

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0263.5
(295 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
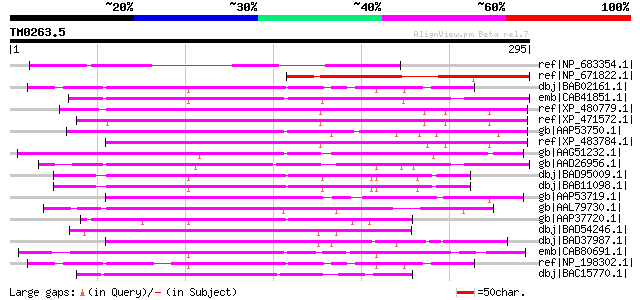
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_683354.1| glycine-rich protein [Arabidopsis thaliana] 126 8e-28
ref|NP_671822.1| hypothetical protein [Arabidopsis thaliana] 113 7e-24
dbj|BAB02161.1| unnamed protein product [Arabidopsis thaliana] g... 98 3e-19
emb|CAB41851.1| hypothetical protein [Arabidopsis thaliana] gi|1... 97 7e-19
ref|XP_480779.1| myb family protein-like [Oryza sativa (japonica... 95 2e-18
ref|XP_471572.1| OSJNBb0032D24.13 [Oryza sativa (japonica cultiv... 95 3e-18
gb|AAP53750.1| putative NAM-like protein [Oryza sativa (japonica... 93 8e-18
ref|XP_483784.1| ribosomal protein-like [Oryza sativa (japonica ... 91 3e-17
gb|AAG51232.1| unknown protein; 59131-63280 [Arabidopsis thalian... 91 4e-17
gb|AAD26956.1| hypothetical protein [Arabidopsis thaliana] gi|61... 91 4e-17
dbj|BAD95009.1| glutathione transferase-like [Arabidopsis thaliana] 90 9e-17
dbj|BAB11098.1| glutathione transferase-like [Arabidopsis thalia... 90 9e-17
gb|AAP53719.1| unknown protein [Oryza sativa (japonica cultivar-... 84 6e-15
gb|AAL79730.1| putative ribosomal protein [Oryza sativa] 84 6e-15
gb|AAP37720.1| At5g41240 [Arabidopsis thaliana] gi|22655095|gb|A... 81 3e-14
dbj|BAD54246.1| myb protein-like [Oryza sativa (japonica cultiva... 80 5e-14
dbj|BAD37987.1| NAM-like protein [Oryza sativa (japonica cultiva... 80 7e-14
emb|CAB80691.1| hypothetical protein [Arabidopsis thaliana] gi|1... 79 2e-13
ref|NP_198302.1| hypothetical protein [Arabidopsis thaliana] 77 8e-13
dbj|BAC15770.1| A [Oryza sativa (japonica cultivar-group)] 76 1e-12
>ref|NP_683354.1| glycine-rich protein [Arabidopsis thaliana]
Length = 268
Score = 126 bits (316), Expect = 8e-28
Identities = 80/211 (37%), Positives = 100/211 (46%), Gaps = 73/211 (34%)
Query: 12 NVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYG 71
NVG T+FPEFST M LG M+ V+E PNEED T ++S W T+QNLVL+S WIKY
Sbjct: 37 NVGATEFPEFSTQMALGVMSGVHETIPNEEDLT--CNHSRSSKWTTDQNLVLLSRWIKYE 94
Query: 72 TSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKR 131
+ +K D KR
Sbjct: 95 QIVSLVETKK---------------------------------------------DNVKR 109
Query: 132 VQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKR 191
+Q SGWS+D +LAKA ++Y+N GN GSGSSGSKR
Sbjct: 110 MQQSGWSKDGVLAKAHELYSNA--------------------------GNTGSGSSGSKR 143
Query: 192 SHESDACGSNSIGSIPRPMGREAAKKKNKKK 222
+HESD +NS+GS RP+G + AKKK KKK
Sbjct: 144 THESDVRDANSVGSSARPIGSDIAKKKTKKK 174
>ref|NP_671822.1| hypothetical protein [Arabidopsis thaliana]
Length = 133
Score = 113 bits (282), Expect = 7e-24
Identities = 69/145 (47%), Positives = 90/145 (61%), Gaps = 30/145 (20%)
Query: 158 PFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKK 217
PF + + L L E Q+GGN SGSSGSKR+HESDA SNS+GS RPM R+AAKK
Sbjct: 12 PFQMQRIQLLLVGEKF---QIGGNTYSGSSGSKRAHESDASDSNSVGSSARPMERDAAKK 68
Query: 218 KNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRL-------EKMKMFLQ 270
K KKK+ QEL+RLEKI+ + E N+L +KMKMF++
Sbjct: 69 KLKKKA--------------------QELDRLEKITMMHSETNQLIKEKTHAKKMKMFIE 108
