Length = 567
Score = 301 bits (772), Expect = 2e-80
Identities = 190/432 (43%), Positives = 251/432 (57%), Gaps = 61/432 (14%)
Query: 23 KFGRKKVVQNGENADKAVGN--------GSSMSVVSNGNVHSK---RSSDMAIYEQLRSQ 71
K G K Q + +AV N G S+S ++ GN + + + L +
Sbjct: 142 KSGDAKSNQKEQPKQEAVNNSHMESADFGLSISGINKGNGEENVRPQKGQIIDFRGLLTD 201
Query: 72 GQNGIHTNDVSPNNMDERPQKS--LLPPFES-----AEMRTLAESLSR------------ 112
G TN++ N+ D P S LL P + +EMR LA +SR
Sbjct: 202 AIKGA-TNELGLNSFDCNPDPSERLLKPLSAFIGMNSEMRELAAMVSRVHESPLYAKLFL 260
Query: 113 ---------------------DIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFT 151
DI +P++KW+ I GL+ AK+L+KEAVV PI+YP+ FT
Sbjct: 261 RKCGNKNVSGWWASLQIHNLKDIYLHNPNIKWDDIIGLDTAKQLVKEAVVYPIRYPQLFT 320
Query: 152 GLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQL 211
G+LSPWKG+LL+GPPGTGKT+LAKAVATECKTTFFNISAS++VSKWRGDSEKLV+VLF+L
Sbjct: 321 GILSPWKGLLLYGPPGTGKTLLAKAVATECKTTFFNISASTIVSKWRGDSEKLVRVLFEL 380
Query: 212 ARHHAPSTIFLDEIDAIISQRGEA-RSEHEASRRLKTELLIQMDGLTRTDELVFVLAATN 270
AR+HAPSTIFLDE+++++SQRG A EHE S R+KTELL+QMDGL R+++LVFVLAA+N
Sbjct: 381 ARYHAPSTIFLDELESVMSQRGTAPGGEHEGSLRMKTELLVQMDGLARSEDLVFVLAASN 440
Query: 271 LPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSD 330
LPWELD AMLRRLEKRILV LP EAR AM LPP ++ +L G
Sbjct: 441 LPWELDCAMLRRLEKRILVDLPSREARRAMIYHWLPPVSKSRAL--ELRTELEYGVLSQH 498
Query: 331 IRLLCKEVAMQPLRRLMSQLEQREDLVPEEELP--KVGPIRPEDIQAALKNTRPSAHLHA 388
+ + V L SQ+ Q + LP ++ + D L +T+PSA
Sbjct: 499 LTAYIRMVVGSFLYCSASQVIQTQ----SSNLPGIQLDTVTTADFLDVLAHTKPSAKNLT 554
Query: 389 HKYDKFNADYGS 400
+Y + +++ S
Sbjct: 555 QRYSAWQSEFES 566
>emb|CAI20705.1| novel protein similar to vertebrate katanin p60 (ATPase-containing)
subunit A 1 (KATNA1) [Danio rerio]
gi|66472538|ref|NP_001018440.1| hypothetical protein
LOC553631 [Danio rerio] gi|63101878|gb|AAH95321.1|
Hypothetical protein LOC553631 [Danio rerio]
Length = 485
Score = 289 bits (740), Expect = 9e-77
Identities = 167/374 (44%), Positives = 237/374 (62%), Gaps = 23/374 (6%)
Query: 41 GNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPN---NMDERPQKSLLPP 97
G+G+ +SV G ++ S +A ++ + Q N P N E + +
Sbjct: 120 GHGNRLSVGPRGQ--ARPSPRVANGDKGKPQKSKEKKENPSKPKEDKNKAEAVETEVKRF 177
Query: 98 FESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPW 157
E + L ++L RDII +P+V W+ I LE AK+LLKEAVV+P+ P++F G+ PW
Sbjct: 178 DRGGEDKDLIDALERDIISQNPNVTWDDIADLEEAKKLLKEAVVLPMWMPEFFKGIRRPW 237
Query: 158 KGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAP 217
KG+L+ GPPGTGKT+LAKAVATEC+TTFFN+S+S++ SK+RG+SEKLV++LF++AR +AP
Sbjct: 238 KGVLMVGPPGTGKTLLAKAVATECRTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAP 297
Query: 218 STIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE-----LVFVLAATNLP 272
+TIF+DEID+I S+RG + EHEASRR+K ELL+QMDG+ T E +V VLAATN P
Sbjct: 298 TTIFIDEIDSICSRRGTS-EEHEASRRVKAELLVQMDGVGGTSENDPSKMVMVLAATNFP 356
Query: 273 WELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIR 332
W++D A+ RRLEKRI +PLP + RV + + L + D + Q EGYSG+DI
Sbjct: 357 WDIDEALRRRLEKRIYIPLPSAKGRVDLLKINLKELDLANDVNMDKIAEQMEGYSGADIT 416
Query: 333 LLCKEVAMQPLRRLMSQLEQREDLVPEE--ELPKVG---PIRPEDIQAALKNTRPS-AHL 386
+C++ ++ +RR + E L PEE LPK P ED + ALK S +
Sbjct: 417 NVCRDASLMAMRRRI------EGLTPEEIRNLPKDEMHMPTTMEDFETALKKVSKSVSAA 470
Query: 387 HAHKYDKFNADYGS 400
KY+K+ A++GS
Sbjct: 471 DLEKYEKWIAEFGS 484
>ref|NP_705800.1| katanin p60 subunit A-like 1 [Mus musculus]
gi|20987888|gb|AAH30434.1| Katanin p60 subunit A-like 1
[Mus musculus] gi|60390206|sp|Q8K0T4|KATL1_MOUSE Katanin
p60 ATPase-containing subunit A-like 1 (Katanin p60
subunit A-like 1) (p60 katanin-like 1)
Length = 488
Score = 287 bits (734), Expect = 5e-76
Identities = 160/372 (43%), Positives = 241/372 (64%), Gaps = 18/372 (4%)
Query: 38 KAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPP 97
K VG G+ +V + SK + + R++G++ D + N+ + S +P
Sbjct: 125 KDVGAGAR-GLVGRAHQISKSDKPASRDKDYRARGRD-----DKARKNVQDGASDSEIPK 178
Query: 98 FESAEM-RTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSP 156
F+ A + L E+L RDI+ +P + W+ I LE AK+LL+EAVV+P+ P +F G+ P
Sbjct: 179 FDGAGYDKDLVEALERDIVSRNPSIHWDDIADLEEAKKLLREAVVLPMWMPDFFKGIRRP 238
Query: 157 WKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHA 216
WKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S+S++ SK+RG+SEKLV++LF++AR +A
Sbjct: 239 WKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYA 298
Query: 217 PSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL------TRTDELVFVLAATN 270
P+TIF+DEID+I S+RG + EHEASRR+K+ELLIQMDG+ ++V VLAATN
Sbjct: 299 PTTIFIDEIDSICSRRGTS-DEHEASRRVKSELLIQMDGVGGALENDDPSKMVMVLAATN 357
Query: 271 LPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSD 330
PW++D A+ RRLEKRI +PLP + R + + L + + + + ++TEGYSG+D
Sbjct: 358 FPWDIDEALRRRLEKRIYIPLPTAKGRAELLKISLREVELDPDVHLEDIADKTEGYSGAD 417
Query: 331 IRLLCKEVAMQPLRRLMSQLEQRE-DLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHA 388
I +C++ ++ +RR ++ L E + +EEL P+ D++ ALK S +
Sbjct: 418 ITNICRDASLMAMRRRINGLSPEEIRALSKEELQM--PVTRGDLELALKKIAKSVSAADL 475
Query: 389 HKYDKFNADYGS 400
KY+K+ ++GS
Sbjct: 476 EKYEKWMVEFGS 487
>ref|NP_001006957.1| katanin p60 subunit A-like 1 [Rattus norvegicus]
gi|53733477|gb|AAH83673.1| Katanin p60 subunit A-like 1
[Rattus norvegicus] gi|60389845|sp|Q5XIK7|KATL1_RAT
Katanin p60 ATPase-containing subunit A-like 1 (Katanin
p60 subunit A-like 1) (p60 katanin-like 1)
Length = 488
Score = 287 bits (734), Expect = 5e-76
Identities = 159/360 (44%), Positives = 233/360 (64%), Gaps = 19/360 (5%)
Query: 52 GNVHSKRSSDMAIY--EQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEM-RTLAE 108
G H SD A + R++G++ D + NM + +P F+ A + L E
Sbjct: 136 GRAHQISKSDKAASRDKDYRARGRD-----DKARKNMQDGASDGEIPKFDGAGYDKDLVE 190
Query: 109 SLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGT 168