Query: 271 LSFEEHLDDRRKELLKQLSQELFEN 295
L+ +E+LDD+ K+LL+QLS +LF N
Sbjct: 109 LTEKENLDDKSKKLLQQLSHDLFGN 133
>dbj|BAB02161.1| unnamed protein product [Arabidopsis thaliana]
gi|15232558|ref|NP_188150.1| hypothetical protein
[Arabidopsis thaliana]
Length = 287
Score = 97.8 bits (242), Expect = 3e-19
Identities = 74/280 (26%), Positives = 127/280 (44%), Gaps = 43/280 (15%)
Query: 11 FNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKY 70
F G T+ P FS+ ++ +++ +E++ P+ ++ W ++++VL+S W+
Sbjct: 4 FASGSTEVPAFSSQLS----EEGSDLEGSEDEVKPKQSISRKK-WTAKEDIVLVSAWLNT 58
Query: 71 GTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDG 128
V+G +Q+G+++W +IA Y S D P RE C++ + ++ + K++G Y
Sbjct: 59 SKDPVIGNDQQGQSFWKRIAAYVAASPSLDGLPKREHAKCKHRWGKVNKSVTKFVGCYKT 118
Query: 129 AKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSG 188
+ SG SEDD++ A +IY N + FTL W L + +C G
Sbjct: 119 TTTHKTSGQSEDDVMKLAYEIYFNDTKKN-FTLDHAWRELRYDQKWCEATS---RKGDKN 174
Query: 189 SKRSHESDACGSNSIGSIP----------RPMGREAAKKKNKKKS--------------R 224
+KR CG + S P RP G +AAK K +K +
Sbjct: 175 AKRK----KCGDGNASSQPIHVEDDSVMSRPPGVKAAKAKARKSATIKEGKKPATVKDDS 230
Query: 225 DAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEK 264
++E N W +L+E++ +R EK S QQ L K
Sbjct: 231 GQSVEHFQNLW----ELKEKDWDRKEKQSKHQQLERLLSK 266
>emb|CAB41851.1| hypothetical protein [Arabidopsis thaliana]
gi|15228231|ref|NP_190352.1| expressed protein
[Arabidopsis thaliana] gi|7487364|pir||T07707
hypothetical protein T23J7.10 - Arabidopsis thaliana
Length = 302
Score = 96.7 bits (239), Expect = 7e-19
Identities = 75/282 (26%), Positives = 130/282 (45%), Gaps = 30/282 (10%)
Query: 34 NEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYC 93
++V+PN+ TP+ ++ + W+ ++LVL+S W+ +V+G QKG +WS+IA Y
Sbjct: 30 DDVSPNQAAHTPKVKRERRK-WSAGEDLVLVSAWLNTSKDAVIGNEQKGYAFWSRIAAYY 88
Query: 94 HEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYT 151
+ RE G + + ++ + K++G+Y+ A + + SG ++DD++A A +IY
Sbjct: 89 GASPKLNGVEKRETGHIKQRWTKINEGVGKFVGSYEAATKQKSSGQNDDDVVALAHEIYN 148
Query: 152 NGKNSHPFTLMKEWLALCNEPHYCS--QVGGNLGSGSSGSKRSHESDACGSNSIGSIPRP 209
+ FTL W L E + S + S S ++ S + ++ RP
Sbjct: 149 SEHGK--FTLEHAWRVLRFEQKWLSAPSTKATVMSKRRKSDKASTSQPQTHEAEEAMSRP 206
Query: 210 MGREAAKKKNKKK---------------SRDAALEVVNNEWSEYKQLREQELERLEKISS 254
+G +AAK K KK + E+ +W ++ REQE E EK
Sbjct: 207 IGVKAAKAKAKKAVSKTTTVEDKGNVMLEIQSIWEIKQKDWELRQKDREQEKEDFEK--- 263
Query: 255 VQQEANRLEKMKMFLQL-SFEEHLDDRRKELLKQLSQELFEN 295
RL + K+ L + +E L D L +L E+ N
Sbjct: 264 ----QERLSRTKLLESLFTKKEPLTDIEVALKNKLINEMLSN 301
>ref|XP_480779.1| myb family protein-like [Oryza sativa (japonica cultivar-group)]
gi|38637186|dbj|BAD03438.1| myb family protein-like
[Oryza sativa (japonica cultivar-group)]
gi|38636855|dbj|BAD03121.1| myb family protein-like
[Oryza sativa (japonica cultivar-group)]
Length = 417
Score = 95.1 bits (235), Expect = 2e-18
Identities = 73/289 (25%), Positives = 132/289 (45%), Gaps = 26/289 (8%)
Query: 29 DMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQ 88
++ + E + N +++ R + W E+NL L+S W+ S+ G ++KGE YW
Sbjct: 131 NLVNIEESSDNSQETGRRGTRVN---WTEEENLRLLSSWLNNSLDSINGNDKKGEYYWRD 187
Query: 89 IA-EYCHEHCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQ 147
+A E+ S + R C+ + + + K+ GAY A+R SG+S+D I+ KA
Sbjct: 188 VAAEFNGNASSNNRKRTVVQCKTHWGGVKKDIAKFCGAYSRARRTWSSGFSDDMIMEKAH 247
Query: 148 DIYTNGKNSHPFTLMKEWLALCNEPHYC-----SQVGGNLGSGSSGSKRSHESDACGSNS 202
+Y + N FTL W L ++P + SG+ S + +
Sbjct: 248 ALYKSENNDKTFTLEYMWRELKDQPKWQRILEEDSKNKRTKISESGAYTSSSNQETEEET 307
Query: 203 IGSIPRPMGREAAKKKNKKKSRDAALEVVNNE-------WSEYKQLREQEL----ERLEK 251
RP G++ AK K K K + A + ++ ++E +LR + + E K
Sbjct: 308 SRKEKRPEGQKKAKAKLKGKGKKPAPSPLGDQPSQDFVLFNEAVKLRAEAVLKSAEATTK 367
Query: 252 ISSVQQEANRLEKMKMFLQL------SFEEHLDDRRKELLKQLSQELFE 294
+ ++E R+EK + +L+L +F + R + +L++L+ EL E
Sbjct: 368 SAEAKKEQTRMEKYQTYLKLLDKDTANFSDAKLKRHEAVLEKLATELAE 416
>ref|XP_471572.1| OSJNBb0032D24.13 [Oryza sativa (japonica cultivar-group)]
gi|38344362|emb|CAE04083.2| OSJNBb0032D24.13 [Oryza
sativa (japonica cultivar-group)]
Length = 399
Score = 94.7 bits (234), Expect = 3e-18
Identities = 75/283 (26%), Positives = 130/283 (45%), Gaps = 27/283 (9%)
Query: 39 NEEDSTPRSRKTQSPA----WNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIA-EYC 93
N E+S+ S++T W E+NL L+S W+ S+ G ++KGE YW +A E+
Sbjct: 116 NIEESSDNSQETGRRGTHVNWTEEENLRLLSSWLNNSLDSINGNDKKGEYYWRDVAAEFN 175
Query: 94 HEHCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNG 153
S + R C+ + + + K+ GAY A+R SG+S+D I+ KA +Y +
Sbjct: 176 GNASSNNRKRTVVQCKTHWGGVKKDIAKFCGAYSRARRTWSSGFSDDMIMEKAHALYKSE 235
Query: 154 KNSHPFTLMKEWLALCNEPHYC-----SQVGGNLGSGSSGSKRSHESDACGSNSIGSIPR 208
N FTL W L ++P + SG+ S + + R
Sbjct: 236 NNDKTFTLEYMWRELKDQPKWRRILEEDSKNKRTKISESGAYTSSSNQETEEETSRKEKR 295
Query: 209 PMGREAAKKKNKKKSRDAALEVVNNE-------WSEYKQLREQEL----ERLEKISSVQQ 257
P G++ AK K K K + A + ++ ++E +LR + + E K + ++
Sbjct: 296 PEGQKKAKAKLKGKGKKPAPSPLGDQPSQDFVLFNEAVKLRAEAVLKSAEATTKSAEAKK 355
Query: 258 EANRLEKMKMFLQL------SFEEHLDDRRKELLKQLSQELFE 294
E R+EK + +L+L +F + R + +L++L+ EL E
Sbjct: 356 EQTRMEKYQTYLKLLDKDTANFSDAKLKRHEAVLEKLATELAE 398
>gb|AAP53750.1| putative NAM-like protein [Oryza sativa (japonica cultivar-group)]
gi|37534322|ref|NP_921463.1| putative NAM-like protein
[Oryza sativa (japonica cultivar-group)]
Length = 370
Score = 93.2 bits (230), Expect = 8e-18
Identities = 73/292 (25%), Positives = 131/292 (44%), Gaps = 33/292 (11%)
Query: 33 VNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEY 92
V E E + R +S +NT ++ +L+S W+ ++ G Q ++YW++I +Y
Sbjct: 78 VGEQPSAEVGGSSRPNNKRSKNFNTAEDQMLVSAWLNTTLDAITGVEQHRDSYWARIHQY 137
Query: 93 CHEHCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTN 152
HE+ +F + + NC+ + Q+NK+ G Y+ SG D + +A +Y +
Sbjct: 138 FHENKNFPSDQNQNSLNNCWGTIQEQVNKFCGYYEQILNGPQSGMVVQDYITQAFTLYKS 197
Query: 153 GKNSHPFTLMKEWLALCNEPHYCSQVGGN----------LGSGSSGSKRSHESDACGSNS 202
+ F+LM W L + P + ++ S S+ S H S N+
Sbjct: 198 AEKK-TFSLMHCWEILHHHPKWNDRLFQKRQKAHVDPLVRPSASTNSSEFHYSP--DINT 254
Query: 203 IGSIPRPMGREAAKKK----NKKKSRDAALEVV---NNEWSEYKQL----REQELERLEK 251
+ RP G++ K K N + VV NN WSE K++ RE+ +RL +
Sbjct: 255 SDPLVRPPGKKVEKAKCPRGNTSSCSSESSPVVIALNNMWSEKKEMSAQSREERNDRLVE 314
Query: 252 ISSVQQEANRLEKMKMFLQLSFEEH---------LDDRRKELLKQLSQELFE 294