+L RDI+ +P + W+ I LE AK+LL+EAVV+P+ P +F G+ PWKG+L+ GPPGT
Sbjct: 191 ALERDIVSRNPSIHWDDIADLEEAKKLLREAVVLPMWMPDFFKGIRRPWKGVLMVGPPGT 250
Query: 169 GKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAI 228
GKTMLAKAVATEC TTFFN+S+S++ SK+RG+SEKLV++LF++AR +AP+TIF+DEID+I
Sbjct: 251 GKTMLAKAVATECGTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAPTTIFIDEIDSI 310
Query: 229 ISQRGEARSEHEASRRLKTELLIQMDGL------TRTDELVFVLAATNLPWELDAAMLRR 282
S+RG + EHEASRR+K+ELLIQMDG+ ++V VLAATN PW++D A+ RR
Sbjct: 311 CSRRGTS-DEHEASRRVKSELLIQMDGVGGALENDDPSKMVMVLAATNFPWDIDEALRRR 369
Query: 283 LEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQP 342
LEKRI +PLP + R + + L + I + + +TEGYSG+DI +C++ ++
Sbjct: 370 LEKRIYIPLPTAKGRAELLKISLREVELDPDIHLEDIAEKTEGYSGADITNICRDASLMA 429
Query: 343 LRRLMSQLEQRE-DLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKYDKFNADYGS 400
+RR ++ L E + +EEL P+ D++ ALK S + KY+K+ ++GS
Sbjct: 430 MRRRINGLSPEEIRALSKEELQM--PVTRGDLELALKKIAKSVSAADLEKYEKWMVEFGS 487
>gb|AAL48764.1| RE17942p [Drosophila melanogaster]
Length = 572
Score = 285 bits (728), Expect = 2e-75
Identities = 164/380 (43%), Positives = 238/380 (62%), Gaps = 18/380 (4%)
Query: 31 QNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVS---PNNMD 87
+N +A G + + + G S +++ A + + G NG D P
Sbjct: 200 RNSTSATAPSGGARTTNGRAGGRKLSTSNTNEARDDDSTAAGNNGGAAGDGENGDPQAAQ 259
Query: 88 ERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYP 147
E +K AE L + L RDI++ P V+W I L +AKRLL+EAVV+P+ P
Sbjct: 260 EEERKFQTNNHIEAE---LVDILERDILQKDPKVRWSDIADLHDAKRLLEEAVVLPMLMP 316
Query: 148 KYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKV 207
YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S++++ SK+RG+SEK+V++
Sbjct: 317 DYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSATLTSKYRGESEKMVRL 376
Query: 208 LFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE---LVF 264
LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+ +E +V
Sbjct: 377 LFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGVGGGEEQAKVVM 435
Query: 265 VLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTE 324
VLAATN PW++D A+ RRLEKRI +PLP E R A+ + L ++S+ + N+ +
Sbjct: 436 VLAATNFPWDIDEALRRRLEKRIYIPLPSDEGREALLKINLREVKVDDSVDLTYVANELK 495
Query: 325 GYSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPEDIQAALKNTR 381
GYSG+DI +C+E +M +RR ++ L EQ L EE +L P+ +D A+
Sbjct: 496 GYSGADITNVCREASMMSMRRKIAGLTPEQIRQLATEEVDL----PVSNKDFNEAMSRCN 551
Query: 382 PS-AHLHAHKYDKFNADYGS 400
S + KY+K+ ++GS
Sbjct: 552 KSVSRADLDKYEKWMREFGS 571
>ref|XP_397402.2| PREDICTED: similar to ENSANGP00000018492 [Apis mellifera]
Length = 506
Score = 284 bits (727), Expect = 3e-75
Identities = 168/419 (40%), Positives = 257/419 (61%), Gaps = 31/419 (7%)
Query: 4 EEPM--PTRWSFQDFKLYYDSKFGR------------KKVVQNGENADKAVGNGSSMSVV 49
EEP PT W++ + ++ R +K ++ +N K S V