+ S+++E ++EK K+ ++ E +D RK +L QE+ +
Sbjct: 315 VISLEKERLKVEKRKLDIETHEREERIMNMDLSAMDGPRKTYYLRLQQEIIK 366
>ref|XP_483784.1| ribosomal protein-like [Oryza sativa (japonica cultivar-group)]
gi|45736169|dbj|BAD13215.1| ribosomal protein-like
[Oryza sativa (japonica cultivar-group)]
Length = 303
Score = 91.3 bits (225), Expect = 3e-17
Identities = 71/263 (26%), Positives = 120/263 (44%), Gaps = 23/263 (8%)
Query: 55 WNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIA-EYCHEHCSFDPPREGGACRNCFN 113
W E+NL L+S W+ S+ G ++KGE YW +A E+ S + R C+ +
Sbjct: 40 WTEEENLRLLSSWLNNSLDSINGNDKKGEYYWRDVAAEFNGNTSSNNRKRTVVQCKTHWG 99
Query: 114 YMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPH 173
+ + K+ GAY A+R SG+S+D I+ KA +Y + N FTL W L ++P
Sbjct: 100 GVKKDIAKFCGAYSRARRTWSSGFSDDMIMEKAHALYKSENNDKTFTLEYMWRELKDQPK 159
Query: 174 YC-----SQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAAL 228
+ SG+ S + + RP G++ AK K K K + A
Sbjct: 160 WRRILEEDSKNKRTKISESGAYTSSSNQETEEETSRKEKRPEGQKKAKVKLKGKGKKHAP 219
Query: 229 EVVNNEWS-------EYKQLREQEL----ERLEKISSVQQEANRLEKMKMFLQL------ 271
+ ++ S E +LR + + E K + ++E R+EK + +L+L
Sbjct: 220 SPLGDQPSQDFVLLNEAVKLRAEAVLKSTEATTKSAEAKKEQTRMEKYQTYLKLLDKDTA 279
Query: 272 SFEEHLDDRRKELLKQLSQELFE 294
+F + R + +L++L+ EL E
Sbjct: 280 NFNDVKLKRHEAILEKLATELAE 302
>gb|AAG51232.1| unknown protein; 59131-63280 [Arabidopsis thaliana]
gi|25405091|pir||E96496 unknown protein, 59131-63280
[imported] - Arabidopsis thaliana
Length = 579
Score = 90.9 bits (224), Expect = 4e-17
Identities = 71/296 (23%), Positives = 132/296 (43%), Gaps = 16/296 (5%)
Query: 5 PSQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLI 64
P+ F +G + P +ST + D + ++ + K W++ +++VLI
Sbjct: 290 PNPYECFELGSANVPVYSTEWSDDDPSEDELPIAGKKGKKGKKAKKPRRNWSSTEDVVLI 349
Query: 65 SGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGG--ACRNCFNYMSNQLNKW 122
SGW+ VVG QKG +W +IA Y + + G C+ ++ +++ +NK+
Sbjct: 350 SGWLNTSKDPVVGNEQKGAAFWERIAVYYNSSPKLKGVEKRGHICCKQRWSKVNDAVNKF 409
Query: 123 IGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNL 182
+G+Y A + Q SG ++DD+++ A I++ FT W + + +Q
Sbjct: 410 VGSYLAASKQQTSGQNDDDVVSLAHQIFSKDYGC-KFTCEHAWRERRYDQKWIAQ----- 463
Query: 183 GSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNNEWSEYK--- 239
+ +R E+D+ RP+G +AAK K K + V E + K
Sbjct: 464 STHGKAKRRKCEADSEPVGVEDKEARPIGVKAAKAAAKAKGKAKLSPVEGEETNALKEIQ 523
Query: 240 ---QLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQEL 292
+++E++ EK+ ++ + NR + + L + E L D EL +L EL
Sbjct: 524 SIWEIKEKDHAAKEKLIIIKDKKNRTKFFERLLGKT--EPLSDIEIELKNKLINEL 577
>gb|AAD26956.1| hypothetical protein [Arabidopsis thaliana]
gi|61676918|gb|AAP21683.2| hypothetical protein
[Arabidopsis thaliana] gi|15226698|ref|NP_179211.1|
expressed protein [Arabidopsis thaliana]
gi|25365951|pir||B84537 hypothetical protein At2g16140
[imported] - Arabidopsis thaliana
Length = 311
Score = 90.9 bits (224), Expect = 4e-17
Identities = 78/306 (25%), Positives = 139/306 (44%), Gaps = 53/306 (17%)
Query: 17 DFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVV 76
D P FST T +P+ E+ P+ ++ + W+ ++++VL+S W+ +V+
Sbjct: 32 DVPIFSTQCT---------DSPSLEEHVPKVKRERMK-WSAKEDMVLVSSWLNTSKDAVI 81