Sbjct: 96 EEPTRDPTLWTYNNSDNSWNQTPARDPDVWPPLTPAEQKNIKPLKNQSKQQNRTSIRKSV 155
Query: 50 SNGNVHSKRSSDMAIYEQ--LRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLA 107
+ G KRS A+ ++ + Q ++ I+ + +D ++ P S R L
Sbjct: 156 TTG----KRSDTKAMVKKDDRKIQKKDDINKEKLETEKIDIEVEERKFEP--SGNDRDLV 209
Query: 108 ESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPG 167
+ L RDI++ +P++ W+ I L AKRLL+EAVV+P+ P +F G+ PWKG+L+ GPPG
Sbjct: 210 DLLERDIVQKNPNIHWDDIADLHEAKRLLEEAVVLPMWMPDFFKGIRRPWKGVLMVGPPG 269
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
TGKTMLAKAVATEC TTFFN+S+S++ SK+RG+SEKLV++LF++AR +APSTIF+DEID+
Sbjct: 270 TGKTMLAKAVATECGTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAPSTIFIDEIDS 329
Query: 228 IISQRGEARSEHEASRRLKTELLIQMDGLTRTDE----LVFVLAATNLPWELDAAMLRRL 283
+ S+RG + SEHEASRR+K+ELL+QMDG++ E +V VLAATN PW++D A+ RRL
Sbjct: 330 LCSRRG-SESEHEASRRVKSELLVQMDGISSNSEDPSKVVMVLAATNFPWDIDEALRRRL 388
Query: 284 EKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPL 343
EKRI +PLP E R A+ + L + S+ + + EGYSG+DI +C++ +M +
Sbjct: 389 EKRIYIPLPNREGREALLKINLREVKVDLSVNLADIAKKLEGYSGADITNVCRDASMMSM 448
Query: 344 RRLMSQLEQRE-DLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKYDKFNADYGS 400
R+ ++ L+ + +P+EEL P+ D A++ S + KY+K+ +++GS
Sbjct: 449 RKKIAGLKPDQIRQLPKEELDL--PVSAADFDEAVERCNKSVSQEDLEKYEKWMSEFGS 505
>gb|AAF52059.2| CG10229-PA [Drosophila melanogaster] gi|24644145|ref|NP_524997.2|
CG10229-PA [Drosophila melanogaster]
Length = 572
Score = 284 bits (726), Expect = 4e-75
Identities = 164/380 (43%), Positives = 238/380 (62%), Gaps = 18/380 (4%)
Query: 31 QNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVS---PNNMD 87
+N +A G + + + G S +++ A + + G NG D P
Sbjct: 200 RNSTSATAPSGGARTTNGRAGGRKLSTSNTNEARDDDSTAAGNNGGAAGDGENGDPQAAQ 259
Query: 88 ERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYP 147
E +K AE L + L RDI++ P V+W I L +AKRLL+EAVV+P+ P
Sbjct: 260 EEERKFQPNNHIEAE---LVDILERDILQKDPKVRWSDIADLHDAKRLLEEAVVLPMLMP 316
Query: 148 KYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKV 207
YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S++++ SK+RG+SEK+V++
Sbjct: 317 DYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSATLTSKYRGESEKMVRL 376
Query: 208 LFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE---LVF 264
LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+ +E +V
Sbjct: 377 LFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGVGGGEEQAKVVM 435
Query: 265 VLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTE 324
VLAATN PW++D A+ RRLEKRI +PLP E R A+ + L ++S+ + N+ +
Sbjct: 436 VLAATNFPWDIDEALRRRLEKRIYIPLPSDEGREALLKINLREVKVDDSVDLTYVANELK 495
Query: 325 GYSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPEDIQAALKNTR 381
GYSG+DI +C+E +M +RR ++ L EQ L EE +L P+ +D A+
Sbjct: 496 GYSGADITNVCREASMMSMRRKIAGLTPEQIRQLATEEVDL----PVSNKDFNEAMSRCN 551
Query: 382 PS-AHLHAHKYDKFNADYGS 400
S + KY+K+ ++GS
Sbjct: 552 KSVSRADLDKYEKWMREFGS 571
>gb|AAF34687.1| putative microtubule severing protein katanin p60 subunit
[Drosophila melanogaster]
Length = 571
Score = 284 bits (726), Expect = 4e-75
Identities = 163/379 (43%), Positives = 238/379 (62%), Gaps = 17/379 (4%)
Query: 31 QNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVS---PNNMD 87
+N +A G + + + G S +++ A + + G NG D P
Sbjct: 200 RNSTSATAPSGGARTTNGRAGGRKLSTSNTNEARDDDSTAAGNNGGAAGDGENGDPQAAQ 259
Query: 88 ERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYP 147
E +K AE L + L RDI++ P V+W I L +AKRLL+EAVV+P+ P
Sbjct: 260 EEERKFQPNNHIEAE---LVDILERDILQKDPKVRWSDIADLHDAKRLLEEAVVLPMLMP 316
Query: 148 KYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKV 207
YF G+ PWKG+L+ GP GTGKTMLAKAVATEC TTFFN+S++++ SK+RG+SEK+V++
Sbjct: 317 DYFKGIRRPWKGVLMVGPSGTGKTMLAKAVATECGTTFFNVSSATLTSKYRGESEKMVRL 376
Query: 208 LFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE--LVFV 265
LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+ R ++ +V V
Sbjct: 377 LFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGVAREEQAKVVMV 435
Query: 266 LAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEG 325
LAATN PW++D A+ RRLEKRI +PLP E R A+ + L ++S+ + N+ +G
Sbjct: 436 LAATNFPWDIDEALRRRLEKRIYIPLPSDEGREALLKINLREVKVDDSVDLTYVANELKG 495
Query: 326 YSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPEDIQAALKNTRP 382
YSG+DI +C+E +M +RR ++ L EQ L EE +L P+ +D A+
Sbjct: 496 YSGADITNVCREASMMSMRRKIAGLTPEQIRQLATEEVDL----PVSNKDFNEAMSRCNK 551
Query: 383 S-AHLHAHKYDKFNADYGS 400
S + KY+K+ ++GS
Sbjct: 552 SVSRADLDKYEKWMREFGS 570
>gb|EAL28551.1| GA10173-PA [Drosophila pseudoobscura]
Length = 578
Score = 283 bits (723), Expect = 9e-75
Identities = 161/377 (42%), Positives = 235/377 (61%), Gaps = 19/377 (5%)
Query: 32 NGENADKAVGNGSSMSVVSNG-NVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERP 90
N A G +S +N N ++ S D + G NG D +
Sbjct: 212 NSARATNGRAGGRKLSTSNNDRNANNNDSKD-----EDSVAGSNGGGAGDGEGGDTQAGE 266
Query: 91 QKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYF 150
++ P E L + L RDI++ P V+W I L++AKRLL+EAVV+P+ P YF
Sbjct: 267 EERKFQPNNHIEAE-LVDILERDILQRDPKVRWSDIADLQDAKRLLEEAVVLPMLMPDYF 325
Query: 151 TGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQ 210
G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S++++ SK+RG+SEK+V++LF+
Sbjct: 326 KGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSATLTSKYRGESEKMVRLLFE 385
Query: 211 LARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE---LVFVLA 267
+AR +APSTIF+DEID++ S+RG + +EHEASRR+K+ELL+QMDG+ +E +V VLA
Sbjct: 386 MARFYAPSTIFIDEIDSLCSRRG-SETEHEASRRVKSELLVQMDGVGGGEEQAKVVMVLA 444
Query: 268 ATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYS 327
ATN PW++D A+ RRLEKRI +PLP E R A+ + L ++++ + N+ +GYS
Sbjct: 445 ATNFPWDIDEALRRRLEKRIYIPLPSDEGREALLKINLREVKVDDTVDLTYVANELKGYS 504