Query: 77 GRNQKGETYWSQIAEYCHEHCSFDPPRE--GGACRNCFNYMSNQLNKWIGAYDGAKRVQG 134
G QK T+WS+IA Y + R+ G + + +++ + K++G+Y+ A R +
Sbjct: 82 GNEQKANTFWSRIAAYYDASPQLNGLRKRMQGNIKQRWAKINDGVCKFVGSYEAASREKS 141
Query: 135 SGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKRSHE 194
SG +++D+++ A +I+ N + F L W L ++ +CSQ +S + +
Sbjct: 142 SGQNDNDVISLAHEIF-NNDYGYKFPLEHAWRVLRHDQKWCSQ--------ASVMSKRRK 192
Query: 195 SDACGSNSIGSIP---------RPMGREAAKKKNKK--------KSRDAAL-------EV 230
D S P RP+G +AAK K KK + + A+ E+
Sbjct: 193 CDKAAQPSTSQPPSHGVEEAMSRPIGVKAAKAKAKKTVTKTTTVEDKGNAMLEIQSIWEI 252
Query: 231 VNNEWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQL-SFEEHLDDRRKELLKQLS 289
+W ++ REQE E EK +RL K + L + +E L D L +L
Sbjct: 253 KQKDWELRQKDREQEKEDFEK-------KDRLSKTTLLESLIAKKEPLTDNEVTLKNKLI 305
Query: 290 QELFEN 295
L E+
Sbjct: 306 DFLLEH 311
>dbj|BAD95009.1| glutathione transferase-like [Arabidopsis thaliana]
Length = 342
Score = 89.7 bits (221), Expect = 9e-17
Identities = 68/262 (25%), Positives = 120/262 (44%), Gaps = 33/262 (12%)
Query: 26 TLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETY 85
T+G+ + + N D RK W+ ++ +LIS W+ +V K +
Sbjct: 1 TIGEKSDDPNLVQNTTDRRKHRRK-----WSRAEDAILISAWLNTSKDPIVDNEHKACAF 55
Query: 86 WSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDIL 143
W +I Y + S P RE C+ ++ +++++ K++G YD A + SG SEDD+
Sbjct: 56 WKRIGAYFNNSASLANLPKREPSHCKQRWSKLNDKVCKFVGCYDQALNQRSSGQSEDDVF 115
Query: 144 AKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCS--QVGGNLGSGSSGSKRSHESDACGSN 201
A +YTN S+ FTL W L + +CS + G GSS + + D S+
Sbjct: 116 QVAYQVYTNNYKSN-FTLEHAWRELRHSKKWCSLYPFENSKGGGSSKRTKLNNGDRVYSS 174
Query: 202 SIG--SIP-----------RPMGREAAKKKNKKKSRDAALEV--------VNNEWSEYKQ 240
S S+P P+G +++K+K KK + +E + N W ++
Sbjct: 175 SSNPESVPIALDEEEQVMDLPLGVKSSKQKEKKVATIITIEEREADSGSRLENLWVLDEE 234
Query: 241 LREQELERLEKISSVQQEANRL 262
EQ ++R + S++Q+ N++
Sbjct: 235 --EQVMDRPLGVKSLEQKENKV 254
>dbj|BAB11098.1| glutathione transferase-like [Arabidopsis thaliana]
gi|15237595|ref|NP_198938.1| glutathione S-transferase,
putative [Arabidopsis thaliana]
Length = 590
Score = 89.7 bits (221), Expect = 9e-17
Identities = 68/262 (25%), Positives = 120/262 (44%), Gaps = 33/262 (12%)
Query: 26 TLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETY 85
T+G+ + + N D RK W+ ++ +LIS W+ +V K +
Sbjct: 249 TIGEKSDDPNLVQNTTDRRKHRRK-----WSRAEDAILISAWLNTSKDPIVDNEHKACAF 303
Query: 86 WSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDIL 143
W +I Y + S P RE C+ ++ +++++ K++G YD A + SG SEDD+
Sbjct: 304 WKRIGAYFNNSASLANLPKREPSHCKQRWSKLNDKVCKFVGCYDQALNQRSSGQSEDDVF 363
Query: 144 AKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCS--QVGGNLGSGSSGSKRSHESDACGSN 201
A +YTN S+ FTL W L + +CS + G GSS + + D S+
Sbjct: 364 QVAYQVYTNNYKSN-FTLEHAWRELRHSKKWCSLYPFENSKGGGSSKRTKLNNGDRVYSS 422
Query: 202 SIG--SIP-----------RPMGREAAKKKNKKKSRDAALEV--------VNNEWSEYKQ 240
S S+P P+G +++K+K KK + +E + N W ++
Sbjct: 423 SSNPESVPIALDEEEQVMDLPLGVKSSKQKEKKVATIITIEEREADSGSRLENLWVLDEE 482
Query: 241 LREQELERLEKISSVQQEANRL 262
EQ ++R + S++Q+ N++
Sbjct: 483 --EQVMDRPLGVKSLEQKENKV 502