Query: 328 GSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPEDIQAALKNTRPS- 383
G+DI +C+E +M +RR ++ L EQ L EE +L P+ +D A+ S
Sbjct: 505 GADITNVCREASMMSMRRKIAGLTPEQIRQLATEEVDL----PVSNKDFNEAMSRCNKSV 560
Query: 384 AHLHAHKYDKFNADYGS 400
+ KY+K+ ++GS
Sbjct: 561 SRADLDKYEKWMKEFGS 577
>ref|NP_999733.1| katanin p60 [Strongylocentrotus purpuratus]
gi|3098603|gb|AAC15706.1| katanin p60 subunit
[Strongylocentrotus purpuratus]
gi|60390159|sp|O61577|KTNA1_STRPU Katanin p60
ATPase-containing subunit (Katanin p60 subunit) (p60
katanin)
Length = 516
Score = 282 bits (721), Expect = 1e-74
Identities = 151/305 (49%), Positives = 208/305 (67%), Gaps = 11/305 (3%)
Query: 104 RTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLF 163
+ L E+L RDI++ +P+V W I GL AKRLL+EAVV+P+ P YF G+ PWKG+L+
Sbjct: 214 KDLVENLERDIVQRNPNVHWADIAGLTEAKRLLEEAVVLPLWMPDYFKGIRRPWKGVLMV 273
Query: 164 GPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLD 223
GPPGTGKTMLAKAVATEC TTFFN+S++S+ SK+ G+SEKLV++LF++AR +APSTIF+D
Sbjct: 274 GPPGTGKTMLAKAVATECGTTFFNVSSASLTSKYHGESEKLVRLLFEMARFYAPSTIFID 333
Query: 224 EIDAIISQRGEARSEHEASRRLKTELLIQMDGLT------RTDELVFVLAATNLPWELDA 277
EID+I S+RG SEHEASRR+K+ELLIQMDG++ + ++V VLAATN PW++D
Sbjct: 334 EIDSICSKRGTG-SEHEASRRVKSELLIQMDGVSGPSAGEESSKMVMVLAATNFPWDIDE 392
Query: 278 AMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKE 337
A+ RRLEKRI +PLPE + R + L P + I + + +GYSG+DI +C++
Sbjct: 393 ALRRRLEKRIYIPLPEIDGREQLLRINLKEVPLADDIDLKSIAEKMDGYSGADITNVCRD 452
Query: 338 VAMQPLRRLMSQLEQRE-DLVPEEELPKVGPIRPEDIQAALKNTRPSAHLH-AHKYDKFN 395
+M +RR + L E +P+EEL + P P D AL+ S KY +
Sbjct: 453 ASMMAMRRRIQGLRPEEIRHIPKEELNQ--PSTPADFLLALQKVSKSVGKEDLVKYMAWM 510
Query: 396 ADYGS 400
++GS
Sbjct: 511 EEFGS 515
>ref|XP_321284.2| ENSANGP00000018492 [Anopheles gambiae str. PEST]
gi|55233566|gb|EAA01173.2| ENSANGP00000018492 [Anopheles
gambiae str. PEST]
Length = 492
Score = 281 bits (720), Expect = 2e-74
Identities = 153/330 (46%), Positives = 225/330 (67%), Gaps = 14/330 (4%)
Query: 79 NDVSPNNMDERPQK--SLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLL 136
+D NN + P++ P A++ L + L RDI++ +P++ W+ I L AKRLL
Sbjct: 168 DDEEGNNGGDTPEEVERKFEPASHADV-DLVDMLERDILQKNPNIHWDDIADLHEAKRLL 226
Query: 137 KEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSK 196
+EAVV+P+ P YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S+S++ SK
Sbjct: 227 EEAVVLPMWMPDYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSSTLTSK 286
Query: 197 WRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL 256
+RG+SEKLV++LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+
Sbjct: 287 YRGESEKLVRLLFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGV 345
Query: 257 TRTD--ELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESI 314
+ + ++V VLAATN PW++D A+ RRLEKRI +PLP E R A+ + L +ES+
Sbjct: 346 SNDEATKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNSEGREALLKINLREVKVDESV 405
Query: 315 PYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPE 371
+ ++ +GYSG+DI +C++ +M +RR ++ L EQ L EE +L P+ +
Sbjct: 406 DMRDIADRLDGYSGADITNVCRDASMMSMRRKIAGLRPEQIRQLAKEELDL----PVSKQ 461
Query: 372 DIQAALKNTRPSAHL-HAHKYDKFNADYGS 400
D + A+ S KY ++ ++GS
Sbjct: 462 DFKEAISKCNKSVSKDDLAKYQQWMKEFGS 491
>emb|CAI13718.1| katanin p60 subunit A-like 1 [Homo sapiens]
gi|12653659|gb|AAH00612.1| Katanin p60 subunit A-like 1
[Homo sapiens] gi|60390214|sp|Q9BW62|KATL1_HUMAN Katanin
p60 ATPase-containing subunit A-like 1 (Katanin p60
subunit A-like 1) (p60 katanin-like 1)
gi|14149767|ref|NP_115492.1| katanin p60 subunit A-like
1 [Homo sapiens] gi|62177112|ref|NP_001014402.1| katanin
p60 subunit A-like 1 [Homo sapiens]
Length = 490
Score = 281 bits (720), Expect = 2e-74
Identities = 151/331 (45%), Positives = 219/331 (65%), Gaps = 12/331 (3%)
Query: 79 NDVSPNNMDERPQKSLLPPFESAEM-RTLAESLSRDIIRGSPDVKWESIKGLENAKRLLK 137
+D NM + +P F+ A + L E+L RDI+ +P + W+ I LE AK+LL+
Sbjct: 162 DDKGRKNMQDGASDGEMPKFDGAGYDKDLVEALERDIVSRNPSIHWDDIADLEEAKKLLR 221
Query: 138 EAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKW 197
EAVV+P+ P +F G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S+S++ SK+
Sbjct: 222 EAVVLPMWMPDFFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSSTLTSKY 281
Query: 198 RGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL- 256
RG+SEKLV++LF++AR +AP+TIF+DEID+I S+RG + EHEASRR+K+ELLIQMDG+
Sbjct: 282 RGESEKLVRLLFEMARFYAPTTIFIDEIDSICSRRGTS-DEHEASRRVKSELLIQMDGVG 340
Query: 257 -----TRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDE 311
++V VLAATN PW++D A+ RRLEKRI +PLP + R + + L +
Sbjct: 341 GALENDDPSKMVMVLAATNFPWDIDEALRRRLEKRIYIPLPTAKGRAELLKINLREVELD 400
Query: 312 ESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRE-DLVPEEELPKVGPIRP 370
I + + + EGYSG+DI +C++ ++ +RR ++ L E + +EEL P+
Sbjct: 401 PDIQLEDIAEKIEGYSGADITNVCRDASLMAMRRRINGLSPEEIRALSKEELQM--PVTK 458
Query: 371 EDIQAALKNTRPS-AHLHAHKYDKFNADYGS 400
D + ALK S + KY+K+ ++GS
Sbjct: 459 GDFELALKKIAKSVSAADLEKYEKWMVEFGS 489
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.133 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 695,762,972
Number of Sequences: 2540612
Number of extensions: 29166906
Number of successful extensions: 112653
Number of sequences better than 10.0: 4415
Number of HSP's better than 10.0 without gapping: 2599
Number of HSP's successfully gapped in prelim test: 1816
Number of HSP's that attempted gapping in prelim test: 102957
Number of HSP's gapped (non-prelim): 5735
length of query: 404
length of database: 863,360,394
effective HSP length: 130
effective length of query: 274
effective length of database: 533,080,834
effective search space: 146064148516
effective search space used: 146064148516
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0265.5