>gb|AAP53719.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37534260|ref|NP_921432.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 300
Score = 83.6 bits (205), Expect = 6e-15
Identities = 67/245 (27%), Positives = 111/245 (44%), Gaps = 21/245 (8%)
Query: 55 WNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFD-PPREGGACRNCFN 113
W E+NL L+S WI Y T V G ++K E YW +A+ + + R + +
Sbjct: 67 WTEEENLHLLSYWIHYSTDPVKGIDRKSEFYWKAVADVFNTSAPTNGHKRSVKQLKTHWG 126
Query: 114 YMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPH 173
+ + K+ G Y K SG S+D ++ A I+ PFTL W + + P
Sbjct: 127 DVKRDITKFCGVYGRLKATWKSGQSDDMVMNSAHAIFEKENKDKPFTLEYMWREVKDLPK 186
Query: 174 YCSQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNN 233
+C V SG+KR+ E +I RP G++ AK K K +DAA + N
Sbjct: 187 WCRIV-----QEDSGNKRTKE------ETISKEKRPDGQKKAKAGLKGKGKDAAPSPLGN 235
Query: 234 EWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQL------SFEEHLDDRRKELLKQ 287
+ S+ L E + ++ ++ + +K + +L+L +F E R + +L Q
Sbjct: 236 QPSQNMILYH---EAMSLKATAMIKSAKEKKYQTYLKLLEKDTSNFSEAHHKRHEGVLDQ 292
Query: 288 LSQEL 292
L++EL
Sbjct: 293 LAKEL 297
>gb|AAL79730.1| putative ribosomal protein [Oryza sativa]
Length = 915
Score = 83.6 bits (205), Expect = 6e-15
Identities = 67/268 (25%), Positives = 123/268 (45%), Gaps = 32/268 (11%)
Query: 20 EFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRN 79
++ST ++ D T VN + D TPR+ K W E+++ ++S W+ T S VG +
Sbjct: 114 DYSTPISARDNTYVNV---DSGDETPRTEKRIF--WTQEEDVRMMSSWLLNSTDSTVGAD 168
Query: 80 QKGETYWSQIAEYCHEHCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSE 139
+K E YW+ + +E R ++ F+ ++ + + A+ A+ + SG+++
Sbjct: 169 RKNEQYWTDVEATYNETTPSHRRRNAKQIKDRFHKVNKWTDLFHSAWLKARMIYTSGYND 228
Query: 140 DDILAKAQDIYTNGK---NSHPFTLMKEWLALCNEPHYCSQVGGNLGS-------GSSGS 189
+ KA Y N PF LM+ W + E + + G + G
Sbjct: 229 QMWIEKAHVFYIKDNEKLNLGPFVLMEVWNTVKTEAKWITYNNGLKAARKRIATKGLGKE 288
Query: 190 KRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNNEWSEYKQLREQELERL 249
K +S + + PRPMG++ AKK ++++ K++ +LE L
Sbjct: 289 KEGEDSSPLYVDELDEQPRPMGQKRAKKL---------------QYAQSKEVDHIDLEEL 333
Query: 250 EKISSVQ--QEANRLEKMKMFLQLSFEE 275
+K S +Q Q ANRL+ +++ +LS E+
Sbjct: 334 DKFSKLQNEQNANRLKVLEIQQKLSSEK 361
>gb|AAP37720.1| At5g41240 [Arabidopsis thaliana] gi|22655095|gb|AAM98138.1|
glutathione transferase [Arabidopsis thaliana]
gi|30693769|ref|NP_198940.3| glutathione S-transferase,
putative [Arabidopsis thaliana]
Length = 591
Score = 81.3 bits (199), Expect = 3e-14
Identities = 58/207 (28%), Positives = 93/207 (44%), Gaps = 21/207 (10%)
Query: 41 EDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSS--VVGRNQKGETYWSQIAEYCHEHCS 98
+D+T RK + W+ +++LIS W+ VV Q+ T+W +I + S
Sbjct: 261 QDTT--DRKARRRKWSPPDDVILISAWLNTSKDRKVVVYDEQQAHTFWKRIGAHVSNSAS 318
Query: 99 FD--PPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNS 156
P RE CR + +++ + K++G YD A + SG SEDD+ A +Y N S
Sbjct: 319 LANLPKREWNHCRQRWRKINDYVCKFVGCYDQALNQRASGQSEDDVFQVAYQLYYNNYMS 378
Query: 157 HPFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKRSH------ESDACGSNSI------- 203
+ F L W L + +CS G SKR+ S +C S+
Sbjct: 379 N-FKLEHAWRELRHNKKWCSTYTSENSKGGGSSKRTKLNGGGVYSSSCNPESVPIALDGE 437
Query: 204 -GSIPRPMGREAAKKKNKKKSRDAALE 229
+ RP+G +++K+K KK + LE
Sbjct: 438 EQVMDRPLGVKSSKQKEKKVATKTMLE 464
>dbj|BAD54246.1| myb protein-like [Oryza sativa (japonica cultivar-group)]
gi|53792023|dbj|BAD54608.1| myb protein-like [Oryza
sativa (japonica cultivar-group)]
Length = 409
Score = 80.5 bits (197), Expect = 5e-14
Identities = 57/202 (28%), Positives = 91/202 (44%), Gaps = 8/202 (3%)
Query: 35 EVTPNEE--DSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIA-E 91
EV EE DS+ R+ W + N+ L+S W+ + G ++K E YW +A E
Sbjct: 141 EVVEVEEASDSSEEGRRGTRINWTEDDNIRLMSSWLNNSVDPIKGNDKKSEQYWKAVARE 200
Query: 92 YCHEHCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYT 151
+ S R CR ++ + + K+ G Y A+ SG+S+D I+ KA++ Y
Sbjct: 201 FNSNMPSNGNKRNPKQCRTHWDNVKRDVTKFCGFYSKARTTFTSGYSDDMIMEKAREWYK 260
Query: 152 NGKNSHPFTLMKEWLALCNEPHY---CSQVGGNLGS--GSSGSKRSHESDACGSNSIGSI 206
N PFTL W L ++P + + N S SG+ S + +
Sbjct: 261 KHNNQKPFTLEYMWKDLKDQPKWRRVLEESSHNKRSKISESGAYTSSSNQDTEEETERKE 320
Query: 207 PRPMGREAAKKKNKKKSRDAAL 228
RP G++AAK++ K K + L
Sbjct: 321 KRPEGQKAAKQRQKGKGAPSPL 342
>dbj|BAD37987.1| NAM-like protein [Oryza sativa (japonica cultivar-group)]
Length = 376
Score = 80.1 bits (196), Expect = 7e-14
Identities = 66/257 (25%), Positives = 117/257 (44%), Gaps = 31/257 (12%)
Query: 55 WNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGGACRNCFNY 114
+ T+++++L+S W+ G + G +Q TYW++I EY H + F+ R + N ++
Sbjct: 112 FTTQEDILLVSAWLNVGMDPIQGVDQSQGTYWARIHEYFHANKEFESTRSESSLLNRWSA 171
Query: 115 MSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSH-PFTLMKEWLALCNEPH 173
+ + +N + G + SG DD +A A +++ H F LM W L ++P
Sbjct: 172 IQHDVNIFCGCMSRIEARNQSGSRVDDKIANACELFKEEDKKHRKFNLMHCWNILKDKPK 231
Query: 174 Y--------CSQVGGN-----LGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKK-- 218
+ C++ N + + S S + D S++ S+ RP G++AAK+K
Sbjct: 232 WMDNRKKVGCAKKPSNKKQKTVANSSPTSVEPADLDVYCSDAQPSV-RPDGKKAAKQKLR 290
Query: 219 ------------NKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEKMK 266
KKK D E+ E + K QE ER ++ ++Q+E N+LE+ K
Sbjct: 291 QGRTIEAVDYLMEKKKEADVVRELKKEEMCK-KAFALQE-ERCKRAFALQEERNKLEREK 348
Query: 267 MFLQLSFEEHLDDRRKE 283
Q E + +E
Sbjct: 349 FEFQKKEAEKAEKVEEE 365
>emb|CAB80691.1| hypothetical protein [Arabidopsis thaliana]
gi|15235196|ref|NP_192107.1| hypothetical protein
[Arabidopsis thaliana] gi|4558567|gb|AAD22660.1|
hypothetical protein [Arabidopsis thaliana]
gi|25365949|pir||D85025 hypothetical protein AT4g01980
[imported] - Arabidopsis thaliana
Length = 302
Score = 78.6 bits (192), Expect = 2e-13
Identities = 76/301 (25%), Positives = 126/301 (41%), Gaps = 38/301 (12%)
Query: 6 SQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLIS 65
++T + VG ++ P F T + E+ W ++++VLIS
Sbjct: 26 NETNQYEVGSSEIPVFGTQ----------RCQESHEERYASRVAIARKKWAAKEDIVLIS 75
Query: 66 GWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWI 123
W+ VVG QK +W +IA Y + D P R C+ + M+ + K++
Sbjct: 76 AWLNTSKDPVVGNEQKAPAFWKRIASYVAANPDLDGVPKRASAQCKQRWAKMNELVMKFV 135
Query: 124 GAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLG 183
G Y + SG +E+D++ A +++ N F+L W L ++ + + N
Sbjct: 136 GCYATTTNQKASGQTENDVMLFANELFENDMKK-KFSLDHAWRLLRHDQKW---LISNAP 191
Query: 184 SGSSGSKRSHESDACGSNSIGSIP----------RPMGREAAKKKNKKKSRDAALEVVNN 233
G +KR G+ + S P RP G +AAK K KK EV +
Sbjct: 192 KGKGIAKR--RKVRVGTQAASSQPIDLEDDDVMRRPPGVKAAKAKAKKTPTVKGEEVNSV 249
Query: 234 E-WSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQEL 292
E + L+E++ +R +K Q + LE + LS E L + L K+L +EL
Sbjct: 250 EHLTSLWDLKERDWDRKDK----QSKNQMLESL-----LSRTEPLSECDLVLKKKLIEEL 300
Query: 293 F 293
F
Sbjct: 301 F 301
>ref|NP_198302.1| hypothetical protein [Arabidopsis thaliana]
Length = 301
Score = 76.6 bits (187), Expect = 8e-13
Identities = 70/280 (25%), Positives = 117/280 (41%), Gaps = 52/280 (18%)
Query: 11 FNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKY 70
F G T+ P FS+ ++ +E+ +E++ P+ ++ W ++++VL+S W+
Sbjct: 27 FASGSTEVPAFSSQLS----EEGSELEGSEDEVKPKQSISRKK-WTAKEDIVLVSAWLNP 81
Query: 71 GTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDG 128
V+G +Q+ Y S D P RE C+ + ++ + K++ Y
Sbjct: 82 SKDPVIGNDQQA---------YVAASPSLDGLPKREHAKCKQRWGKVNKSVTKFVACYKT 132
Query: 129 AKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSG 188
+ SG SEDD++ A +IY N + FTL W L + +C G
Sbjct: 133 TTTHKTSGQSEDDVMKLAYEIYFNDTKKN-FTLDHAWRELRYDQKWCEATS---RKGDEN 188
Query: 189 SKRSHESDACGSNSIGSIP----------RPMGREAAKKKNKKKS--------------R 224
+KR CG + S P RP G +AAK K +K +
Sbjct: 189 AKRR----KCGDGNASSHPIHVEDDSIMSRPPGVKAAKAKGRKSAIVKEGKKPATVKDDS 244
Query: 225 DAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEK 264
++E N W +L+E++ +R EK S QQ L K
Sbjct: 245 GQSVEHFQNLW----ELKEKDWDRKEKQSKHQQLERLLSK 280
>dbj|BAC15770.1| A [Oryza sativa (japonica cultivar-group)]
Length = 352
Score = 76.3 bits (186), Expect = 1e-12
Identities = 50/192 (26%), Positives = 95/192 (49%), Gaps = 15/192 (7%)
Query: 39 NEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCS 98
+EED+ SR + W E+ L S W+ S+ G ++KG+T+W ++ + ++ +
Sbjct: 89 DEEDNNDDSRAAKK-RWTHEEEERLASAWLNASKDSIHGNDKKGDTFWKEVTDEFNKKGN 147
Query: 99 FDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHP 158
RE + ++ + + ++++ + ++ SG+S+D + +AQ +Y N + P
Sbjct: 148 GKRRREINQLKVHWSRLKSAISEFNDYWSTVTQMHTSGYSDDMLEKEAQRLYAN-RFGKP 206
Query: 159 FTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAK-K 217
F L+ W L +EP +C+Q K E DA RP+GREAAK +
Sbjct: 207 FALVHWWKILKDEPKWCAQF--------ESEKDKSEMDAVPEQQ----SRPIGREAAKSE 254
Query: 218 KNKKKSRDAALE 229
+N K+ ++ +E
Sbjct: 255 RNGKRKKENVME 266
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.312 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 549,183,156
Number of Sequences: 2540612
Number of extensions: 24189821
Number of successful extensions: 81190
Number of sequences better than 10.0: 340
Number of HSP's better than 10.0 without gapping: 108
Number of HSP's successfully gapped in prelim test: 237
Number of HSP's that attempted gapping in prelim test: 80335
Number of HSP's gapped (non-prelim): 836
length of query: 295
length of database: 863,360,394
effective HSP length: 127
effective length of query: 168
effective length of database: 540,702,670
effective search space: 90838048560
effective search space used: 90838048560
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0263.5


